IMMIGRATE: A Margin-Based Feature Selection Method with Interaction Terms
Abstract
1. Introduction
2. Review: The Relief Algorithm
| Algorithm 1 The Original Relief Algorithm |
| N: the number of training instances. |
| A: the number of features (i.e., attributes). |
| M: the number of randomly chosen training samples to update feature weight . |
| Input: a training dataset . |
| Initialization: Initialize all feature weights to 0: . |
| for i = 1 to M do |
| Randomly select an instance and find its and . |
| Update the feature weights by , |
| where the square operation is element-wise. |
| Return: . |
3. IMMIGRATE Algorithm
3.1. Hypothesis-Margin
3.2. Entropy to Measure Margin Stability
3.3. Quadratic-Manhattan Measurement
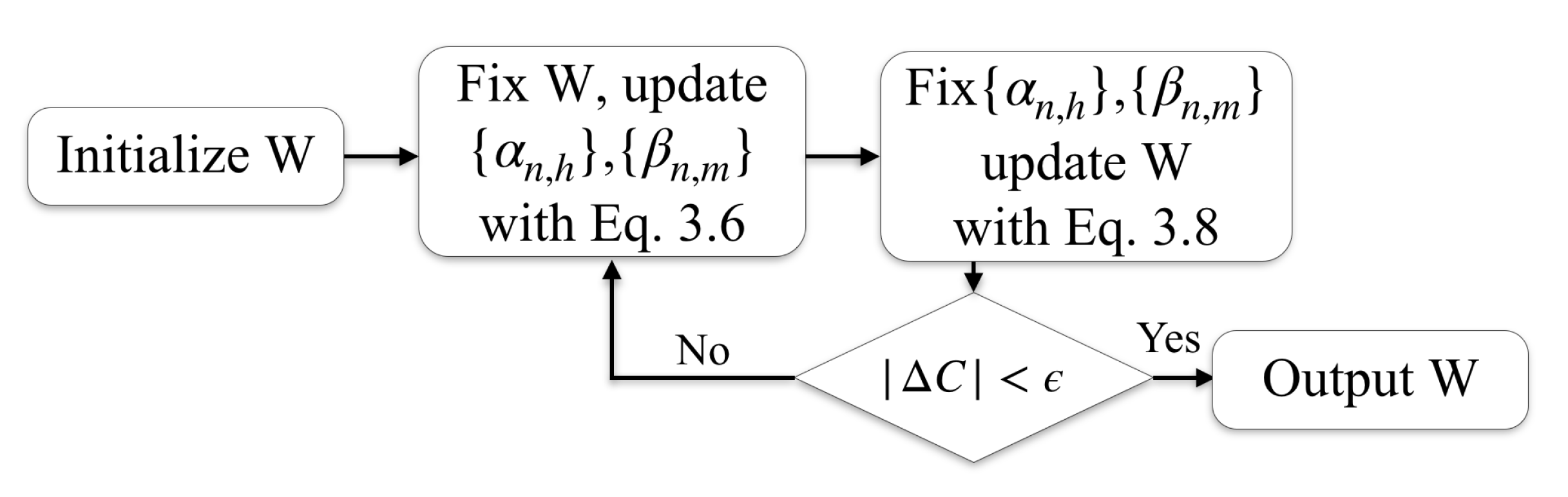
3.4. IMMIGRATE
| Algorithm 2 The IMMIGRATE Algorithm |
| Input: a training dataset . |
| Initialization: Let , randomly initialize satisfying , , . |
| repeat |
| Calculate , with Equation (6). |
| Calculate with Theorem 1, Equation (8). |
| . |
| until the change of C in Equation (5) is small enough or the iteration indicator t reaches a preset limit. |
| Output: . |
3.4.1. Step 1: Fix , Update and
3.4.2. Step 2: Fix and , Update
3.4.3. Weight Pruning
3.4.4. Predict New Samples
3.5. IMMIGRATE in Ensemble Learning
| Algorithm 3 The BIM Algorithm |
| T: the number of classifiers for BIM. |
| Input: a training dataset . |
| Initialization: for each , set . |
| for t: = 1 to T do |
| Limit max number of iteration of IMMIGRATE less than preset. |
| Train weak IMMIGRATE classifier using a chosen and weights by Equation (19). |
| Compute the error rate as . |
| if or then |
| Discard , and continue. |
| Set . |
| Update : For each , |
| . |
| Normalize , so that . |
| Output: . |
3.6. IMMIGRATE for High-Dimensional Data Space
4. Experiments
4.1. Synthetic Dataset
4.2. Real Datasets
4.2.1. Gene Expression Datasets
4.2.2. UCI Datasets
5. Related Works
6. Conclusions and Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| NH | Nearest Hit |
| NM | Nearest Miss |
| IM4E | Iterative Margin-Maximization under Max-Min Entropy algorithm |
| IMMIGRATE | Iterative Max-MIn entropy marGin-maximization with inteRAction TErms algorithm |
Appendix A. Information of the Real Datasets
