The Effect of Threshold Values and Weighting Factors on the Association between Entropy Measures and Mortality after Myocardial Infarction in the Cardiac Arrhythmia Suppression Trial (CAST)
Abstract
:1. Introduction
2. Methods
2.1. Data
2.2. Entropy Measures
- The approximate entropy as introduced by Pincus et al. [12] is calculated aswhere is the number of points found within the distance r for any point within the points , divided by .
- Another modification of in order to correct its bias by self-matches is the sample entropy as introduced by Richman and Moorman [13]. It is defined ashowever, this time, does not count self-matches.
- To soften the effects of a hard threshold r, Chen et al. [14] replaced it with the fuzzy membership functionThe factor of 0.69 was incorporated to get a value of 0.5 for , which is important for comparisons between rectangular and fuzzy membership functions. Finally, withwhere d is the Chebyshev distance, the fuzzy entropy is defined as:
- The fuzzy measure entropy , proposed by Liu et al. [15], introduces a distinction between local and global similarity based on :
2.3. Application of Entropy Measures to CAST Data
2.4. Statistical Analysis
3. Results
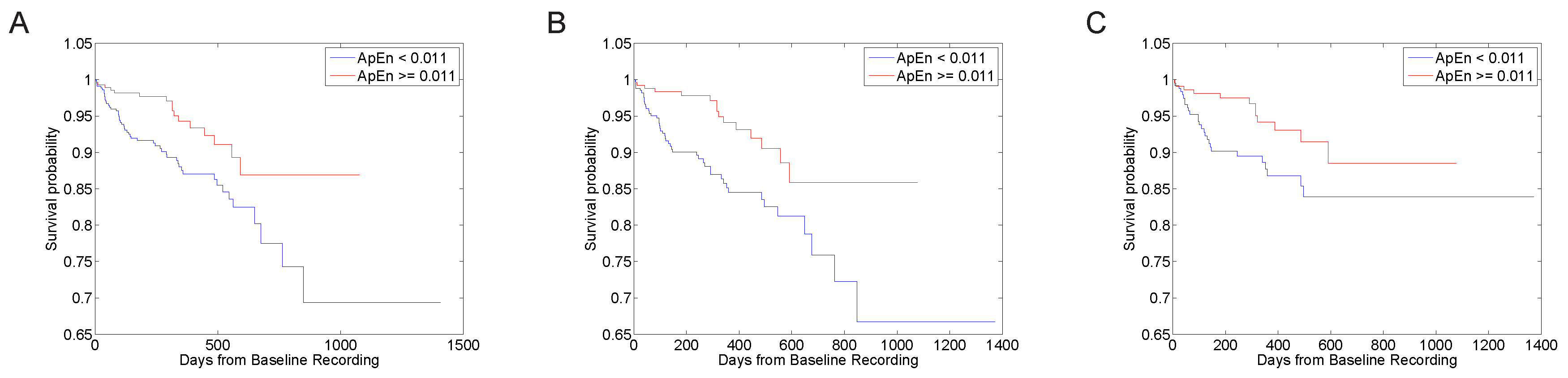
3.1. Predictive Value with Standard Parameter Sets
3.2. Parameter Selection Process
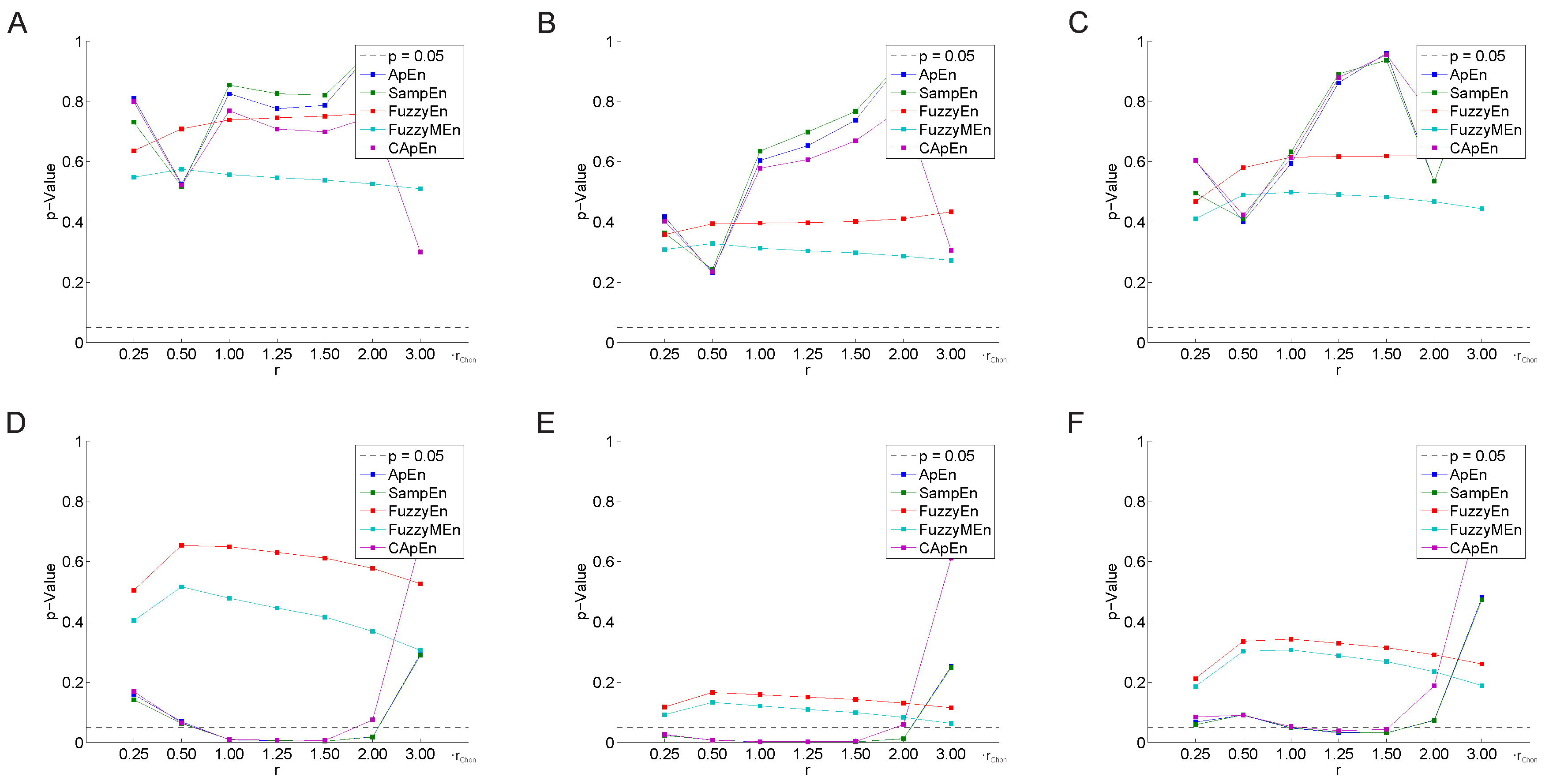
3.2.1. Variation of the Threshold Value
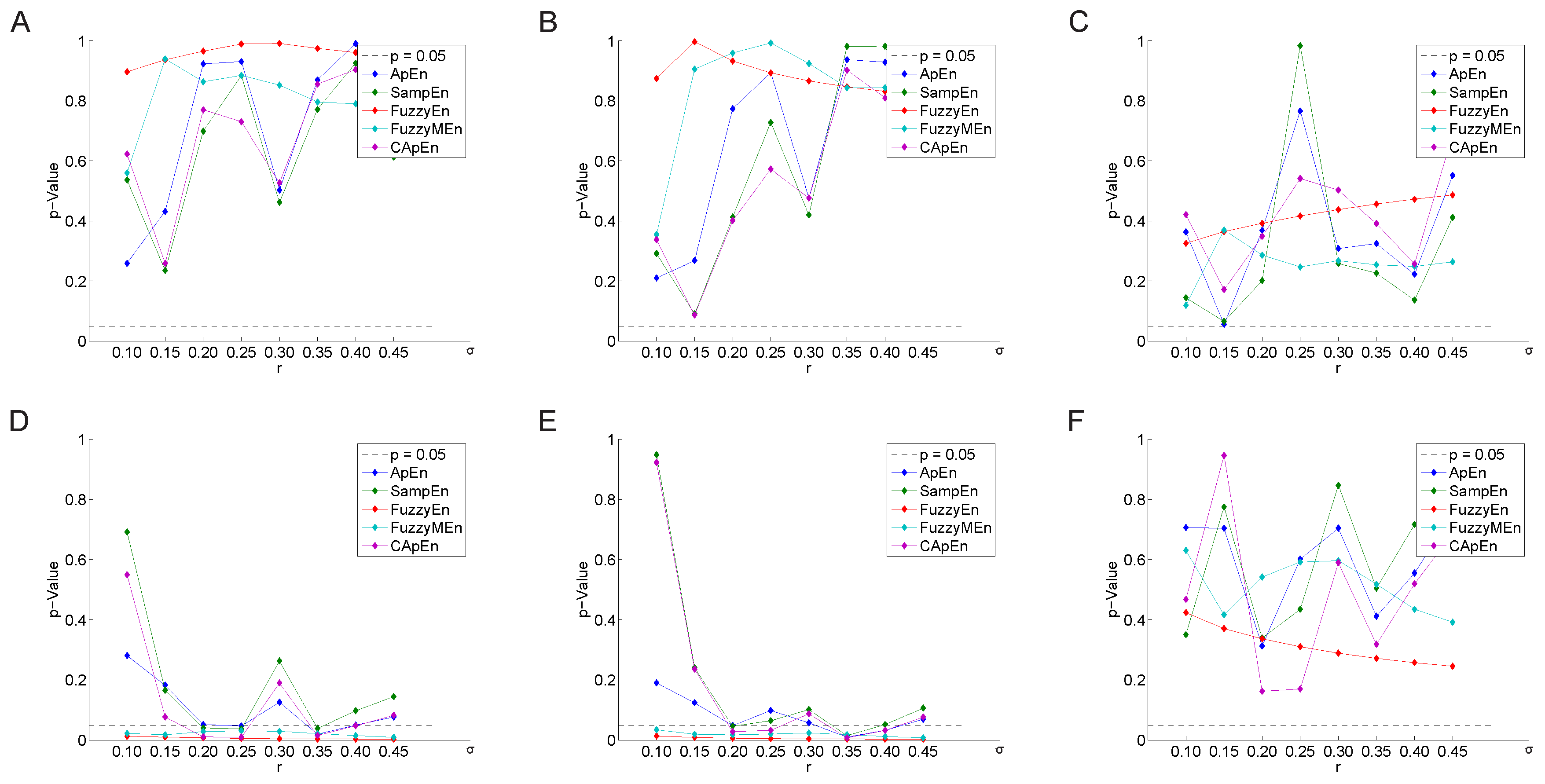
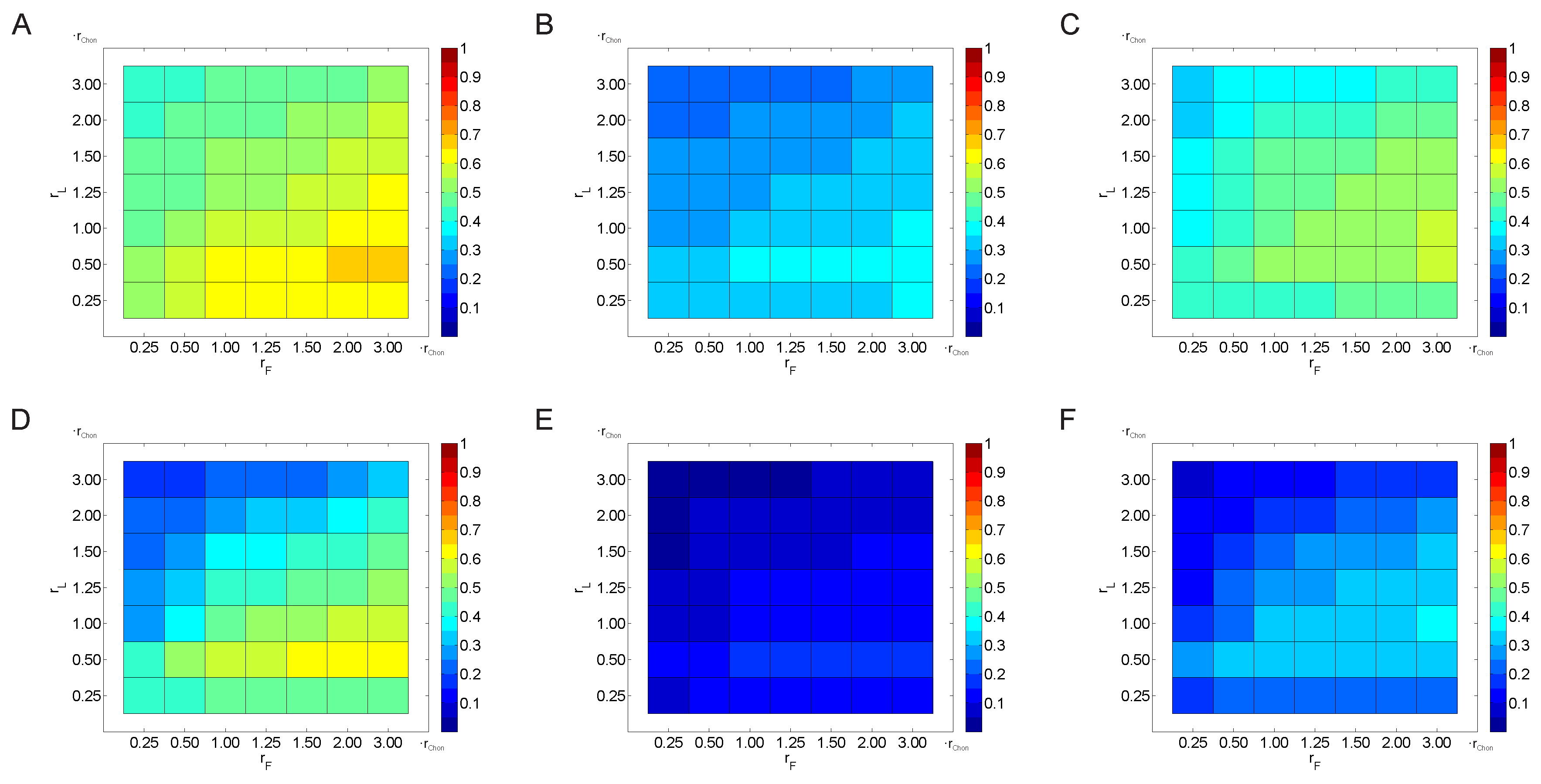
