Measures of Morphological Complexity of Gray Matter on Magnetic Resonance Imaging for Control Age Grouping
Abstract
:1. Introduction
2. Methods
2.1. Approximate Entropy and Sample Entropy
2.2. Regularity Dimension
2.3. Recurrence Plots
2.4. Largest Lyapunov Exponent
2.5. Dynamic-Time Warping
- Boundary constraint by imposing , and . This condition is also known as endpoint constraints.
- Monotonicity property such that , and . This means a valid path must follow a monotonic order with respect to time.
- Continuity condition by setting and , . This is also known as step-size constraints.
- Warping window: By setting , where ω is a positive integer representing the window bandwidth. This restriction means that only features within a warping-path window are considered.
- Slope constraint: By introducing a slope-weighting vector , where , , and are the weights for the horizontal, vertical, and diagonal directions, respectively. The purpose of this slope constraint is to avoid having a warping path that is either too steep or too shallow, and prevent matching very short segments with very long ones.
2.6. Phylogenetic Tree Reconstruction
- Given dataset , , , where is the Kronecker delta: 0 if and 1 if (each is a singleton cluster at the number of clusters , , where is the c-partition of ).
- At step k, , using to directly solve the measure of hard-cluster similarity (hard clustering means that each data point is a member of one and only one cluster) by minimizing the following function to identify the minimum distance as the similarity between any two data points in :where , denote the j-th and k-th rows of , , and is any measure of dissimilarity on , and d was used as a spectral-distortion measure in this study.
- Let solve Equation (39). Merge and , thus constructing from the updated partition , record .
- If , go to Step 2; if , . Merge the two remaining clusters, set , compute , and stop.
3. MRI Data and Preprocessing
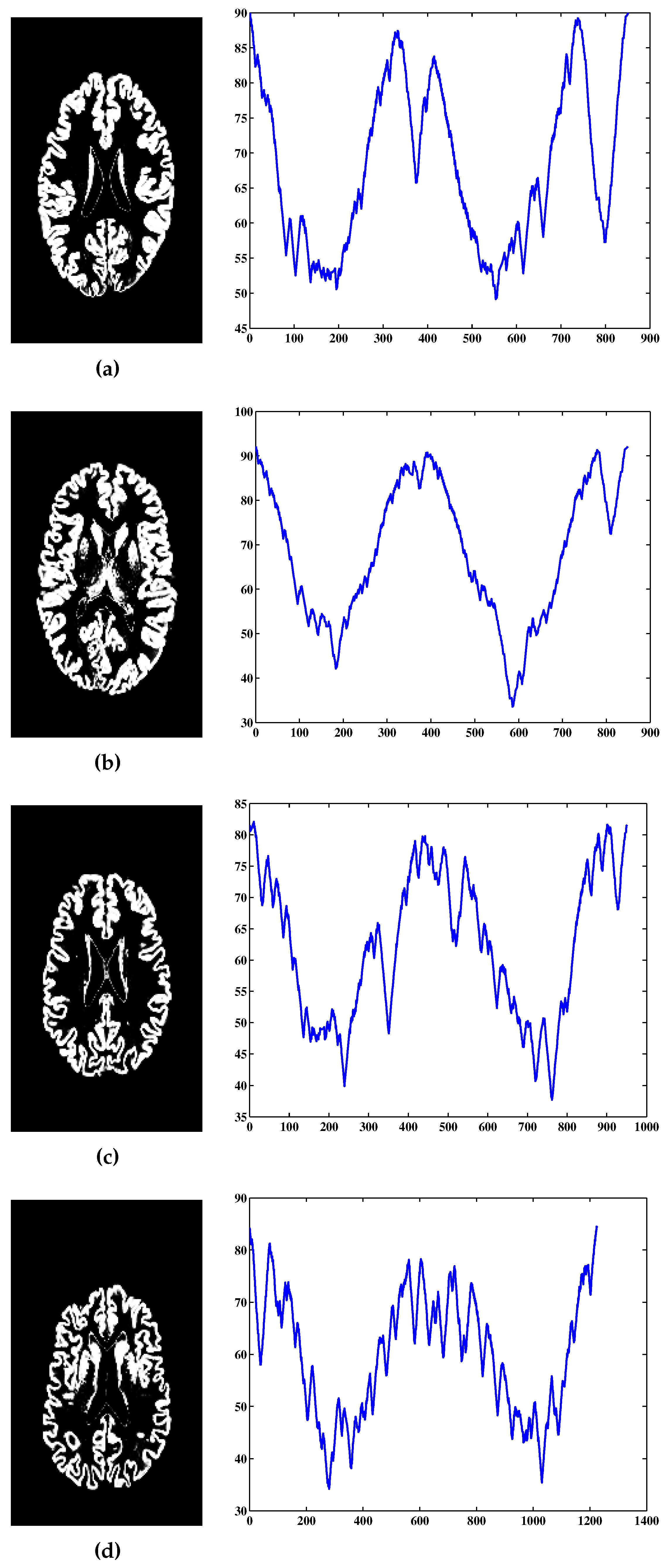
4. Results and Discussion
| Age Group | m = 1 | m = 2 | m = 3 |
|---|---|---|---|
| 20–29 | (4.5079, 0.0173) | (1.5254, 0.0032) | (1.2375, 0.0031) |
| 30–39 | (4.5115, 0.0139) | (1.5424, 0.0028) | (1.2400, 0.0026) |
| 40–49 | (4.5540, 0.0145) | (1.5653, 0.0024) | (1.2504, 0.0023) |
