Influence of Conformational Entropy on the Protein Folding Rate
Abstract
:1. Introduction
84 proteins | multi 26 proteins | two 58 proteins | ||
|---|---|---|---|---|
| Parameters of protein size and average size of protein loop | ||||
| L | -0.64 ± 0.06 | -0.58 ± 0.08 | -0.61 ± 0.11 | |
| L1/2 | -0.73 ± 0.05 | -0.61 ± 0.12 | -0.71 ± 0.06 | |
| L2/3 | -0.69 ± 0.06 | -0.60 ± 0.13 | -0.67 ± 0.07 | |
| ln L | -0.77 ± 0.04 | -0.64 ± 0.12 | -0.76 ± 0.06 | |
| AbsCO | -0.77 ± 0.04 | -0.65 ± 0.11 | -0.78 ± 0.05 | |
| Relative contact order is the parameter of an average size of the protein loop normalized to exclude dependence on protein size. | ||||
| CO | 0.00 ± 0.11 | -0.01 ± 0.20 | -0.17 ± 0.13 | |
2. Results and Discussion
2.1. Entropy Capacity for Proteins with Given Topology
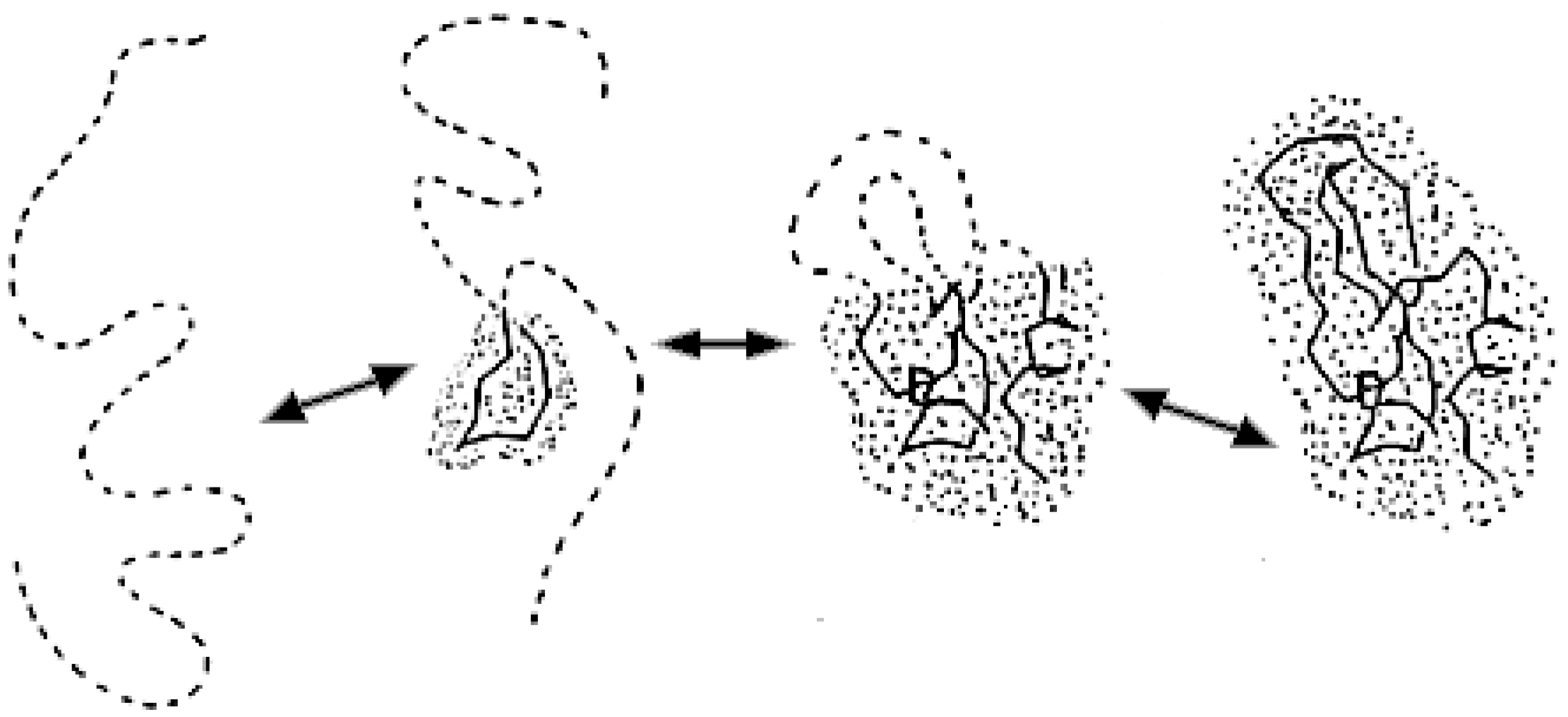
2.2. Optimal Value of Entropy Capacity for Fast Protein Folding
2.3. Statistical Analysis of Average Conformational Entropy and Average Number of Contacts per Residue for Different Classes of Proteins
| Type of residue | ALA | ARG | ASN | ASP | CYS | GLN | GLU | GLY | HIS | ILE |
| Degrees of freedom | 2 | 6 | 4 | 4 | 3 | 5 | 5 | 3* | 4 | 4 |
| Type of residue | LEU | LYS | MET | PHE | PRO | SER | THR | TRP | TYR | VAL |
| Degrees of freedom | 4 | 6 | 5 | 4 | 1 | 3 | 3 | 4 | 4 | 3 |
| SCOP 1.61 (5829 proteins) | ||||
|---|---|---|---|---|
| Class a (1133) | Class b (1644) | Class c (1617) | Class d (1435) | |
| <L> | 151 ± 3 | 156 ± 2 | 256 ± 3 | 163 ± 2 |
| <m(8 Å)> | 18.33 ± 0.09 | 19.48 ± 0.06 | 21.56 ± 0.06 | 19.39 ± 0.07 |
| <ν> | 3.87 ± 0.01 | 3.71 ± 0.01 | 3.76 ± 0.01 | 3.80 ± 0.01 |
| <Cmod=ν/m> | 0.219 ± 0.001 | 0.196 ± 0.002 | 0.178 ± 0.001 | 0.200 ± 0.001 |
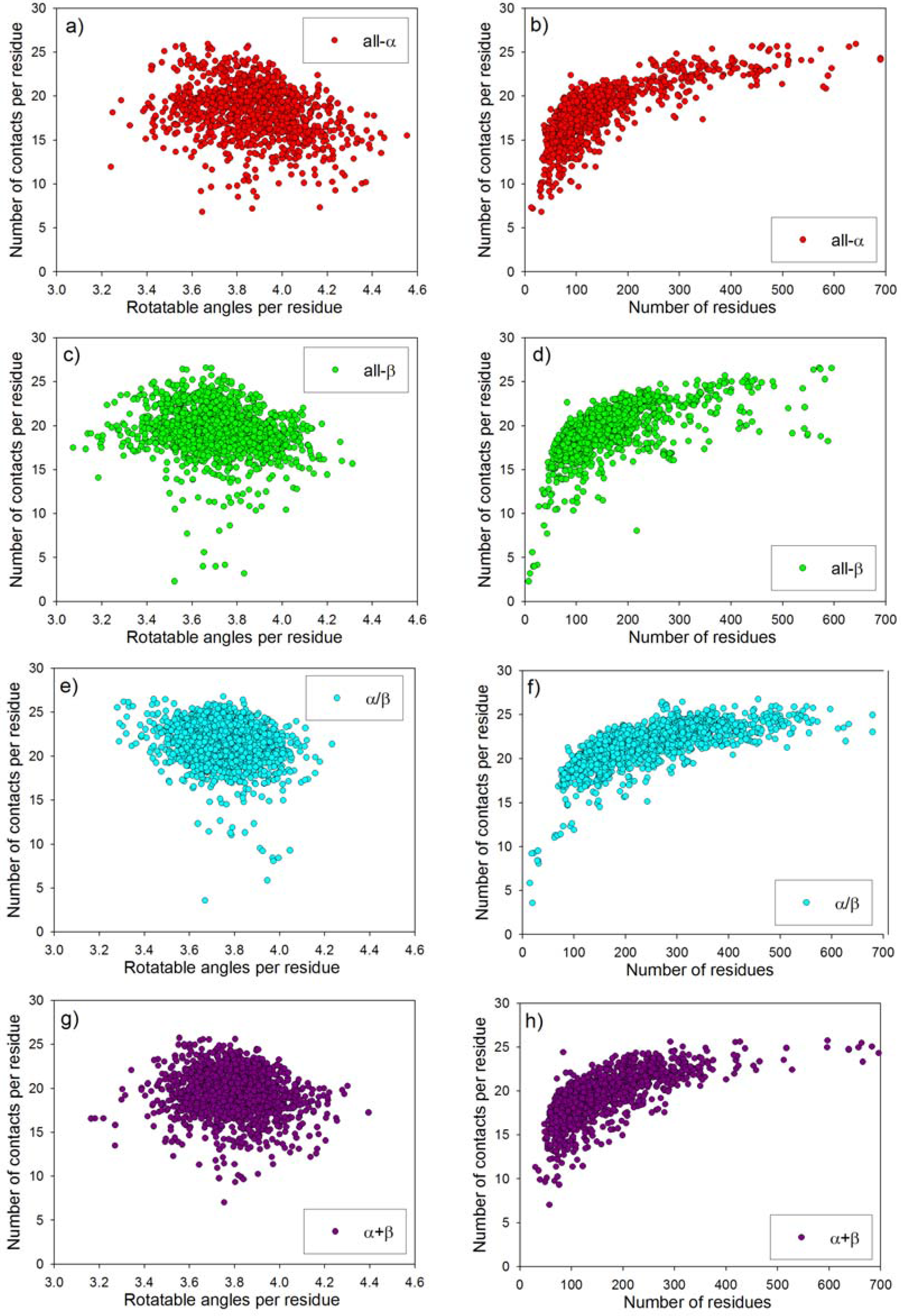
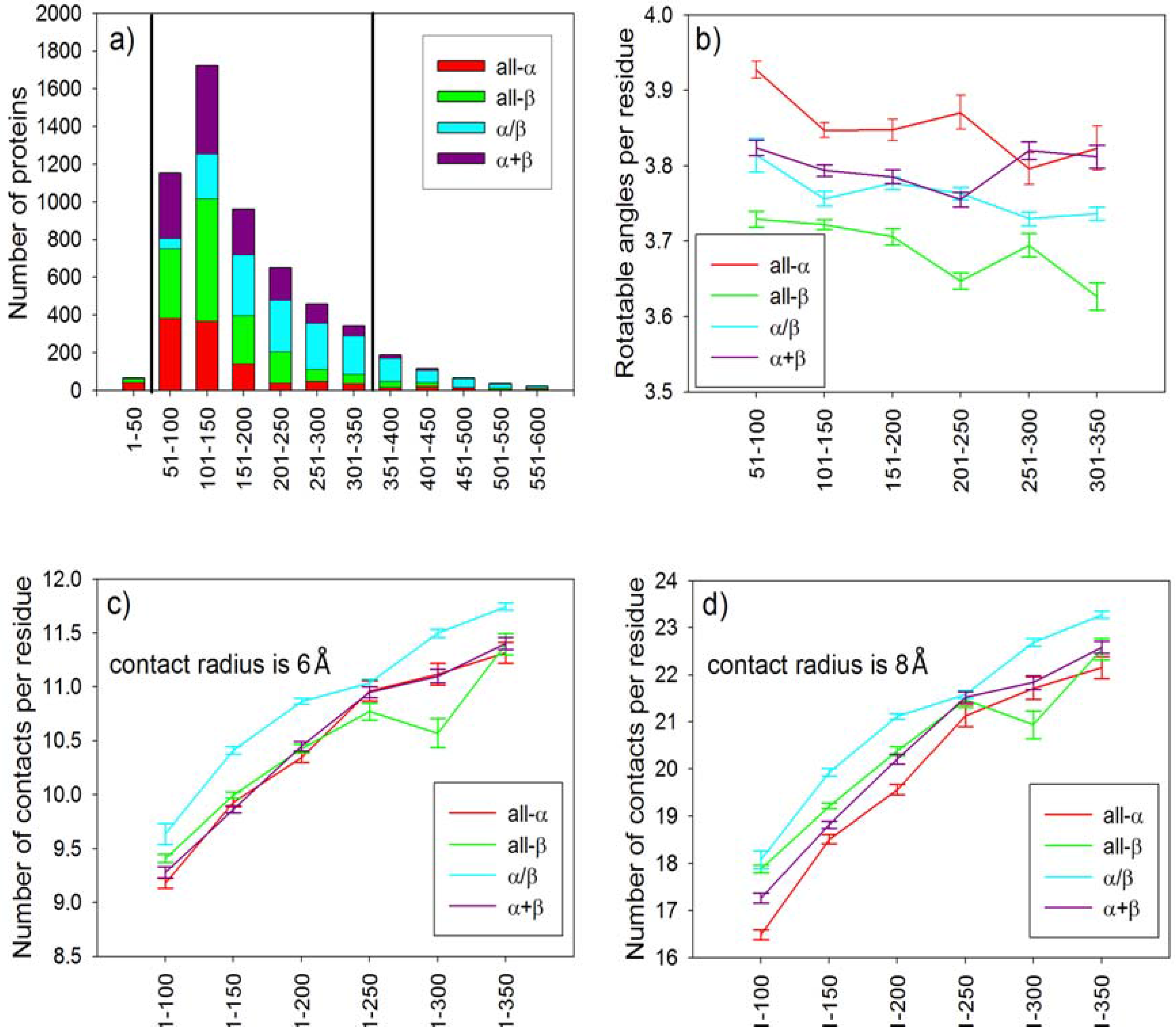
| SCOP fold | Name of protein | PDB entry | Degrees of freedom per residue | Number of contacts per residue | Cmod=ν/m (8 Å) | kf in water, s-1 |
|---|---|---|---|---|---|---|
| b.1 | Tenascin (short form) [43] | 1TEN | 3.74 | 18.52 | 0.202 | 3 |
