Entropy and the Complexity of Graphs Revisited
Abstract
:1. Introduction
2. Taxonomy
- The deterministic category encompasses the encoding, substructure count and generative approaches. Dominant in the encoding approach is Kolmogorov complexity. The second includes measures which count the number of substructures of a specified kind [20]. Generative approaches consist of measures based on operations required to generate a graph [21].
- The probabilistic category includes measures that apply an entropy function to a probability distribution associated with a graph. This category is subdivided into intrinsic and extrinsic subcategories. Intrinsic measures use structural features of a graph to partition the graph (usually the set of vertices or edges) and thereby determine a probability distribution over the components of the partition. Extrinsic measures impose an arbitrary probability distribution on graph elements [22]. Both of these categories employ the probability distribution to compute an entropy value. Shannon’s entropy function is most commonly used, but several different families of entropy functions have been considered [23]. In the next section, we provide a brief overview of the main subcategories of the deterministic class of complexity measures. The probabilistic category is our main concern and will be examined in more detail in subsequent sections.
| Deterministic Measures | Probabilistic Measures |
|---|---|
| Encoding | Intrinsic (probability distribution derived from structural features) |
| Substructure Count | Extrinsic (probability distribution externally imposed) |
| Generative |
3. Deterministic Complexity Measures
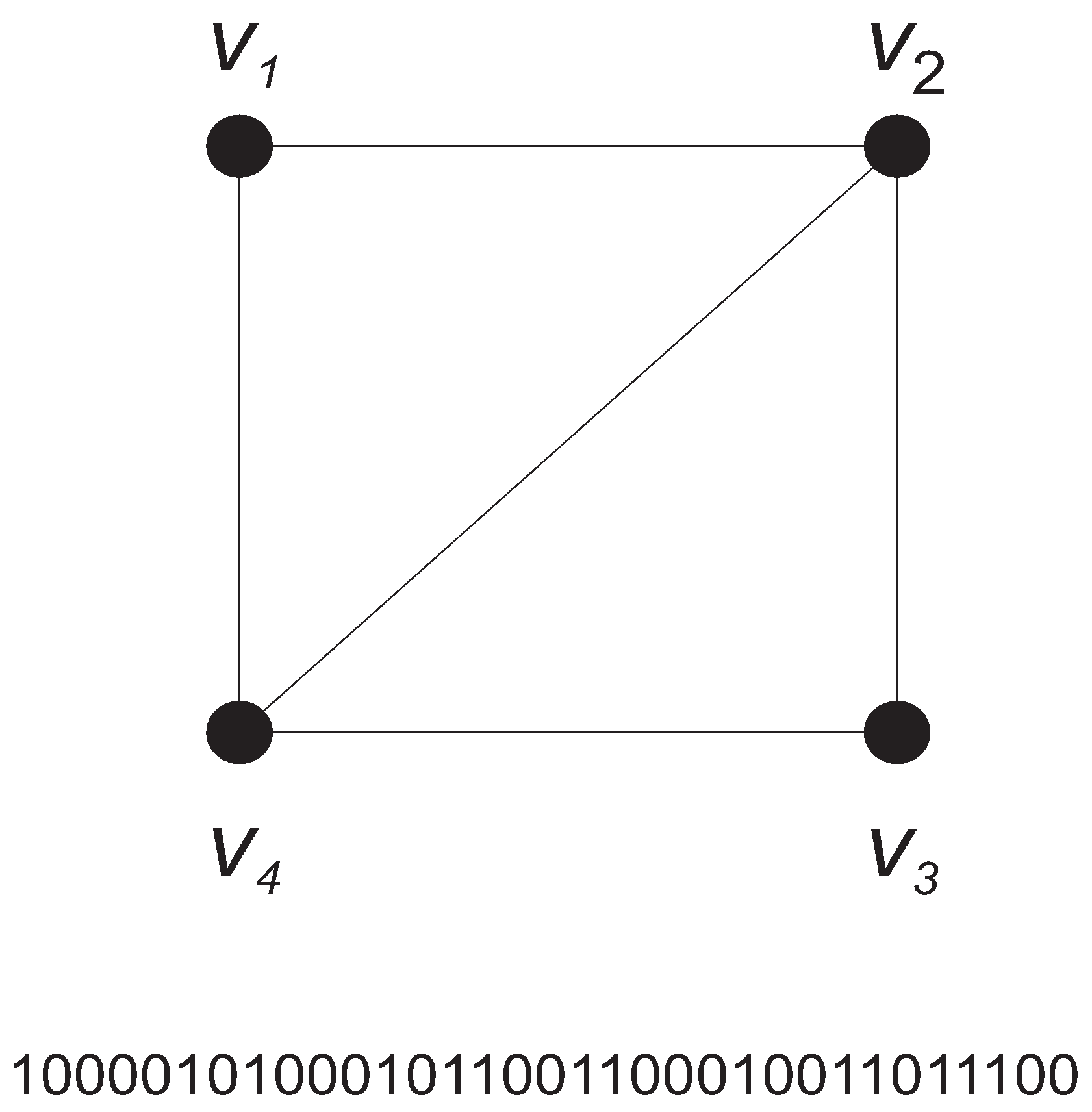
4. Probabilistic Measures of Graph Complexity
4.1. Classical Graph Entropies
4.2. Körner Entropy
4.3. Parametric Graph Entropies
4.4. Non-Parametric Graph Entropies