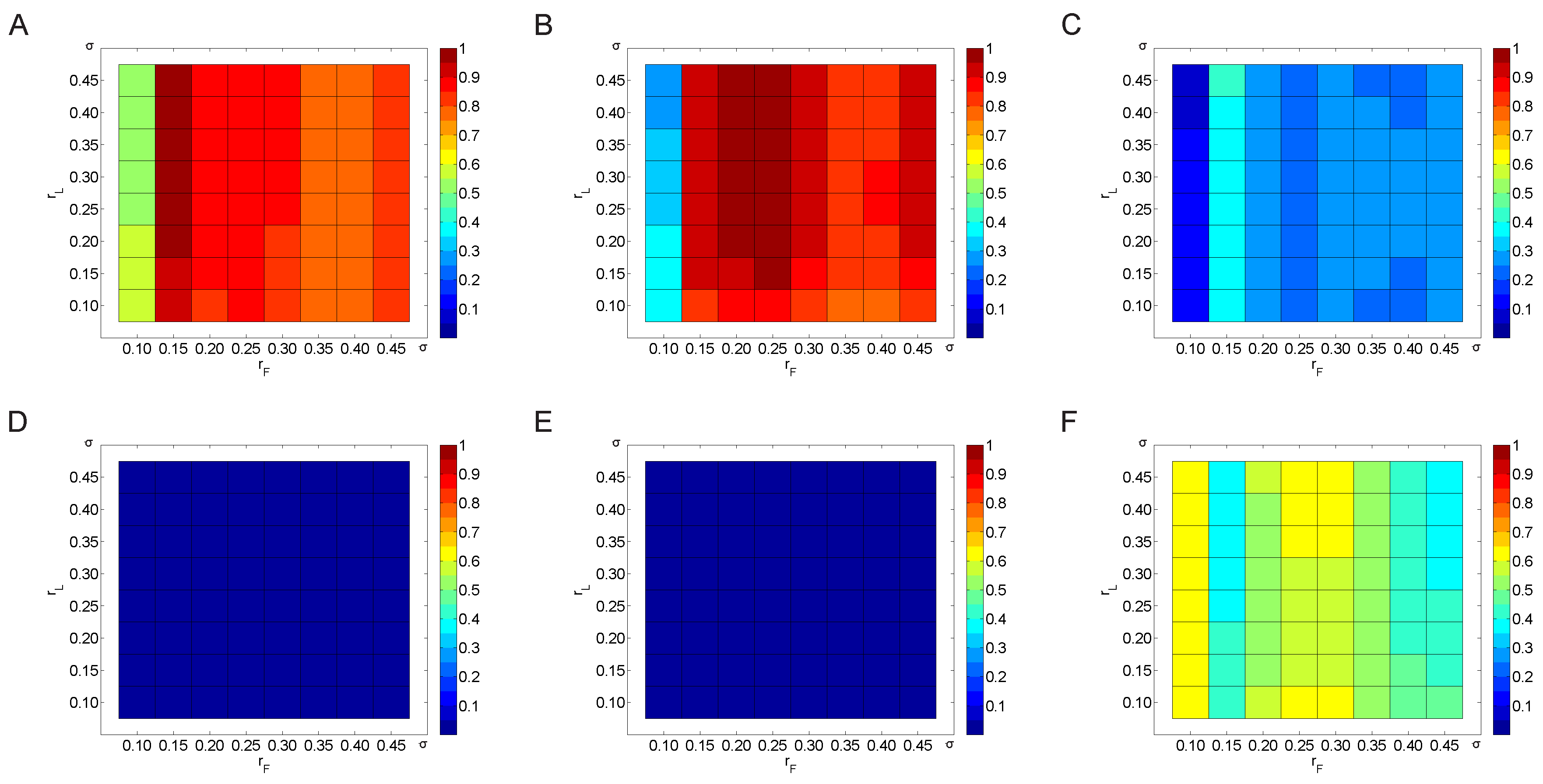
3.2.2. Variation of the Weighting Factor
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Conflicts of Interest
References
- Hon, E.H.; Lee, S. The fetal electrocardiogram. Am. J. Obstet. Gynecol. 1963, 87, 804–813. [Google Scholar] [CrossRef]
- Huikuri, H.V.; Stein, P.K. Heart Rate Variability in Risk Stratification of Cardiac Patients. Progr. Cardiovasc. Dis. 2013, 56, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Rajendra Acharya, U.; Paul Joseph, K.; Kannathal, N.; Lim, C.M.; Suri, J.S. Heart rate variability: A review. Med. Biol. Eng. Comput. 2006, 44, 1031–1051. [Google Scholar] [CrossRef] [PubMed]
- American Heart Association Inc.; European Society of Cardiology. Guidelines—Heart rate variability. Eur. Heart J. 1996, 17, 354–381. [Google Scholar]
- Sassi, R.; Cerutti, S.; Lombardi, F.; Malik, M.; Huikuri, H.V.; Peng, C.K.; Schmidt, G.; Yamamoto, Y.; Gorenek, B.; Lip, G.H.; et al. Advances in Heart Rate Variability Signal Analysis: Joint Position Statement by the E-Cardiology ESC Working Group and the European Heart Rhythm Association Co-Endorsed by the Asia Pacific Heart Rhythm Society. Europace 2015. [Google Scholar] [CrossRef] [PubMed]
- Stein, P.K.; Domitrovich, P.P.; Huikuri, H.V.; Kleiger, R.E.; Investigators, C. Traditional and Nonlinear Heart Rate Variability Are Each Independently Associated with Mortality after Myocardial Infarction. J. Cardiovasc. Electrophysiol. 