| Fibronectin 10th FN3 module [44] | 1FNF | 3.36 | 18.43 | 0.182 | 244.7 | |
| b.34 | SH3 domain (PI3 kinase) [45] | 1PNJ | 3.91 | 16.53 | 0.236 | 0.3 |
| SH3 domain (c-src protein tyrosine kinase) [46] | 1SRL | 3.70 | 16.79 | 0.220 | 54.6 | |
| b.40 | CspA (Escherichia coli) [47] | 1MJC | 3.67 | 18.00 | 0.204 | 200.3 |
| CspB (Bacillus subtilis) [48,49] | 1CSP | 3.90 | 17.67 | 0.220 | 897.8 | |
| d.58 | HypF-N [25] | 1GXT | 3.82 | 19.35 | 0.197 | 81.5 |
| Spliceosomal protein U1A [50] | 1URN | 4.00 | 19.38 | 0.206 | 330.3 |
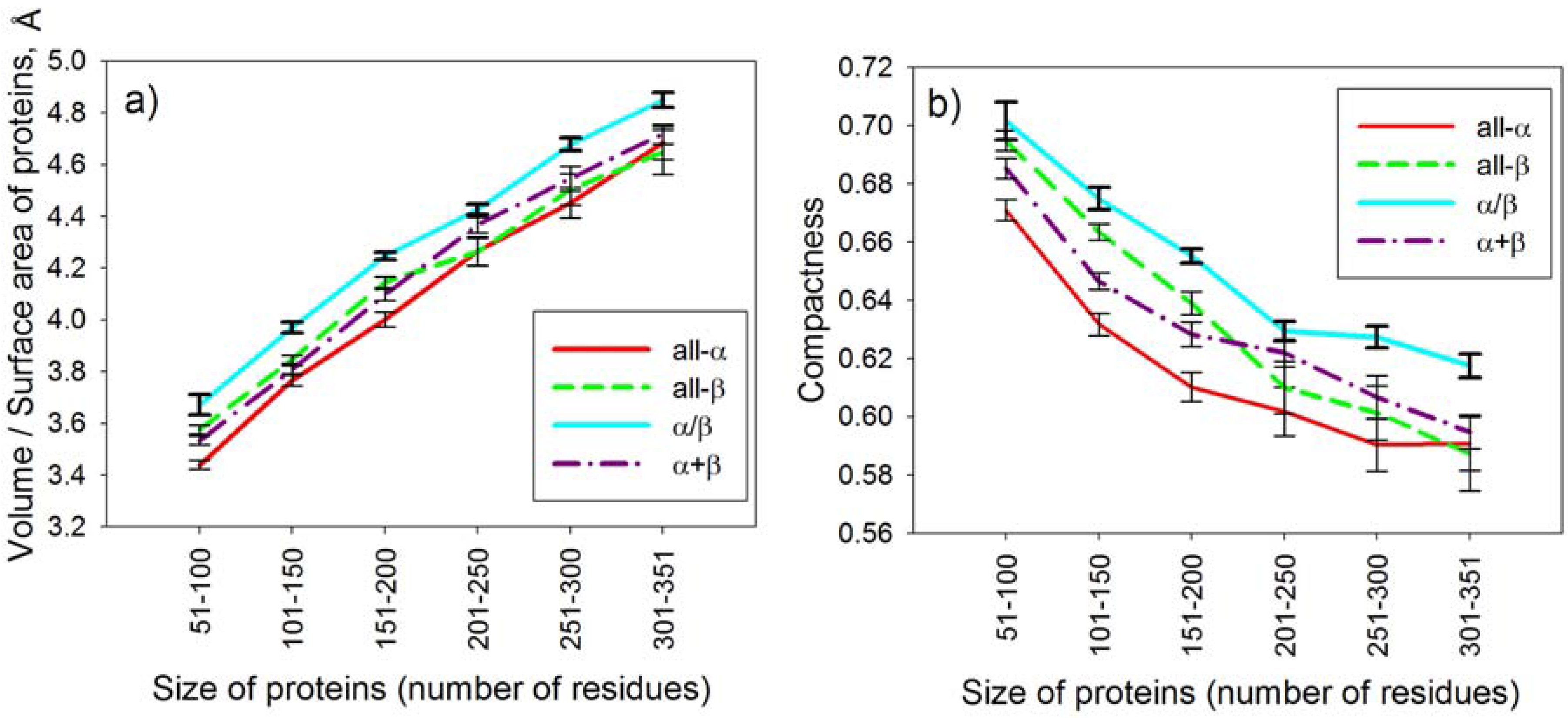
2.4. Statistical Analysis of Average Conformational Entropy and Average Number of Contacts per Residue for Different Classes of Proteins
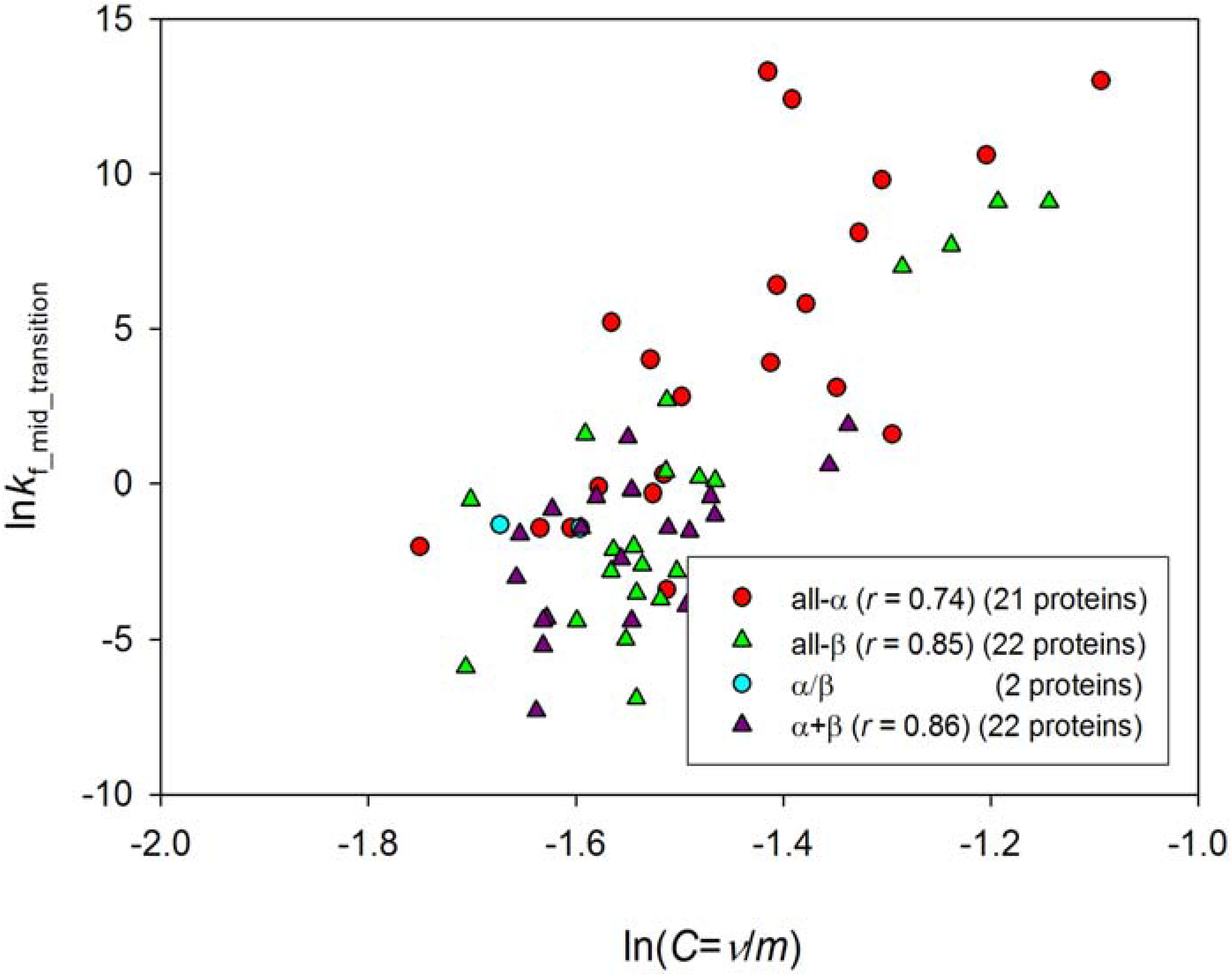
| ln kmt 67 proteins | ln kmta 21 proteins | ln kmtb 22 proteins | ln kmtd 22 proteins | |
|---|---|---|---|---|
| L | -0.69 ± 0.06 | -0.70 ± 0.11 | -0.76 ± 0.09 | -0.72 ± 0.10 |
| Ln(L) | -0.76 ± 0.05 | -0.73 ± 0.10 | -0.85 ± 0.06 | -0.82 ± 0.07 |
| L1/2 | -0.73 ± 0.06 | -0.72 ± 0.11 | -0.82 ± 0.07 | -0.77 ± 0.09 |
| L2/3 | -0.72 ± 0.06 | -0.72 ± 0.11 | -0.80 ± 0.08 | -0.75 ± 0.09 |
| AbsCO | -0.81 ± 0.04 | -0.76 ± 0.09 | -0.88 ± 0.05 | -0.74 ± 0.10 |
| RelativeCO | -0.12 ± 0.12 | 0.37 ± 0.19 | 0.44 ± 0.17 | -0.38 ± 0.18 |
| Normalized radius of gyration, Rg/Rg* [18] | 0.08 ± 0.12 | -0.34 ± 0.19 | 0.41 ± 0.18 | 0.32 ± 0.19 |
| Radius of cross-section,V/S | -0.53 ± 0.09 | -0.47 ± 0.17 | -0.55 ± 0.15 | -0.48 ± 0.16 |
| Ln(Entropy capacity) | 0.78 ± 0.05 | 0.74 ± 0.10 | 0.85 ± 0.06 | 0.86 ± 0.06 |
| Ln(t1/2), Monte-Carlo steps | -0.82 ± 0.04 | -0.74 ± 0.10 | -0.83 ± 0.07 | -0.84 ± 0.06 |
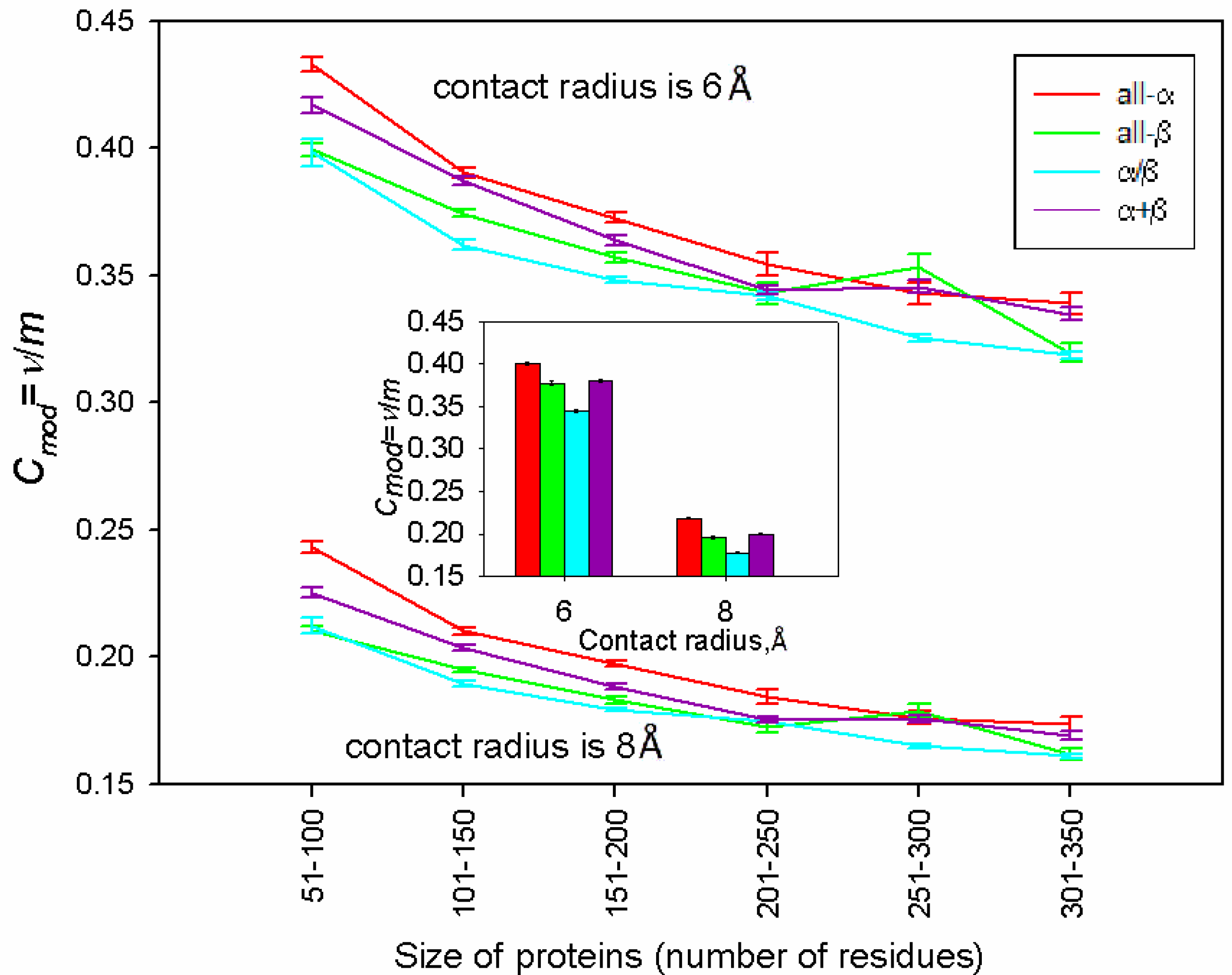
| Class a | Class b | Class c | Class d | |
|---|---|---|---|---|
| Class a | 1.3 × 10-25 | 8.4 × 10-109 | 5.3 × 10-27 | |
| Class b | 1.3 × 10-25 | 1.5 × 10-25 | 4.3 × 10-02 | |
| Class c | 8.4 × 10-109 | 1.5 × 10-25 | 2.1 × 10-64 | |
| Class d | 5.3 × 10-27 | 4.3 × 10-02 | 2.1 × 10-64 |
2.5. Behavior of Proteins with a High and Low Number of Contacts and Side-Chain Entropy
