HDACs and Their Inhibitors on Post-Translational Modifications: The Regulation of Cardiovascular Disease
Abstract
1. Introduction
1.1. Cardiovascular Disease
1.2. Post-Translational Modifications of Proteins
1.3. Histone Deacetylases (HDACs)
1.4. Histone Deacetylase Inhibitors (HDACis)
2. HDACs and Post-Translational Modifications Are Closely Associated with Cardiovascular Diseases
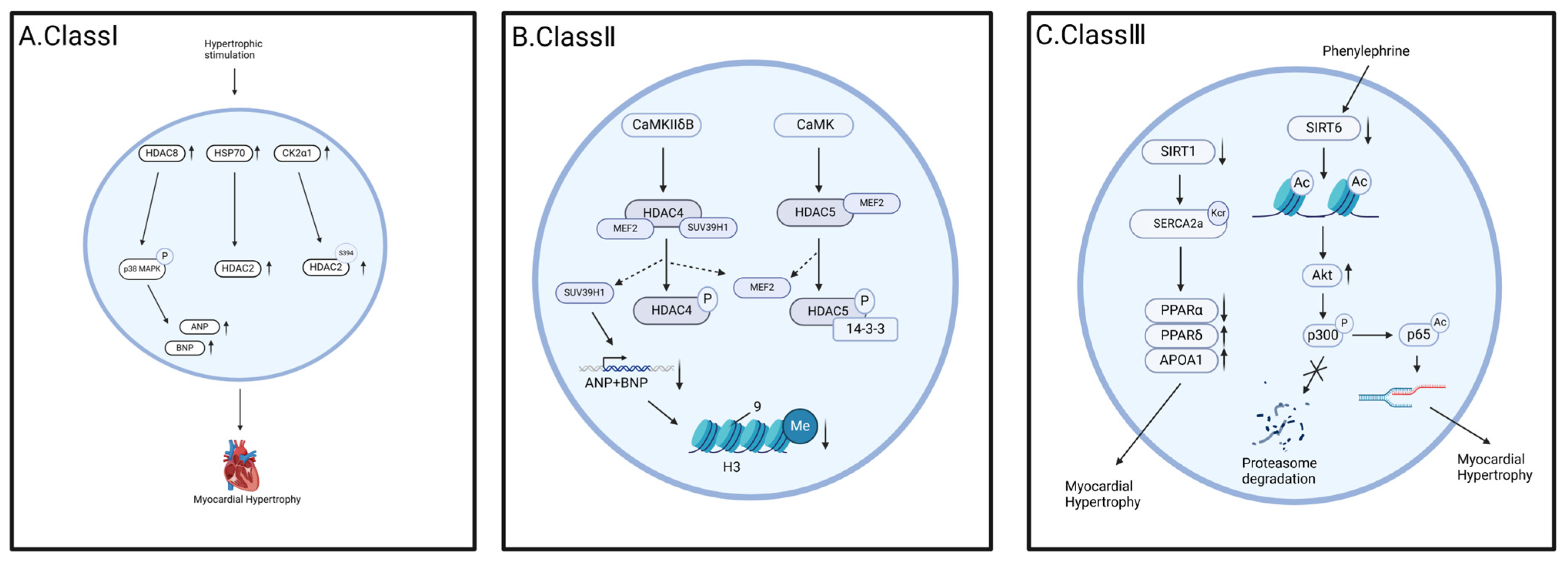
2.1. Cardiac Hypertrophy
2.2. Heart Failure
2.3. Atherosclerosis
2.4. Myocardial Ischemia/Reperfusion (I/R) Injury
2.5. Other Diseases
3. Regulation of Cardiovascular Disease by HDACis Targeting Post-Translational Protein Acetylation
4. Directions for Future Research
5. Summary and Outlook
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chong, B.; Jayabaskaran, J.; Jauhari, S.M.; Chan, S.P.; Goh, R.; Kueh, M.T.W.; Li, H.; Chin, Y.H.; Kong, G.; Anand, V.V.; et al. Global burden of cardiovascular diseases: Projections from 2025 to 2050. Eur. J. Prev. Cardiol. 2024, zwae281, Online ahead of print. [Google Scholar] [CrossRef] [PubMed]
- Partridge, L.; Deelen, J.; Slagboom, P.E. Facing up to the global challenges of ageing. Nature 2018, 561, 45–56. [Google Scholar] [CrossRef] [PubMed]
- Paneni, F.; Diaz Cañestro, C.; Libby, P.; Lüscher, T.F.; Camici, G.G. The Aging Cardiovascular System: Understanding It at the Cellular and Clinical Levels. J. Am. Coll. Cardiol. 2017, 69, 1952–1967. [Google Scholar] [CrossRef] [PubMed]
- Malhotra, A.; Redberg, R.F.; Meier, P. Saturated fat does not clog the arteries: Coronary heart disease is a chronic inflammatory condition, the risk of which can be effectively reduced from healthy lifestyle interventions. Br. J. Sports Med. 2017, 51, 1111–1112. [Google Scholar] [CrossRef] [PubMed]
- Cheng, X.; Wang, K.; Zhao, Y.; Wang, K. Research progress on post-translational modification of proteins and cardiovascular diseases. Cell Death Discov. 2023, 9, 275. [Google Scholar] [CrossRef] [PubMed]
- Marquez, J.; Lee, S.R.; Kim, N.; Han, J. Post-Translational Modifications of Cardiac Mitochondrial Proteins in Cardiovascular Disease: Not Lost in Translation. Korean Circ. J. 2016, 46, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Jiang, G.; Li, C.; Lu, M.; Lu, K.; Li, H. Protein lysine crotonylation: Past, present, perspective. Cell Death Dis. 2021, 12, 703. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Wang, G. Protein Modification and Autophagy Activation. Adv. Exp. Med. Biol. 2019, 1206, 237–259. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Fan, X.; Yu, W. Regulatory Mechanism of Protein Crotonylation and Its Relationship with Cancer. Cells 2024, 13, 1812. [Google Scholar] [CrossRef] [PubMed]
- Grotenbreg, G.; Ploegh, H. Chemical biology: Dressed-up proteins. Nature 2007, 446, 993–995. [Google Scholar] [CrossRef] [PubMed]
- Geiss-Friedlander, R.; Melchior, F. Concepts in sumoylation: A decade on. Nat. Rev. Mol. Cell Biol. 2007, 8, 947–956. [Google Scholar] [CrossRef] [PubMed]
- Morrison, R.S.; Kinoshita, Y.; Johnson, M.D.; Uo, T.; Ho, J.T.; McBee, J.K.; Conrads, T.P.; Veenstra, T.D. Proteomic analysis in the neurosciences. Mol. Cell. Proteom. 2002, 1, 553–560. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, H.; Sano, N.; Muto, S.; Horikoshi, M. Simple histone acetylation plays a complex role in the regulation of gene expression. Brief. Funct. Genom. Proteomic 2006, 5, 190–208. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Foley, E.A.; Kawashima, S.A.; Molloy, K.R.; Li, Y.; Chait, B.T.; Kapoor, T.M. Examining post-translational modification-mediated protein-protein interactions using a chemical proteomics approach. Protein Sci. 2013, 22, 287–295. [Google Scholar] [CrossRef] [PubMed]
- Chavez, J.D.; Weisbrod, C.R.; Zheng, C.; Eng, J.K.; Bruce, J.E. Protein interactions, post-translational modifications and topologies in human cells. Mol. Cell. Proteom. 2013, 12, 1451–1467. [Google Scholar] [CrossRef] [PubMed]
- Czuba, L.C.; Hillgren, K.M.; Swaan, P.W. Post-translational modifications of transporters. Pharmacol. Ther. 2018, 192, 88–99. [Google Scholar] [CrossRef] [PubMed]
- You, H.; Li, S.; Chen, Y.; Lin, J.; Wang, Z.; Dennis, M.; Li, C.; Yang, D. Global proteome and lysine succinylation analyses provide insights into the secondary metabolism in Salvia miltiorrhiza. J. Proteom. 2023, 288, 104959. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.; Larsen, B.; Lin, Z.Y.; Breitkreutz, A.; Mellacheruvu, D.; Fermin, D.; Qin, Z.S.; Tyers, M.; Gingras, A.C.; Nesvizhskii, A.I. SAINT: Probabilistic scoring of affinity purification-mass spectrometry data. Nat. Methods 2011, 8, 70–73. [Google Scholar] [CrossRef] [PubMed]
- Henikoff, S. Mechanisms of Nucleosome Dynamics In Vivo. Cold Spring Harb. Perspect. Med. 2016, 6, a026666. [Google Scholar] [CrossRef] [PubMed]
- Gregoretti, I.V.; Lee, Y.M.; Goodson, H.V. Molecular evolution of the histone deacetylase family: Functional implications of phylogenetic analysis. J. Mol. Biol. 2004, 338, 17–31. [Google Scholar] [CrossRef] [PubMed]
- Bradner, J.E.; West, N.; Grachan, M.L.; Greenberg, E.F.; Haggarty, S.J.; Warnow, T.; Mazitschek, R. Chemical phylogenetics of histone deacetylases. Nat. Chem. Biol. 2010, 6, 238–243. [Google Scholar] [CrossRef] [PubMed]
- Thiagalingam, S.; Cheng, K.H.; Lee, H.J.; Mineva, N.; Thiagalingam, A.; Ponte, J.F. Histone deacetylases: Unique players in shaping the epigenetic histone code. Ann. N. Y. Acad. Sci. 2003, 983, 84–100. [Google Scholar] [CrossRef] [PubMed]
- Hsu, K.C.; Liu, C.Y.; Lin, T.E.; Hsieh, J.H.; Sung, T.Y.; Tseng, H.J.; Yang, J.M.; Huang, W.J. Novel Class IIa-Selective Histone Deacetylase Inhibitors Discovered Using an in Silico Virtual Screening Approach. Sci. Rep. 2017, 7, 3228. [Google Scholar] [CrossRef] [PubMed]
- Demos-Davies, K.M.; Ferguson, B.S.; Cavasin, M.A.; Mahaffey, J.H.; Williams, S.M.; Spiltoir, J.I.; Schuetze, K.B.; Horn, T.R.; Chen, B.; Ferrara, C.; et al. HDAC6 contributes to pathological responses of heart and skeletal muscle to chronic angiotensin-II signaling. Am. J. Physiol. Heart Circ. Physiol. 2014, 307, H252–H258. [Google Scholar] [CrossRef] [PubMed]
- Cheng, F.; Su, L.; Yao, C.; Liu, L.; Shen, J.; Liu, C.; Chen, X.; Luo, Y.; Jiang, L.; Shan, J.; et al. SIRT1 promotes epithelial-mesenchymal transition and metastasis in colorectal cancer by regulating Fra-1 expression. Cancer Lett. 2016, 375, 274–283. [Google Scholar] [CrossRef] [PubMed]
- Rine, J.; Strathern, J.N.; Hicks, J.B.; Herskowitz, I. A suppressor of mating-type locus mutations in Saccharomyces cerevisiae: Evidence for and identification of cryptic mating-type loci. Genetics 1979, 93, 877–901. [Google Scholar] [CrossRef] [PubMed]
- Tanno, M.; Sakamoto, J.; Miura, T.; Shimamoto, K.; Horio, Y. Nucleocytoplasmic shuttling of the NAD+-dependent histone deacetylase SIRT1. J. Biol. Chem. 2007, 282, 6823–6832. [Google Scholar] [CrossRef] [PubMed]