Appendix B. Heat Maps
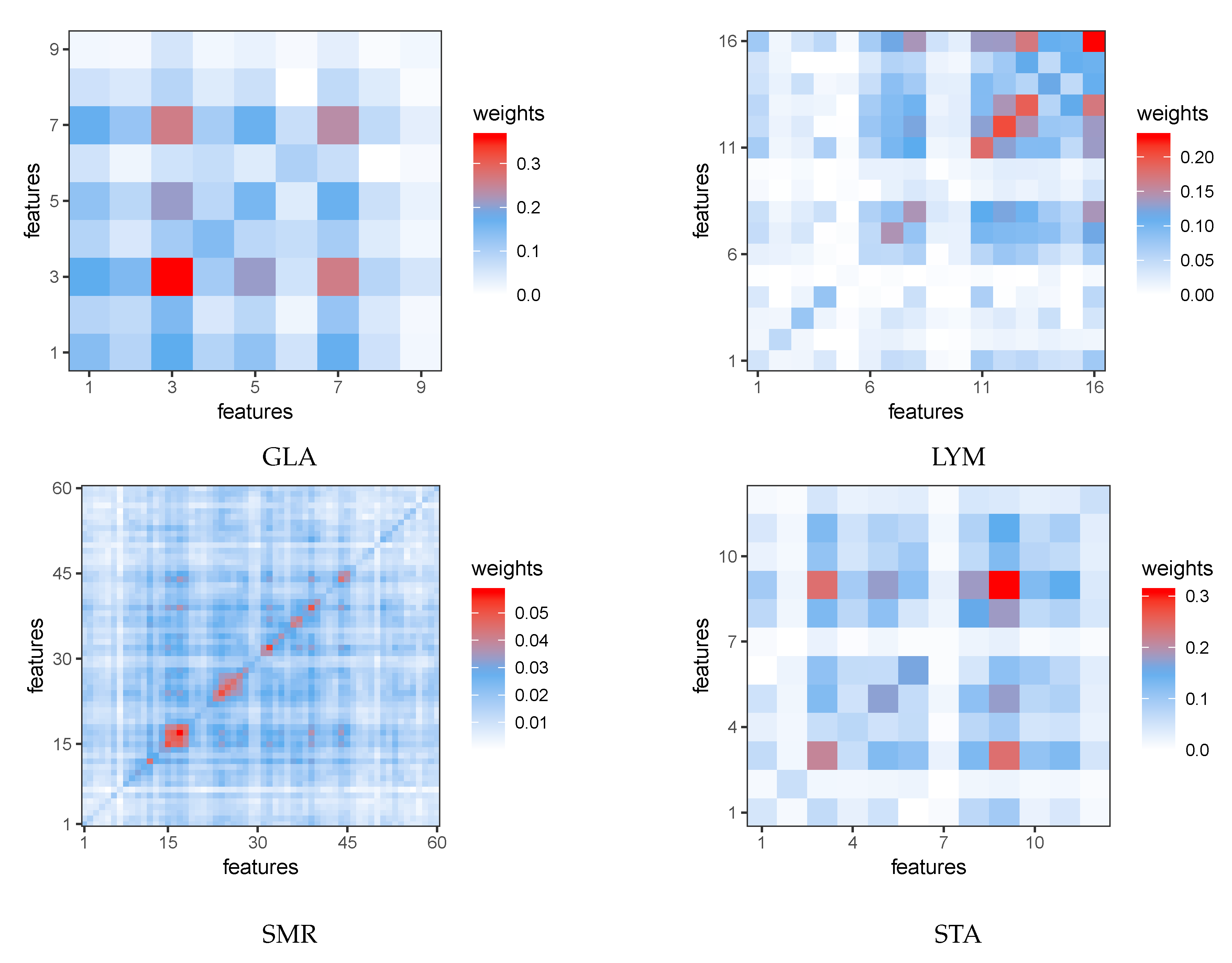
References
- Fukunaga, K. Introduction to Statistical Pattern Recognition; Elsevier: Amsterdam, The Netherlands, 2013. [Google Scholar]
- Kira, K.; Rendell, L.A. A practical approach to feature selection. In Machine Learning Proceedings 1992; Morgan Kaufmann: Burlington, MA, USA, 1992; pp. 249–256. [Google Scholar]
- Gilad-Bachrach, R.; Navot, A.; Tishby, N. Margin based feature selection-theory and algorithms. In Proceedings of the 21st International Conference on Machine Learning, Banff, AB, Canada, 4–8 July 2004; p. 43. [Google Scholar]
- Kononenko, I. Estimating attributes: Analysis and extensions of RELIEF. In European Conference on Machine Learning; Springer: Berlin, Germany, 1994; pp. 171–182. [Google Scholar]
- Yang, M.; Wang, F.; Yang, P. A Novel Feature Selection Algorithm Based on Hypothesis-Margin. JCP 2008, 3, 27–34. [Google Scholar] [CrossRef]
- Sun, Y.; Li, J. Iterative RELIEF for feature weighting. In Proceedings of the 23rd International Conference on Machine Learning, Pittsburgh, PA, USA, 25–29 June 2006; pp. 913–920. [Google Scholar]
- Sun, Y.; Wu, D. A relief based feature extraction algorithm. In Proceedings of the 2008 SIAM International Conference on Data Mining, Atlanta, GA, USA, 24–26 April 2008; pp. 188–195. [Google Scholar]
- Bei, Y.; Hong, P. Maximizing margin quality and quantity. In Proceedings of the 2015 IEEE 25th International Workshop on Machine Learning for Signal Processing (MLSP), Boston, MA, USA, 17–20 September 2015; pp. 1–6. [Google Scholar]
- Urbanowicz, R.J.; Meeker, M.; La Cava, W.; Olson, R.S.; Moore, J.H. Relief-based feature selection: Introduction and review. J. Biomed. Inform. 2018, 85, 189–203. [Google Scholar] [CrossRef] [PubMed]
- Schapire, R.E. The strength of weak learnability. Mach. Learn. 1990, 5, 197–227. [Google Scholar] [CrossRef]
- Kuhn, H.W.; Tucker, A.W. Nonlinear programming. In Traces and Emergence of Nonlinear Programming; Springer: Berlin, Germany, 2014; pp. 247–258. [Google Scholar]
- Li, Y.; Liu, J.S. Robust variable and interaction selection for logistic regression and general index models. J. Am. Stat. Assoc. 2018, 114, 1–16. [Google Scholar] [CrossRef]