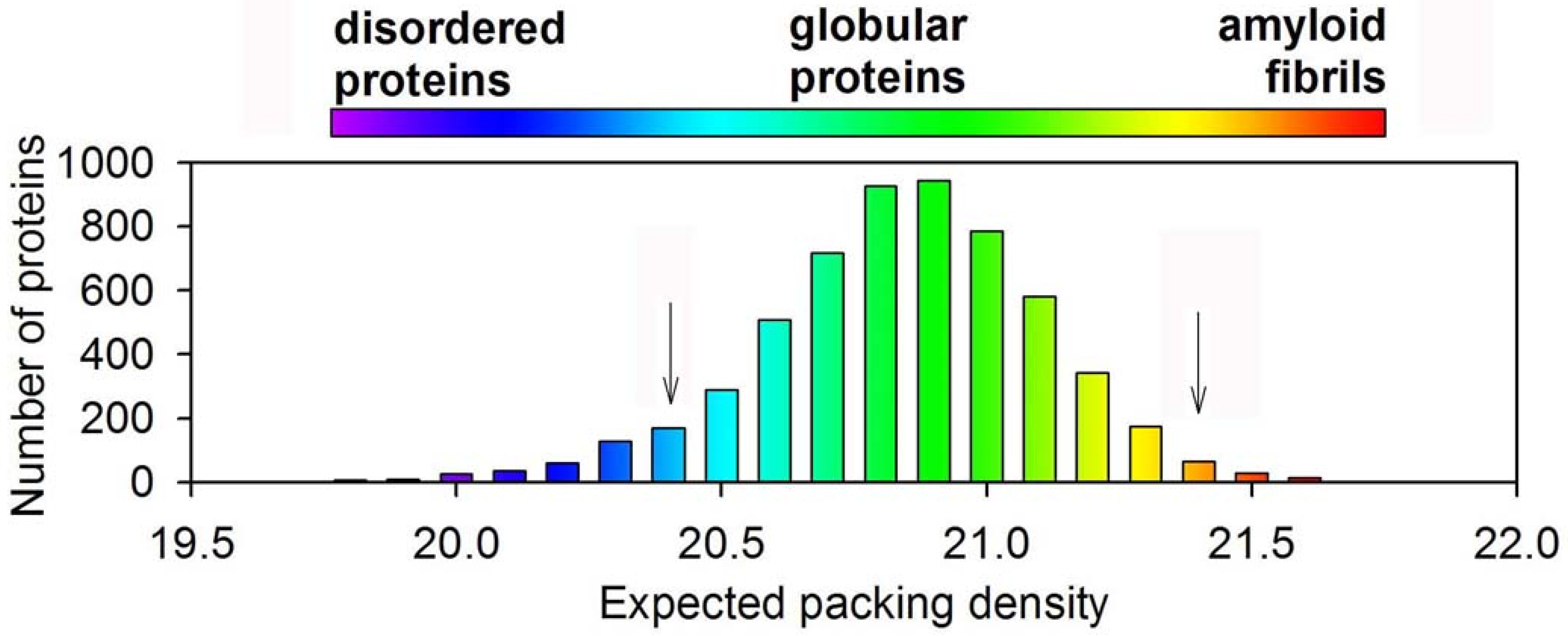
3. Conclusion
Acknowledgements
References and Notes
- Finkelstein, A.V.; Badretdinov, A.Ya. Rate of protein folding near the point of thermodynamic equilibrium between the coil and the most stable chain fold. Fold. Des. 1997, 2, 115–121. [Google Scholar] [CrossRef]
- Jackson, S.E. How do small single-domain proteins fold? Fold. Des. 1998, 3, R81–R91. [Google Scholar] [CrossRef]
- Plaxco, K.W.; Simons, K.W.; Baker, D. Contact order, transition state placement and the refolding rates of single domain proteins. J. Mol. Biol. 1998, 277, 985–994. [Google Scholar] [CrossRef] [PubMed]
- De Sancho, D.; Doshi, U.; Munoz, V. Protein folding rates and stability: How much is there beyond size? J. Am. Chem. Soc. 2009, 131, 2074–2075. [Google Scholar]
- Galzitskaya, O.V.; Garbuzynskiy, S.O.; Ivankov, D.N.; Finkelstein, A.V. Chain length is the main determinant of the folding rate for proteins with three-state folding kinetics. Proteins 2003, 51, 162–166. [Google Scholar] [CrossRef] [PubMed]
- Kuznetsov, I.B.; Rackovsky, S. Class-Specific correlations between protein folding rate, structure-derived, and sequence-derived descriptors. Proteins 2004, 54, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Finkelstein, A.V.; Galzitskaya, O.V. Physics of protein folding. Phys. Life Rev. 2004, 1, 23–56. [Google Scholar] [CrossRef]
- Guijarro, J.I.; Morton, C.J.; Plaxco, K.W.; Campbell, I.D.; Dobson, C.M. Folding kinetics of the SH3 domain of PI3 kinase by Real-Time NMR combined with optical spectroscopy. J. Mol. Biol. 1998, 276, 657–667. [Google Scholar] [CrossRef] [PubMed]
- Plaxco, K.W.; Guijarro, J.I.; Morton, C.J.; Pitkeathly, M.; Campbell, I.D.; Dobson, C.M. The folding kinetics and thermodynamics of the Fyn-SH3 domain. Biochemistry 1998, 37, 2529–2537. [Google Scholar] [CrossRef] [PubMed]
- Perl, D.; Welker, Ch.; Schindler, Th.; Schroder, K.; Marahiel, M.A.; Jaenicke, R.; Schmid, F.X. Conservation of rapid two-state folding in Mesophilic, Thermophilic and Hyperthermophilic cold shock proteins. Nature Struct. Biol. 1998, 5, 229–235. [Google Scholar] [CrossRef] [PubMed]