- Movahedi Naini, S.; Sheridan, A.M.; Force, T.; Shah, J.V.; Bonventre, J.V. Group IVA Cytosolic Phospholipase A2 Regulates the G2-to-M Transition by Modulating the Activity of Tumor Suppressor SIRT2. Mol. Cell. Biol. 2015, 35, 3768–3784. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Nakagawa, T.; Lomb, D.J.; Haigis, M.C.; Guarente, L. SIRT5 Deacetylates carbamoyl phosphate synthetase 1 and regulates the urea cycle. Cell 2009, 137, 560–570. [Google Scholar] [CrossRef] [PubMed]
- Seto, E.; Yoshida, M. Erasers of histone acetylation: The histone deacetylase enzymes. Cold Spring Harb. Perspect. Biol. 2014, 6, a018713. [Google Scholar] [CrossRef] [PubMed]
- Bagchi, R.A.; Ferguson, B.S.; Stratton, M.S.; Hu, T.; Cavasin, M.A.; Sun, L.; Lin, Y.H.; Liu, D.; Londono, P.; Song, K.; et al. HDAC11 suppresses the thermogenic program of adipose tissue via BRD2. JCI Insight 2018, 3, e120159. [Google Scholar] [CrossRef] [PubMed]
- Marks, P.A.; Richon, V.M.; Breslow, R.; Rifkind, R.A. Histone deacetylase inhibitors as new cancer drugs. Curr. Opin. Oncol. 2001, 13, 477–483. [Google Scholar] [CrossRef] [PubMed]
- Glauben, R.; Siegmund, B. Molecular basis of histone deacetylase inhibitors as new drugs for the treatment of inflammatory diseases and cancer. Methods Mol. Biol. 2009, 512, 365–376. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Zhao, D.; Wu, C.; Wu, Z.; Zhang, W.; Chen, S.; Zhao, X.; Wu, S. Role of histone deacetylase inhibitors in non-neoplastic diseases. Heliyon 2024, 10, e33997. [Google Scholar] [CrossRef] [PubMed]
- Singh, B.N.; Zhang, G.; Hwa, Y.L.; Li, J.; Dowdy, S.C.; Jiang, S.W. Nonhistone protein acetylation as cancer therapy targets. Expert. Rev. Anticancer Ther. 2010, 10, 935–954. [Google Scholar] [CrossRef] [PubMed]
- Bush, E.W.; McKinsey, T.A. Protein acetylation in the cardiorenal axis: The promise of histone deacetylase inhibitors. Circ. Res. 2010, 106, 272–284. [Google Scholar] [CrossRef] [PubMed]
- Hsing, C.H.; Hung, S.K.; Chen, Y.C.; Wei, T.S.; Sun, D.P.; Wang, J.J.; Yeh, C.H. Histone Deacetylase Inhibitor Trichostatin A Ameliorated Endotoxin-Induced Neuroinflammation and Cognitive Dysfunction. Mediat. Inflamm. 2015, 2015, 163140. [Google Scholar] [CrossRef] [PubMed]
- Trapani, D.; Esposito, A.; Criscitiello, C.; Mazzarella, L.; Locatelli, M.; Minchella, I.; Minucci, S.; Curigliano, G. Entinostat for the treatment of breast cancer. Expert. Opin. Investig. Drugs 2017, 26, 965–971. [Google Scholar] [CrossRef] [PubMed]
- Ruwizhi, N.; Aderibigbe, B.A. Cinnamic Acid Derivatives and Their Biological Efficacy. Int. J. Mol. Sci. 2020, 21, 5712. [Google Scholar] [CrossRef] [PubMed]
- Weeks, K.L.; Gao, X.; Du, X.J.; Boey, E.J.; Matsumoto, A.; Bernardo, B.C.; Kiriazis, H.; Cemerlang, N.; Tan, J.W.; Tham, Y.K.; et al. Phosphoinositide 3-kinase p110α is a master regulator of exercise-induced cardioprotection and PI3K gene therapy rescues cardiac dysfunction. Circ. Heart Fail. 2012, 5, 523–534. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, M.; Sadoshima, J. Mechanisms of physiological and pathological cardiac hypertrophy. Nat. Rev. Cardiol. 2018, 15, 387–407. [Google Scholar] [CrossRef] [PubMed]
- Grimm, M.; Brown, J.H. Beta-adrenergic receptor signaling in the heart: Role of CaMKII. J. Mol. Cell. Cardiol. 2010, 48, 322–330. [Google Scholar] [CrossRef] [PubMed]
- Erickson, J.R.; Joiner, M.L.; Guan, X.; Kutschke, W.; Yang, J.; Oddis, C.V.; Bartlett, R.K.; Lowe, J.S.; O’Donnell, S.E.; Aykin-Burns, N.; et al. A dynamic pathway for calcium-independent activation of CaMKII by methionine oxidation. Cell 2008, 133, 462–474. [Google Scholar] [CrossRef] [PubMed]
- Feng, N.; Anderson, M.E. CaMKII is a nodal signal for multiple programmed cell death pathways in heart. J. Mol. Cell. Cardiol. 2017, 103, 102–109. [Google Scholar] [CrossRef] [PubMed]
- Backs, J.; Backs, T.; Neef, S.; Kreusser, M.M.; Lehmann, L.H.; Patrick, D.M.; Grueter, C.E.; Qi, X.; Richardson, J.A.; Hill, J.A.; et al. The delta isoform of CaM kinase II is required for pathological cardiac hypertrophy and remodeling after pressure overload. Proc. Natl. Acad. Sci. USA 2009, 106, 2342–2347. [Google Scholar] [CrossRef] [PubMed]
- He, B.J.; Joiner, M.L.; Singh, M.V.; Luczak, E.D.; Swaminathan, P.D.; Koval, O.M.; Kutschke, W.; Allamargot, C.; Yang, J.; Guan, X.; et al. Oxidation of CaMKII determines the cardiotoxic effects of aldosterone. Nat. Med. 2011, 17, 1610–1618. [Google Scholar] [CrossRef] [PubMed]
- Kee, H.J.; Eom, G.H.; Joung, H.; Shin, S.; Kim, J.R.; Cho, Y.K.; Choe, N.; Sim, B.W.; Jo, D.; Jeong, M.H.; et al. Activation of histone deacetylase 2 by inducible heat shock protein 70 in cardiac hypertrophy. Circ. Res. 2008, 103, 1259–1269. [Google Scholar] [CrossRef] [PubMed]
- Eom, G.H.; Cho, Y.K.; Ko, J.H.; Shin, S.; Choe, N.; Kim, Y.; Joung, H.; Kim, H.S.; Nam, K.I.; Kee, H.J.; et al. Casein kinase-2α1 induces hypertrophic response by phosphorylation of histone deacetylase 2 S394 and its activation in the heart. Circulation 2011, 123, 2392–2403. [Google Scholar] [CrossRef] [PubMed]
- Grégoire, S.; Xiao, L.; Nie, J.; Zhang, X.; Xu, M.; Li, J.; Wong, J.; Seto, E.; Yang, X.J. Histone deacetylase 3 interacts with and deacetylates myocyte enhancer factor 2. Mol. Cell. Biol. 2007, 27, 1280–1295. [Google Scholar] [CrossRef] [PubMed]
- Montgomery, R.L.; Potthoff, M.J.; Haberland, M.; Qi, X.; Matsuzaki, S.; Humphries, K.M.; Richardson, J.A.; Bassel-Duby, R.; Olson, E.N. Maintenance of cardiac energy metabolism by histone deacetylase 3 in mice. J. Clin. Investig. 2008, 118, 3588–3597. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.H.; Warren, C.M.; Li, J.; McKinsey, T.A.; Russell, B. Myofibril growth during cardiac hypertrophy is regulated through dual phosphorylation and acetylation of the actin capping protein CapZ. Cell Signal 2016, 28, 1015–1024. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Kee, H.J.; Bai, L.; Kim, M.K.; Kee, S.J.; Jeong, M.H. Selective HDAC8 Inhibition Attenuates Isoproterenol-Induced Cardiac Hypertrophy and Fibrosis via p38 MAPK Pathway. Front. Pharmacol. 2021, 12, 677757. [Google Scholar] [CrossRef] [PubMed]
- Karamboulas, C.; Swedani, A.; Ward, C.; Al-Madhoun, A.S.; Wilton, S.; Boisvenue, S.; Ridgeway, A.G.; Skerjanc, I.S. HDAC activity regulates entry of mesoderm cells into the cardiac muscle lineage. J. Cell Sci. 2006, 119, 4305–4314. [Google Scholar] [CrossRef] [PubMed]