| 50–59 | (4.5838, 0.0143) | (1.5897, 0.0013) | (1.2673, 0.0015) |
| 60–69 | (4.6531, 0.0148) | (1.6067, 0.0008) | (1.2773, 0.0015) |
| 70–79 | (4.7003, 0.0089) | (1.6197, 0.0006) | (1.2812, 0.0010) |
| 80–86 | (4.7765, 0.0018) | (1.6158, 0.0006) | (1.3029, 0.0006) |
| Age Group | m = 1 | m = 2 | m = 3 |
|---|---|---|---|
| 20–29 | (1.0482, 0.0002) | (0.7409, 0.0006) | (0.6907, 0.0020) |
| 30–39 | (1.0475, 0.0001) | (0.7448, 0.0005) | (0.6666, 0.0024) |
| 40–49 | (1.0519, 0.0001) | (0.7572, 0.0003) | (0.6441, 0.0024) |
| 50–59 | (1.0528, 0.0001) | (0.7664, 0.0002) | (0.6094, 0.0027) |
| 60–69 | (1.0524, 0.0001) | (0.7733, 0.0002) | (0.5997, 0.0024) |
| 70–79 | (1.0521, 0.0001) | (0.7753, 0.0002) | (0.5786, 0.0036) |
| 80–86 | (1.0516, 0.0001) | (0.7783, 0.0002) | (0.6282, 0.0008) |
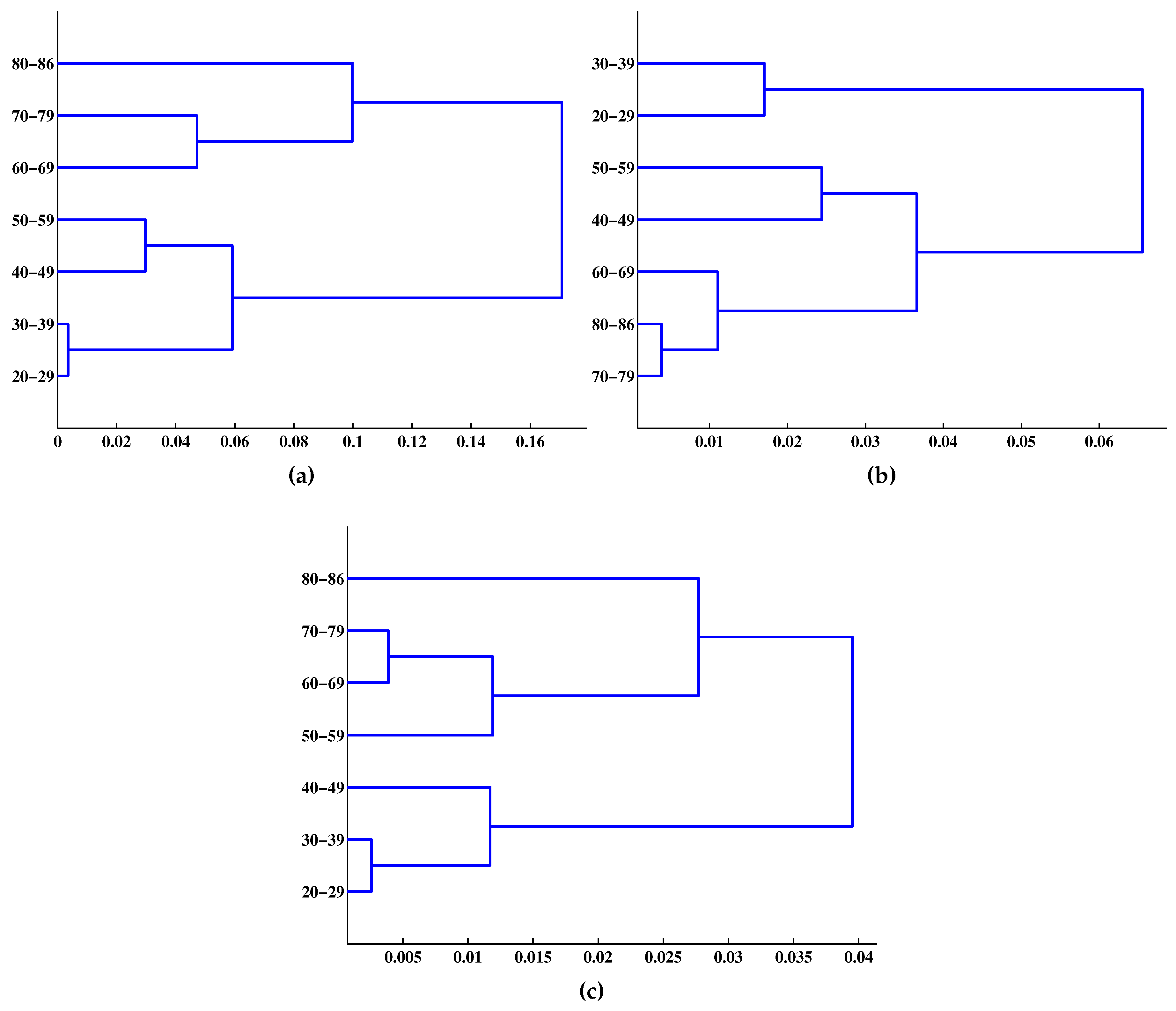
| Age Group | m = 1 | m = 2 | m = 3 |
|---|---|---|---|
| 20–29 | (0.0187, 0.0000) | (0.0161, 0.0000) | (0.0154, 0.0000) |
| 30–39 | (0.0192, 0.0000) | (0.0167, 0.0000) | (0.0159, 0.0000) |
| 40–49 | (0.0189, 0.0000) | (0.0165, 0.0000) | (0.0158, 0.0000) |
| 50–59 | (0.0189, 0.0000) | (0.0166, 0.0000) | (0.0159, 0.0000) |