- Bien, J.; Taylor, J.; Tibshirani, R. A lasso for hierarchical interactions. Ann. Stat. 2013, 41, 1111. [Google Scholar] [CrossRef]
- Freund, Y.; Schapire, R.E. Experiments with a new boosting algorithm. Icml 1996, 96, 148–156. [Google Scholar]
- Freund, Y.; Mason, L. The alternating decision tree learning algorithm. Icml 1999, 99, 124–133. [Google Scholar]
- Soentpiet, R. Advances in Kernel Methods: Support Vector Learning; MIT Press: Cambridge, MA, USA, 1999. [Google Scholar]
- Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. 1996, 58, 267–288. [Google Scholar] [CrossRef]
- John, G.H.; Langley, P. Estimating continuous distributions in Bayesian classifiers. In Proceedings of the Eleventh Conference on Uncertainty in Artificial Intelligence; Morgan Kaufmann Publishers Inc.: Burlington, MA, USA, 1995; pp. 338–345. [Google Scholar]
- Haykin, S. Neural Networks: A Comprehensive Foundation; Prentice Hall PTR: Upper Saddle River, NJ, USA, 1994. [Google Scholar]
- Aha, D.W.; Kibler, D.; Albert, M.K. Instance-based learning algorithms. Mach. Learn. 1991, 6, 37–66. [Google Scholar] [CrossRef]
- Weinberger, K.Q.; Saul, L.K. Distance metric learning for large margin nearest neighbor classification. J. Mach. Learn. Res. 2009, 10, 207–244. [Google Scholar]
- Robnik-Šikonja, M.; Kononenko, I. Theoretical and empirical analysis of ReliefF and RReliefF. Mach. Learn. 2003, 53, 23–69. [Google Scholar] [CrossRef]
- Fisher, R.A. The use of multiple measurements in taxonomic problems. Ann. Eugen. 1936, 7, 179–188. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. Xgboost: A scalable tree boosting system. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 785–794. [Google Scholar]
- Freije, W.A.; Castro-Vargas, F.E.; Fang, Z.; Horvath, S.; Cloughesy, T.; Liau, L.M.; Mischel, P.S.; Nelson, S.F. Gene expression profiling of gliomas strongly predicts survival. Cancer Res. 2004, 64, 6503–6510. [Google Scholar] [CrossRef] [PubMed]
- Alon, U.; Barkai, N.; Notterman, D.A.; Gish, K.; Ybarra, S.; Mack, D.; Levine, A.J. Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proc. Natl. Acad. Sci. USA 1999, 96, 6745–6750. [Google Scholar] [CrossRef]
