On a Robust MaxEnt Process Regression Model with Sample-Selection
Abstract
1. Introduction
2. Robust Sample-Selection GPR Model
2.1. MaxEnt Process Regression Model
2.2. Proposed Model
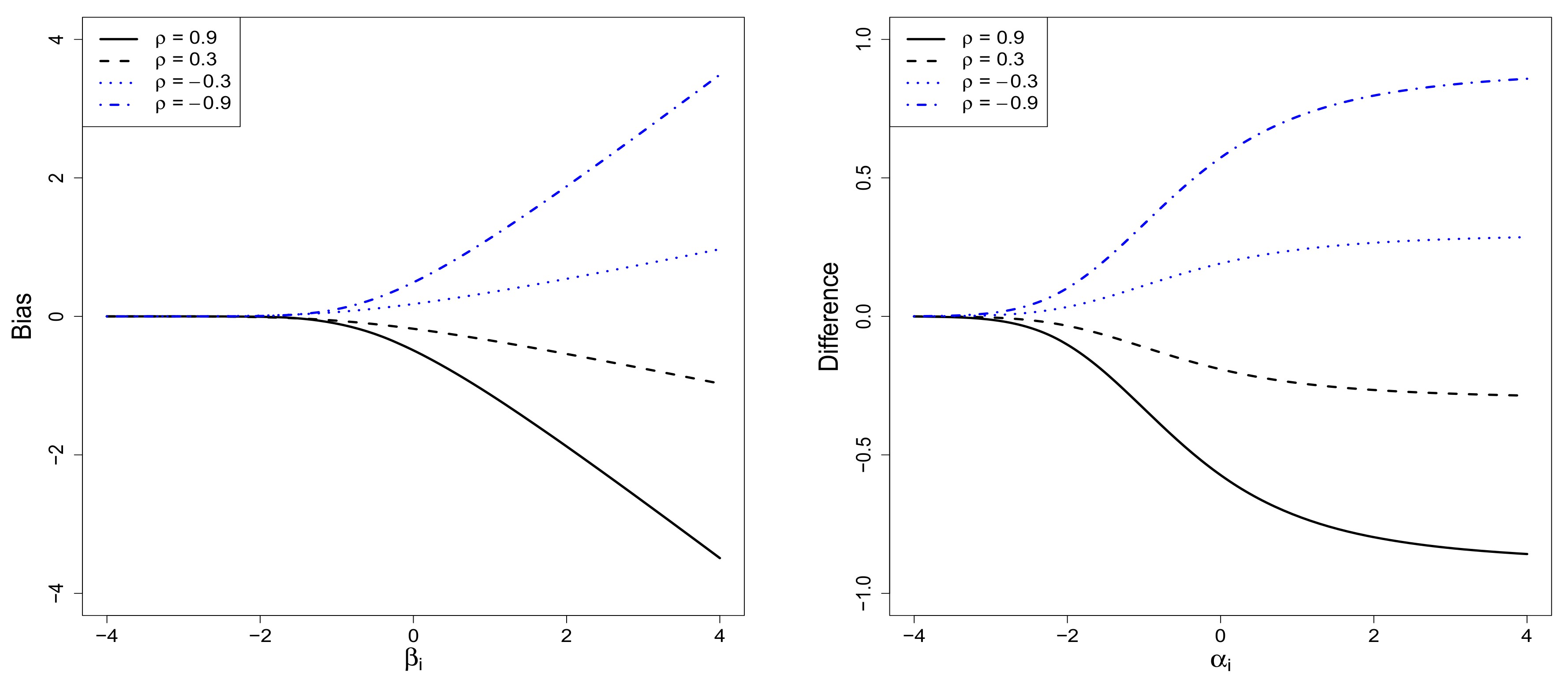
2.3. The Sample-Selection Bias
3. Bayesian Hierarchical Methodology
3.1. Hierarchical Representation of the RSGPR Model
3.2. Full Conditional Posteriors
- (1)
- The full conditional posterior distribution of is given bywhere and
- (2)
- The full conditional posterior distribution of is an inverse Gamma distribution:
- (3)
- The full conditional posterior distribution of is a normal distribution:where
- (4)
- The full conditional posterior distributions of ’s are independent and their distributions are given byfor where
- (5)
- The full conditional posterior density of is:where and
- (6)
- The full conditional posterior densities of ’s are independent and they are given by
3.3. Markov Chain Monte Carlo Method
- (1)
- (2)
- Gibbs samples of and can be obtained by using those of and .