| 60–69 | (0.0181, 0.0000) | (0.0158, 0.0000) | (0.0152, 0.0000) |
| 70–79 | (0.0176, 0.0000) | (0.0154, 0.0000) | (0.0147, 0.0000) |
| 80–86 | (0.0164, 0.0000) | (0.0142, 0.0000) | (0.0136, 0.0000) |
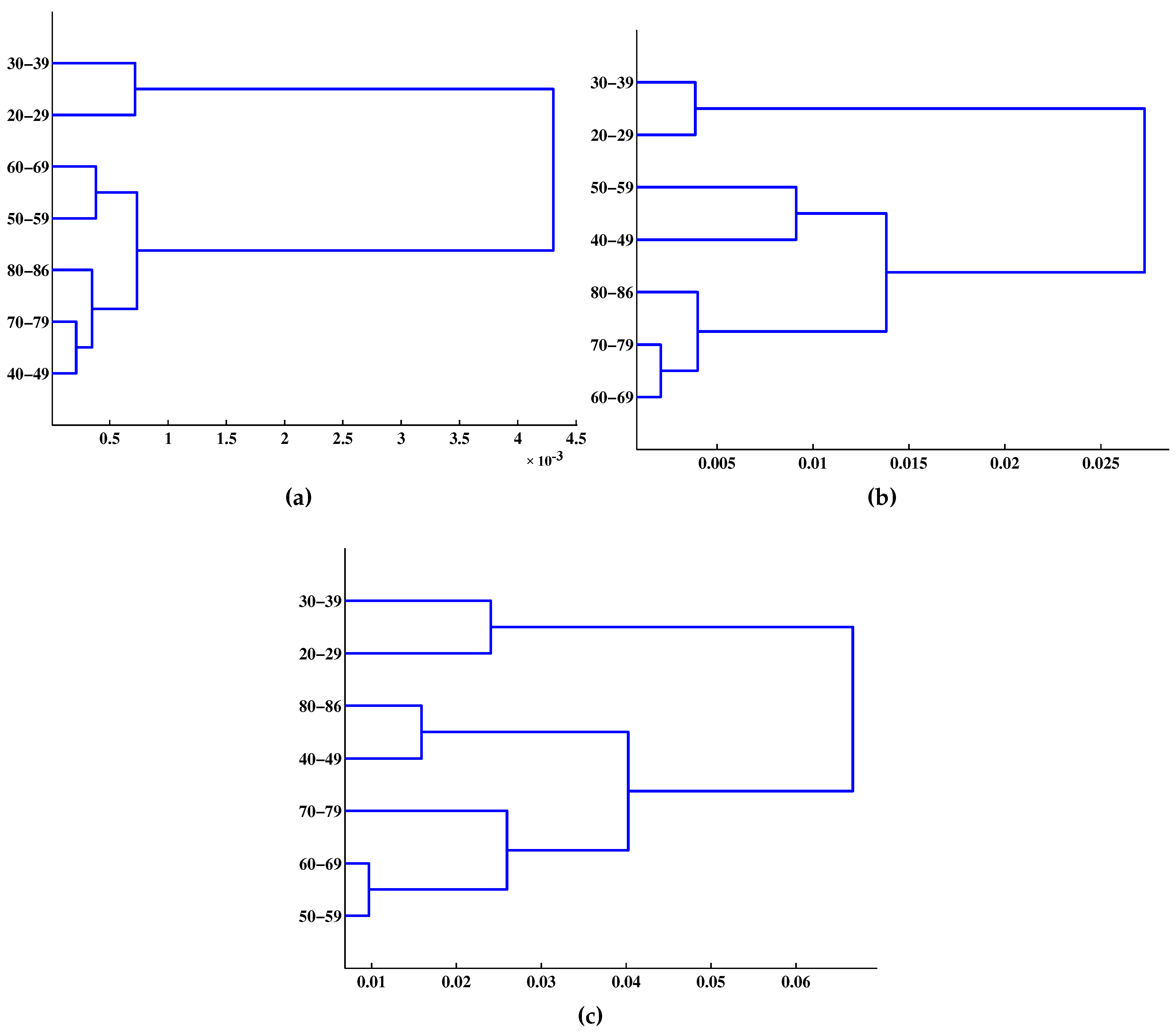
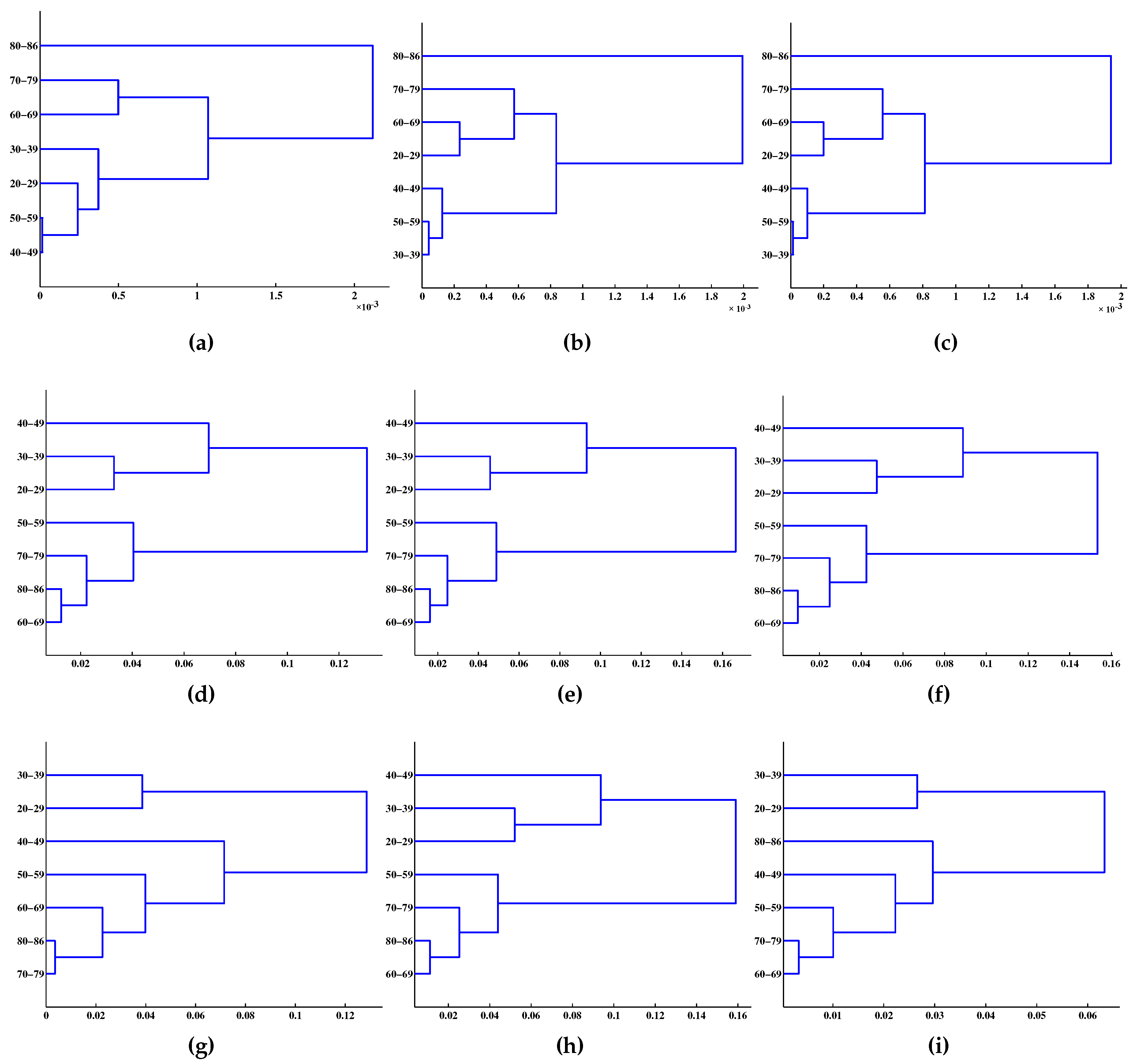
| Age Group | m = 1 | m = 2 | m = 3 |
|---|---|---|---|
| 20–29 | (5.1295, 0.0134) | (4.6164, 0.0202) | (4.3049, 0.0224) |
| 30–39 | (5.0966, 0.0123) | (4.5690, 0.0170) | (4.2528, 0.0184) |
| 40–49 | (5.0435, 0.0090) | (4.5039, 0.0118) | (4.1852, 0.0125) |
| 50–59 | (4.9894, 0.0066) | (4.4417, 0.0087) | (4.1217, 0.0091) |
| 60–69 | (4.9627, 0.0056) | (4.4123, 0.0070) | (4.0917, 0.0071) |
| 70–79 | (4.9341, 0.0041) | (4.3826, 0.0053) | (4.0609, 0.0054) |
| 80–86 | (4.9502, 0.0027) | (4.4027, 0.0031) | (4.0806, 0.0033) |
| Age Group | m = 1 | m = 2 | m = 3 |
|---|---|---|---|
| 20–29 | (4.9974, 0.0243) | (4.4715, 0.0157) | (4.2885, 0.0068) |
| 30–39 | (4.9517, 0.0210) | (4.4329, 0.0114) | (4.2620, 0.0042) |
| 40–49 | (4.8813, 0.0142) | (4.3807, 0.0075) | (4.2346, 0.0027) |
| 50–59 | (4.8136, 0.0098) | (4.3391, 0.0043) | (4.2190, 0.0014) |