- Davis, F.J.; Gupta, M.; Camoretti-Mercado, B.; Schwartz, R.J.; Gupta, M.P. Calcium/calmodulin-dependent protein kinase activates serum response factor transcription activity by its dissociation from histone deacetylase, HDAC4. Implications in cardiac muscle gene regulation during hypertrophy. J. Biol. Chem. 2003, 278, 20047–20058. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.L.; McKinsey, T.A.; Chang, S.; Antos, C.L.; Hill, J.A.; Olson, E.N. Class II histone deacetylases act as signal-responsive repressors of cardiac hypertrophy. Cell 2002, 110, 479–488. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, R.S.; Ferreira, J.C.; Gomes, E.R.; Paixão, N.A.; Rolim, N.P.; Medeiros, A.; Guatimosim, S.; Brum, P.C. Cardiac anti-remodelling effect of aerobic training is associated with a reduction in the calcineurin/NFAT signalling pathway in heart failure mice. J. Physiol. 2009, 587, 3899–3910. [Google Scholar] [CrossRef] [PubMed]
- Miska, E.A.; Karlsson, C.; Langley, E.; Nielsen, S.J.; Pines, J.; Kouzarides, T. HDAC4 deacetylase associates with and represses the MEF2 transcription factor. EMBO J. 1999, 18, 5099–5107. [Google Scholar] [CrossRef] [PubMed]
- Madugula, K.; Mulherkar, R.; Khan, Z.K.; Chigbu, D.I.; Patel, D.; Harhaj, E.W.; Jain, P. MEF-2 isoforms’ (A-D) roles in development and tumorigenesis. Oncotarget 2019, 10, 2755–2787. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Xu, J.; Gong, N.L.; Bodi, I.; Aronow, B.J.; Backx, P.H.; Molkentin, J.D. Myocyte enhancer factors 2A and 2C induce dilated cardiomyopathy in transgenic mice. J. Biol. Chem. 2006, 281, 9152–9162. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, Y.; Nishikimi, T.; Kuwahara, K. Atrial and brain natriuretic peptides: Hormones secreted from the heart. Peptides 2019, 111, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Hohl, M.; Wagner, M.; Reil, J.C.; Müller, S.A.; Tauchnitz, M.; Zimmer, A.M.; Lehmann, L.H.; Thiel, G.; Böhm, M.; Backs, J.; et al. HDAC4 controls histone methylation in response to elevated cardiac load. J. Clin. Investig. 2013, 123, 1359–1370. [Google Scholar] [CrossRef] [PubMed]
- McKinsey, T.A.; Zhang, C.L.; Olson, E.N. Activation of the myocyte enhancer factor-2 transcription factor by calcium/calmodulin-dependent protein kinase-stimulated binding of 14-3-3 to histone deacetylase 5. Proc. Natl. Acad. Sci. USA 2000, 97, 14400–14405. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.; McKinsey, T.A.; Zhang, C.L.; Richardson, J.A.; Hill, J.A.; Olson, E.N. Histone deacetylases 5 and 9 govern responsiveness of the heart to a subset of stress signals and play redundant roles in heart development. Mol. Cell. Biol. 2004, 24, 8467–8476. [Google Scholar] [CrossRef] [PubMed]
- Vega, R.B.; Harrison, B.C.; Meadows, E.; Roberts, C.R.; Papst, P.J.; Olson, E.N.; McKinsey, T.A. Protein kinases C and D mediate agonist-dependent cardiac hypertrophy through nuclear export of histone deacetylase 5. Mol. Cell. Biol. 2004, 24, 8374–8385. [Google Scholar] [CrossRef] [PubMed]
- McKinsey, T.A.; Zhang, C.L.; Olson, E.N. Identification of a signal-responsive nuclear export sequence in class II histone deacetylases. Mol. Cell. Biol. 2001, 21, 6312–6321. [Google Scholar] [CrossRef] [PubMed]
- Harrison, B.C.; Kim, M.S.; van Rooij, E.; Plato, C.F.; Papst, P.J.; Vega, R.B.; McAnally, J.A.; Richardson, J.A.; Bassel-Duby, R.; Olson, E.N.; et al. Regulation of cardiac stress signaling by protein kinase d1. Mol. Cell. Biol. 2006, 26, 3875–3888. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Deng, M.; Lu, A.; Chen, Y.; Chen, Y.; Wu, C.; Tan, Z.; Boini, K.M.; Yang, T.; Zhu, Q.; et al. Sodium butyrate attenuates angiotensin II-induced cardiac hypertrophy by inhibiting COX2/PGE2 pathway via a HDAC5/HDAC6-dependent mechanism. J. Cell. Mol. Med. 2019, 23, 8139–8150. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Chan, K.T.; Huang, X.; Cerra, C.; Blake, S.; Trigos, A.S.; Anderson, D.; Creek, D.J.; De Souza, D.P.; Wang, X.; et al. Cystathionine-β-synthase is essential for AKT-induced senescence and suppresses the development of gastric cancers with PI3K/AKT activation. Elife 2022, 11, e71929. [Google Scholar] [CrossRef] [PubMed]
- Borradaile, N.M.; Pickering, J.G. NAD(+), sirtuins, and cardiovascular disease. Curr. Pharm. Des. 2009, 15, 110–117. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wang, C.; Turdi, S.; Richmond, K.L.; Zhang, Y.; Ren, J. ALDH2 protects against high fat diet-induced obesity cardiomyopathy and defective autophagy: Role of CaM kinase II, histone H3K9 methyltransferase SUV39H, Sirt1, and PGC-1α deacetylation. Int. J. Obes. 2018, 42, 1073–1087. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Liu, C.; Chen, Q.; Liu, N.; Yan, Y.; Liu, B. SIRT3: A New Regulator of Cardiovascular Diseases. Oxid. Med. Cell. Longev. 2018, 2018, 7293861. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Yin, A.; Zhang, Q.; Zhong, T.; O’Rourke, S.T.; Sun, C. Angiotensin-(1-7) attenuates angiotensin II-induced cardiac hypertrophy via a Sirt3-dependent mechanism. Am. J. Physiol. Heart Circ. Physiol. 2017, 312, H980–H991. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Gong, B.; Duan, W.; Fan, C.; Zhang, J.; Li, Z.; Xue, X.; Xu, Y.; Meng, D.; Li, B.; et al. Melatonin ameliorates myocardial ischemia/reperfusion injury in type 1 diabetic rats by preserving mitochondrial function: Role of AMPK-PGC-1α-SIRT3 signaling. Sci. Rep. 2017, 7, 41337. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.X.; Wang, X.C.; Hou, H.T.; Wang, J.; Yang, Q.; Chen, Y.L.; Chen, H.Z.; He, G.W. Lysine crotonylation of SERCA2a correlates to cardiac dysfunction and arrhythmia in Sirt1 cardiac-specific knockout mice. Int. J. Biol. Macromol. 2023, 242, 125151. [Google Scholar] [CrossRef] [PubMed]
- Sadhukhan, S.; Liu, X.; Ryu, D.; Nelson, O.D.; Stupinski, J.A.; Li, Z.; Chen, W.; Zhang, S.; Weiss, R.S.; Locasale, J.W.; et al. Metabolomics-assisted proteomics identifies succinylation and SIRT5 as important regulators of cardiac function. Proc. Natl. Acad. Sci. USA 2016, 113, 4320–4325. [Google Scholar] [CrossRef] [PubMed]
- Pillai, V.B.; Samant, S.; Sundaresan, N.R.; Raghuraman, H.; Kim, G.; Bonner, M.Y.; Arbiser, J.L.; Walker, D.I.; Jones, D.P.; Gius, D.; et al. Honokiol blocks and reverses cardiac hypertrophy in mice by activating mitochondrial Sirt3. Nat. Commun. 2015, 6, 6656. [Google Scholar] [CrossRef] [PubMed]
- Reed, S.M.; Quelle, D.E. p53 Acetylation: Regulation and Consequences. Cancers 2014, 7, 30–69. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Ozden, O.; Jiang, H.; Cha, Y.I.; Pennington, J.D.; Aykin-Burns, N.; Spitz, D.R.; Gius, D.; Kim, H.S. Sirt3, mitochondrial ROS, ageing, and carcinogenesis. Int. J. Mol. Sci. 2011, 12, 6226–6239. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Ren, D.; Fedorova, J.; He, Z.; Li, J. SIRT1/SIRT3 Modulates Redox Homeostasis during Ischemia/Reperfusion in the Aging Heart. Antioxidants 2020, 9, 858. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.X.; Tang, X.; An, X.Z.; Xie, X.M.; Chen, X.F.; Zhao, X.; Hao, D.L.; Chen, H.Z.; Liu, D.P. SIRT4 accelerates Ang II-induced pathological cardiac hypertrophy by inhibiting manganese superoxide dismutase activity. Eur. Heart J. 2017, 38, 1389–1398. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Chen, X.F.; Wang, N.Y.; Wang, X.M.; Liang, S.T.; Zheng, W.; Lu, Y.B.; Zhao, X.; Hao, D.L.; Zhang, Z.Q.; et al. SIRT2 Acts as a Cardioprotective Deacetylase in Pathological Cardiac Hypertrophy. Circulation 2017, 136, 2051–2067. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Zhang, X.; Guo, Z.; Zhong, Y.; Wang, P.; Li, J.; Li, Z.; Liu, P. SIRT6 Suppresses NFATc4 Expression and Activation in Cardiomyocyte Hypertrophy. Front. Pharmacol. 2018, 9, 1519. [Google Scholar] [CrossRef] [PubMed]