- (3)
- If degenerates at the RSGPR model can be reduced to the SGPR model. In this case, the MCMC procedure excludes the Gibbs sampling of ’s by using the posterior distribution (19).
- (4)
- (5)
- When and the RSGPR model becomes the SGPR model. For generating ’s, we may use the following posteriorswhere Except for the SGPR and SGPR we need to adopt the Metropolis–Hastings algorithm within the Gibbs sampler because the conditional posterior density of does not have explicit form of known distribution. See [26,27] for the algorithm for sampling from the posterior density.
- (6)
- When the squared exponential covariance function in Equation (4) is chosen with unknown hyperparameters and we need to elicit the priors of and for the full Bayes methods based on the MCMC method. The priors considered by [28] can be used for this assessment as follows. The prior distributions are a conjugate and Here denotes the half-Cauchy distribution with the p.d.f. location parameter and scale parameter See [28], for compatibility with to elicit the prior information on
- (7)
- Full conditional posterior distributions of and arewhere and Note that the conditional posterior density of does not have explicit form of known distribution. This implies the use of the Metropolis–Hastings algorithm within the Gibbs sampler to generate from the posterior density.
- (8)
- After obtaining the Gibbs samples of we can use them for Monte Carlo estimation of regression function and missing observations They can be also used for predicting regression functions and s evaluated at new predictors (see, e.g., [26]).
3.4. Prediction with Bias Corrected Regression Function
4. Numerical Illustrations
4.1. Simulation Scheme
4.2. Performance of the RSGPR Model
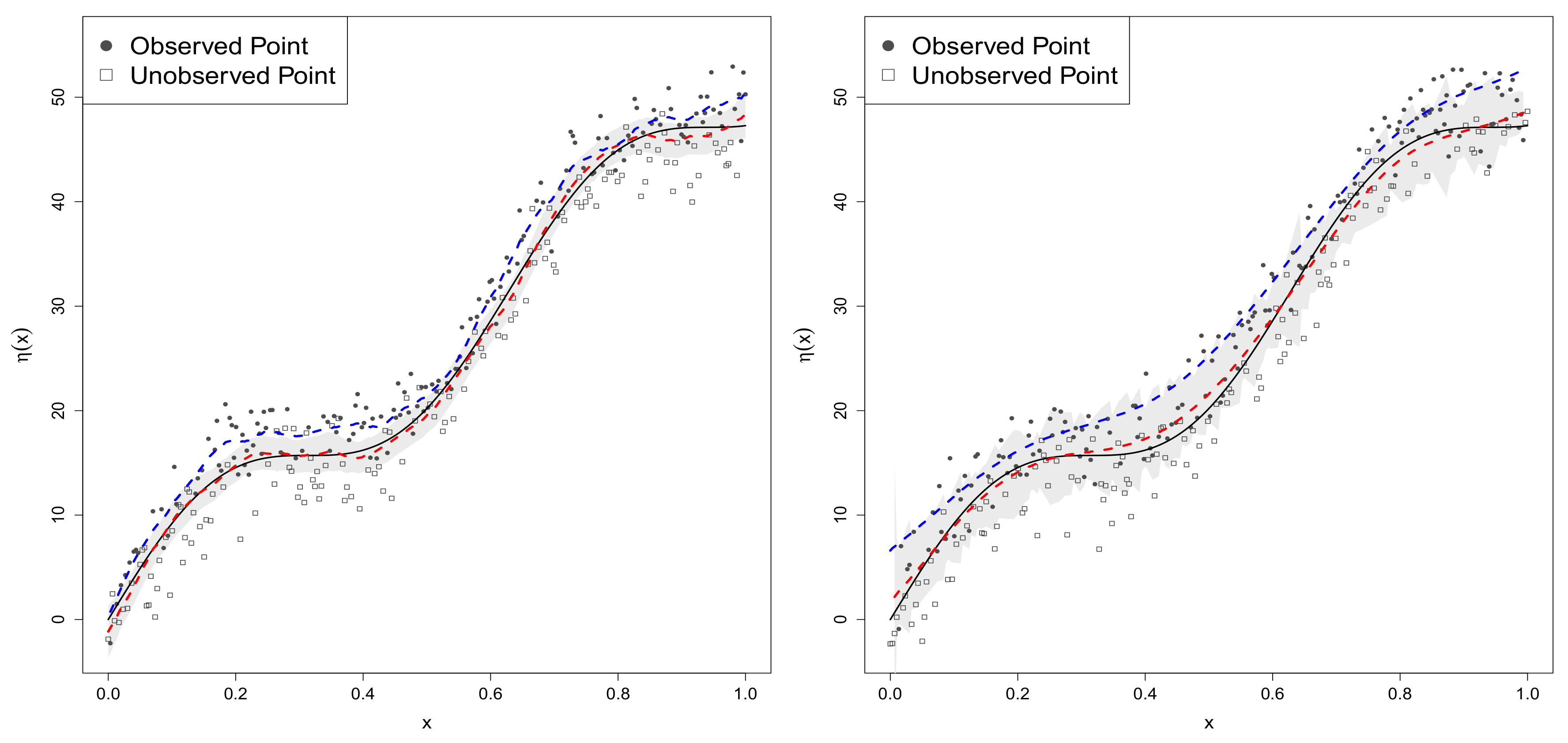
4.2.1. Sample-Selection Data Generated from Model 1
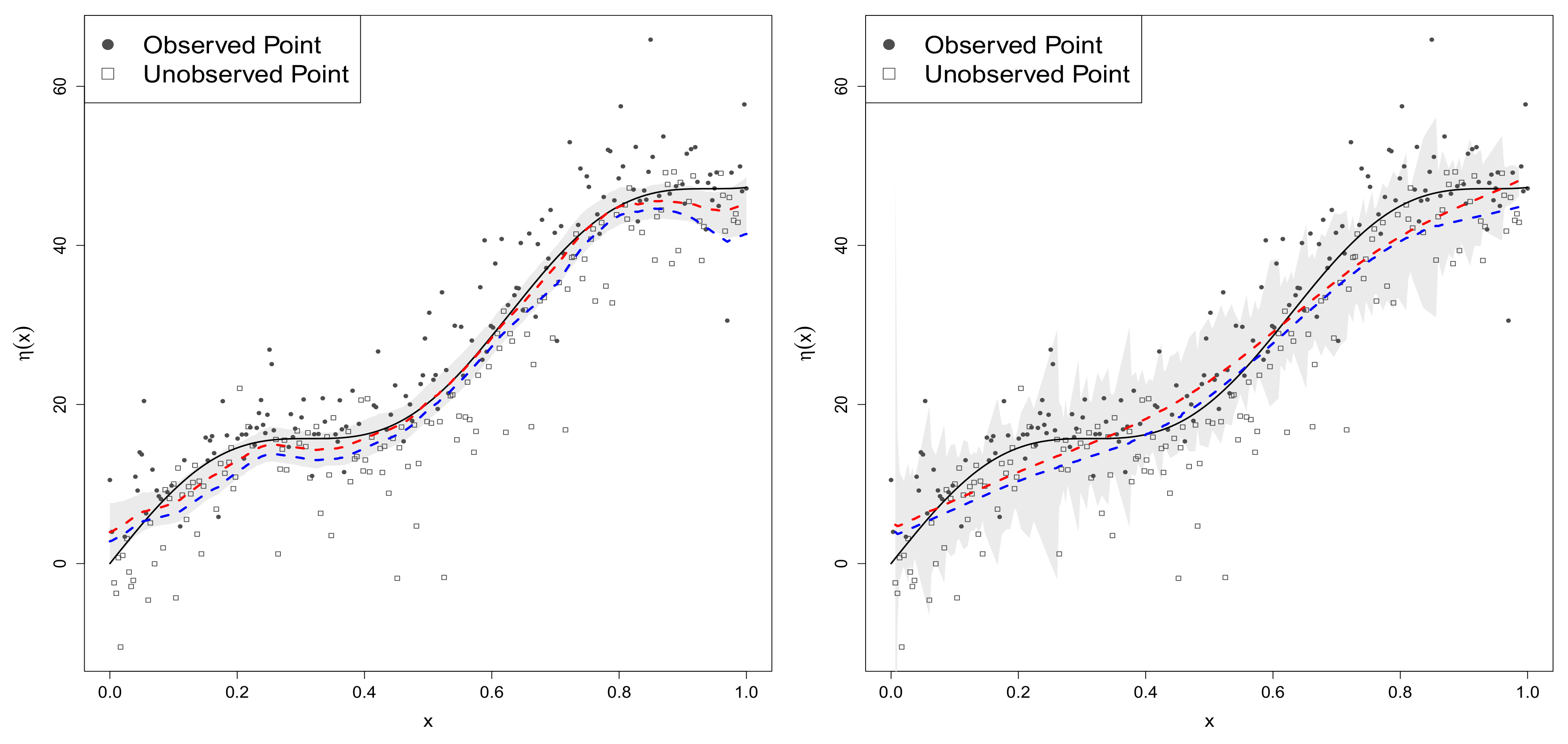
4.2.2. Data Generated from Model 2
4.2.3. Data Generated from Model 3 with Normal Mixture Errors
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Appendix A
Appendix A.1. Proof of Lemma 1
Appendix A.2. Proof of Lemma 2
Appendix A.3. Proof of Lemma 3
Appendix A.4. Proof of Corollary 2
Appendix A.5. Proof of Corollary 3
Appendix A.6. Proof of Theorem 1
Appendix A.7. Derivation of Conditional Posterior Distributions
- (1)
- Full conditional distribution of : Equation (13) states that the full conditional density of iswhich is the kernel of distribution.