2005, 16, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Stein, P.K.; Kleiger, R.E.; Domitrovich, P.P.; Schechtman, K.B.; Rottman, J.N. Clinical and demographic determinants of heart rate variability in patients post myocardial infarction: Insights from the cardiac arrhythmia suppression trial (CAST). Clin. Cardiol. 2000, 23, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Jordan, M.I.; Mitchell, T.M. Machine learning: Trends, perspectives, and prospects. Science 2015, 349, 255–260. [Google Scholar] [CrossRef] [PubMed]
- Holzinger, A.; Hörtenhuber, M.; Mayer, C.; Bachler, M.; Wassertheurer, S.; Pinho, A.J.; Koslicki, D. On Entropy-Based Data Mining. In Interactive Knowledge Discovery and Data Mining in Biomedical Informatics; Holzinger, A., Jurisica, I., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; Volume 8401, pp. 209–226. [Google Scholar]
- Pincus, S. Assessing serial irregularity and its implications for health. Ann. N. Y. Acad. Sci. 2001, 954, 245–267. [Google Scholar] [CrossRef] [PubMed]
- Hornero, R.; Aboy, M.; Abasolo, D.; McNames, J.; Goldstein, B. Interpretation of approximate entropy: Analysis of intracranial pressure approximate entropy during acute intracranial hypertension. IEEE Trans. Biomed. Eng. 2005, 52, 1671–1680. [Google Scholar] [CrossRef] [PubMed]
- Pincus, S.M. Approximate entropy as a measure of system complexity. Proc. Natl. Acad. Sci. USA 1991, 88, 2297–2301. [Google Scholar] [CrossRef] [PubMed]