- Sundaresan, N.R.; Vasudevan, P.; Zhong, L.; Kim, G.; Samant, S.; Parekh, V.; Pillai, V.B.; Ravindra, P.V.; Gupta, M.; Jeevanandam, V.; et al. The sirtuin SIRT6 blocks IGF-Akt signaling and development of cardiac hypertrophy by targeting c-Jun. Nat. Med. 2012, 18, 1643–1650. [Google Scholar] [CrossRef] [PubMed]
- Shen, P.; Feng, X.; Zhang, X.; Huang, X.; Liu, S.; Lu, X.; Li, J.; You, J.; Lu, J.; Li, Z.; et al. SIRT6 suppresses phenylephrine-induced cardiomyocyte hypertrophy though inhibiting p300. J. Pharmacol. Sci. 2016, 132, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Vakhrusheva, O.; Smolka, C.; Gajawada, P.; Kostin, S.; Boettger, T.; Kubin, T.; Braun, T.; Bober, E. Sirt7 increases stress resistance of cardiomyocytes and prevents apoptosis and inflammatory cardiomyopathy in mice. Circ. Res. 2008, 102, 703–710. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Yu, S.S.; Chen, S.R.; Pi, R.B.; Gao, S.; Li, H.; Ye, J.T.; Liu, P.Q. Nmnat2 protects cardiomyocytes from hypertrophy via activation of SIRT6. FEBS Lett. 2012, 586, 866–874. [Google Scholar] [CrossRef] [PubMed]
- Abate, G.; Bastonini, E.; Braun, K.A.; Verdone, L.; Young, E.T.; Caserta, M. Snf1/AMPK regulates Gcn5 occupancy, H3 acetylation and chromatin remodelling at S. cerevisiae ADY2 promoter. Biochim. Biophys. Acta 2012, 1819, 419–427. [Google Scholar] [CrossRef] [PubMed]
- Tanai, E.; Frantz, S. Pathophysiology of Heart Failure. Compr. Physiol. 2015, 6, 187–214. [Google Scholar] [CrossRef] [PubMed]
- Lüscher, T.F. Risk factors for and management of heart failure. Eur. Heart J. 2015, 36, 2267–2269. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Savarese, G.; Becher, P.M.; Lund, L.H.; Seferovic, P.; Rosano, G.M.C.; Coats, A.J.S. Global burden of heart failure: A comprehensive and updated review of epidemiology. Cardiovasc. Res. 2023, 118, 3272–3287. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Zhong, M.; Lin, Y.; Zhang, W.; Zhu, Y.; Chen, L.; Huang, Z.; Luo, K.; Lu, Z.; Huang, Z.; et al. METTL7B-induced histone lactylation prevents heart failure by ameliorating cardiac remodelling. J. Mol. Cell. Cardiol. 2025, 202, 64–80. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.Y.; Gao, B.; Yang, Y.; Jia, S.B.; Ma, X.P.; Zhang, M.H.; Wang, L.J.; Ma, A.Q.; Zhang, Q.N. Histone deacetylase 3 suppresses the expression of SHP-1 via deacetylation of DNMT1 to promote heart failure. Life Sci. 2022, 292, 119552. [Google Scholar] [CrossRef] [PubMed]
- Travers, J.G.; Hu, T.; McKinsey, T.A. The black sheep of class IIa: HDAC7 SIKens the heart. J. Clin. Investig. 2020, 130, 2811–2813. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Qian, S.; Tang, B.; Kang, P.; Zhang, H.; Shi, C. Resveratrol inhibits ferroptosis and decelerates heart failure progression via Sirt1/p53 pathway activation. J. Cell. Mol. Med. 2023, 27, 3075–3089. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Li, S.; Zhu, J.; You, A.; Huang, X.; Yi, X.; Xue, M. Mangiferin prevents myocardial infarction-induced apoptosis and heart failure in mice by activating the Sirt1/FoxO3a pathway. J. Cell. Mol. Med. 2021, 25, 2944–2955. [Google Scholar] [CrossRef] [PubMed]
- Lin, B.; Zhao, H.; Li, L.; Zhang, Z.; Jiang, N.; Yang, X.; Zhang, T.; Lian, B.; Liu, Y.; Zhang, C.; et al. Sirt1 improves heart failure through modulating the NF-κB p65/microRNA-155/BNDF signaling cascade. Aging 2020, 13, 14482–14498. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.; Wang, C.; Weng, J. MicroRNA-138-5p drives the progression of heart failure via inhibiting sirtuin 1 signaling. Mol. Med. Rep. 2021, 23, 11915. [Google Scholar] [CrossRef] [PubMed]
- Lu, T.M.; Tsai, J.Y.; Chen, Y.C.; Huang, C.Y.; Hsu, H.L.; Weng, C.F.; Shih, C.C.; Hsu, C.P. Downregulation of Sirt1 as aging change in advanced heart failure. J. Biomed. Sci. 2014, 21, 57. [Google Scholar] [CrossRef] [PubMed]
- Gorski, P.A.; Jang, S.P.; Jeong, D.; Lee, A.; Lee, P.; Oh, J.G.; Chepurko, V.; Yang, D.K.; Kwak, T.H.; Eom, S.H.; et al. Role of SIRT1 in Modulating Acetylation of the Sarco-Endoplasmic Reticulum Ca(2+)-ATPase in Heart Failure. Circ. Res. 2019, 124, e63–e80. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Cao, Y.; Zheng, Q.; Tu, J.; Zhou, W.; He, J.; Zhong, J.; Chen, Y.; Wang, J.; Cai, R.; et al. SENP1-Sirt3 Signaling Controls Mitochondrial Protein Acetylation and Metabolism. Mol. Cell 2019, 75, 823–834.e825. [Google Scholar] [CrossRef] [PubMed]
- Sundaresan, N.R.; Bindu, S.; Pillai, V.B.; Samant, S.; Pan, Y.; Huang, J.Y.; Gupta, M.; Nagalingam, R.S.; Wolfgeher, D.; Verdin, E.; et al. SIRT3 Blocks Aging-Associated Tissue Fibrosis in Mice by Deacetylating and Activating Glycogen Synthase Kinase 3β. Mol. Cell. Biol. 2015, 36, 678–692. [Google Scholar] [CrossRef] [PubMed]
- Su, H.; Cantrell, A.C.; Chen, J.X.; Gu, W.; Zeng, H. SIRT3 Deficiency Enhances Ferroptosis and Promotes Cardiac Fibrosis via p53 Acetylation. Cells 2023, 12, 1428. [Google Scholar] [CrossRef] [PubMed]
- Castillo, E.C.; Morales, J.A.; Chapoy-Villanueva, H.; Silva-Platas, C.; Treviño-Saldaña, N.; Guerrero-Beltrán, C.E.; Bernal-Ramírez, J.; Torres-Quintanilla, A.; García, N.; Youker, K.; et al. Mitochondrial Hyperacetylation in the Failing Hearts of Obese Patients Mediated Partly by a Reduction in SIRT3: The Involvement of the Mitochondrial Permeability Transition Pore. Cell Physiol. Biochem. 2019, 53, 465–479. [Google Scholar] [CrossRef] [PubMed]
- Chang, X.; Zhang, T.; Wang, J.; Liu, Y.; Yan, P.; Meng, Q.; Yin, Y.; Wang, S. SIRT5-Related Desuccinylation Modification Contributes to Quercetin-Induced Protection against Heart Failure and High-Glucose-Prompted Cardiomyocytes Injured through Regulation of Mitochondrial Quality Surveillance. Oxid. Med. Cell. Longev. 2021, 2021, 5876841. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.S.; Wu, F.; Jin, Y.M.; Chang, W.Q.; Xu, T.M. HDAC11: A rising star in epigenetics. Biomed. Pharmacother. 2020, 131, 110607. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Chang, J.; Li, X.; Wang, J.; Wu, X.; Liu, X.; Zhu, Y.; Yu, X.Y. Association of DNA methylation and transcriptome reveals epigenetic etiology of heart failure. Funct. Integr. Genom. 2022, 22, 89–112. [Google Scholar] [CrossRef] [PubMed]