5. Conclusions
Acknowledgements
References and Notes
- Rashevsky, N. Life, information theory, and topology. Bull. Math. Biophys. 1955, 17, 229–235. [Google Scholar] [CrossRef]
- Albert, R.; Jeong, H.; Barabási, A.L. Diameter of the world wide web. Nature 1999, 401, 130–131. [Google Scholar]
- Albert, R.; Barabási, A.L. Statistical mechanics of complex networks. Rev. Mod. Phys. 2002, 74, 47. [Google Scholar] [CrossRef]
- Bonchev, D.; Trinajstić, N. Information theory, distance matrix and molecular branching. J. Chem. Phys. 1977, 67, 4517–4533. [Google Scholar] [CrossRef]
- Bonchev, D. Information Theoretic Indices for Characterization of Chemical Structures; Research Studies Press: Chichester, West Sussex, UK, 1983. [Google Scholar]
- Butts, C.T. The complexity of social networks: Theoretical and empirical findings. Soc. Network. 2001, 23, 31–71. [Google Scholar] [CrossRef]
- Mehler, A.; Weiß, P.; Lücking, A. A network model of interpersonal alignment. Entropy 2010, 12, 1440–1483. [Google Scholar] [CrossRef]
- Solé, R.V.; Montoya, J.M. Complexity and fragility in ecological networks. Proc. Roy. Soc. Lond. B Biol. Sci. 2001, 268, 2039–2045. [Google Scholar] [CrossRef] [PubMed]
- Ulanowicz, R.E. Quantitative methods for ecological network analysis. Comput. Biol. Chem. 2004, 28, 321–339. [Google Scholar] [CrossRef]
- Dehmer, M.; Barbarini, N.; Varmuza, K.; Graber, A. Novel topological descriptors for analyzing biological networks. BMC Struct. Biol. 2010, 10. [Google Scholar] [CrossRef]
- Basak, S.C.; Balaban, A.T.; Grunwald, G.D.; Gute, B.D. Topological indices: Their nature and mutual relatedness. J. Chem. Inf. Comput. Sci. 2000, 40, 891–898. [Google Scholar] [CrossRef] [PubMed]
- Bonchev, D.; Rouvray, D.H. (Eds.) Complexity in Chemistry, Biology, and Ecology; Mathematical and Computational Chemistry; Springer: New York, NY, USA, 2005.
