Spatial Organization of Five-Fold Morphology as a Source of Geometrical Constraint in Biology
Abstract
1. Introduction
2. Methods
2.1. Statistics of Spatial Organization for Shapes Γ
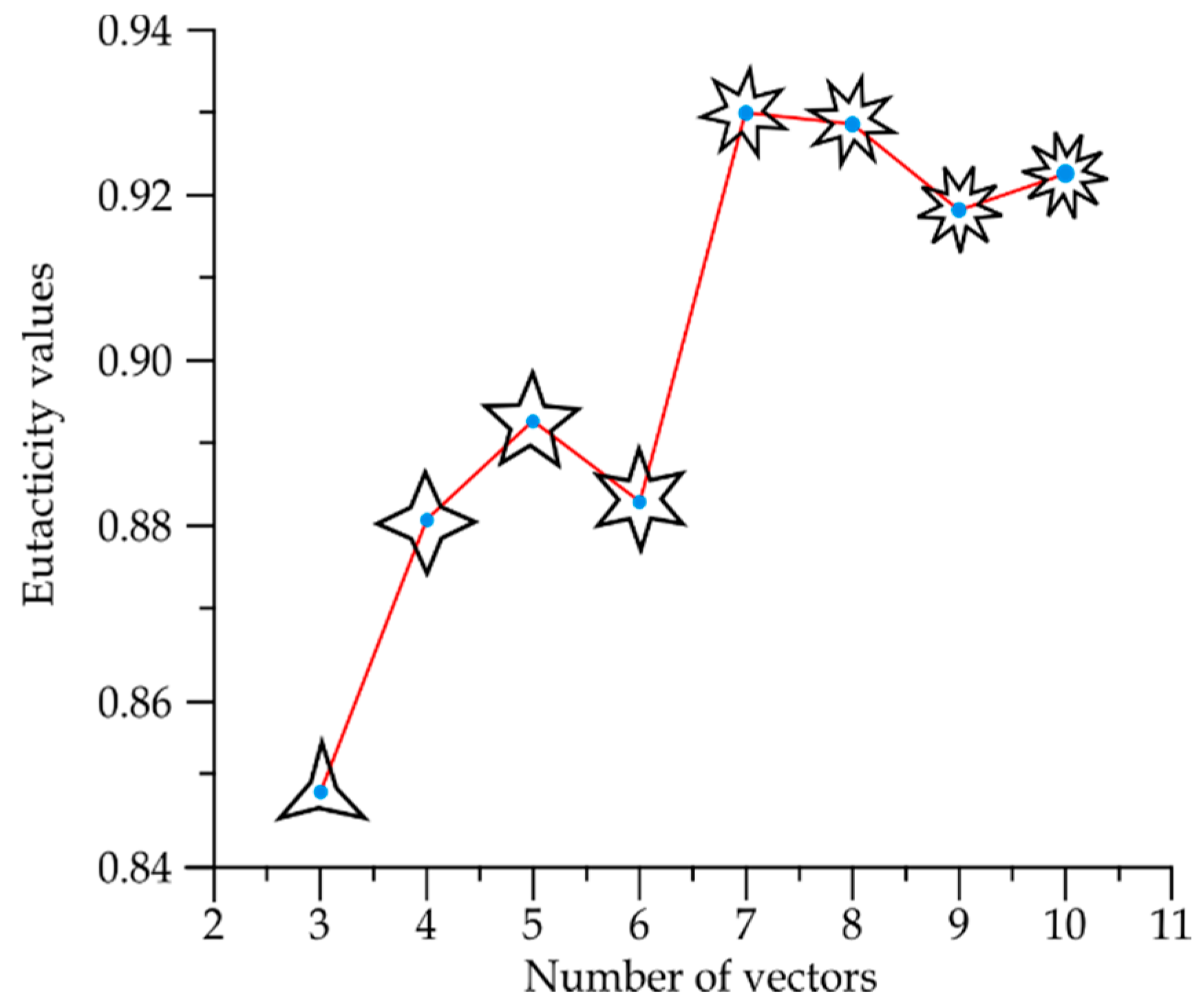
2.2. Mathematical Basis of Eutacticity
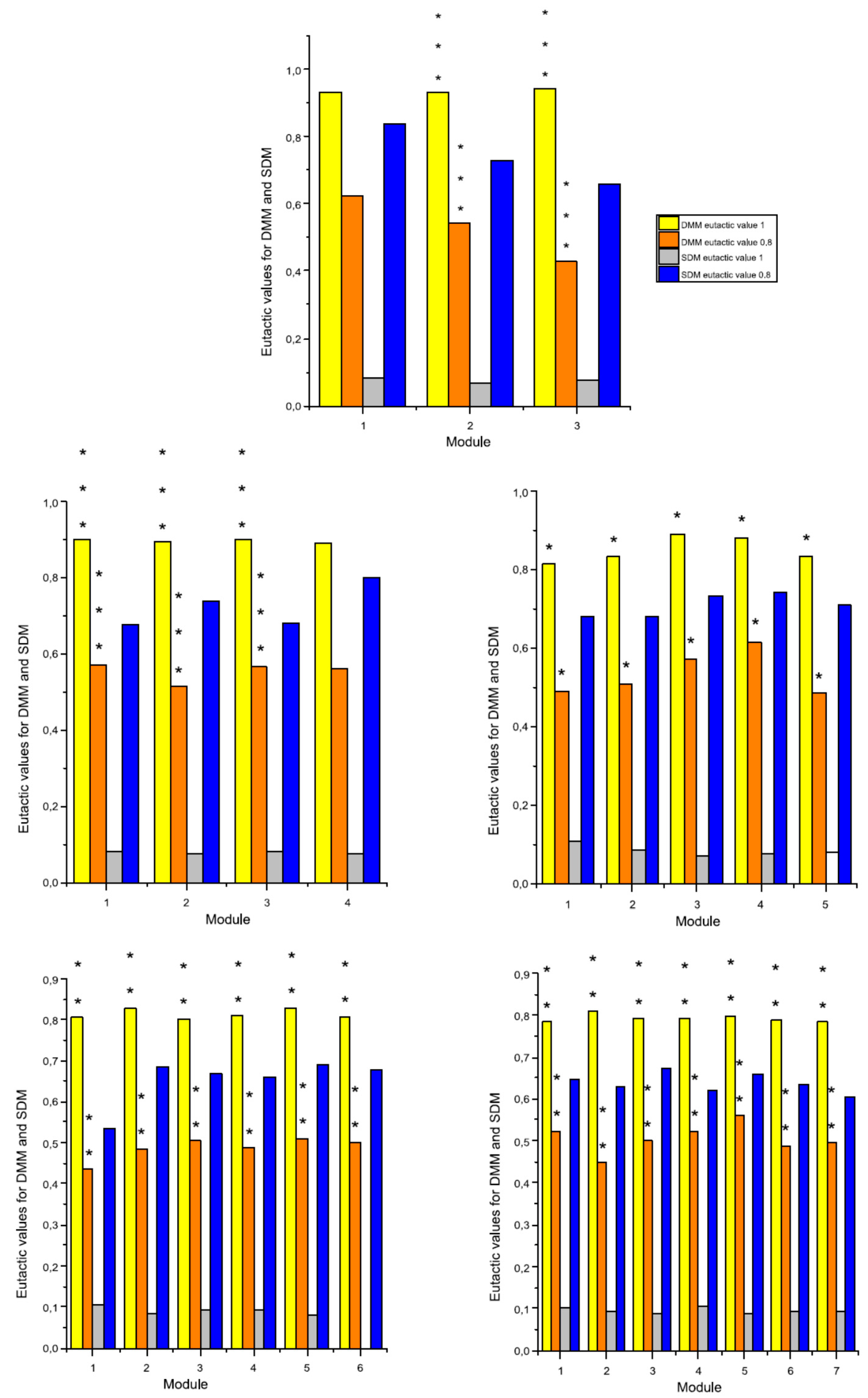
2.3. The Eutacticity and the Standard Deviation of Dispersion Mean of a Module
2.4. Standard Deviation of Partition Variability
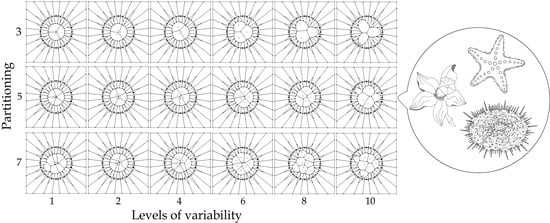
- Partitioning number (pn) defines the number of partitions inside a disc (ranging from three to ten).