| 60–69 | (4.7811, 0.0081) | (4.3144, 0.0031) | (4.2106, 0.0008) |
| 70–79 | (4.7483, 0.0060) | (4.2935, 0.0016) | (4.2073, 0.0004) |
| 80–86 | (4.7649, 0.0038) | (4.2899, 0.0020) | (4.1883, 0.0002) |
| Age Group | m = 1 | m = 2 | m = 3 |
|---|---|---|---|
| 20–29 | (0.3230, 0.0002) | (0.3546, 0.0001) | (0.3516, 0.0001) |
| 30–39 | (0.3267, 0.0001) | (0.3599, 0.0001) | (0.3566, 0.0001) |
| 40–49 | (0.3322, 0.0001) | (0.3642, 0.0001) | (0.3601, 0.0001) |
| 50–59 | (0.3328, 0.0001) | (0.3671, 0.0001) | (0.3624, 0.0000) |
| 60–69 | (0.3357, 0.0001) | (0.3677, 0.0000) | (0.3626, 0.0000) |
| 70–79 | (0.3361, 0.0000) | (0.3685, 0.0000) | (0.3628, 0.0000) |
| 80–86 | (0.3408, 0.0000) | (0.3722, 0.0000) | (0.3652, 0.0000) |
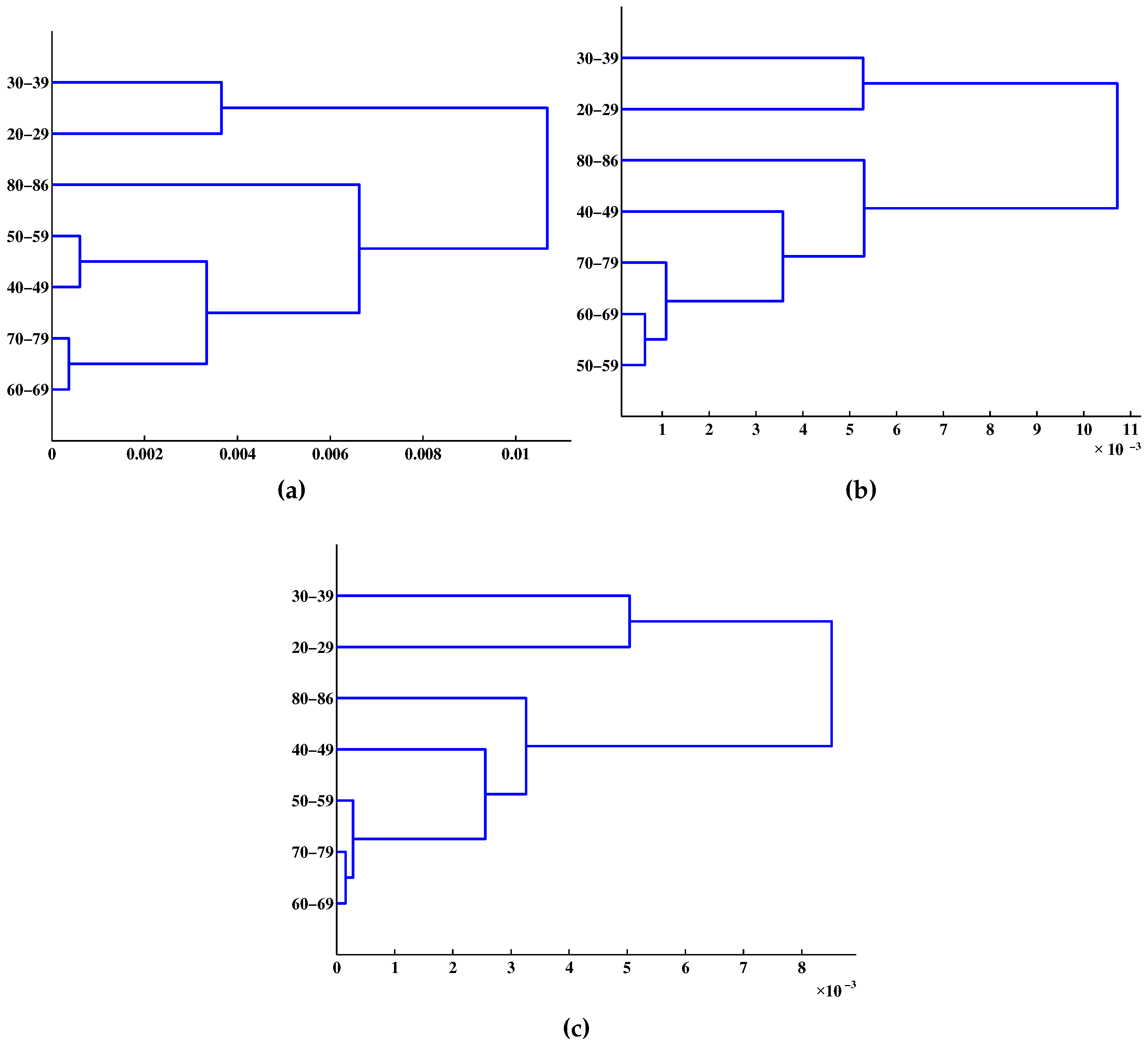
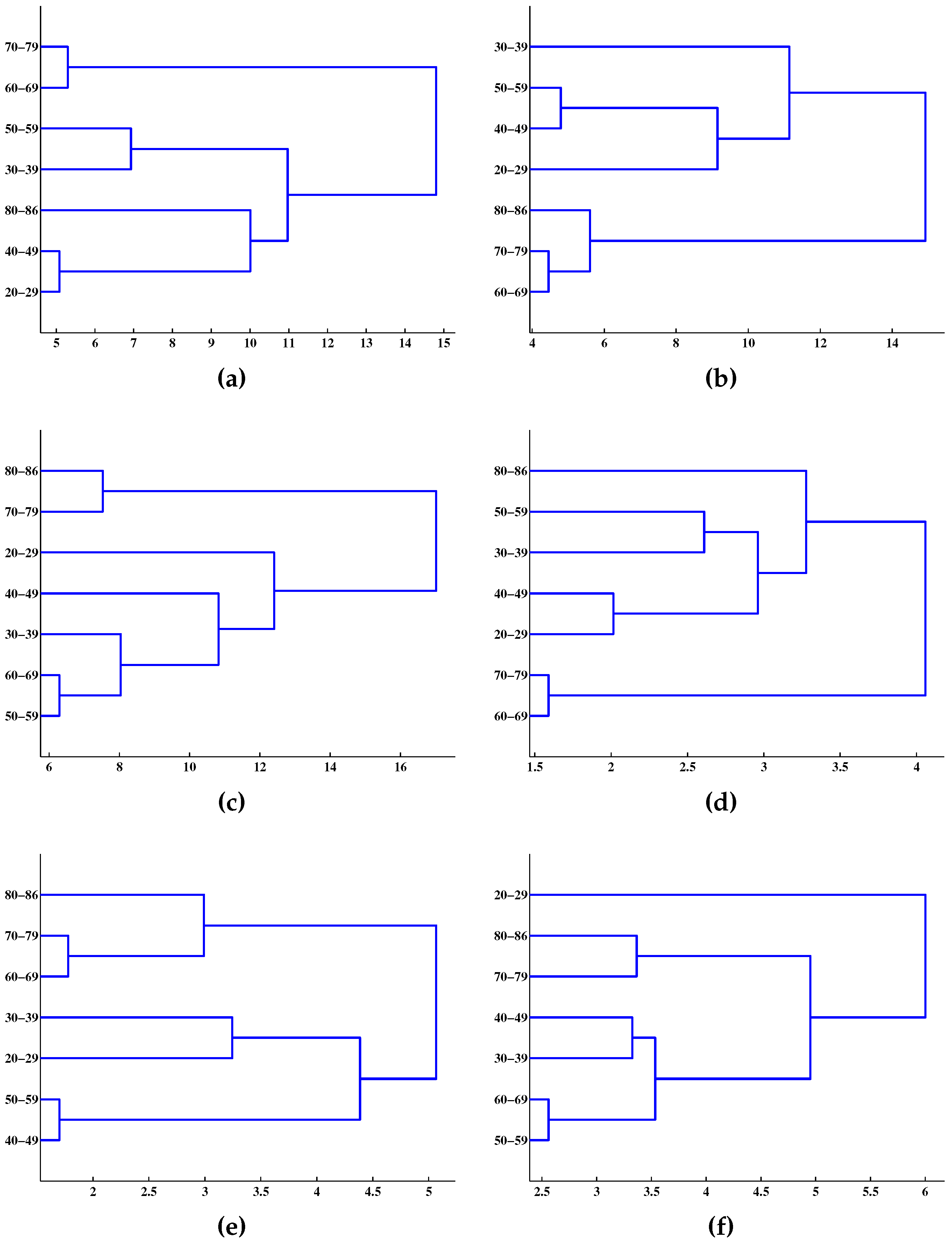
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Teverovskiy, L.A.; Becker, J.T.; Lopez, O.L.; Liu, Y. Quantified brain asymmetry for age estimation of normal and AD/MCI subjects. In Proceedings of the 5th IEEE International Symposium on Biomedical Imaging: From Nano to Macro (ISBI 2008), Paris, France, 14–17 May 2008; pp. 1509–1512.