- (2)
- Full conditional distribution of : We see from Equation (13) that the full conditional posterior density isThis is the kernel of distribution.
- (3)
- Full conditional distribution of : Equation (13) gives the full conditional density of given bywhich is the kernel of distribution.
- (4)
- Full conditional distributions of ’s: Equation (13) indicates that the full conditional posterior densities of s are mutually independent, and that for each i,Since the support of is for while for we have the truncated normal distributions.
- (5)
- Full conditional distribution of : The full conditional posterior density of is given bywhich is the kernel of distribution.
References
- Cox, G.; Kachergis, G.; Shiffrin, R. Gaussian process regression for trajectory analysis. In Proceedings of the Annual Meeting of the Cognitive Science Society, Sapporo, Japan, 1–4 August 2012; Volume 34. [Google Scholar]
- Rasmussen, C.E.; Nickisch, H. Gaussian process for machine learning (gpml) toolbox. J. Mach. Learn. Res. 2010, 11, 3011–3015. [Google Scholar]
- Liutkus, A.; Badeau, R.; Richard, G. Gaussian processes for underdetermined source separation. IEEE Trans. Signal Process. 2011, 59, 3155–3167. [Google Scholar] [CrossRef]
- Caywood, M.S.; Roberts, D.M.; Colombe, J.B.; Greenward, H.S.; Weiland, M.Z. Gaussian Process Regression for Predictive But Interpretable Machine Learning Models: An Example of Predicting Mental Worklosd across Tasks. Front. Hum. Neurosci. 2017, 10, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Contreras-Reyes, J.E.; Arellano-Valle, R.B.; Canales, T.M. Comparing growth curves with asymmetric heavy-tailed errors: Application to the southern blue whiting (Micromesistius australis). Fish. Res. 2014, 159, 88–94. [Google Scholar] [CrossRef]
- Heckman, J.J. Sample selection bias as a specification error. Econometrica 1979, 47, 153–161. [Google Scholar] [CrossRef]
- Marchenko, Y.V.; Genton, M.G. A Heckman selection-t model. J. Am. Stat. Assoc. 2012, 107, 304–317. [Google Scholar] [CrossRef]
- Ding, P. Bayesian robust inference of sample selection using selection t-models. J. Multivar. Anal. 2014, 124, 451–464. [Google Scholar] [CrossRef]
- Hasselt, V.M. Bayesian inference in a sample selection model. J. Econ. 2011, 165, 221–232. [Google Scholar] [CrossRef]
- Arellano-Valle, R.B.; Contreras-Reyes, J.E.; Stehlík, M. Generalized skew-normal negentropy and its application to fish condition factor time series. Entropy 2017, 19, 528. [Google Scholar] [CrossRef]
- Kim, H.-J.; Kim, H.-M. Elliptical regression models for multivariate sample-selection bias correction. J. Korean Stat. Soc. 2016, 45, 422–438. [Google Scholar] [CrossRef]
- Kim, H.-J. Bayesian hierarchical robust factor analysis models for partially observed sample-selection data. J. Multivar. Anal. 2018, 164, 65–82. [Google Scholar] [CrossRef]
- Kim, H.-J. A class of weighted multivariate normal distributions and its properties. J. Multivar. Anal. 2008, 99, 1758–1771. [Google Scholar] [CrossRef]
- Lenk, P.J. Bayesian inference for semiparametric regression using a Fourier representation. J. R. Stat. Soc. Ser. B. 1999, 61, 863–879. [Google Scholar] [CrossRef][Green Version]
- Fahrmeir, L.; Kneib, T. Bayesian Smoothing and Regression for Longitudial, Spatial and Event History Data; Oxford Statistical Science Series; Oxford University Press: Oxford, UK, 2011; Volume 36. [Google Scholar]
- Chakraborty, S.; Ghosh, M.; Mallick, B.K. Bayesian nonlinear regression for large p and small n problems. J. Multivar. Anal. 2012, 108, 28–40. [Google Scholar] [CrossRef]
