Proteomic Determination of Low-Molecular-Weight Glutenin Subunit Composition in Aroona Near-Isogenic Lines and Standard Wheat Cultivars
Abstract
:1. Introduction
2. Results
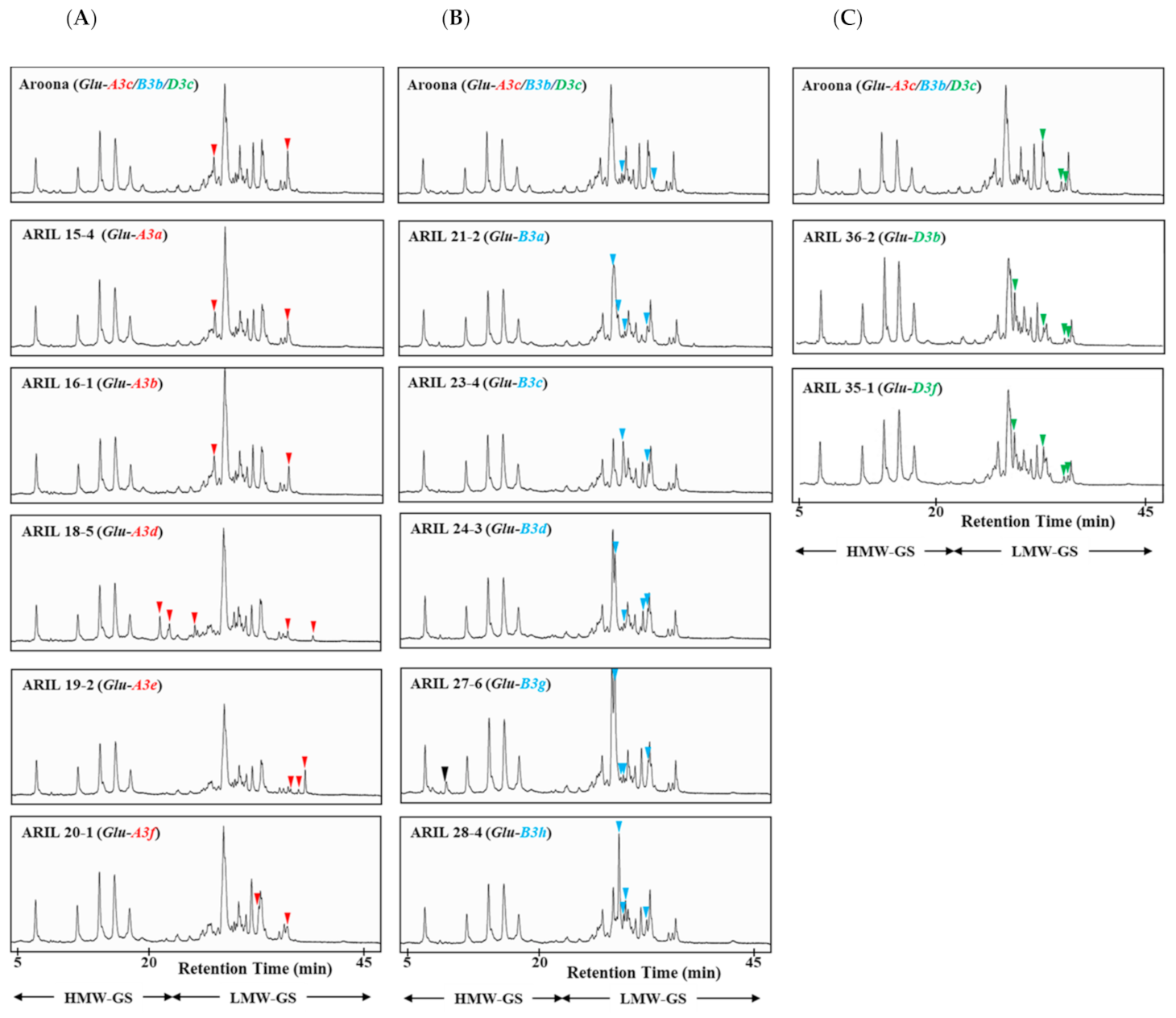
2.1. Identification of LMW-GS Alleles Using RP-HPLC
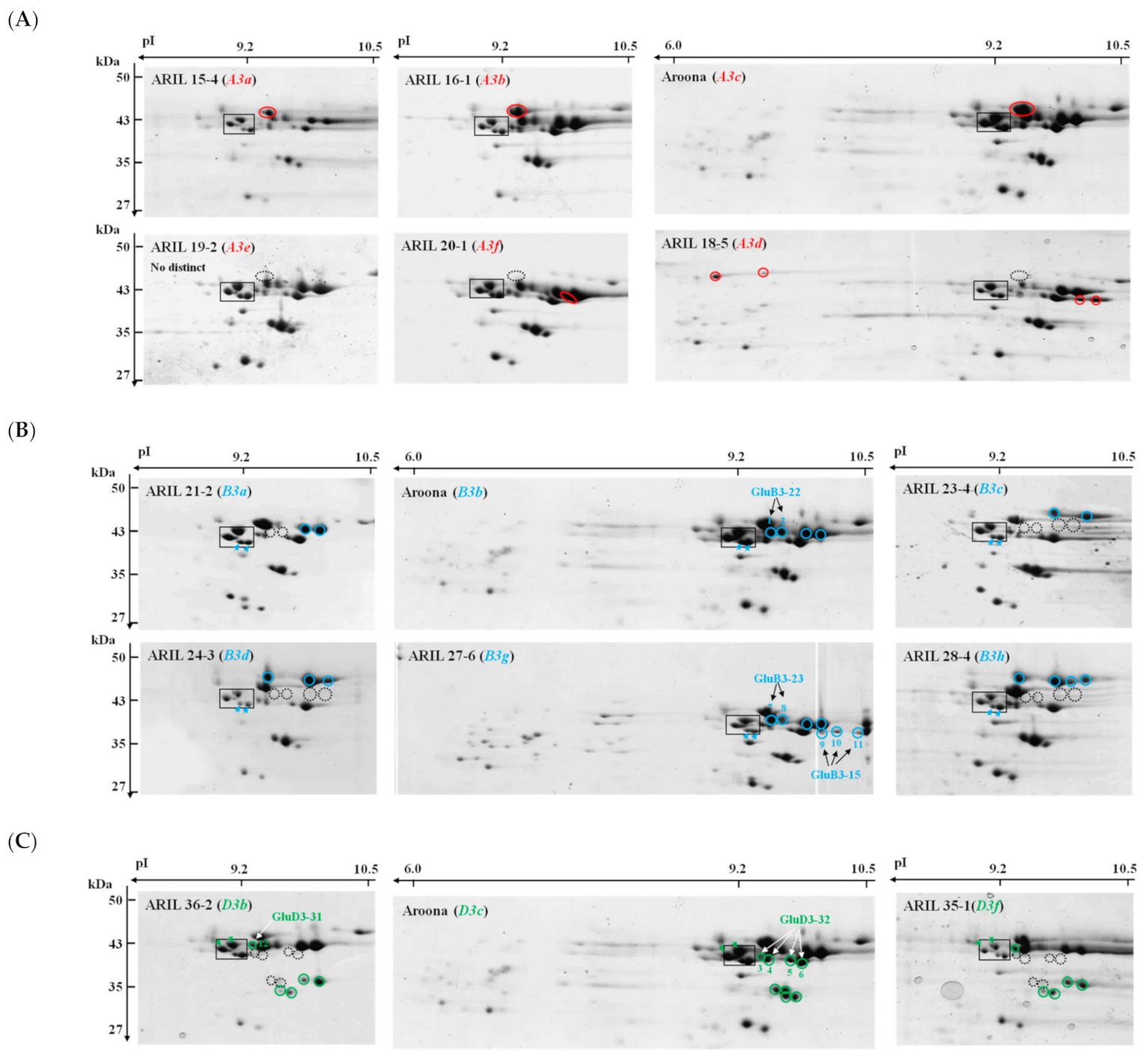
2.2. Comparison of LMW-GSs among ‘Aroona’ and Its Near-Isogenic Lines Using 2-DGE
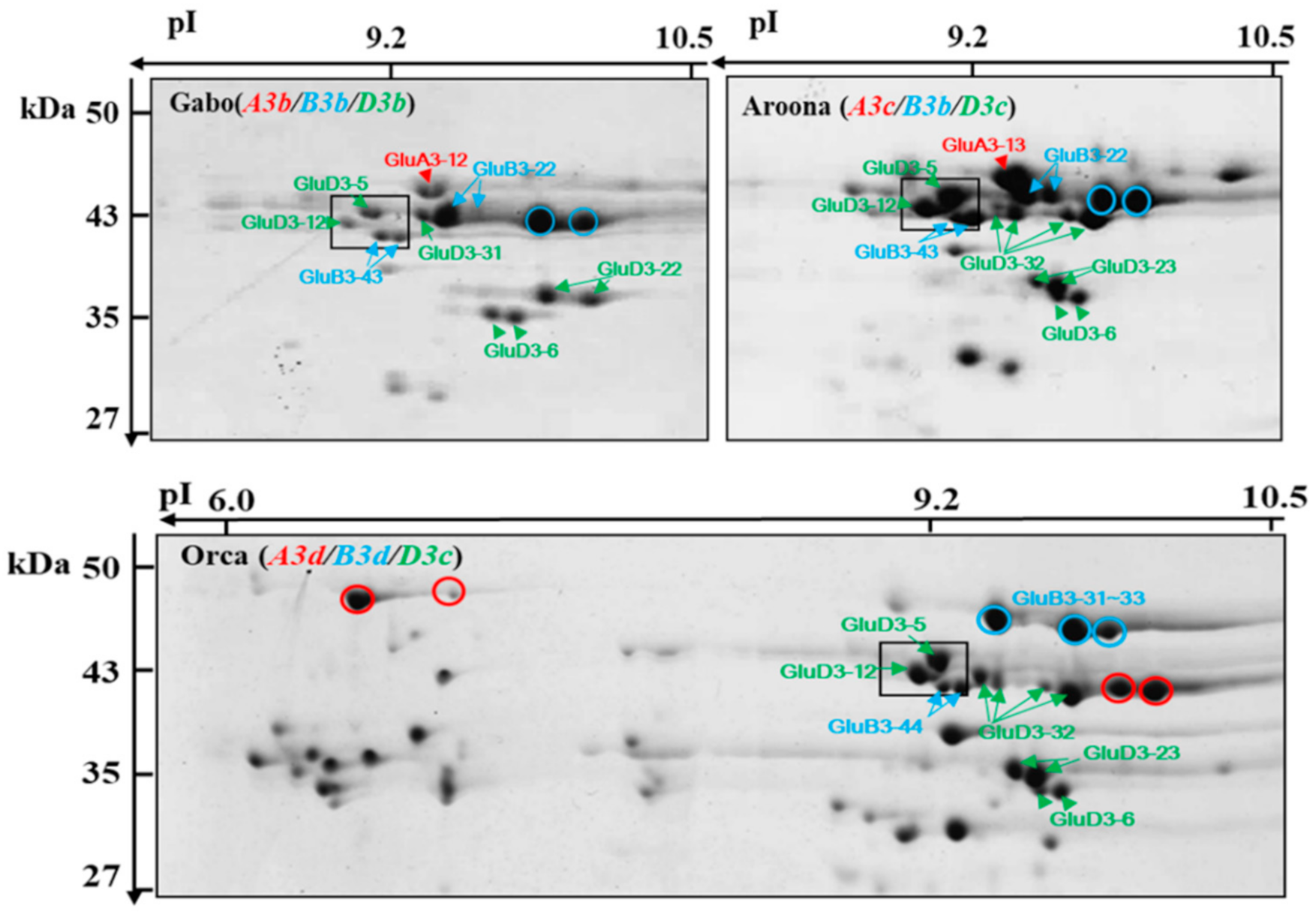
2.3. Identification of LMW-GSs Using LC–MS/MS Analysis
3. Discussion
3.1. Glu-A3 Alleles
3.2. Glu-B3 Alleles
3.3. Glu-D3 Alleles
4. Materials and Methods
4.1. Plant Materials
4.2. Glutenin Extraction
4.3. Two-Dimensional Gel Electrophoresis (2-DGE)
4.4. Identification of LMW-GSs Using UPLC–MS/MS
4.5. Separation of LMW-GSs Using RP-HPLC
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Shewry, P.R.; Tatham, A.S.; Barro, F.; Barcelo, P.; Lazzeri, P. Biotechnology of breadmaking: Unraveling and manipulating the multi-protein gluten complex. Biotechnology 1995, 13, 1185–1190. [Google Scholar] [CrossRef]
- Weegels, P.L.; Pijpekamp, A.M.; Graveland, A.; Hamer, R.J.; Schofield, J.D. Depolymerisation and re-polymerisation of wheat glutenin during dough processing. 1. Relationships between glutenin macropolymer content and quality parameters. J. Cereal Sci. 1996, 23, 103–111. [Google Scholar] [CrossRef]
- Payne, P.I. Genetics of wheat storage proteins and the effect of allelic variation on bread-making quality. Annu. Rev. Plant Physiol. 1987, 38, 141–153. [Google Scholar] [CrossRef]
- Wieser, H. Chemistry of gluten proteins. Food Microbiol. 2007, 24, 115–119. [Google Scholar] [CrossRef] [PubMed]
- Wrigley, C.W. Giant proteins with flour power. Nature 1996, 381, 738–739. [Google Scholar] [CrossRef] [PubMed]