- Foks, A.C.; Bot, I. Preface: Pathology and Pharmacology of Atherosclerosis. Eur. J. Pharmacol. 2017, 816, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Famularo, G.; Trinchieri, V.; Santini, G.; De Simone, C. Infections, atherosclerosis, and coronary heart disease. Ann. Ital. Med. Int. 2000, 15, 144–155. [Google Scholar] [PubMed]
- Holmstedt, C.A.; Turan, T.N.; Chimowitz, M.I. Atherosclerotic intracranial arterial stenosis: Risk factors, diagnosis, and treatment. Lancet Neurol. 2013, 12, 1106–1114. [Google Scholar] [CrossRef] [PubMed]
- Wolberg, A.S.; Rosendaal, F.R.; Weitz, J.I.; Jaffer, I.H.; Agnelli, G.; Baglin, T.; Mackman, N. Venous thrombosis. Nat. Rev. Dis. Primers 2015, 1, 15006. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.; Peng, K.; Wu, Q.; Wang, Y.; Fan, X.; Zhang, D.M.; Passerini, A.G.; Sun, C. HDAC1 and 2 regulate endothelial VCAM-1 expression and atherogenesis by suppressing methylation of the GATA6 promoter. Theranostics 2021, 11, 5605–5619. [Google Scholar] [CrossRef] [PubMed]
- Desvignes, T.; Loher, P.; Eilbeck, K.; Ma, J.; Urgese, G.; Fromm, B.; Sydes, J.; Aparicio-Puerta, E.; Barrera, V.; Espín, R.; et al. Unification of miRNA and isomiR research: The mirGFF3 format and the mirtop API. Bioinformatics 2020, 36, 698–703. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Lu, Z.T.; Wu, G.; Yang, Z.C.; Wu, X.; Wang, D.; Nie, Z.M.; Sheng, Q. Gastrodia elata specific miRNA attenuates neuroinflammation via modulating NF-κB signaling pathway. Int. J. Neurosci. 2024, 134, 1652–1662. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Wu, F. A new miRNA regulator, miR-672, reduces cardiac hypertrophy by inhibiting JUN expression. Gene 2018, 648, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Tajbakhsh, A.; Bianconi, V.; Pirro, M.; Gheibi Hayat, S.M.; Johnston, T.P.; Sahebkar, A. Efferocytosis and Atherosclerosis: Regulation of Phagocyte Function by MicroRNAs. Trends Endocrinol. Metab. 2019, 30, 672–683. [Google Scholar] [CrossRef] [PubMed]
- Fasolo, F.; Di Gregoli, K.; Maegdefessel, L.; Johnson, J.L. Non-coding RNAs in cardiovascular cell biology and atherosclerosis. Cardiovasc. Res. 2019, 115, 1732–1756. [Google Scholar] [CrossRef] [PubMed]
- Stojkovic, S.; Nossent, A.Y.; Haller, P.; Jäger, B.; Vargas, K.G.; Wojta, J.; Huber, K. MicroRNAs as Regulators and Biomarkers of Platelet Function and Activity in Coronary Artery Disease. Thromb. Haemost. 2019, 119, 1563–1572. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Sugimoto, K.; Lu, H.; Yang, W.Y.; Liu, J.Y.; Yang, H.Y.; Song, Y.B.; Yan, D.; Zou, T.Y.; Shen, S. HDAC1-mediated deacetylation of HIF1α prevents atherosclerosis progression by promoting miR-224-3p-mediated inhibition of FOSL2. Mol. Ther. Nucleic Acids 2021, 23, 577–591. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Li, S.; Li, N.; Yang, X.; Ma, S.; Yang, A.; Zhang, H.; Yang, S.; Mao, C.; Xu, L.; et al. miR-34a Targets HDAC1-Regulated H3K9 Acetylation on Lipid Accumulation Induced by Homocysteine in Foam Cells. J. Cell Biochem. 2017, 118, 4617–4627. [Google Scholar] [CrossRef] [PubMed]
- Mittelstadt, M.L.; Patel, R.C. AP-1 mediated transcriptional repression of matrix metalloproteinase-9 by recruitment of histone deacetylase 1 in response to interferon β. PLoS ONE 2012, 7, e42152. [Google Scholar] [CrossRef] [PubMed]
- Kong, X.; Fang, M.; Li, P.; Fang, F.; Xu, Y. HDAC2 deacetylates class II transactivator and suppresses its activity in macrophages and smooth muscle cells. J. Mol. Cell. Cardiol. 2009, 46, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Ahn, Y.J.; Asmis, R. Inhibition of myeloid HDAC2 upregulates glutaredoxin 1 expression, improves protein thiol redox state and protects against high-calorie diet-induced monocyte dysfunction and atherosclerosis. Atherosclerosis 2021, 328, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Zheng, B.; Han, M.; Shu, Y.N.; Li, Y.J.; Miao, S.B.; Zhang, X.H.; Shi, H.J.; Zhang, T.; Wen, J.K. HDAC2 phosphorylation-dependent Klf5 deacetylation and RARα acetylation induced by RAR agonist switch the transcription regulatory programs of p21 in VSMCs. Cell Res. 2011, 21, 1487–1508. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Chen, M.; Chen, X.; He, X.; Li, X.; Wei, H.; Tan, Y.; Min, J.; Azam, T.; Xue, M.; et al. TRAP1 drives smooth muscle cell senescence and promotes atherosclerosis via HDAC3-primed histone H4 lysine 12 lactylation. Eur. Heart J. 2024, 45, 4219–4235. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Li, J.; Zheng, T.; Chen, T.; Yuan, Z. KLF7 Alleviates Atherosclerotic Lesions and Inhibits Glucose Metabolic Reprogramming in Macrophages by Regulating HDAC4/miR-148b-3p/NCOR1. Gerontology 2022, 68, 1291–1310. [Google Scholar] [CrossRef] [PubMed]
- Lee, G.H.; Park, J.S.; Jin, S.W.; Pham, T.H.; Thai, T.N.; Kim, J.Y.; Kim, C.Y.; Choi, J.H.; Han, E.H.; Jeong, H.G. Betulinic Acid Induces eNOS Expression via the AMPK-Dependent KLF2 Signaling Pathway. J. Agric. Food Chem. 2020, 68, 14523–14530. [Google Scholar] [CrossRef] [PubMed]
- Kai, H.; Wu, Q.; Yin, R.; Tang, X.; Shi, H.; Wang, T.; Zhang, M.; Pan, C. LncRNA NORAD Promotes Vascular Endothelial Cell Injury and Atherosclerosis Through Suppressing VEGF Gene Transcription via Enhancing H3K9 Deacetylation by Recruiting HDAC6. Front. Cell Dev. Biol. 2021, 9, 701628. [Google Scholar] [CrossRef] [PubMed]
- Gorenne, I.; Kumar, S.; Gray, K.; Figg, N.; Yu, H.; Mercer, J.; Bennett, M. Vascular smooth muscle cell sirtuin 1 protects against DNA damage and inhibits atherosclerosis. Circulation 2013, 127, 386–396. [Google Scholar] [CrossRef] [PubMed]
- Yan, P.; Li, Z.; Xiong, J.; Geng, Z.; Wei, W.; Zhang, Y.; Wu, G.; Zhuang, T.; Tian, X.; Liu, Z.; et al. LARP7 ameliorates cellular senescence and aging by allosterically enhancing SIRT1 deacetylase activity. Cell Rep. 2021, 37, 110038. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Liu, S.; Cheng, L.; Huang, T.; Guo, H.; Wang, D.; Xia, M.; Ling, W.; Xiao, Y. Epigenetic Upregulation of H19 and AMPK Inhibition Concurrently Contribute to S-Adenosylhomocysteine Hydrolase Deficiency-Promoted Atherosclerotic Calcification. Circ. Res. 2022, 130, 1565–1582. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; He, F.; Liang, B.; Jing, Y.; Zhang, P.; Liu, W.; Zhu, B.; Dou, D. LincRNA-p21 alleviates atherosclerosis progression through regulating the miR-221/SIRT1/Pcsk9 axis. J. Cell. Mol. Med. 2021, 25, 9141–9153. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Zhang, B.; Guo, R.; Li, S.; Xu, Y. Enhancement in efferocytosis of oxidized low-density lipoprotein-induced apoptotic RAW264.7 cells through Sirt1-mediated autophagy. Int. J. Mol. Med. 2014, 33, 523–533. [Google Scholar] [CrossRef] [PubMed]