- Emmert-Streib, F.; Dehmer, M. Information theoretic measures of UHG graphs with low computational complexity. Appl. Math. Comput. 2007, 190, 1783–1794. [Google Scholar] [CrossRef]
- Dehmer, M.; Mowshowitz, A. A history of graph entropy measures. Inform. Sci. 2011, 1, 57–78. [Google Scholar] [CrossRef]
- Mowshowitz, A. Entropy and the complexity of the graphs: I. An index of the relative complexity of a graph. Bull. Math. Biophys. 1968, 30, 175–204. [Google Scholar] [CrossRef]
- Anand, K.; Bianconi, G. Entropy measures for networks: Toward an information theory of complex topologies. Phys. Rev. E 2009, 80, 045102(R). [Google Scholar] [CrossRef]
- Solé, R.V.; Valverde, S. Information theory of complex networks: On evolution and architectural constraints. Lect. Notes Phys. 2004, 650, 189–207. [Google Scholar]
- Thurner, S. Statistical mechanics of complex networks. In Analysis of Complex Networks: From Biology to Linguistics; Dehmer, M., Emmert-Streib, F., Eds.; Wiley-VCH: Weinheim, Germany, 2009; pp. 23–45. [Google Scholar]
- Wilhelm, T.; Hollunder, J. Information theoretic description of networks. Physica A 2007, 388, 385–396. [Google Scholar] [CrossRef]
- Constantine, G. Graph complexity and the Laplacian matrix in blocked experiments. Linear and Multilinear Algebra 1990, 28, 49–56. [Google Scholar] [CrossRef]
- Bonchev, D. Kolmogorov’s information, Shannon’s entropy, and topological complexity of molecules. Bulg. Chem. Commun. 1995, 28, 567–582. [Google Scholar]
- Dehmer, M. Information processing in complex networks: Graph entropy and information functionals. Appl. Math. Comput. 2008, 201, 82–94. [Google Scholar] [CrossRef]
- Dehmer, M.; Mowshowitz, A. Generalized graph entropies. Complexity 2011, 17, 45–50. [Google Scholar] [CrossRef]
- Kolmogorov, A.N. Three approaches to the definition of information (in Russian). Probl. Peredaci Inform. 1965, 1, 3–11. [Google Scholar]
- Li, M.; Vitányi, P. An Introduction to Kolmogorov Complexity and Its Applications; Springer: New York, NY, USA, 1997. [Google Scholar]
- Bertz, S.H.; Sommmer, T.J. Rigorous mathematical approaches to strategic bonds and synthetic analysis based on conceptually simple new complexity indices. Chem. Commun. 1997. [Google Scholar] [CrossRef]
- Bonchev, D. Complexity analysis of yeast proteome network. Chem. Biodivers. 2004, 1, 312–326. [Google Scholar] [CrossRef]
- Platt, J.R. Influence of neighbor bonds on additive bond properties in paraffins. J. Chem. Phys. 1947, 15, 419–420. [Google Scholar] [CrossRef]
- Bonchev, D. Overall connectivities and topological complexities: A new powerful tool for QSPR/QSAR. J. Chem. Inf. Comput. Sci. 2000, 40, 934–941. [Google Scholar] [CrossRef] [PubMed]
- Bonchev, D. The overall Wiener index—A new tool for characterization of molecular topology. J. Chem. Inf. Comput. Sci. 2001, 41, 582–592. [Google Scholar] [CrossRef] [PubMed]
- Bonchev, D. The overall topological complexity indices. In Advances in Computational Methods in Science and Engineering; Simos, T., Maroulis, G., Eds.; VSP Publications: Melville, NY, USA, 2005; Volume 4B, pp. 1554–1557. [Google Scholar]
- Pudlák, P.; Rödl, V.; Savickiý, P. Graph complexity. Acta Informatica 1988, 25, 515–535. [Google Scholar]
- Shannon, C.E.; Weaver, W. The Mathematical Theory of Communication; University of Illinois Press: Urbana, IL, USA, 1949. [Google Scholar]
- Cover, T.M.; Thomas, J.A. Elements of Information Theory; Wiley Series in Telecommunications and Signal Processing; Wiley & Sons: Hoboken, NJ, USA, 2006. [Google Scholar]
- Butts, C.T. An axiomatic approach to network complexity. J. Math. Sociol. 2000, 24, 273–301. [Google Scholar] [CrossRef]
- Mowshowitz, A. Entropy and the complexity of graphs II: The information content of digraphs and infinite graphs. Bull. Math. Biophys. 1968, 30, 225–240. [Google Scholar] [CrossRef] [PubMed]
- Bonchev, D. Information theoretic measures of complexity. In Encyclopedia of Complexity and System Science; Meyers, R., Ed.; Springer: New York, NY, USA, 2009; Volume 5, pp. 4820–4838. [Google Scholar]
- Todeschini, R.; Consonni, V.; Mannhold, R. Handbook of Molecular Descriptors; Wiley-VCH: Weinheim, Germany, 2002. [Google Scholar]
- Bang-Jensen, J.; Gutin, G. Digraphs, Theory, Algorithms and Applications; Springer: London, UK, 2002. [Google Scholar]
- Körner, J. Coding of an information source having ambiguous alphabet and the entropy of graphs. In Proceedings of the 6th Prague Conference on Information Theory, Prague, Czech Republic, 1973; pp. 411–425.