- Partition variability (pv) determines multiple levels of variability (ten) inside discs by using random points, which in turn define the Voronoi diagrams. These levels of variability will be defined below.
- We initially consider a disk with a unitary radius where a second inscribed disk will be partitioned into a pn with a pv during the experiment (steps 4 and 5 of this algorithm, respectively). These discs are defined by particular features each: (a) The first disc is the external limit of the second and their coordinates are constant during the experiment; (b) the second one is constantly changing to obtain a pv (step 5 of this algorithm) and it is obtained by establishing a Voronoi tessellation. These two features (a) and (b) are described in the next steps (2) and (3) of this algorithm.
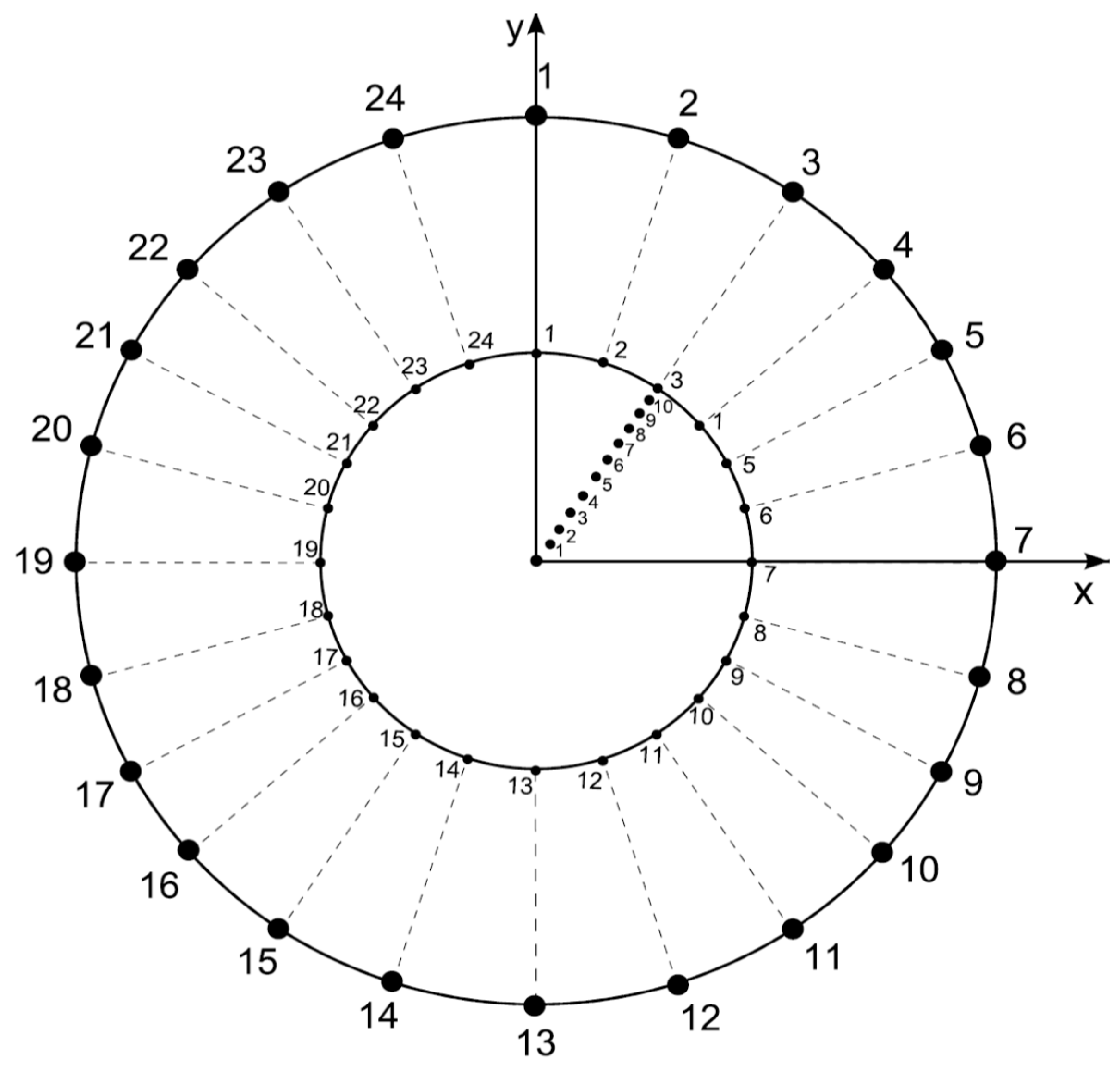
- Features of external disc. The boundaries of the external limit are defined by 24 fixed points generated as follows: The radius of the external disk is set to r = 1 and consecutive points are separated by an angle θ/24. The functionality of this feature lies in the establishment of a fixed limit of reference to maintain a constant area during variation of partitions.
- Features of internal disc. The boundaries of the internal limit are defined by 24 fixed points generated as follows: The radius of the internal disk is initially set to r = 0.53 ± 0.4 (established by the first level of variability step 6 of this algorithm) with 24 points consecutively separated by an angle θ/24. These radii are derived from a Voronoi tessellation whose points are the 24 points established before in this step besides the points derived from step 5. The functionality of this feature lies in the establishment of an internal limit able to change, providing statistical variation determining levels of variability of areas inside discs.