- Mattagajasingh, I.; Kim, C.S.; Naqvi, A.; Yamamori, T.; Hoffman, T.A.; Jung, S.B.; DeRicco, J.; Kasuno, K.; Irani, K. SIRT1 promotes endothelium-dependent vascular relaxation by activating endothelial nitric oxide synthase. Proc. Natl. Acad. Sci. USA 2007, 104, 14855–14860. [Google Scholar] [CrossRef] [PubMed]
- Jung, S.B.; Kim, C.S.; Naqvi, A.; Yamamori, T.; Mattagajasingh, I.; Hoffman, T.A.; Cole, M.P.; Kumar, A.; Dericco, J.S.; Jeon, B.H.; et al. Histone deacetylase 3 antagonizes aspirin-stimulated endothelial nitric oxide production by reversing aspirin-induced lysine acetylation of endothelial nitric oxide synthase. Circ. Res. 2010, 107, 877–887. [Google Scholar] [CrossRef] [PubMed]
- Schermuly, R.T.; Ghofrani, H.A.; Wilkins, M.R.; Grimminger, F. Mechanisms of disease: Pulmonary arterial hypertension. Nat. Rev. Cardiol. 2011, 8, 443–455. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Wang, S.; Xiao, M.; Lin, Y.; Zhou, L.; Lei, Q.; Xiong, Y.; Guan, K.L.; Zhao, S. Acetylation regulates gluconeogenesis by promoting PEPCK1 degradation via recruiting the UBR5 ubiquitin ligase. Mol. Cell 2011, 43, 33–44. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Zhang, L.; Yin, K.; Zang, G.; Qian, Y.; Mao, X.; Li, L.; Jing, Q.; Wang, Z. SIRT3-and FAK-mediated acetylation-phosphorylation crosstalk of NFATc1 regulates N(ε)-carboxymethyl-lysine-induced vascular calcification in diabetes mellitus. Atherosclerosis 2023, 377, 43–59. [Google Scholar] [CrossRef] [PubMed]
- Velpuri, P.; Patel, P.; Yazdani, A.; Abdi, A.; Rai, V.; Agrawal, D.K. Increased Oxidative Stress and Decreased Sirtuin-3 and FOXO3 Expression Following Carotid Artery Intimal Injury in Hyperlipidemic Yucatan Microswine. Cardiol. Cardiovasc. Med. 2024, 8, 33–42. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Huang, Y.; Zhu, L.; Yang, K.; Liang, K.; Tan, J.; Yu, B. SIRT6 promotes angiogenesis and hemorrhage of carotid plaque via regulating HIF-1α and reactive oxygen species. Cell Death Dis. 2021, 12, 77. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Miao, C.; Cui, J.; Bian, X. miR-92a-3p promotes ox-LDL induced-apoptosis in HUVECs via targeting SIRT6 and activating MAPK signaling pathway. Braz. J. Med. Biol. Res. 2021, 54, e9386. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Shao, T.; Liu, H.; Wang, Q.; Li, T.; Zhao, Q. SIRT6 mediates MRTF-A deacetylation in vascular endothelial cells to antagonize oxLDL-induced ICAM-1 transcription. Cell Death Discov. 2022, 8, 96. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Dong, S.; Wu, Y.; Yi, H.; Zhang, W.; Ai, X. Sirtuin 6 Deacetylates Apoptosis-Associated Speck-Like Protein (ASC) to Inhibit Endothelial Cell Pyroptosis in Atherosclerosis. Int. Heart J. 2024, 65, 466–474. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, T.L.; Michishita, E.; Adler, A.S.; Damian, M.; Berber, E.; Lin, M.; McCord, R.A.; Ongaigui, K.C.; Boxer, L.D.; Chang, H.Y.; et al. SIRT6 links histone H3 lysine 9 deacetylation to NF-kappaB-dependent gene expression and organismal life span. Cell 2009, 136, 62–74. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.F.; Zhao, M.; Zhao, L.N.; Yun, H.L.; Yang, G.; Geng, Y.; Wang, Y.F.; Zheng, W.; Yuan, Y.; Song, T.Q.; et al. PRMT5 confers lipid metabolism reprogramming, tumour growth and metastasis depending on the SIRT7-mediated desuccinylation of PRMT5 K387 in tumours. Acta Pharmacol. Sin. 2022, 43, 2373–2385. [Google Scholar] [CrossRef] [PubMed]
- Luan, Y.; Liu, H.; Luan, Y.; Yang, Y.; Yang, J.; Ren, K.D. New Insight in HDACs: Potential Therapeutic Targets for the Treatment of Atherosclerosis. Front. Pharmacol. 2022, 13, 863677. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.L.; Morrow, D.A. Acute Myocardial Infarction. N. Engl. J. Med. 2017, 376, 2053–2064. [Google Scholar] [CrossRef] [PubMed]
- Yap, J.; Cabrera-Fuentes, H.A.; Irei, J.; Hausenloy, D.J.; Boisvert, W.A. Role of Macrophages in Cardioprotection. Int. J. Mol. Sci. 2019, 20, 2474. [Google Scholar] [CrossRef] [PubMed]
- Ibanez, B.; James, S.; Agewall, S.; Antunes, M.J.; Bucciarelli-Ducci, C.; Bueno, H.; Caforio, A.L.P.; Crea, F.; Goudevenos, J.A.; Halvorsen, S.; et al. 2017 ESC Guidelines for the management of acute myocardial infarction in patients presenting with ST-segment elevation: The Task Force for the management of acute myocardial infarction in patients presenting with ST-segment elevation of the European Society of Cardiology (ESC). Eur. Heart J. 2018, 39, 119–177. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Zhang, L.; Zhuang, S.; Qin, G.J.; Zhao, T.C. HDAC4 degradation mediates HDAC inhibition-induced protective effects against hypoxia/reoxygenation injury. J. Cell. Physiol. 2015, 230, 1321–1331. [Google Scholar] [CrossRef] [PubMed]
- Leng, Y.; Wu, Y.; Lei, S.; Zhou, B.; Qiu, Z.; Wang, K.; Xia, Z. Inhibition of HDAC6 Activity Alleviates Myocardial Ischemia/Reperfusion Injury in Diabetic Rats: Potential Role of Peroxiredoxin 1 Acetylation and Redox Regulation. Oxid. Med. Cell. Longev. 2018, 2018, 9494052. [Google Scholar] [CrossRef] [PubMed]
- Ju, J.; Li, X.M.; Zhao, X.M.; Li, F.H.; Wang, S.C.; Wang, K.; Li, R.F.; Zhou, L.Y.; Liang, L.; Wang, Y.; et al. Circular RNA FEACR inhibits ferroptosis and alleviates myocardial ischemia/reperfusion injury by interacting with NAMPT. J. Biomed. Sci. 2023, 30, 45. [Google Scholar] [CrossRef] [PubMed]
- Eid, R.A.; Bin-Meferij, M.M.; El-Kott, A.F.; Eleawa, S.M.; Zaki, M.S.A.; Al-Shraim, M.; El-Sayed, F.; Eldeen, M.A.; Alkhateeb, M.A.; Alharbi, S.A.; et al. Exendin-4 Protects Against Myocardial Ischemia-Reperfusion Injury by Upregulation of SIRT1 and SIRT3 and Activation of AMPK. J. Cardiovasc. Transl. Res. 2021, 14, 619–635. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.J.; Cui, J.; Lin, Q.; Chen, X.Y.; Zhang, J.; Gao, E.H.; Wei, B.; Zhao, W. Protection of the enhanced Nrf2 deacetylation and its downstream transcriptional activity by SIRT1 in myocardial ischemia/reperfusion injury. Int. J. Cardiol. 2021, 342, 82–93. [Google Scholar] [CrossRef] [PubMed]
- Tao, A.; Xu, X.; Kvietys, P.; Kao, R.; Martin, C.; Rui, T. Experimental diabetes mellitus exacerbates ischemia/reperfusion-induced myocardial injury by promoting mitochondrial fission: Role of down-regulation of myocardial Sirt1 and subsequent Akt/Drp1 interaction. Int. J. Biochem. Cell Biol. 2018, 105, 94–103. [Google Scholar] [CrossRef] [PubMed]
- Ning, S.; Li, Z.; Ji, Z.; Fan, D.; Wang, K.; Wang, Q.; Hua, L.; Zhang, J.; Meng, X.; Yuan, Y. MicroRNA-494 suppresses hypoxia/reoxygenation-induced cardiomyocyte apoptosis and autophagy via the PI3K/AKT/mTOR signaling pathway by targeting SIRT1. Mol. Med. Rep. 2020, 22, 5231–5242. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Shi, H.; Li, Y.; Gao, W.; Guo, J.; Zhu, J.; Dong, Z.; Sun, A.; Zou, Y.; Ge, J. Hypertrophic preconditioning attenuates myocardial ischemia/reperfusion injury through the deacetylation of isocitrate dehydrogenase 2. Sci. Bull. 2021, 66, 2099–2114. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; He, Z.; Fedorova, J.; Logan, C.; Bates, L.; Davitt, K.; Le, V.; Murphy, J.; Li, M.; Wang, M.; et al. Alterations in mitochondrial dynamics with age-related Sirtuin1/Sirtuin3 deficiency impair cardiomyocyte contractility. Aging Cell 2021, 20, e13419. [Google Scholar] [CrossRef] [PubMed]