- Payne, P.I.; Law, C.N.; Mudd, E.E. Control by homoeologous group 1 chromosomes of the high-molecular-weight subunits of glutenin, a major protein of wheat endosperm. Theor. Appl. Genet. 1980, 58, 113–120. [Google Scholar] [CrossRef]
- Jackson, E.A.; Morel, M.H.; Sontag Strohm, T.; Branlard, G.; Metakovsky, E.; Redaelli, R. Proposal for combining the classification systems of alleles of Gli-1 and Glu-3 loci in bread wheat (Triticum aestivum L.). J. Genet. Breed. 1996, 50, 321–336. [Google Scholar]
- Nagamine, T.; Kai, Y.; Takayama, T.; Yanagisawa, T.; Taya, S. Allelic variation at the Glu-1 and Glu-3 loci in southern Japanese wheats, and its effects on gluten properties. J. Cereal Sci. 2000, 32, 129–135. [Google Scholar] [CrossRef]
- D’Ovidio, R.; Masci, S. The low-molecular-weight glutenin subunits of wheat gluten. J. Cereal Sci. 2004, 39, 321–339. [Google Scholar] [CrossRef]
- Gupta, R.B.; Singh, N.K.; Shepherd, K. The cumulative effect of allelic variation in LMW and HMW glutenin subunits on dough properties in the progeny of two bread wheats. Theor. Appl. Genet. 1989, 77, 57–64. [Google Scholar] [CrossRef]
- Cornish, G.B.; Bekes, F.; Allen, H.; Martin, D. Flour proteins linked to quality traits in an Australian doubled haploid wheat population. Aust. J. Agric. Res. 2001, 52, 1339–1348. [Google Scholar] [CrossRef]
- Gianibelli, M.; Larroque, O.; MacRitchie, F.; Wrigley, C. Biochemical, genetic, and molecular characterization of wheat endosperm proteins. Cereal Chem. 2001, 78, 635–646. [Google Scholar] [CrossRef]
- Wieser, H.; Kieffer, R. Correlations of the amount of gluten protein types to the technological properties of wheat flours determined on a micro-scale. J. Cereal Sci. 2001, 34, 19–27. [Google Scholar] [CrossRef]
- Shewry, P.R.; Halford, N.G.; Tatham, A.S.; Popineau, Y.; Lafiandra, D.; Belton, P.S. The high molecular weightsubunitsofwheatgluteninandtheirroleindetermining wheat processing properties. Adv. Food Nutr. Res. 2003, 45, 219–302. [Google Scholar]