- Now, we define partition numbering (pn) inside the disk. Once the number of partitions is defined, say n (where 3 ≤ n ≤ 10 and ) to define a Voronoi tessellation, points are located in the disk at angles 2π/n ± 0.069 radians but at different radius. These radius values will define the pv, as described in the next item.
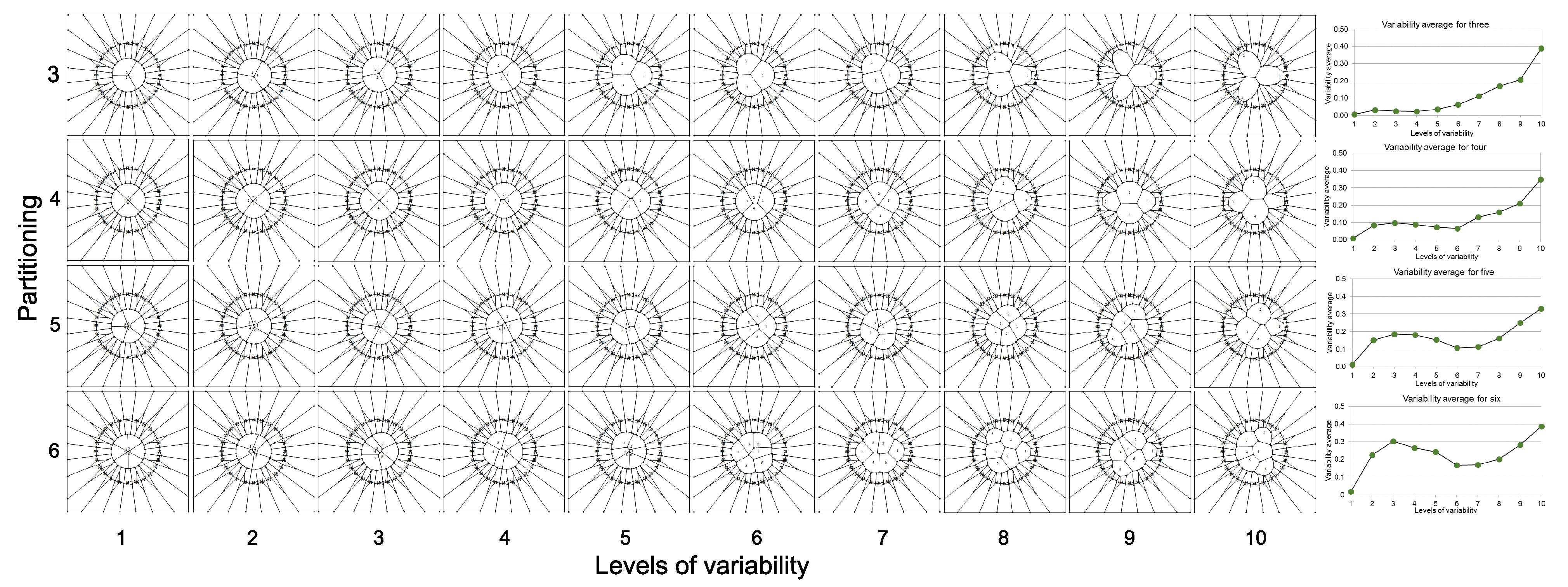
- Partition variability (pv). For each angular region defined above, 10 points are located at radius (between r = 0 and r = 10) at different positions to define different degrees of variability using Voronoi tessellations. The first point (first level of variability) is at r = 1. After the second point, all of them are located at random radius between 1 to 10. Hence, each level of variability (10) is given by radii ranges except 1 which is fixed at 1 (Figure 3); (a) 1, (b) 1–2, (c) 1–3, (d) 1–4, (e) 1–5, (f) 1–6, (g) 1–7, (h) 1–8, (i) 1–9 and (j) 1–10. Partition variability will define the broad spectrum of possibilities for area distribution inside discs without losing partitioning number. According to Equation (1), the average of areas requires a summation of sub-localities areas which were derived from partitions.
- Once the partition areas inside discs were obtained and Equation (1) was solved, Equation (2) is used to get standard deviations of variability for each disc. In order to normalize the level of variability for each pn, an index dividing the standard deviation of partitions and the particular area average of each partition was obtained (variability average; supporting information 2). There are eight particular area averages of partitions since we have a sample of 8 discs with different pn (from 3 to 10). These particular area averages are derived from a value n/(≈108.5 ± 