- Franke, K.; Ziegler, G.; Kloppel, S.; Gaser, C. Estimating the age of healthy subjects from T1-weighted MRI scans using kernel methods: Exploring the influence of various parameters. NeuroImage 2010, 50, 883–892. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Pham, T.D. MRI-based age prediction using hidden Markov models. J. Neurosci. Methods 2011, 199, 140–145. [Google Scholar] [CrossRef] [PubMed]
- Dukart, J.; Schroeter, M.L.; Mueller, K. Age correction in dementia–matching to a healthy brain. PLoS ONE 2011, 6, e22193. [Google Scholar] [CrossRef] [PubMed]
- Kandel, B.M.; Wolk, D.A.; Gee, J.C.; Avants, B. Predicting cognitive data from medical images using sparse linear regression. In Information Processing in Medical Imaging; Gee, J.C., Joshi, S., Pohl, K.M., Wells, W.M., ZÃũllei, L., Eds.; Springer: Berlin/Heidelberg, Germany, 2013; Volume 7917, pp. 86–97. [Google Scholar]
- Irimia, A.; Torgerson, C.M.; Goh, S.Y.; van Horn, J.D. Statistical estimation of physiological brain age as a descriptor of senescence rate during adulthood. Brain Imaging Behav. 2015, 9, 678–689. [Google Scholar] [CrossRef] [PubMed]
- Cole, J.H.; Leech, R.; Sharp, D.J. Prediction of brain age suggests accelerated atrophy after traumatic brain injury. Ann. Neurol. 2015, 77, 571–581. [Google Scholar] [CrossRef] [PubMed]
- Spulber, G.; Niskanen, E.; MacDonald, S.; Smilovici, O.; Chen, K.; Reimanet, E.M.; Jauhiainen, A.M.; Hallikainen, M.; Tervo, S.; Wahlund, L.-O.; et al. Whole brain atrophy rate predicts progression from MCI to Alzheimer’s disease. Neurobiol. Aging 2010, 31, 1601–1605. [Google Scholar] [CrossRef] [PubMed]
- Pham, T.D.; Salvetti, F.; Wang, B.; Diani, M.; Heindel, W.; Knecht, S.; Wersching, H.; Baune, B.T.; Berger, K. The hidden-Markov brain: Comparison and inference of white matter hyperintensities on magnetic resonance imaging (MRI). J. Neural Eng. 2011, 8, 016004. [Google Scholar] [CrossRef] [PubMed]
- Su, L.; Wang, L.; Hu, D. Predicting the age of healthy adults from structural MRI by sparse representation. In Intelligent Science and Intelligent Data Engineering; Yang, J., Fang, F., Sun, C., Eds.; Springer: Berlin/Heidelberg, Germany, 2013; Volume 7751, pp. 271–279. [Google Scholar]
- Gaser, C.; Franke, K.; Kloppel, S.; Koutsouleris, N.; Sauer, H. BrainAGE in mild cognitive impaired patients: Predicting the conversion to Alzheimer’s disease. PLoS ONE 2013, 8, e67346. [Google Scholar] [CrossRef] [PubMed]
- Bigler, E.D. Traumatic brain injury, neuroimaging, and neurodegeneration. Front. Hum. Neurosci. 2013, 7. [Google Scholar] [CrossRef] [PubMed]
- Sowell, E.R.; Peterson, B.S.; Thompson, P.M. Mapping cortical change across the human life span. Nat. Neurosci. 2003, 6, 309–315. [Google Scholar] [CrossRef] [PubMed]
- Raz, N.; Rodrigue, K.M. Differential aging of the brain: Patterns, cognitive correlates and modifiers. Neurosci. Biobehav. Rev. 2006, 30, 730–748. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Pham, T.D. HMM-based brain age interpolation using kriging estimator. In Proceedings of the IEEE International Symposium on Image and Signal Processing and Analysis, Dubrovnik, Croatia, 4–6 September 2011; pp. 704–708.
- Chen, Y.; Pham, T.D. Entropy and regularity dimension in complexity analysis of cortical surface structure in early Alzheimer’s disease and aging. J. Neurosci. Methods 2013, 215, 210–217. [Google Scholar] [CrossRef] [PubMed]
- What is Alzheimer’s? Available online: http://www.alz.org/alzheimers_disease_what_is_alzheimers.asp (accessed on 16 August 2015).