- Payne, P.; Holt, L.; Jarvis, M.; Jackson, E. Two-dimensional fractionation of the endosperm proteins of bread wheat. Cereal Chem. 1985, 62, 319–326. [Google Scholar]
- Masci, S.; Rovelli, L.; Kasarda, D.; Vensel, W.; Lafiandra, D. Characterisation and chromosomal localisation of C-type low-molecular-weight glutenin subunits in the bread wheat cultivar Chinese Spring. Theor. Appl. Genet. 2002, 104, 422–428. [Google Scholar] [CrossRef]
- Ruiz, M.; Carrillo, J. Linkage relationships between prolamin genes on chromosomes 1A and 1B of durum wheat. Theor. Appl. Genet. 1993, 87, 353–360. [Google Scholar] [CrossRef]
- Liu, C.-Y.; Shepherd, K. Inheritance of B subunits of glutenin and ω-and γ-gliadins in tetraploid wheats. Theor. Appl. Genet. 1995, 90, 1149–1157. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Jin, H.; Zhang, Y.; Liu, D.; Li, G.; Xia, X.; He, Z.; Zhang, A. Composition and functional analysis of low-molecular-weight glutenin alleles with Aroona near-isogenic lines of bread wheat. BMC Plant. Biol. 2012, 12, 243. [Google Scholar] [CrossRef] [Green Version]
- Dong, K.; Hao, C.; Wang, A.; Cai, M.; Yan, Y. Characterization of HMW glutenin subunits in bread and tetraploid wheats by reversed-phase high-performance liquid chromatography. Cereal Res. Commun. 2009, 37, 65–73. [Google Scholar] [CrossRef]
- Gao, L.; Ma, W.; Chen, J.; Wang, K.; Li, J.; Wang, S.; Bekes, F.; Appels, R.; Tan, Y. Characterization and comparative analysis of wheat high molecular weight glutenin subunits by SDS-PAGE, RP-HPLC, HPCE, and MALDI-TOF-MS. J. Agric. Food Chem. 2010, 58, 2777–2786. [Google Scholar] [CrossRef]