1.5) which are n values obtained from the first level of variability (pv) at r = 1. It is important to say that the radius of the external disc (1) and the radius of the internal disc (r = 0.53 ± 0.4) was modified in order to get the particular area averages. However, in spite of the modification, the index between external discs and the internal ones remains constant. A sample of 20 discs to get 20 standard deviations was generated for each pn, and also for each level of pv (10) giving a sample of 200 discs for each pn. An average of standard deviations (; variability average) was derived for each level of variability.Once the partition areas inside discs were obtained and Equation (1) was solved, Equation (2) is used to get standard deviations of variability for each disc. In order to normalize the level of variability for each pn, an index dividing the standard deviation of partitions and the particular area average of each partition was obtained (variability average; supporting information 2). There are eight particular area averages of partitions since we have a sample of 8 discs with different pn (from 3 to 10). These particular area averages are derived from a value n/(≈108.5 ± 1.5) which are n values obtained from the first level of variability (pv) at r = 1. It is important to say that the radius of the external disc (1) and the radius of the internal disc (r = 0.53 ± 0.4) was modified in order to get the particular area averages. However, in spite of the modification, the index between external discs and the internal ones remains constant. A sample of 20 discs to get 20 standard deviations was generated for each pn, and also for each level of pv (10) giving a sample of 200 discs for each pn. An average of standard deviations ; variability average) was derived for each level of variability.