- Richman, J.S.; Moorman, J.R. Physiological time-series analysis using approximate entropy and sample entropy. Am. J. Physiol. Heart Circ. Physiol. 2000, 278, H2039–H2049. [Google Scholar] [PubMed]
- Chen, W.; Zhuang, J.; Yu, W.; Wang, Z. Measuring complexity using FuzzyEn, ApEn, and SampEn. Med. Eng. Phys. 2009, 31, 61–68. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Li, K.; Zhao, L.; Liu, F.; Zheng, D.; Liu, C.; Liu, S. Analysis of heart rate variability using fuzzy measure entropy. Comput. Biol. Med. 2013, 43, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Porta, A.; Gnecchi-Ruscone, T.; Tobaldini, E.; Guzzetti, S.; Furlan, R.; Montano, N. Progressive decrease of heart period variability entropy-based complexity during graded head-up tilt. J. Appl. Physiol. 2007, 103, 1143–1149. [Google Scholar] [CrossRef] [PubMed]
- Mayer, C.; Bachler, M.; Hörtenhuber, M.; Stocker, C.; Holzinger, A.; Wassertheurer, S. Selection of entropy-measure parameters for knowledge discovery in heart rate variability data. BMC Bioinform. 2014, 15. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Wei, S.; Zhang, C.; Zhang, Y.; Jiang, X.; Liu, F.; Liu, C. Determination of Sample Entropy and Fuzzy Measure Entropy Parameters for Distinguishing Congestive Heart Failure from Normal Sinus Rhythm Subjects. Entropy 2015, 17, 6270–6288. [Google Scholar] [CrossRef]
- Bachler, M.; Hörtenhuber, M.; Mayer, C.; Holzinger, A.; Wassertheurer, S. Entropy-Based Data Mining on the Example of Cardiac Arrhythmia Suppression. In Brain Informatics and Health; Ślȩzak, D., Tan, A.-H., Peters, J., Schwabe, L., Eds.; Springer International Publishing: Berlin/Heidelberg, Germany, 2014; Volume 8609, pp. 574–585. [Google Scholar]