- Liu, L.; Ikeda, T.M.; Branlard, G.; Peña, R.J.; Rogers, W.J.; Lerner, S.E.; Kolman, M.A.; Xia, X.; Wang, L.; Ma, W.; et al. Comparison of low molecular weight glutenin subunits identified by SDS-PAGE, 2-DE, MALDI-TOF-MS and PCR in common wheat. BMC Plant. Biol. 2010, 10, 124. [Google Scholar] [CrossRef] [Green Version]
- Lee, J.Y.; Jang, Y.R.; Beom, H.R.; Altenbach, S.B.; Lim, S.H.; Lee, C.K. Allelic analysis of low molecular weight glutenin subunits using 2-DGE in Korean wheat cultivars. Breed. Sci. 2017, 67, 398–407. [Google Scholar] [CrossRef] [Green Version]
- Yu, Z.; Han, C.; Yan, X.; Li, X.; Jiang, G.; Yan, Y. Rapid characterization of wheat low molecular weight glutenin subunits by ultraperformance liquid chromatography (UPLC). J. Agric. Food Chem. 2013, 61, 4026–4034. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.B.; Shepherd, K. Two-step one-dimensional SDS-PAGE analysis of LMW subunits of glutelin. Theor. Appl. Genet. 1990, 80, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Dong, L.; Zhang, X.; Liu, D.; Fan, H.; Sun, J.; Zhang, Z.; Qin, H.; Li, B.; Hao, S.; Li, Z.; et al. New insights into the organization, recombination, expression and functional mechanism of low molecular weight glutenin subunit genes in bread wheat. PLoS ONE 2010, 5, e13548. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beom, H.-R.; Kim, J.S.; Jang, Y.-R.; Lim, S.-H.; Kim, C.-K.; Lee, C.K.; Lee, J.-Y. Proteomic analysis of low-molecular-weight glutenin subunits and relationship with their genes in a common wheat variety. 3 Biotech. 2018, 8, 56. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Beom, H.R.; Altenbach, S.B.; Lim, S.H.; Kim, Y.T.; Kang, C.S.; Yoon, U.-H.; Gupta, R.; Kim, S.-T.; Anh, S.-N.; et al. Comprehensive identification of LMW-GS genes and their protein products in a common wheat variety. Funct. Integr. Genom. 2016, 16, 269–279. [Google Scholar] [CrossRef]