- Finally, a standard deviation of all variability averages is obtained for each pn.
3. Results
3.1. Star Morphospace for Shapes Γ
3.2. Experimental Evidence
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hargittai, I. Fivefold Symmetry, 1st ed.; World Scientific: Farrer Road, Singapore, 1994; pp. 18–20. [Google Scholar]
- Darvas, G. Symmetry: Cultural-Historical and Ontological Aspects of Science-Arts Relations; The Natural and Man-Made World in an Interdisciplinary Approach, 1st ed.; Birkhauser: Berlin, Germany, 2007; pp. 1–35. [Google Scholar]
- Arenas-Mena, C.; Cameron, A.R.; Davidson, E.H. Spatial expression of Hox cluster genes in the ontogeny of a sea urchin. Development 2000, 127, 4631–4643. [Google Scholar] [PubMed]
- Mooi, R.; David, B. Radial symmetry, the anterior/posterior axis, and echinoderm Hox genes. Annu. Rev. Ecol. Evol. Syst. 2008, 39, 43–62. [Google Scholar] [CrossRef]
- Brown, W.M.; Price, M.E.; Kang, J.S.; Pound, N.; Zhao, Y.; Yu, H. Fluctuating asymmetry and preferences for sex-typical bodily characteristics. Proc. Natl. Acad. Sci. USA 2008, 105, 12938–12943. [Google Scholar] [CrossRef] [PubMed]
- Hotchkiss, F.H.C. On the number of rays in starfish. Am. Zool. 2000, 40, 340–354. [Google Scholar] [CrossRef]
- Martindale, M.Q.; Finnerty, J.R.; Henry, J.Q. The Radiata and the evolutionary origins of the bilaterian body plan. Mol. Phylogenet. Evol. 2002, 24, 358–365. [Google Scholar] [CrossRef]
- Moller, A.P.; Eriksson, M. Pollinator preference for symmetrical flowers and sexual selection in plants. Oikos 1995, 73, 15–22. [Google Scholar] [CrossRef]
- Moller, A.P.; Sorci, G. Insect preference for symmetrical artificial flowers. Oecologia 1998, 114, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Popodi, E.; Raff, R.A. Hox genes in a pentameral animal. Bioessays 2001, 23, 211–214. [Google Scholar] [CrossRef]
- Smith, A.B. Deuterostomes in a twist: The origins of a radical new body plan. Evol. Dev. 2008, 10, 493–503. [Google Scholar] [CrossRef] [PubMed]