- Neeb, H.; Zilles, K.; Shah, N.J. Fully-automated detection of cerebral water content changes: Study of age- and gender-related H2O patterns with quantitative MRI. NeuroImage 2006, 29, 910–922. [Google Scholar] [CrossRef] [PubMed]
- Ashburner, J. A fast diffeomorphic image registration algorithm. NeuroImage 2007, 38, 95–113. [Google Scholar] [CrossRef] [PubMed]
- Brown, T.A. Genomics, 2nd ed.; Wiley: New York, NY, USA, 2002. [Google Scholar]
- Radford, A.; Atkinson, M.; Britain, D.; Clahsen, H.; Spencer, A. Linguistics: An Introduction, 2nd ed.; Cambridge University Press: Cambridge, UK, 1999. [Google Scholar]
- Douaud, G.; Refsum, H.; de Jager, C.A.; Jacoby, R.; Nichols, T.E.; Smith, S.M.; Smith, A.D. Preventing Alzheimer’s disease-related gray matter atrophy by B-vitamin treatment. Proc. Natl. Acad. Sci. USA 2013, 110, 9523–9528. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Riklin-Raviv, T.; Bouix, S. Shape analysis, a field in need of careful validation. Hum. Brain Mapp. 2014, 35, 4965–4978. [Google Scholar] [CrossRef] [PubMed]
- The Brain Geek. Available online: http://thebraingeek.blogspot.jp/2012/04/folds-of-brain.html (accessed on 7 September 2015).
- Geschwind, D.H.; Rakic, P. Cortical evolution: Judge the brain by its cover. Neuron 2013, 80, 633–647. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Hevner, R.F. Growth and folding of the mammalian cerebral cortex: From molecules to malformations. Nat. Rev. Neurosci. 2014, 15, 217–232. [Google Scholar] [CrossRef] [PubMed]
- Keogh, E.; Wei, L.; Xi, X.; Lee, S.H.; Vlachos, M. LB_Keogh supports exact indexing of shapes under rotation invariance with arbitrary representations and distance measures. In Proceedings of the 32nd International Conference on Very Large Data Bases, Seoul, Korea, 12–15 September 2006; pp. 882–893.
- Tak, Y.S.; Hwang, E. A leaf image retrieval scheme based on partial dynamic time warping and two-level filtering. In Proceedings of the 7th IEEE International Conference on Computer and Information Technology, Fukushima, Japan, 16–19 October 2007; pp. 633–638.
- Bartolini, I.; Ciaccia, P.; Patella, M. WARP: Accurate retrieval of shapes using phase of Fourier descriptors and time warping distance. IEEE Trans. Pattern Anal. Mach. Intell. 2005, 27, 142–147. [Google Scholar] [CrossRef] [PubMed]
- Skarda, C.A.; Freeman, W.J. Chaos and the new science of the brain. Concepts Neurosci. 1990, 1, 275–285. [Google Scholar]
- Liebovitch, L.S. Fractals and Chaos Simplified for the Life Science; Oxford University Press: New York, NY, USA, 1998. [Google Scholar]
- Van Straaten, E.C.W.; Stam, C.J. Structure out of chaos: Functional brain network analysis with EEG, MEG, and functional MRI. Eur. Neuropsychopharmacol. 2013, 23, 7–18. [Google Scholar] [CrossRef] [PubMed]
- Strogatz, S.H. Nonlinear Dynamics and Chaos: With Applications to Physics, Biology, Chemistry, and Engineering, 2nd ed.; Westview: Cambridge, MA, USA, 2014. [Google Scholar]
- Pham, T.D. Classification of complex biological aging images using fuzzy Kolmogorov-Sinai entropy. J. Phys. D Appl. Phys. 2014, 47. [Google Scholar] [CrossRef]
- Gutierrez-Tobal, G.C.; Alvarez, D.; Gomez-Pilar, J.; del Campo, F.; Hornero, R. Assessment of time and frequency domain entropies to detect sleep apnoea in heart rate variability recordings from men and women. Entropy 2015, 17, 123–141. [Google Scholar] [CrossRef]
- Pan, W.Y.; Su, M.C.; Wu, H.T.; Lin, M.C.; Tsai, I.T.; Sun, C.K. Multiscale entropy analysis of heart rate variability for assessing the severity of sleep disordered breathing. Entropy 2015, 17, 231–243. [Google Scholar] [CrossRef]
- Pincus, S.M. Approximate entropy (ApEn) as a complexity measure. Chaos 1995, 5, 110–117. [Google Scholar] [CrossRef] [PubMed]
- Richman, J.S.; Moorman, J.R. Physiological time-series analysis using approximate entropy and sample entropy. Am. J. Physiol. Heart Circ. Physiol. 2000, 278, H2039–H2049. [Google Scholar] [PubMed]
- Pham, T.D. Regularity dimension of sequences and its application to phylogenetic tree reconstruction. Chaos Soliton. Fract. 2012, 45, 879–887. [Google Scholar] [CrossRef]
- Eckmann, J.P.; Kamphorst, S.O.; Ruelle, D. Recurrence plots of dynamical systems. EPL Europhys. Lett. 1987, 4, 973–977. [Google Scholar] [CrossRef]
- Rosenstein, M.T.; Collins, J.J.; DeLuca, C.J. A practical method for calculating largest Lyapunov exponents from small data sets. Phys. D Nonlinear Phenom. 1993, 65, 117–134. [Google Scholar] [CrossRef]
- Williams, G.P. Chaos Theory Tamed; Joseph Henry Press: Washington, DC, USA, 1997. [Google Scholar]
- Sakoe, H.; Chiba, S. Dynamic programming algorithm optimization for spoken word recognition. IEEE Trans. Acoust. Speech Signal Process 1978, 26, 43–49. [Google Scholar] [CrossRef]
- Rabiner, L.R.; Juang, B. Fundamentals of Speech Recognition; Prentice-Hall: Upper Saddle River, NJ, USA, 1993. [Google Scholar]