- Luo, C.; Griffin, W.; Branlard, G.; McNeil, D. Comparison of low-and high molecular-weight wheat glutenin allele effects on flour quality. Theor. Appl. Genet. 2001, 102, 1088–1098. [Google Scholar] [CrossRef]
- Branlard, G.; Dardevet, M.; Saccomano, R.; Lagoutte, F.; Gourdon, J. Genetic diversity of wheat storage proteins and bread wheat quality. Euphytica 2001, 119, 59–67. [Google Scholar] [CrossRef]
- Rasheed, A.; Xia, X.; Yan, Y.; Appels, R.; Mahmood, T.; He, Z. Wheat seed storage proteins: Advances in molecular genetics, diversity and breeding applications. J. Cereal Sci. 2014, 60, 11–24. [Google Scholar] [CrossRef] [Green Version]
- Jin, H.; Zhang, Y.; Li, G.; Mu, P.; Fan, Z.; Xia, X.; He, Z. Effects of allelic variation of HMW-GS and LMW-GS on mixograph properties and Chinese noodle and steamed bread qualities in a set of Aroona near-isogenic wheat lines. J. Cereal Sci. 2013, 57, 146–152. [Google Scholar] [CrossRef]
- Jang, Y.R.; Beom, H.R.; Altenbach, S.B.; Lee, M.K.; Lim, S.H.; Lee, J.Y. Improved Method for Reliable HMW-GS Identification by RP-HPLC and SDS-PAGE in Common Wheat Cultivars. Molecules 2017, 22, 1055. [Google Scholar] [CrossRef]
- Wang, L.H.; Zhao, X.L.; He, Z.H.; Ma, W.; Appels, R.; Pena, R.J.; Xia, X.C. Characterization of low-molecular-weight glutenin subunit Glu-B3 genes and development of STS markers in common wheat (Triticum aestivum L.). Theor. Appl. Genet. 2009, 118, 525–539. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.L.; Xia, X.C.; He, Z.H.; Lei, Z.S.; Appels, R.; Yang, Y.; Sun, Y.Y.; Ma, W. Novel DNA variations to characterize low molecular weight glutenin Glu-D3 genes and develop STS markers in common wheat. Theor. Appl. Genet. 2007, 114, 451–460. [Google Scholar] [CrossRef]