- Bochaton, T.; Crola-Da-Silva, C.; Pillot, B.; Villedieu, C.; Ferreras, L.; Alam, M.R.; Thibault, H.; Strina, M.; Gharib, A.; Ovize, M.; et al. Inhibition of myocardial reperfusion injury by ischemic postconditioning requires sirtuin 3-mediated deacetylation of cyclophilin D. J. Mol. Cell. Cardiol. 2015, 84, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Chouchani, E.T.; Pell, V.R.; Gaude, E.; Aksentijević, D.; Sundier, S.Y.; Robb, E.L.; Logan, A.; Nadtochiy, S.M.; Ord, E.N.J.; Smith, A.C.; et al. Ischaemic accumulation of succinate controls reperfusion injury through mitochondrial ROS. Nature 2014, 515, 431–435. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Tian, X.; Zhao, Y.; Wang, Z.; Lin, M.; Sun, R.; Wang, Y.; Wang, Z.; Li, G.; Zheng, S.; et al. Sirtuin 5 Alleviates Liver Ischemia/Reperfusion Injury by Regulating Mitochondrial Succinylation and Oxidative Stress. Antioxid. Redox Signal 2024, 40, 616–631. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Chen, X.; Xu, X.; Shi, R.; Suo, S.; Cheng, K.; Zheng, Z.; Wang, M.; Wang, L.; Zhao, Y.; et al. Lysine Acetylation and Succinylation in HeLa Cells and their Essential Roles in Response to UV-induced Stress. Sci. Rep. 2016, 6, 30212. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Liu, L.; Jiang, W.; Liu, M.; Wang, Y.; Ma, H.; Mu, N.; Wang, H. SIRT6 Protects Against Myocardial Ischemia-Reperfusion Injury by Attenuating Aging-Related CHMP2B Accumulation. J. Cardiovasc. Transl. Res. 2022, 15, 740–753. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Zeng, H.; Chen, J.X. Emerging role of SIRT3 in endothelial metabolism, angiogenesis, and cardiovascular disease. J. Cell. Physiol. 2019, 234, 2252–2265. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Zeng, H.; Chen, S.T.; Roman, R.J.; Aschner, J.L.; Didion, S.; Chen, J.X. Endothelial specific SIRT3 deletion impairs glycolysis and angiogenesis and causes diastolic dysfunction. J. Mol. Cell. Cardiol. 2017, 112, 104–113. [Google Scholar] [CrossRef] [PubMed]
- Palomer, X.; Román-Azcona, M.S.; Pizarro-Delgado, J.; Planavila, A.; Villarroya, F.; Valenzuela-Alcaraz, B.; Crispi, F.; Sepúlveda-Martínez, Á.; Miguel-Escalada, I.; Ferrer, J.; et al. SIRT3-mediated inhibition of FOS through histone H3 deacetylation prevents cardiac fibrosis and inflammation. Signal Transduct. Target. Ther. 2020, 5, 14. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.Z.; Xu, W.; Zang, Y.X.; Lou, Q.; Hang, P.Z.; Gao, Q.; Shi, H.; Liu, Q.Y.; Wang, H.; Sun, X.; et al. Honokiol Inhibits Atrial Metabolic Remodeling in Atrial Fibrillation Through Sirt3 Pathway. Front. Pharmacol. 2022, 13, 813272. [Google Scholar] [CrossRef] [PubMed]
- Qi, J.; Wang, F.; Yang, P.; Wang, X.; Xu, R.; Chen, J.; Yuan, Y.; Lu, Z.; Duan, J. Mitochondrial Fission Is Required for Angiotensin II-Induced Cardiomyocyte Apoptosis Mediated by a Sirt1-p53 Signaling Pathway. Front. Pharmacol. 2018, 9, 176. [Google Scholar] [CrossRef] [PubMed]
- Kong, X.; Guan, J.; Li, J.; Wei, J.; Wang, R. P66(Shc)-SIRT1 Regulation of Oxidative Stress Protects Against Cardio-cerebral Vascular Disease. Mol. Neurobiol. 2017, 54, 5277–5285. [Google Scholar] [CrossRef] [PubMed]
- Oshikawa, J.; Kim, S.J.; Furuta, E.; Caliceti, C.; Chen, G.F.; McKinney, R.D.; Kuhr, F.; Levitan, I.; Fukai, T.; Ushio-Fukai, M. Novel role of p66Shc in ROS-dependent VEGF signaling and angiogenesis in endothelial cells. Am. J. Physiol. Heart Circ. Physiol. 2012, 302, H724–H732. [Google Scholar] [CrossRef] [PubMed]
- Paneni, F.; Mocharla, P.; Akhmedov, A.; Costantino, S.; Osto, E.; Volpe, M.; Lüscher, T.F.; Cosentino, F. Gene silencing of the mitochondrial adaptor p66(Shc) suppresses vascular hyperglycemic memory in diabetes. Circ. Res. 2012, 111, 278–289. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Kim, Y.R.; Vikram, A.; Naqvi, A.; Li, Q.; Kassan, M.; Kumar, V.; Bachschmid, M.M.; Jacobs, J.S.; Kumar, A.; et al. Sirtuin1-regulated lysine acetylation of p66Shc governs diabetes-induced vascular oxidative stress and endothelial dysfunction. Proc. Natl. Acad. Sci. USA 2017, 114, 1714–1719. [Google Scholar] [CrossRef] [PubMed]
- Pereira, M.; Cruz, M.T.; Fortuna, A.; Bicker, J. Restoring the epigenome in Alzheimer’s disease: Advancing HDAC inhibitors as therapeutic agents. Drug Discov. Today 2024, 29, 104052. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Lv, J.; Guo, N.; Guo, Q.; Zeng, S.; Fang, Y.; Chen, W.; Wang, Z. Dose-dependent Effects of PRC2 and HDAC Inhibitors on Cardiomyocyte Hypertrophy Induced by Phenylephrine. Curr. Drug Targets 2023, 24, 371–378. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.H.; Seok, Y.M.; Song, M.J.; Lee, H.A.; Kurz, T.; Kim, I. Histone deacetylase inhibition attenuates cardiac hypertrophy and fibrosis through acetylation of mineralocorticoid receptor in spontaneously hypertensive rats. Mol. Pharmacol. 2015, 87, 782–791. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhu, Z.; Xue, M.; Yi, X.; Liang, J.; Niu, C.; Chen, G.; Shen, Y.; Zhang, H.; Zheng, J.; et al. Fibroblast growth factor 21 protects the heart from angiotensin II-induced cardiac hypertrophy and dysfunction via SIRT1. Biochim. Biophys. Acta Mol. Basis Dis. 2019, 1865, 1241–1252. [Google Scholar] [CrossRef] [PubMed]
- Gao, Q.; Wei, A.; Chen, F.; Chen, X.; Ding, W.; Ding, Z.; Wu, Z.; Du, R.; Cao, W. Enhancing PPARγ by HDAC inhibition reduces foam cell formation and atherosclerosis in ApoE deficient mice. Pharmacol. Res. 2020, 160, 105059. [Google Scholar] [CrossRef] [PubMed]
- Herr, D.J.; Baarine, M.; Aune, S.E.; Li, X.; Ball, L.E.; Lemasters, J.J.; Beeson, C.C.; Chou, J.C.; Menick, D.R. HDAC1 localizes to the mitochondria of cardiac myocytes and contributes to early cardiac reperfusion injury. J. Mol. Cell. Cardiol. 2018, 114, 309–319. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Yin, Y.; Guo, X.; Liu, M.; Ma, F.; Tian, L.; Zheng, M.; Liu, G. The histone deacetylase inhibitor SAHA exerts a protective effect against myocardial ischemia/reperfusion injury by inhibiting sodium-calcium exchanger. Biochem. Biophys. Res. Commun. 2023, 671, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Guerra-Ojeda, S.; Suarez, A.; Belmonte, B.; Marchio, P.; Genovés, P.; Arias, O.J.; Aldasoro, M.; Vila, J.M.; Serna, E.; Mauricio, M.D. Sodium valproate treatment reverses endothelial dysfunction in aorta from rabbits with acute myocardial infarction. Eur. J. Pharmacol. 2024, 970, 176475. [Google Scholar] [CrossRef] [PubMed]