- Van Nuland, N.A.J.; Chiti, F.; Taddei, N.; Raugei, G.; Ramponi, G.; Dobson, C.M. Slow folding of muscle acylphosphatase in the absence of intermediates. J. Mol. Biol. 1998, 283, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Zerovnik, E.; Virden, R.; Jerala, R.; Turk, V.; Waltho, J.P. On the mechanism of human Stefin B folding: I. comparison to homologous Stefin A. influence of Ph and Trifluoroethanol on the fast and slow folding phases. Proteins 1998, 32, 296–303. [Google Scholar] [CrossRef]
- Thirumalai, D. From minimal models to real proteins: Time Scales for Protein Folding Kinetics. J. Phys. Orsay Fr. 1995, 5, 1457–1467. [Google Scholar] [CrossRef]
- Gutin, A.M.; Abkevich, V.I.; Shakhnovich, E.I. Chain length scaling of protein folding time. Phys. Rev. Lett. 1996, 77, 5433–5456. [Google Scholar] [CrossRef] [PubMed]
- Finkelstein, A.V.; Badretdinov, A.Ya. Physical reasons for fast folding of stable spatial structure of proteins: A solution of the Levinthal Paradox. Mol. Biol. (Russia) Engl. Edition 1997, 31, 391–398. [Google Scholar]
- Koga, N.; Takada, S. Roles of native topology and chain-length scaling in protein folding: A simulation study with a Gō-Like model. J. Mol. Biol. 2001, 313, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Galzitskaia, O.V.; Ivankov, D.N.; Finkelstein, A.V. Folding nuclei in proteins. FEBS Lett. 2001, 489, 113–118. [Google Scholar] [CrossRef]
- Ivankov, D.N.; Bogatyreva, N.S.; Lobanov, M.Yu.; Galzitskaya, O.V. Coupling between properties of the protein shape and the rate of protein folding. PLoS ONE 2009, 4, e6476. [Google Scholar] [CrossRef] [PubMed]
- Fersht, A.R. Transition-state structure as a unifying basis in protein-folding mechanisms: contact order, chain topology, stability, and the extended nucleus mechanism. Proc. Natl. Acad. Sci. USA 2000, 97, 1525–1529. [Google Scholar] [CrossRef] [PubMed]
- Ivankov, D.N.; Garbuzynskiy, S.O.; Alm, E.; Plaxco, K.W.; Baker, D.; Finkelstein, A.V. Contact order revisited: influence of protein size on the folding rate. Protein Sci. 2003, 12, 2057–2062. [Google Scholar] [CrossRef] [PubMed]