- Breder, C.M. Observations on the occurrence and attributes of pentagonal symmetry. Bull. Am. Mus. Nat. Hist. 1955, 106, 173–220. [Google Scholar]
- Smith, J.M.; Burian, R.; Kauffman, S.; Alberch, P.; Campbell, J.; Goodwin, B.; Lande, R.; Raup, D.; Wolpert, L. Developmental constraints and evolution. Q. Rev. Biol. 1985, 60, 265–287. [Google Scholar] [CrossRef]
- Raff, R.A. The Shape of Life: Genes, Development, and the Evolution of Animal Form, 1st ed.; The University of Chicago Press: Chicago, IL, USA, 1996; pp. 38–61. [Google Scholar]
- Stephenson, D.G. Pentameral symmetry in echinoderms. Nature 1967, 216, 994. [Google Scholar] [CrossRef]
- Grabowsky, G.L. Symmetry, locomotion, and the evolution of an anterior end: A lesson from sea-urchins. Evolution 1994, 48, 1130–1146. [Google Scholar] [CrossRef] [PubMed]
- Aragon, J.L.; Torres, M.; Gil, D.; Barrio, R.A.; Maini, P.K. Turing patterns with pentagonal symmetry. Phys. Rev. E 2002, 65, 051913. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Sauceda, J.; Rueda-Contreras, M.D. A method to categorize 2-dimensional patterns using statistics of spatial organization. Evol. Bioinf. 2017, 13. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Sauceda, J.; Aragon, J.L. Eutacticity in sea urchin evolution. Bull. Math. Biol. 2008, 70, 625–634. [Google Scholar] [CrossRef] [PubMed]
- Coxeter, H.S.M. Regular Polytopes, 2nd ed.; Dover: New York, NY, USA, 1973; pp. 15–20. [Google Scholar]
- Chen, E.R.; Klotsa, D.; Engel, M.; Damasceno, P.F.; Glotzer, S.C. Complexity in surfaces of densest packings for families of polyhedra. Phys. Rev. X 2014, 4, 011024. [Google Scholar] [CrossRef]
- Damasceno, P.F.; Engel, M.; Glotzer, S.C. Predictive self-assembly of polyhedra into complex structures. Science 2012, 337, 453–457. [Google Scholar] [CrossRef] [PubMed]
- Wagner, G.P. Homologues, natural kinds and the evolution of modularity. Am. Zool. 1996, 36, 36–43. [Google Scholar] [CrossRef]
- Fontana, W.; Buss, L.W. The arrival of the fittest-toward a theory of biological organization. Bull. Math. Biol. 1994, 56, 1–64. [Google Scholar]