- Grassberger, P.; Procaccia, L. Estimation of the Kolmogorov entropy from a chaotic signal. Phys. Rev. A 1983, 28, 2591–2593. [Google Scholar] [CrossRef]
- Eckmann, J.P.; Ruelle, D. Ergodic theory of chaos and strange attractors. Rev. Mod. Phys. 1985, 57, 617–656. [Google Scholar] [CrossRef]
- Schroeder, M. Fractals, Chaos, Power Laws: Minutes from an Infinite Paradise; W.H. Freeman: New York, NY, USA, 1991. [Google Scholar]
- Casdagli, M.C. Recurrence plots revisited. Phys. D Nonlinear Phenom. 1997, 108, 12–44. [Google Scholar] [CrossRef]
- Marwan, N.; Romano, M.C.; Thiel, M.; Kurths, J. Recurrence plots for the analysis of complex systems. Phys. Rep. 2007, 438, 237–329. [Google Scholar] [CrossRef]
- Facchini, A.; Mocenni, C.; Vicino, A. Generalized recurrence plots for the analysis of images from spatially distributed systems. Phys. D Nonlinear Phenom. 2009, 238, 162–169. [Google Scholar] [CrossRef]
- Dingwell, J.B. Lyapunov exponents. In Wiley Encyclopedia of Biomedical Engineering; Metin, A., Ed.; John Wiley & Sons: New York, NY, USA, 2006. [Google Scholar]
- Pham, T.D. The butterfly effect in ER dynamics and ER-mitochondrial contacts. Chaos Soliton. Fract. 2014, 65, 5–19. [Google Scholar] [CrossRef]
- Pham, T.D. Validation of computer models for evaluating the efficacy of cognitive stimulation therapy. Wirel. Pers. Commun. 2015. [Google Scholar] [CrossRef]
- Takens, F. Detecting strange attractors in turbulence. Lect. Notes Math. 1981, 898, 366–381. [Google Scholar]
- Ecker, J.G.; Kupferschmid, M. Introduction to Operations Research; John Wiley & Sons: New York, NY, USA, 1988. [Google Scholar]
- Pham, T.D.; Oyama-Higa, M.; Truong, C.T.; Okamoto, K.; Futaba, F.; Kanemoto, S.; Sugiyama, M.; Lampe, L. Computerized assessment of communication for cognitive stimulation for people with cognitive decline using spectral-distortion measures and phylogenetic inference. PLoS ONE 2015, 10, e0118739. [Google Scholar] [CrossRef] [PubMed]
- Michener, C.D.; Sokal, R.R. A quantitative approach to a problem in classification. Evolution 1957, 11, 130–162. [Google Scholar] [CrossRef]
- Bezdek, J.C. Pattern Recognition with Fuzzy Objective Function Algorithms; Plenum: New York, NY, USA, 1981. [Google Scholar]
- IXI (Information eXtraction from Images) Dataset. Available online: http://www.brain-development.org (accessed on 4 December 2015).
- Giorgio, A.; Santelli, L.; Tomassini, V.; Bosnell, R.; Smith, S.; de Stefano, N.; Johansen-Berg, H. Age-related changes in grey and white matter structure throughout adulthood. Neuroimage 2010, 51, 943–951. [Google Scholar] [CrossRef] [PubMed]
- SPM: Statistical Parametric Mapping. Available online: http://www.fil.ion.ucl.ac.uk/spm (accessed on 11 March 2015).
- Ashburner, J.; Friston, K.J. Unified segmentation. Neuroimage 2005, 26, 839–851. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Pham, T.D. Development of a brain MRI-based hidden Markov model for dementia recognition. BioMed. Eng. Online 2013, 12, S2. [Google Scholar] [CrossRef] [PubMed]
- Theodoridis, S.; Pikrakis, A.; Koutroumbas, K.; Cavouras, D. Introduction to Pattern Recognition: A Matlab Approach; Academic Press: New York, NY, USA, 2010. [Google Scholar]
- Sprott, J.C. Chaos and Time-Series Analysis; Oxford University Press: Oxford, UK, 2003. [Google Scholar]
- Cardoza, J.D.; Goldstein, R.B.; Filly, R.A. Exclusion of fetal ventriculomegaly with a single measurement: The width of the lateral ventricular atrium. Radiology 1988, 169, 711–714. [Google Scholar] [CrossRef] [PubMed]
- Raz, N.; Lindenberger, U.; Rodrigue, K.M.; Kennedy, K.M.; Head, D.; Williamson, A.; Dahle, C.; Gerstorf, D.; Acker, J.D. Regional brain changes in aging healthy adults: General trends, individual differences and modifiers. Cereb. Cortex 2005, 15, 1676–1689. [Google Scholar] [CrossRef] [PubMed]
- Craik, F.I.M.; Salthouse, T.A. The Handbook of Aging and Cognition, 3rd ed.; Psychology Press: New York, NY, USA, 2008. [Google Scholar]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pham, T.D.; Abe, T.; Oka, R.; Chen, Y.-F. Measures of Morphological Complexity of Gray Matter on Magnetic Resonance Imaging for Control Age Grouping. Entropy 2015, 17, 8130-8151. https://doi.org/10.3390/e17127868
Pham TD, Abe T, Oka R, Chen Y-F. Measures of Morphological Complexity of Gray Matter on Magnetic Resonance Imaging for Control Age Grouping. Entropy. 2015; 17(12):8130-8151. https://doi.org/10.3390/e17127868
Chicago/Turabian StylePham, Tuan D., Taishi Abe, Ryuichi Oka, and Yung-Fu Chen. 2015. "Measures of Morphological Complexity of Gray Matter on Magnetic Resonance Imaging for Control Age Grouping" Entropy 17, no. 12: 8130-8151. https://doi.org/10.3390/e17127868
APA StylePham, T. D., Abe, T., Oka, R., & Chen, Y.-F. (2015). Measures of Morphological Complexity of Gray Matter on Magnetic Resonance Imaging for Control Age Grouping. Entropy, 17(12), 8130-8151. https://doi.org/10.3390/e17127868