- Goldberger, A.L.; Amaral, L.A.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation 2000, 101, E215–E220. [Google Scholar] [CrossRef] [PubMed]
- The Cardiac Arrhythmia Suppression Trial (CAST) Investigators. Preliminary report: Effect of encainide and flecainide on mortality in a randomized trial of arrhythmia suppression after myocardial infarction. N. Engl. J. Med. 1989, 321, 406–412. [Google Scholar]
- Epstein, A.E.; Bigger, J.T.; Wyse, D.G.; Romhilt, D.W.; Reynolds-Haertle, R.A.; Hallstrom, A.P. Events in the Cardiac Arrhythmia Suppression Trial (CAST): Mortality in the entire population enrolled. J. Am. Coll. Cardiol. 1991, 18, 14–19. [Google Scholar] [CrossRef]
- Silva, I.; Moody, G. An open-source toolbox for analysing and processing physionet databases in matlab and octave. J. Open Res. Softw. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Chon, K.; Scully, C.G.; Lu, S. Approximate entropy for all signals. IEEE Eng. Med. Biol. Mag. 2009, 28, 18–23. [Google Scholar] [CrossRef] [PubMed]
- Pincus, S.M.; Goldberger, A.L. Physiological time-series analysis: what does regularity quantify? Am. J. Physiol. 1994, 266, H1643–H1656. [Google Scholar] [PubMed]
- Yentes, J.M.; Hunt, N.; Schmid, K.; Kaipust, J.; McGrath, D.; Stergiou, N. The Appropriate Use of Approximate Entropy and Sample Entropy with Short Data Sets. Ann. Biomed. Eng. 2013, 41, 349–365. [Google Scholar] [CrossRef] [PubMed]