- Makarov, D.E.; Keller, C.A.; Plaxco, K.W.; Metiu, H. How the folding rate constant of simple, single-domain proteins depends on the number of native contacts. Proc. Natl. Acad. Sci. USA 2002, 9, 3535–3539. [Google Scholar] [CrossRef] [PubMed]
- Punta, M.; Rost, B. Protein folding rates estimated from contact predictions. J. Mol. Biol. 2005, 348, 507–512. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Isom, D.G.; Srinivasan, R.; Rose, G.D. Local secondary structure content predicts folding rates for simple, two-state proteins. J. Mol. Biol. 2003, 327, 1149–1154. [Google Scholar] [CrossRef]
- Shao, H.; Peng, Y.; Zeng, Z.-H. A simple parameter relating sequences with folding rates of small alpha helical proteins. Protein Pept. Lett. 2003, 10, 277–280. [Google Scholar] [CrossRef] [PubMed]
- Calloni, G.; Taddei, N.; Plaxco, K.W.; Ramponi, G.; Stefani, M.; Chiti, F. Comparison of the folding processes of distantly related proteins. Importance of hydrophobic content in folding. J. Mol. Biol. 2003, 330, 577–591. [Google Scholar] [CrossRef]
- Naganathan, A.N.; Munoz, V. Scaling of folding times with protein size. J. Am. Chem. Soc. 2005, 127, 480–481. [Google Scholar] [CrossRef] [PubMed]
- Galzitskaya, O.V.; Surin, A.K.; Nakamura, H. Optimal region of average side-chain entropy for fast protein folding. Protein Sci. 2000, 9, 580–586. [Google Scholar] [CrossRef] [PubMed]
- Murzin, A.G.; Brenner, S.E.; Hubbard, T.; Chothia, C. SCOP: A structural classification of proteins database for the investigation of sequences and structures. J. Mol. Biol. 1995, 247, 536–540. [Google Scholar] [CrossRef]
- Galzitskaya, O.V.; Garbuzynskiy, S.O. Entropy capacity determines protein folding. Proteins 2006, 63, 144–154. [Google Scholar] [CrossRef] [PubMed]
- Galzitskaya, O.V.; Danielle, C.; Reifsnyder, D.C.; Bogatyreva, N.S.; Ivankov, D.N.; Garbuzynskiy, S.O. More compact protein globules exhibit slower folding rates. Proteins 2008, 70, 329–332. [Google Scholar] [CrossRef] [PubMed]