- Zhang, X.; Liu, D.; Zhang, J.; Jiang, W.; Luo, G.; Yang, W.; Sun, J.; Tong, Y.; Cui, D.; Zhang, A. Novel insights into the composition, variation, organization, and expression of the low-molecular-weight glutenin subunit gene family in common wheat. J. Exp. Bot. 2013, 64, 2027–2040. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.H.; Li, G.Y.; Pena, R.J.; Xia, X.C.; He, Z.H. Development of STS markers and establishment of multiplex PCR for Glu-A3 alleles in common wheat (Triticum aestivum L.). J. Cereal Sci. 2010, 51, 305–312. [Google Scholar] [CrossRef]
- Zhen, S.; Han, C.; Ma, C.; Gu, A.; Zhang, M.; Shen, X.; Li, X.; Yan, Y. Deletion of the low-molecular-weight glutenin subunit allele Glu-A3a of wheat (Triticum aestivum L.) significantly reduces dough strength and breadmaking quality. BMC Plant Biol. 2014, 14, 367. [Google Scholar] [CrossRef] [Green Version]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Altenbach, S.B.; Chang, H.-C.; Simon-Buss, A.; Mohr, T.; Huo, N.; Gu, Y.Q. Exploiting the reference genome sequence of hexaploid wheat: A proteomic study of flour proteins from the cultivar Chinese Spring. Funct. Integr. Genom. 2020, 20, 1–16. [Google Scholar] [CrossRef] [Green Version]
| Line | Glu-A3 | Glu-B3 | Glu-D3 | Donor Parent |
|---|---|---|---|---|
| Aroona | c | b | c | Aroona |
| ARIL 15-4 | a | b | c | Chinese spring |
| ARIL 16-1 | b | b | c | Gabo |
| ARIL 18-5 | d | b | c | Orca |
| ARIL 19-2 | e | b | c | Lerma Rojo |
| ARIL 20-1 | f | b | c | Bungulla |
| ARIL 21-2 | c | a | c | Chinese spring |
| ARIL 23-4 | c | c | c | Halberd |
| ARIL 24-3 | c | d | c | Orca |
| ARIL 27-6 | c | g | c | Millewa |
| ARIL 28-4 | c | h | c | Sonalika |
| ARIL 36-2 | c | b | b | Bungulla |
| ARIL 35-1 | c | b | f | India 115 |
| # Spot | Cultivars | Gene | MS/MS Identification | a Gene Haplotype | N-Terminal Sequence | # A.A | b Putative Corresponding Genes | |