- Leonard, T.; Hsu, J.S.J. Bayesian Methods: An Analysis for Statisticians and Interdisciplinary Researchers; Cambridge University Press: New York, NY, USA, 1999. [Google Scholar]
- Kim, H.-J. A two-stage maximum entropy prior of location parameter with a stochastic multivariate interval constraint and its properties. Entropy 2016, 18, 188. [Google Scholar] [CrossRef]
- Shi, J.; Choi, T. Monographs on Statistics and Applied Probability, Gaussian Process Regression Analysis for Functional Data; Chapman & Hall: London, UK, 2011. [Google Scholar]
- Rasmussen, C.E.; Williams, C.K.I. Gaussian Process for Machine Learning; The MIT Press: Cambridge, MA, USA, 2006. [Google Scholar]
- Andrews, D.F.; Mallows, C.L. Scale mixtures of normal distributions. J. R. Stat. Soc. Ser. B 1974, 36, 99–102. [Google Scholar]
- Lachos, V.H.; Labra, F.V.; Bolfarine, H.; Ghosh, P. Multivariate measurement error models based on scale mixtures of the skew-normal distribution. Statistics 2010, 44, 541–556. [Google Scholar] [CrossRef]
- Arellano-Valle, R.B.; Branco, M.D.; Genton, M.G. A unified view on skewed distributions arising from selection. Can. J. Stat. 2006, 34, 581–601. [Google Scholar] [CrossRef]
- Kim, H.J.; Choi, T.; Lee, S. A hierarchical Bayesian regression model for the uncertain functional constraint using screened scale mixture of Gaussian distributions. Statistics 2016, 50, 350–376. [Google Scholar] [CrossRef]
- Rubin, D.B. Inference and missing data. Biometrika 1976, 63, 581–592. [Google Scholar] [CrossRef]
- Ntzoufras, I. Bayesian Modeling Using WinBUGS; Wiley: New York, NY, USA, 2009. [Google Scholar]
- Chib, S.; Greenberg, E. Understanding the Metropolis-Hastings algorithm. Am. Stat. 1995, 49, 327–335. [Google Scholar]
- Gelman, A. Prior distributions for variance parameters in hierarchical models (comment on article by Browne and Draper). Bayesian Anal. 2006, 1, 515–534. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017; ISBN 3-900051-07-0. [Google Scholar]
- Spiegelhalter, D.; Best, N.; Carlin, B.; van der Linde, A. Bayesian measure of model complexity and fit (with discussion). J. R. Stat. Soc. Ser. B 2002, 64, 583–639. [Google Scholar] [CrossRef]
- Johnson, N.L.; Kotz, S.; Balakrishnan, N. Distribution in Statistics: Continuous Univariate Distributions, 2nd ed.; John Wiley & Son: New York, NY, USA, 1994; Volume 1. [Google Scholar]
| True Value | Mean | s.d. | SGPR Model | MC Error | Mean | s.d. | GPR Model | MC Error | ||
|---|---|---|---|---|---|---|---|---|---|---|
| RMSE | MAB | RMSE | MAB | |||||||
| 2.831 | 0.308 | 0.351 | 0.426 | 0.018 | 2.094 | 0.104 | 0.912 | 0.800 | 0.002 | |
| 0.380 | 0.376 | 0.563 | 0.287 | 0.064 | NA | NA | NA | NA | NA | |
| SGPR Model | GPR Model | |||||||||
| 2.880 | 0.974 | 0.509 | 0.515 | 0.050 | 2.130 | 0.109 | 0.876 | 0.800 | 0.003 | |
| 0.435 | 0.275 | 0.627 | 0.422 | 0.032 | NA | NA | NA | NA | NA | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, H.-J.; Bae, M.; Jin, D. On a Robust MaxEnt Process Regression Model with Sample-Selection. Entropy 2018, 20, 262. https://doi.org/10.3390/e20040262
Kim H-J, Bae M, Jin D. On a Robust MaxEnt Process Regression Model with Sample-Selection. Entropy. 2018; 20(4):262. https://doi.org/10.3390/e20040262
Chicago/Turabian StyleKim, Hea-Jung, Mihyang Bae, and Daehwa Jin. 2018. "On a Robust MaxEnt Process Regression Model with Sample-Selection" Entropy 20, no. 4: 262. https://doi.org/10.3390/e20040262
APA StyleKim, H.-J., Bae, M., & Jin, D. (2018). On a Robust MaxEnt Process Regression Model with Sample-Selection. Entropy, 20(4), 262. https://doi.org/10.3390/e20040262