- Galzitskaya, O.V.; Bogatyreva, N.S.; Ivankov, D.N. Compactness determines protein folding type. J. Bioinform. Comput. Biol. 2008, 6, 667–680. [Google Scholar] [CrossRef] [PubMed]
- Taketomi, H.; Ueda, Y.; Gō, N. Studies on protein folding, unfolding and fluctuations by computer simulation. I. The effect of specific amino acid sequence represented by specific inter-unit interactions. Int. J. Pept. Protein Res. 1975, 7, 445–459. [Google Scholar] [CrossRef] [PubMed]
- Lifshits, E.M.; Pitaevskii, L.P. Physical Kinetics; Pergamon: Oxford, UK, 1991. [Google Scholar]
- Pande, V.S.; Grosberg, A.Yu.; Tanaka, T.; Rokhsar, D.S. Pathways for protein folding: Is a new view needed? Curr. Opin. Struct. Biol. 1998, 8, 68–79. [Google Scholar] [CrossRef]
- Fersht, A.R. Characterizing transition states in protein folding: An essential step in the puzzle. Curr. Opin. Struct. Biol. 1995, 5, 79–84. [Google Scholar] [CrossRef]
- Fersht, A.R. Nucleation mechanisms in protein folding. Curr. Opin. Struct. Biol. 1997, 7, 3–9. [Google Scholar] [CrossRef]
- Matouschek, J.T.; Kellis, J.T.Jr.; Serrano, L.; Fersht, A.R. Mapping the transition state and pathway of protein folding by protein engineering. Nature 1989, 340, 122–126. [Google Scholar] [CrossRef] [PubMed]
- Matouschek, A.; Kellis, J.T.Jr.; Serrano, L.; Bycroft, M.; Fersht, A.R. Transient folding intermediates characterized by protein engineering. Nature 1990, 346, 440–445. [Google Scholar] [CrossRef] [PubMed]
- Fersht, A.R.; Matouschek, A.; Serrano, L. The folding of an enzyme. I. Theory of protein engineering analysis of stability and pathway of protein folding. J. Mol. Biol. 1992, 224, 771–782. [Google Scholar] [CrossRef]
- Abkevich, V.I.; Gutin, A.M.; Shakhnovich, E.I. Specific nucleus as a transition state for protein folding: an evidence from lattice model. Biochemistry 1994, 33, 10026–10036. [Google Scholar] [CrossRef] [PubMed]
- Wolynes, P.G. Folding funnels and energy landscapes of larger proteins within the capillarity approximation. Proc. Natl. Acad. Sci. USA 1997, 94, 6170–6175. [Google Scholar] [CrossRef] [PubMed]