- Alcaraz, R.; Abásolo, D.; Hornero, R.; Rieta, J.J. Optimal parameters study for sample entropy-based atrial fibrillation organization analysis. Comput. Methods Progr. Biomed. 2010, 99, 124–132. [Google Scholar] [CrossRef] [PubMed]
- Aktaruzzaman, M.; Sassi, R. Parametric estimation of sample entropy in heart rate variability analysis. Biomed. Signal Process. Control 2014, 14, 141–147. [Google Scholar] [CrossRef]
- Cox, D.R.; Oakes, D. Analysis of Survival Data; CRC Press: Boca Raton, FL, USA, 1984; Volume 21. [Google Scholar]
- Altman, D.G. Practical Statistics for Medical Research; CRC Press: Boca Raton, FL, USA, 1990. [Google Scholar]
- Box, G.E.P.; Cox, D.R. An Analysis of Transformations. J. R. Stat. Soc. Series B Methodol. 1964, 26, 211–252. [Google Scholar]
- Sakia, R. The Box–Cox transformation technique: A review. The Statistician 1992, 41, 169–178. [Google Scholar] [CrossRef]
- Mantel, N. Evaluation of survival data and two new rank order statistics arising in its consideration. Cancer Chemother. Rep. Part 1 1966, 50, 163–170. [Google Scholar]
- Stein, P.K.; Domitrovich, P.P.; Kleiger, R.E. Including patients with diabetes mellitus or coronary artery bypass grafting decreases the association between heart rate variability and mortality after myocardial infarction. Am. Heart J. 2004, 147, 309–316. [Google Scholar] [CrossRef]