|---|---|---|---|---|---|---|---|---|
| Accession No. (Identity) | Gene | |||||||
| 1 | AROONA | Glu-B3b | ACA63873 | GluB3-22 | MENSHIP | 349 | EU369721 (100%) | B3-621a |
| 2 | AROONA | Glu-B3b | ACA63873 | GluB3-22 | MENSHIP | 349 | EU369721 (100%) | B3-621a |
| 3 | AROONA | Glu-D3c | ABC84367 | GluD3-32 | IENSHIP | 334 | FJ755316 (100%) | D3-578a |
| 4 | AROONA | Glu-D3c | ABC84367 | GluD3-32 | IENSHIP | 334 | FJ755316 (100%) | D3-578a |
| 5 | AROONA | Glu-D3c | ABC84367 | GluD3-32 | IENSHIP | 334 | FJ755316 (100%) | D3-578a |
| 6 | AROONA | Glu-D3c | ABC84367 | GluD3-32 | IENSHIP | 334 | FJ755316 (100%) | D3-578a |
| 7 | ARIL 27-6 | Glu-B3g | ACA63857 | GluB3-23 | MENSHIP | 349 | EU369705 (100%) | B3-621b |
| 8 | ARIL 27-6 | Glu-B3g | ACA63857 | GluB3-23 | MENSHIP | 349 | EU369705 (100%) | B3-621b |
| 9 | ARIL 27-6 | Glu-B3g | ABY58126 | GluB3-15 | MENSHIP | 323 | EU369703 (100%) | B3-544 |
| 10 | ARIL 27-6 | Glu-B3g | ABY58126 | GluB3-15 | MENSHIP | 323 | EU369703 (100%) | B3-544 |
| 11 | ARIL 27-6 | Glu-B3g | ACZ59817 | - | MENSHIP | 324 | EU369703 (100%) | B3-544 |
| 12 | ARIL 36-2 | Glu-D3b | ABC84366 | GluD3-31 | MENSHIP | 344 | JX878006 (100%) | D3-578b |
| Line | Alleles | a Active LMW-GS Genes (b Gene Name in Chinese Wheat Germplasm) | |||||
|---|---|---|---|---|---|---|---|
| At Glu-3A | |||||||
| ARIL15-4 | Glu-A3a | GluA3-11 (A3-620) | |||||
| ARIL16-1 | Glu-A3b | GluA3-12 (A3-643) | |||||
| Aroona | Glu-A3c | GluA3-13 (A3-620) | |||||
| ARIL18-5 | Glu-A3d | GluA3-23 (A3-402) | - (A3-568) | GluA3-4 (A3-662) | |||
| ARIL19-2 | Glu-A3e | GluA3-15/17 (A3-646) | |||||
| ARIL20-1 | Glu-A3f | GluA3-16 (A-573) | |||||
| At Glu-3B | |||||||
| ARIL21-2 | Glu-B3a | GluB3-44 (B3-530a) | GluB3-11 (B3-593) | GluB3-21 (B3-624) | |||
| Aroona | Glu-B3b | GluB3-43 (B3-530b) | GluB3-12 (B3-607) | GluB3-22 (B3-621a) | |||