- Tian, E.; Zhan, F.; Walker, R.; Rasmussen, E.; Ma, Y.; Barlogie, B.; Shaughnessy, J.D., Jr. The role of the Wnt-signaling antagonist DKK1 in the development of osteolytic lesions in multiple myeloma. N. Engl. J. Med. 2003, 349, 2483–2494. [Google Scholar] [CrossRef] [PubMed]
- Van’t Veer, L.J.; Dai, H.; Van De Vijver, M.J.; He, Y.D.; Hart, A.A.; Mao, M.; Peterse, H.L.; Van Der Kooy, K.; Marton, M.J.; Witteveen, A.T.; et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 2002, 415, 530. [Google Scholar] [CrossRef]
- Singh, D.; Febbo, P.G.; Ross, K.; Jackson, D.G.; Manola, J.; Ladd, C.; Tamayo, P.; Renshaw, A.A.; D’Amico, A.V.; Richie, J.P.; et al. Gene expression correlates of clinical prostate cancer behavior. Cancer Cell 2002, 1, 203–209. [Google Scholar] [CrossRef]
- Frank, A.; Asuncion, A. UCI Machine Learning Repository. Available online: http://archive.ics.uci.edu/ml (accessed on 1 August 2019).
- Garcia, S.; Derrac, J.; Cano, J.; Herrera, F. Prototype selection for nearest neighbor classification: Taxonomy and empirical study. IEEE Trans. Pattern Anal. Mach. Intell. 2012, 34, 417–435. [Google Scholar] [CrossRef]
- Yu, S.; Giraldo, L.G.S.; Jenssen, R.; Principe, J.C. Multivariate Extension of Matrix-based Renyi’s α-order Entropy Functional. IEEE Trans. Pattern Anal. Mach. Intell. 2019. [Google Scholar] [CrossRef]
- Vinh, N.X.; Zhou, S.; Chan, J.; Bailey, J. Can high-order dependencies improve mutual information based feature selection? Pattern Recognit. 2016, 53, 46–58. [Google Scholar] [CrossRef]



| Data | SV1 | SV2 | LAS | DT | NBC | 1NN | 3NN | SOD | RF | XGB | IM4 | EGT | B4G |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GLI | 85.1 | 86.0 | 85.2 | 83.8 | 83.0 | 88.7 | 87.7 | 88.7 | 87.6 | 86.3 | 87.5 | 89.1 | 89.9 |
| COL | 73.7 | 82.0 | 80.6 | 69.2 | 71.1 | 72.1 | 77.9 | 78.1 | 82.6 | 79.5 | 84.3 | 78.6 | 82.5 |
| ELO | 72.9 | 90.2 | 74.6 | 77.3 | 76.3 | 85.6 | 91.3 | 86.9 | 79.2 | 77.9 | 88.9 | 88.6 | 88.4 |
| BRE | 76.0 | 88.7 | 91.4 | 76.4 | 69.4 | 83.0 | 73.6 | 82.6 | 86.3 | 87.3 | 88.1 | 90.2 | 91.5 |
| PRO | 71.3 | 69.9 | 87.9 | 86.4 | 68.0 | 83.2 | 82.7 | 83.2 | 91.8 | 90.5 | 88.0 | 89.5 | 89.7 |
| W,T,L | 5,0,0 | 4,0,1 | 4,1,0 | 5,0,0 | 5,0,0 | 5,0,0 | 4,0,1 | 5,0,0 | 3,1,1 | 4,0,1 | 3,1,1 | -,-,- | -,-,- |