- Altman, D.G.; Machin, D.; Bryant, T.N.; Gardner, M.J.; Gardner, S.; Bird, S.; Campbell, M.; Daly, L.E.; Morris, J. Statistics with Confidence: Confidence Intervals and Statistical Guidelines; BMJ Books: London, UK, 2000. [Google Scholar]
- Echt, D.S.; Liebson, P.R.; Mitchell, L.B.; Peters, R.W.; Obias-Manno, D.; Barker, A.H.; Arensberg, D.; Baker, A.; Friedman, L.; Greene, H.L.; et al. Mortality and morbidity in patients receiving encainide, flecainide, or placebo: The Cardiac Arrhythmia Suppression Trial. N. Engl. J. Med. 1991, 324, 781–788. [Google Scholar] [CrossRef] [PubMed]
- The Cardiac Arrhythmia Suppression Trial II Investigators. Effect of the antiarrhythmic agent moricizine on survival after myocardial infarction. N. Engl. J. Med. 1992, 327, 227–233. [Google Scholar]
- Lu, S.; Chen, X.; Kanters, J.K.; Solomon, I.C.; Chon, K.H. Automatic selection of the threshold value r for approximate entropy. IEEE Trans. Biomed. Eng. 2008, 55, 1966–1972. [Google Scholar] [PubMed]
- Liu, C.; Liu, C.; Shao, P.; Li, L.; Sun, X.; Wang, X.; Liu, F. Comparison of different threshold values r for approximate entropy: Application to investigate the heart rate variability between heart failure and healthy control groups. Physiol. Meas. 2011, 32, 167–180. [Google Scholar] [CrossRef] [PubMed]
- Castiglioni, P.; Di Rienzo, M. How the threshold “r” influences approximate entropy analysis of heart-rate variability. In Proceedings of Computers in Cardiology, Bologna, Italy, 14–17 September 2008; pp. 561–564.