- McGhee, G. The Geometry of Evolution: Adaptative Landscapes and Theoretical Morphospaces, 1st ed.; Cambridge University Press: Cambridge, UK, 2006; pp. 57–65. [Google Scholar]
- Torres, M.; Aragon, J.L.; Dominguez, P.; Gil, D. Regularity in irregular echinoids. J. Math. Biol. 2002, 44, 330–340. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Sauceda, J.; Malda-Barrera, J.; Laguarda-Figueras, A.; Solis-Marin, F.; Aragon, J.L. Influence of modularity and regularity on disparity of atelostomata sea urchins. Evol. Bioinf. 2014, 10, 97–105. [Google Scholar] [CrossRef] [PubMed]
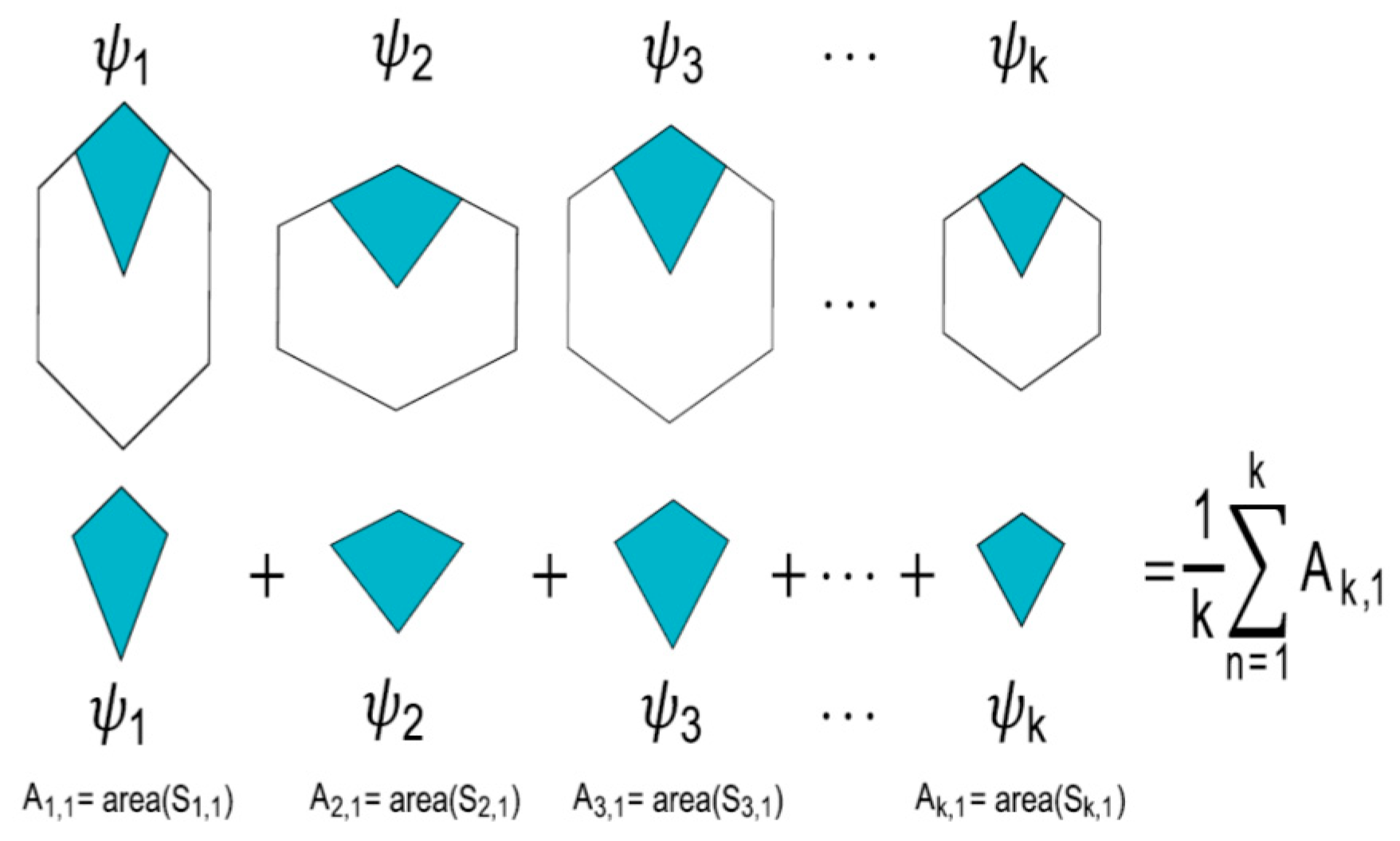

| Set of Random Points Defining the Associated Areas for Sub-Locality 1 (Algorithm Defined in Reference [24]) | Summation of Areas for Star | |||||
|---|---|---|---|---|---|---|
| Stars | … | |||||
| … | ⇒ | |||||
| … | ⇒ | |||||
| . . . | . . . | . . . | . . . | . . . | . . . | . . . |
| … | ⇒ | |||||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
López-Sauceda, J.; López-Ortega, J.; Laguna Sánchez, G.A.; Sandoval Gutiérrez, J.; Rojas Meza, A.P.; Aragón, J.L. Spatial Organization of Five-Fold Morphology as a Source of Geometrical Constraint in Biology. Entropy 2018, 20, 705. https://doi.org/10.3390/e20090705
López-Sauceda J, López-Ortega J, Laguna Sánchez GA, Sandoval Gutiérrez J, Rojas Meza AP, Aragón JL. Spatial Organization of Five-Fold Morphology as a Source of Geometrical Constraint in Biology. Entropy. 2018; 20(9):705. https://doi.org/10.3390/e20090705
Chicago/Turabian StyleLópez-Sauceda, Juan, Jorge López-Ortega, Gerardo Abel Laguna Sánchez, Jacobo Sandoval Gutiérrez, Ana Paola Rojas Meza, and José Luis Aragón. 2018. "Spatial Organization of Five-Fold Morphology as a Source of Geometrical Constraint in Biology" Entropy 20, no. 9: 705. https://doi.org/10.3390/e20090705
APA StyleLópez-Sauceda, J., López-Ortega, J., Laguna Sánchez, G. A., Sandoval Gutiérrez, J., Rojas Meza, A. P., & Aragón, J. L. (2018). Spatial Organization of Five-Fold Morphology as a Source of Geometrical Constraint in Biology. Entropy, 20(9), 705. https://doi.org/10.3390/e20090705