| ARIL23-4 | Glu-B3c | GluB3-44 (B3-530a) | GluB3-34 (B3-688a) | ||||
| ARIL24-3 | Glu-B3d | GluB3-44 (B3-530a) | GluB3-31–33 (B3-688b) | ||||
| ARIL27-6 | Glu-B3g | GluB3-41 (B3-530c) | GluB3-15 (B3-544) | GluB3-23 (B3-621b) | |||
| ARIL28-4 | Glu-B3h | GluB3-43 (B3-530b) | GluB3-61–64 (B3-688c) | ||||
| At Glu-3D | |||||||
| ARIL36-2 | Glu-D3b | GluD3-6 (D3-385) | GluD3-4 (D3-394) | GluD3-22 (D3-441) | GluD3-12 (D3-528) | GluD3-5 (D3-575) | GluD3-31 (D3-578b) |
| Aroona | Glu-D3c | GluD3-6 (D3-385) | GluD3-4 (D3-394) | GluD3-23 (D3-432) | GluD3-12 (D3-528) | GluD3-5 (D3-575) | GluD3-32 (D3-578a) |
| ARIL35-1 | Glu-D3f | GluD3-6 (D3-385) | GluD3-4 (D3-394) | GluD3-22 (D3-441) | GluD3-12 (D3-528) | GluD3-5 (D3-575) | GluD3-31 (D3-578b) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cho, K.; Jang, Y.-R.; Lim, S.-H.; Altenbach, S.B.; Gu, Y.Q.; Simon-Buss, A.; Lee, J.-Y. Proteomic Determination of Low-Molecular-Weight Glutenin Subunit Composition in Aroona Near-Isogenic Lines and Standard Wheat Cultivars. Int. J. Mol. Sci. 2021, 22, 7709. https://doi.org/10.3390/ijms22147709
Cho K, Jang Y-R, Lim S-H, Altenbach SB, Gu YQ, Simon-Buss A, Lee J-Y. Proteomic Determination of Low-Molecular-Weight Glutenin Subunit Composition in Aroona Near-Isogenic Lines and Standard Wheat Cultivars. International Journal of Molecular Sciences. 2021; 22(14):7709. https://doi.org/10.3390/ijms22147709
Chicago/Turabian StyleCho, Kyoungwon, You-Ran Jang, Sun-Hyung Lim, Susan B. Altenbach, Yong Q. Gu, Annamaria Simon-Buss, and Jong-Yeol Lee. 2021. "Proteomic Determination of Low-Molecular-Weight Glutenin Subunit Composition in Aroona Near-Isogenic Lines and Standard Wheat Cultivars" International Journal of Molecular Sciences 22, no. 14: 7709. https://doi.org/10.3390/ijms22147709