- Creamer, T.P.; Rose, G.D. Side-chain entropy opposes alpha-Helix formation but rationalizes experimentally determined Helix-forming propensities. Proc. Natl. Acad. Sci. USA 1992, 89, 5937–5941. [Google Scholar] [CrossRef] [PubMed]
- Clarke, J.; Hamill, S.J.; Johnson, C.M. Folding and stability of a fibronectin type III domain of human tenascin. J. Mol. Biol. 1997, 270, 771–778. [Google Scholar] [CrossRef] [PubMed]
- Cota, E.; Clarke, J. Folding of beta-sandwich proteins: Three-state transition of a Fibronectin type III module. Protein Sci. 2000, 9, 112–120. [Google Scholar] [CrossRef] [PubMed]
- Guijarro, J.I.; Morton, C.J.; Plaxco, K.W.; Campbell, I.D.; Dobson, C.M. Folding kinetics of the SH3 domain of PI3 kinase by Real-Time NMR combined with optical spectroscopy. J. Mol. Biol. 1998, 276, 657–667. [Google Scholar] [CrossRef] [PubMed]
- Grantcharova, V.P.; Baker, D. Folding dynamics of the Src SH3 domain. Biochemistry 1997, 36, 15685–15692. [Google Scholar] [CrossRef] [PubMed]
- Reid, K.L.; Rodriguez, H.M.; Hillier, B.J.; Gregoret, L.M. Stability and folding properties of a model beta-sheet protein, Escherichia coli Cspa. Protein Sci. 1998, 7, 470–479. [Google Scholar] [CrossRef] [PubMed]
- Perl, D.; Welker, C.; Schindler, T.; Schroder, K.; Marahiel, M.A.; Jaenicke, R.; Schmid, F.X. Conservation of rapid two-state folding in Mesophilic, Thermophilic and Hyperthermophilic cold shock proteins. Nature Struct. Biol. 1998, 5, 229–235. [Google Scholar] [CrossRef] [PubMed]
- Schindler, T.; Herrler, M.; Marahiel, M.A.; Schmid, F.X. Extremely rapid protein folding in the absence of intermediates. Nature Struct. Biol. 1995, 2, 663–673. [Google Scholar] [CrossRef] [PubMed]
- Silow, M.; Oliveberg, M. High-energy channeling in protein folding. Biochemistry 1997, 36, 7633–7637. [Google Scholar] [CrossRef] [PubMed]
- Galzitskaya, O.V. Estimation of protein folding rate from Monte-Carlo simulations and entropy capacity. Curr. Protein Pept. Sci. In press.