- Simonyi, G. Graph entropy: A survey. In Combinatorial Optimization; Cook, W., Lovász, L., Seymour, P., Eds.; DIMACS Series in Discrete Mathematics and Theoretical Computer Science; American Mathematical Society: Providence, RI, USA, 1995; Volume 20, pp. 399–441. [Google Scholar]
- Cardinal, L.; Fiorini, S.; Assche, G.V. On minimum entropy graph colorings. In Proceedings of the 2004 IEEE International Symposium on Information Theory, Piscataway, NJ, USA, 2004; p. 43.
- Dehmer, M.; Varmuza, K.; Borgert, S.; Emmert-Streib, F. On entropy-based molecular descriptors: Statistical analysis of real and synthetic chemical structures. J. Chem. Inf. Model. 2009, 49, 1655–1663. [Google Scholar] [CrossRef] [PubMed]
- Dehmer, M.; Mowshowitz, A.; Emmert-Streib, F. Connections between classical and parametric network entropies. PLoS One 2011, 6, e15733. [Google Scholar] [CrossRef]
- Dehmer, M.; Sivakumar, L.; Varmuza, K. Uniquely discriminating molecular structures using novel eigenvalue-based descriptors. MATCH Commun. Math. Comput. Chem. 2012, 67, 147–172. [Google Scholar]
- Polansky, O.E.; Bonchev, D. The Wiener number of graphs. I. General theory and changes due to graph operations. MATCH MATCH Commun. Math. Comput. Chem. 1986, 21, 133–186. [Google Scholar]
- Hosoya, H. On some counting polynomials. Discrete Appl. Math. 1988, 19, 239–257. [Google Scholar] [CrossRef]
- Balaban, A.T.; Harary, F. The characteristic polynomial does not uniquely determined the topology of a molecule. J. Chem. Doc. 1971, 11, 258–259. [Google Scholar] [CrossRef]
- Randić, M.; Vracko, M.; Novic, M. Eigenvalues as molecular descriptors. In QSPR/QSAR Studies by Molecular Descriptors; Diudea, M.V., Ed.; Nova Publishing: Huntington, NJ, USA, 2001; pp. 93–120. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mowshowitz, A.; Dehmer, M. Entropy and the Complexity of Graphs Revisited. Entropy 2012, 14, 559-570. https://doi.org/10.3390/e14030559
Mowshowitz A, Dehmer M. Entropy and the Complexity of Graphs Revisited. Entropy. 2012; 14(3):559-570. https://doi.org/10.3390/e14030559
Chicago/Turabian StyleMowshowitz, Abbe, and Matthias Dehmer. 2012. "Entropy and the Complexity of Graphs Revisited" Entropy 14, no. 3: 559-570. https://doi.org/10.3390/e14030559
APA StyleMowshowitz, A., & Dehmer, M. (2012). Entropy and the Complexity of Graphs Revisited. Entropy, 14(3), 559-570. https://doi.org/10.3390/e14030559