| Data | SV1 | SV2 | LAS | DT | NBC | RBF | 1NN | 3NN | LMN | REL | RFF | SIM | LFE | LDA | SOD | hIN | IM4 | IGT |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BCW | 61.4 | 66.6 | 71.4 | 70.5 | 62.4 | 56.9 | 68.2 | 72.2 | 69.5 | 66.4 | 67.1 | 67.7 | 67.1 | 73.9 | 65.2 | 71.8 | 66.4 | 74.5 |
| CRY | 72.9 | 90.6 | 87.4 | 85.3 | 84.4 | 89.7 | 89.1 | 85.4 | 87.8 | 73.8 | 77.2 | 79.7 | 86.0 | 88.6 | 86.0 | 87.9 | 86.2 | 89.8 |
| CUS | 86.5 | 88.9 | 89.6 | 89.6 | 89.5 | 86.8 | 86.5 | 88.7 | 88.8 | 82.1 | 84.7 | 84.3 | 86.4 | 90.3 | 90.8 | 90.3 | 87.5 | 90.1 |
| ECO | 92.9 | 96.9 | 98.6 | 98.6 | 97.8 | 94.6 | 96.0 | 97.8 | 97.8 | 89.0 | 90.7 | 91.2 | 93.1 | 99.0 | 97.9 | 98.7 | 97.5 | 98.2 |
| GLA | 64.2 | 76.7 | 72.3 | 79.4 | 69.5 | 73.0 | 81.1 | 78.1 | 79.4 | 64.1 | 63.5 | 67.1 | 81.2 | 72.0 | 75.3 | 75.0 | 78.0 | 87.5 |
| HMS | 63.8 | 64.5 | 67.7 | 72.5 | 67.2 | 66.8 | 66.0 | 69.3 | 71.2 | 65.3 | 66.0 | 65.7 | 64.9 | 69.0 | 67.4 | 69.4 | 66.6 | 69.2 |
| IMM | 74.3 | 70.6 | 74.4 | 84.1 | 77.9 | 67.3 | 69.4 | 77.9 | 76.7 | 69.9 | 71.8 | 69.0 | 75.0 | 75.2 | 72.3 | 70.2 | 80.7 | 83.8 |
| ION | 80.5 | 93.5 | 83.6 | 87.4 | 89.4 | 79.9 | 86.7 | 84.1 | 84.5 | 85.8 | 86.2 | 84.2 | 91.0 | 83.3 | 90.3 | 92.6 | 88.3 | 92.9 |
| LYM | 83.6 | 81.5 | 85.2 | 75.2 | 83.6 | 71.1 | 77.2 | 82.8 | 86.6 | 64.9 | 71.0 | 70.4 | 79.6 | 85.2 | 79.3 | 84.8 | 83.3 | 87.2 |
| MON | 74.4 | 91.7 | 75.0 | 86.4 | 74.0 | 68.2 | 75.1 | 84.4 | 84.9 | 61.4 | 61.8 | 65.0 | 64.8 | 74.4 | 91.9 | 97.2 | 75.6 | 99.5 |
| PAR | 72.7 | 72.5 | 77.1 | 84.8 | 74.1 | 71.5 | 94.6 | 91.4 | 91.8 | 87.3 | 90.3 | 84.6 | 94.0 | 85.6 | 88.2 | 89.5 | 83.2 | 93.8 |
| PID | 65.6 | 73.1 | 74.7 | 74.3 | 71.2 | 70.3 | 70.3 | 73.5 | 74.0 | 64.8 | 68.0 | 67.0 | 67.8 | 74.5 | 75.7 | 74.1 | 72.1 | 74.7 |
| SMR | 73.5 | 83.9 | 73.6 | 72.3 | 70.3 | 67.1 | 86.9 | 84.7 | 86.1 | 69.5 | 78.3 | 81.0 | 84.3 | 73.1 | 70.5 | 83.0 | 76.4 | 86.5 |
| STA | 69.8 | 71.6 | 70.8 | 68.9 | 71.0 | 69.5 | 67.8 | 70.8 | 71.3 | 59.7 | 64.0 | 63.0 | 66.7 | 71.3 | 71.8 | 69.2 | 70.8 | 75.9 |
| URB | 85.2 | 87.9 | 88.1 | 82.6 | 85.8 | 75.3 | 87.2 | 87.5 | 87.9 | 81.9 | 83.2 | 73.0 | 87.9 | 73.0 | 87.9 | 88.3 | 87.4 | 89.9 |
| USE | 95.7 | 95.2 | 97.2 | 93.2 | 90.6 | 84.9 | 90.5 | 91.5 | 92.0 | 54.5 | 63.7 | 69.5 | 85.8 | 96.9 | 96.2 | 96.5 | 94.1 | 96.4 |