- Winstanley, H.F.; Abeln, S.; Deane, C.M. How old is your fold? Bioinformatics 2005, 21, i449–i458. [Google Scholar] [CrossRef] [PubMed]
- Tiana, G.; Shakhnovich, B.E.; Dokholyan, N.V.; Shakhnovich, E.I. Imprint of evolution on protein structures. Proc. Natl. Acad. Sci. USA 2004, 101, 2846–2851. [Google Scholar] [CrossRef] [PubMed]
- Shakhnovich, B.E.; Deeds, E.; Delisi, C.; Shakhnovich, E. Protein structure and evolutionary history determine sequence space topology. Genome. Res. 2005, 15, 385–392. [Google Scholar] [CrossRef] [PubMed]
- England, J.L.; Shakhnovich, B.E.; Shakhnovich, E.I. Natural selection of more designable folds: A mechanism for thermophilic adaptation. Proc. Natl. Acad. Sci. USA 2003, 100, 8727–8731. [Google Scholar] [CrossRef] [PubMed]
- McIntosh, P.B.; Frenkiel, T.A.; Wollborn, U.; McCormick, J.E.; Klempnauer, K.H.; Feeney, J.; Carr, M.D. Solution structure of the B-Myb DNA-binding domain: A possible role for conformational instability of the protein in DNA binding and control of gene expression. Biochemistry 1998, 37, 9619–9629. [Google Scholar] [CrossRef] [PubMed]
- Ippel, H.; Larsson, G.; Behravan, G.; Zdunek, J.; Lundqvist, M.; Schleucher, J.; Lycksell, P.O.; Wijmenga, S. The solution structure of the homeodomain of the rat insulin-gene enhancer protein Isl-1. Comparison with other homeodomains. J. Mol. Biol. 1999, 288, 689–703. [Google Scholar] [CrossRef] [PubMed]
- Baker, D. Metastable states and folding free energy barriers. Nature Struct. Biol. 1998, 5, 1021–1024. [Google Scholar] [CrossRef] [PubMed]
- Sohl, J.L.; Jaswal, S.S.; Agard, D.A. Unfolded conformations of α-Lytic protease are more stable than its native state. Nature 1998, 395, 817–819. [Google Scholar] [PubMed]
- James, T.L.; Liu, H.; Ulyanov, N.B.; Farr-Jones, S.; Zhang, H.; Donne, D.G.; Kaneko, K.; Groth, D.; Mehlhorn, I.; Prusiner, S.B.; Cohen, F.E. Solution structure of a 142-residue recombinant prion protein corresponding to the infectious fragment of the scrapie isoform. Proc. Natl. Acad. Sci. USA 1997, 94, 10086–10091. [Google Scholar] [CrossRef] [PubMed]
- Taylor, K.L.; Cheng, N.; Williams, R.W.; Steven, A.C.; Wickner, R.B. Prion domain initiation of amyloid formation in vitro from native Ure2p. Science 1999, 283, 1339–1343. [Google Scholar] [CrossRef] [PubMed]
- Wright, P.E.; Dyson, H.J. Intrinsically unstructured proteins: Reassessing the protein structure–function paradigm. J. Mol. Biol. 1999, 293, 321–331. [Google Scholar] [CrossRef] [PubMed]
- Romero, P.; Obradovic, Z.; Dunker, A.K. Natively disordered proteins: Functions and predictions. Appl. Bioinformatics 2004, 3, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Fink, A.L. Natively unfolded proteins. Curr. Opin. Struct. Biol. 2005, 15, 35–41. [Google Scholar] [CrossRef] [PubMed]
- Dunker, A.K.; Cortese, M.S.; Romero, P.; Iakoucheva, L.M.; Uversky, V.N. Flexible Nets. The roles of intrinsic disorder in protein interaction networks. FEBS J. 2005, 272, 5129–5148. [Google Scholar] [CrossRef] [PubMed]
- Galzitskaya, O.V.; Garbuzynskiy, S.O.; Lobanov, M.Y. Prediction of natively unfolded regions in protein chain. Mol. Biol. (Moscow) 2006, 40, 341–348. [Google Scholar] [CrossRef]
- Galzitskaya, O.V.; Garbuzynskiy, S.O.; Lobanov, M.Y. FoldUnfold: web server for the prediction of disordered regions in protein chain. Bioinformatics 2006, 22, 2948–2949. [Google Scholar] [CrossRef] [PubMed]
- Dyson, H.J.; Wright, P.E. Intrinsically unstructured proteins and their functions. Nat. Rev. Mol. Cell. Biol. 2005, 6, 197–208. [Google Scholar] [CrossRef] [PubMed]
- Ptitsyn, O.B. Molten globule and protein folding. Adv. Protein Chem. 1995, 47, 83–229. [Google Scholar] [PubMed]
- Chiti, F.; Webster, P.; Taddei, N.; Clark, A.; Stefani, M.; Ramponi, G.; Dobson, C.M. Designing conditions for in vitro formation of amyloid protofilaments and fibrils. Proc. Natl. Acad. Sci. USA 1999, 96, 3590–3594. [Google Scholar] [CrossRef] [PubMed]
- Harrison, P.M.; Chan, H.S.; Prusiner, S.B.; Cohen, F.E. Thermodynamics of model prions and its implications for the problem of prion protein folding. J. Mol. Biol. 1999, 286, 593–606. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, J.L.; Guijarro, J.I.; Orlova, E.; Zurdo, J.; Dobson, C.M.; Sunde, M.; Saibil, H.R. Cryo-electron microscopy structure of an SH3 amyloid fibril and model of the molecular packing. EMBO J. 1999, 18, 815–821. [Google Scholar] [CrossRef] [PubMed]
- Galzitskaya, O.V.; Garbuzynskiy, S.O.; Lobanov, M.Y. Prediction of amyloidogenic and disordered regions in protein chains. PLoS Comput. Biol. 2006, 2, e177. [Google Scholar] [CrossRef] [PubMed]
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Galzitskaya, O.V. Influence of Conformational Entropy on the Protein Folding Rate. Entropy 2010, 12, 961-982. https://doi.org/10.3390/e12040961
Galzitskaya OV. Influence of Conformational Entropy on the Protein Folding Rate. Entropy. 2010; 12(4):961-982. https://doi.org/10.3390/e12040961
Chicago/Turabian StyleGalzitskaya, Oxana V. 2010. "Influence of Conformational Entropy on the Protein Folding Rate" Entropy 12, no. 4: 961-982. https://doi.org/10.3390/e12040961
APA StyleGalzitskaya, O. V. (2010). Influence of Conformational Entropy on the Protein Folding Rate. Entropy, 12(4), 961-982. https://doi.org/10.3390/e12040961