- Beckers, F.; Verheyden, B.; Aubert, A.E. Aging and nonlinear heart rate control in a healthy population. Am. J. Physiol.-Heart Circulatory Physiol. 2006, 290, H2560–H2570. [Google Scholar] [CrossRef] [PubMed]
- Ryan, S.M.; Goldberger, A.L.; Pincus, S.M.; Mietus, J.; Lipsitz, L.A. Gender- and age-related differences in heart rate dynamics: Are women more complex than men? J. Am. Coll. Cardiol. 1994, 24, 1700–1707. [Google Scholar] [CrossRef]
| Before Treatment | After Treatment | |
|---|---|---|
| Sample size | 760 | 740 |
| Age (years) | ||
| Sex (m/f) | ||
| Time after MI (days) | ||
| CABG | 141 | 137 |
| DM | 160 | 155 |
| CABG & DM | 270 | 262 |
| Follow-up time (days) |
| No. | Source | m | N | Entropy Type | |||
|---|---|---|---|---|---|---|---|
| 1 | Mayer et al. [17] | 2 | 1200 | 2 | 1 | all | |
| 2 | Mayer et al. [17] | 2 | 1200 | 1 | 3 | all | |
| 3 | Zhao et al. [18] | 2 | 1000 | - | - | 0.15 | |
| 4 | Zhao et al. [18] | 2 | 1000 | 3 | 2 | 0.15 |
| No. | Variable | λ | All | w/o CABG | w/o CABG, w/o DM | |||
|---|---|---|---|---|---|---|---|---|
| p | p | p | ||||||
| 1 | ApEn | |||||||
| CApEn | ||||||||
| SampEn | ||||||||
| FuzzyEn | ||||||||
| FuzzyMEn | ||||||||
| 2 | ApEn | |||||||
| CApEn | ||||||||
| SampEn | ||||||||
| FuzzyEn | ||||||||
| FuzzyMEn | ||||||||
| 3 | SampEn | |||||||
| 4 | FuzzyMEn | |||||||
| No. | Variable | λ | All | w/o CABG | w/o CABG, w/o DM | |||
|---|---|---|---|---|---|---|---|---|
| p | p | p | ||||||
| 1 | ApEn | |||||||
| CApEn | ||||||||
| SampEn | ||||||||
| FuzzyEn | ||||||||
| FuzzyMEn | ||||||||
| 2 | ApEn | |||||||
| CApEn | ||||||||
| SampEn | ||||||||
| FuzzyEn | ||||||||
| FuzzyMEn | ||||||||
| 3 | SampEn | |||||||
| 4 | FuzzyMEn | |||||||
| All | w/o CABG | w/o CABG, w/o DM | ||||
|---|---|---|---|---|---|---|
| Variable | p | p | p | |||
| AVGNN | ||||||
| SDNN | ||||||
| Ln SDANN | ||||||
| Ln SDNNIDX | ||||||
| LN rMSSD | ||||||
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mayer, C.; Bachler, M.; Holzinger, A.; Stein, P.K.; Wassertheurer, S. The Effect of Threshold Values and Weighting Factors on the Association between Entropy Measures and Mortality after Myocardial Infarction in the Cardiac Arrhythmia Suppression Trial (CAST). Entropy 2016, 18, 129. https://doi.org/10.3390/e18040129
Mayer C, Bachler M, Holzinger A, Stein PK, Wassertheurer S. The Effect of Threshold Values and Weighting Factors on the Association between Entropy Measures and Mortality after Myocardial Infarction in the Cardiac Arrhythmia Suppression Trial (CAST). Entropy. 2016; 18(4):129. https://doi.org/10.3390/e18040129
Chicago/Turabian StyleMayer, Christopher, Martin Bachler, Andreas Holzinger, Phyllis K. Stein, and Siegfried Wassertheurer. 2016. "The Effect of Threshold Values and Weighting Factors on the Association between Entropy Measures and Mortality after Myocardial Infarction in the Cardiac Arrhythmia Suppression Trial (CAST)" Entropy 18, no. 4: 129. https://doi.org/10.3390/e18040129
APA StyleMayer, C., Bachler, M., Holzinger, A., Stein, P. K., & Wassertheurer, S. (2016). The Effect of Threshold Values and Weighting Factors on the Association between Entropy Measures and Mortality after Myocardial Infarction in the Cardiac Arrhythmia Suppression Trial (CAST). Entropy, 18(4), 129. https://doi.org/10.3390/e18040129