| WIN | 98.3 | 99.3 | 98.6 | 93.1 | 97.3 | 97.2 | 96.4 | 96.6 | 96.5 | 87.2 | 95.0 | 95.0 | 93.8 | 99.7 | 92.9 | 98.9 | 98.2 | 99.0 |
| CRO * | 75.4 | 97.5 | 89.9 | 91.0 | 88.8 | 75.4 | 98.4 | 98.5 | 98.6 | 98.5 | 98.7 | 95.1 | 98.6 | 89.1 | 95.2 | 95.5 | 81.9 | 98.2 |
| ELE * | 72.3 | 95.7 | 79.9 | 80.0 | 82.5 | 70.8 | 81.1 | 83.9 | 89.7 | 64.6 | 75.4 | 76.2 | 79.8 | 79.9 | 93.7 | 93.6 | 83.2 | 93.7 |
| WAV * | 90.0 | 91.9 | 92.2 | 86.2 | 91.4 | 84.0 | 86.5 | 88.3 | 88.8 | 77.6 | 80.0 | 83.6 | 84.7 | 91.8 | 92.0 | 92.1 | 91.1 | 92.4 |
| W,T,L | 20,0,0 | 16,2,2 | 15,4,1 | 16,3,1 | 19,1,0 | 20,0,0 | 17,2,1 | 18,2,0 | 16,3,1 | 19,1,0 | 19,1,0 | 19,1,0 | 18,2,0 | 15,4,1 | 13,4,3 | 12,7,1 | 19,0,1 | -,-,- |
| Data | ADB | RF | XGB | BIM |
|---|---|---|---|---|
| BCW | 78.2 | 78.6 | 78.6 | 78.3 |
| CRY | 90.4 | 92.9 | 89.9 | 91.5 |
| CUS | 90.8 | 91.1 | 91.4 | 91.0 |
| ECO | 98.0 | 98.9 | 98.2 | 98.6 |
| GLA | 85.0 | 87.0 | 87.9 | 86.8 |
| HMS | 65.8 | 72.1 | 70.0 | 72.0 |
| IMM | 77.2 | 84.2 | 81.7 | 86.1 |
| ION | 92.1 | 93.5 | 92.5 | 93.1 |
| LYM | 84.8 | 87.0 | 87.4 | 88.1 |
| MON | 98.4 | 95.8 | 99.1 | 99.7 |
| PAR | 90.5 | 91.0 | 91.9 | 93.2 |
| PID | 73.5 | 76.0 | 75.1 | 76.2 |
| SMR | 81.4 | 82.8 | 83.3 | 86.6 |
| STA | 69.0 | 71.3 | 69.5 | 74.1 |
| URB | 87.9 | 88.6 | 88.8 | 91.4 |
| USE | 96.0 | 95.3 | 94.9 | 96.1 |
| WIN | 97.5 | 99.1 | 98.2 | 99.1 |
| CRO * | 97.3 | 97.4 | 98.5 | 98.6 |
| ELE * | 91.1 | 92.3 | 95.2 | 94.1 |
| WAV * | 89.5 | 91.2 | 90.8 | 93.3 |
| W,T,L | 17,3,0 | 11,8,1 | 14,4,2 | -,-,- |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, R.; Hong, P.; Liu, J.S. IMMIGRATE: A Margin-Based Feature Selection Method with Interaction Terms. Entropy 2020, 22, 291. https://doi.org/10.3390/e22030291
Zhao R, Hong P, Liu JS. IMMIGRATE: A Margin-Based Feature Selection Method with Interaction Terms. Entropy. 2020; 22(3):291. https://doi.org/10.3390/e22030291
Chicago/Turabian StyleZhao, Ruzhang, Pengyu Hong, and Jun S. Liu. 2020. "IMMIGRATE: A Margin-Based Feature Selection Method with Interaction Terms" Entropy 22, no. 3: 291. https://doi.org/10.3390/e22030291
APA StyleZhao, R., Hong, P., & Liu, J. S. (2020). IMMIGRATE: A Margin-Based Feature Selection Method with Interaction Terms. Entropy, 22(3), 291. https://doi.org/10.3390/e22030291