- Williams, A.M.; He, W.; Li, Y.; Bhatti, U.F.; Nikolian, V.C.; Chang, P.; Chang, Z.; Halaweish, I.; Liu, B.; Cheng, X.; et al. Histone Deacetylase Inhibition Attenuates Cardiomyocyte Hypoxia-Reoxygenation Injury. Curr. Mol. Med. 2018, 18, 711–718. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.C.; Cheng, G.; Zhang, L.X.; Tseng, Y.T.; Padbury, J.F. Inhibition of histone deacetylases triggers pharmacologic preconditioning effects against myocardial ischemic injury. Cardiovasc. Res. 2007, 76, 473–481. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Fang, C.; Zong, N.C.; Liem, D.A.; Cadeiras, M.; Scruggs, S.B.; Yu, H.; Kim, A.K.; Yang, P.; Deng, M.; et al. Regulation of acetylation restores proteolytic function of diseased myocardium in mouse and human. Mol. Cell. Proteom. 2013, 12, 3793–3802. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Liu, Q.; Chen, L.; Zhang, L.; Gu, E. Remifentanil postconditioning ameliorates histone H3 acetylation modification in H9c2 cardiomyoblasts after hypoxia/reoxygenation via attenuating endoplasmic reticulum stress. Apoptosis 2017, 22, 662–671. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Patel, D.; Zhang, X.; Veenstra, R.D. Changes in cardiac Nav1.5 expression, function, and acetylation by pan-histone deacetylase inhibitors. Am. J. Physiol. Heart Circ. Physiol. 2016, 311, H1139–H1149. [Google Scholar] [CrossRef] [PubMed]
- Colussi, C.; Berni, R.; Rosati, J.; Straino, S.; Vitale, S.; Spallotta, F.; Baruffi, S.; Bocchi, L.; Delucchi, F.; Rossi, S.; et al. The histone deacetylase inhibitor suberoylanilide hydroxamic acid reduces cardiac arrhythmias in dystrophic mice. Cardiovasc. Res. 2010, 87, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.F.; Hsu, K.C.; HuangFu, W.C.; Lin, T.E.; Huang, H.L.; Pan, S.L. Investigating the potential effects of selective histone deacetylase 6 inhibitor ACY1215 on infarct size in rats with cardiac ischemia-reperfusion injury. BMC Pharmacol. Toxicol. 2020, 21, 21. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, P.; Wang, L.; Zhao, J.; Zhong, Z.; Wang, Y.; Xu, J. Inhibition of Histone Deacetylases Prevents Cardiac Remodeling After Myocardial Infarction by Restoring Autophagosome Processing in Cardiac Fibroblasts. Cell Physiol. Biochem. 2018, 49, 1999–2011. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Tao, H.; Cao, W.; Cao, L.; Lin, Y.; Zhao, S.M.; Xu, W.; Cao, J.; Zhao, J.Y. Ketogenic diets inhibit mitochondrial biogenesis and induce cardiac fibrosis. Signal Transduct. Target. Ther. 2021, 6, 54. [Google Scholar] [CrossRef] [PubMed]
- David, G.; Neptune, M.A.; DePinho, R.A. SUMO-1 modification of histone deacetylase 1 (HDAC1) modulates its biological activities. J. Biol. Chem. 2002, 277, 23658–23663. [Google Scholar] [CrossRef] [PubMed]
- Brandl, A.; Wagner, T.; Uhlig, K.M.; Knauer, S.K.; Stauber, R.H.; Melchior, F.; Schneider, G.; Heinzel, T.; Krämer, O.H. Dynamically regulated sumoylation of HDAC2 controls p53 deacetylation and restricts apoptosis following genotoxic stress. J. Mol. Cell. Biol. 2012, 4, 284–293. [Google Scholar] [CrossRef] [PubMed]
- Nott, A.; Watson, P.M.; Robinson, J.D.; Crepaldi, L.; Riccio, A. S-Nitrosylation of histone deacetylase 2 induces chromatin remodelling in neurons. Nature 2008, 455, 411–415. [Google Scholar] [CrossRef] [PubMed]
- Colussi, C.; Mozzetta, C.; Gurtner, A.; Illi, B.; Rosati, J.; Straino, S.; Ragone, G.; Pescatori, M.; Zaccagnini, G.; Antonini, A.; et al. HDAC2 blockade by nitric oxide and histone deacetylase inhibitors reveals a common target in Duchenne muscular dystrophy treatment. Proc. Natl. Acad. Sci. USA 2008, 105, 19183–19187. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.D.; Wang, L.N.; Wen, P.P.; Shi, S.P.; Qiu, J.D. Site-Specific Systematic Analysis of Lysine Modification Crosstalk. Proteomics 2018, 18, e1700292. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, T.G.; Möller, A.; Sirma, H.; Zentgraf, H.; Taya, Y.; Dröge, W.; Will, H.; Schmitz, M.L. Regulation of p53 activity by its interaction with homeodomain-interacting protein kinase-2. Nat. Cell Biol. 2002, 4, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Sakaguchi, K.; Herrera, J.E.; Saito, S.; Miki, T.; Bustin, M.; Vassilev, A.; Anderson, C.W.; Appella, E. DNA damage activates p53 through a phosphorylation-acetylation cascade. Genes. Dev. 1998, 12, 2831–2841. [Google Scholar] [CrossRef] [PubMed]
- Appella, E.; Anderson, C.W. Signaling to p53: Breaking the posttranslational modification code. Pathol. Biol. 2000, 48, 227–245. [Google Scholar] [PubMed]
- Ferguson, B.S.; Harrison, B.C.; Jeong, M.Y.; Reid, B.G.; Wempe, M.F.; Wagner, F.F.; Holson, E.B.; McKinsey, T.A. Signal-dependent repression of DUSP5 by class I HDACs controls nuclear ERK activity and cardiomyocyte hypertrophy. Proc. Natl. Acad. Sci. USA 2013, 110, 9806–9811. [Google Scholar] [CrossRef] [PubMed]
- Guttzeit, S.; Backs, J. Post-translational modifications talk and crosstalk to class IIa histone deacetylases. J. Mol. Cell. Cardiol. 2022, 162, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Yang, X.; Cao, J. Application of Single-Cell Genomics in Cardiovascular Research. Cardiol. Ther. 2023, 12, 101–125. [Google Scholar] [CrossRef] [PubMed]
- Chaffin, M.; Papangeli, I.; Simonson, B.; Akkad, A.D.; Hill, M.C.; Arduini, A.; Fleming, S.J.; Melanson, M.; Hayat, S.; Kost-Alimova, M.; et al. Single-nucleus profiling of human dilated and hypertrophic cardiomyopathy. Nature 2022, 608, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Yu, P.; Zhou, B.; Song, J.; Li, Z.; Zhang, M.; Guo, G.; Wang, Y.; Chen, X.; Han, L.; et al. Single-cell reconstruction of the adult human heart during heart failure and recovery reveals the cellular landscape underlying cardiac function. Nat. Cell Biol. 2020, 22, 108–119. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.; Shim, D.; Lee, J.S.; Zaitsev, K.; Williams, J.W.; Kim, K.W.; Jang, M.Y.; Seok Jang, H.; Yun, T.J.; Lee, S.H.; et al. Transcriptome Analysis Reveals Nonfoamy Rather Than Foamy Plaque Macrophages Are Proinflammatory in Atherosclerotic Murine Models. Circ. Res. 2018, 123, 1127–1142. [Google Scholar] [CrossRef] [PubMed]
- Gladka, M.M.; Molenaar, B.; de Ruiter, H.; van der Elst, S.; Tsui, H.; Versteeg, D.; Lacraz, G.P.A.; Huibers, M.M.H.; van Oudenaarden, A.; van Rooij, E. Single-Cell Sequencing of the Healthy and Diseased Heart Reveals Cytoskeleton-Associated Protein 4 as a New Modulator of Fibroblasts Activation. Circulation 2018, 138, 166–180. [Google Scholar] [CrossRef] [PubMed]
- Piekarz, R.L.; Frye, R.; Turner, M.; Wright, J.J.; Allen, S.L.; Kirschbaum, M.H.; Zain, J.; Prince, H.M.; Leonard, J.P.; Geskin, L.J.; et al. Phase II multi-institutional trial of the histone deacetylase inhibitor romidepsin as monotherapy for patients with cutaneous T-cell lymphoma. J. Clin. Oncol. 2009, 27, 5410–5417. [Google Scholar] [CrossRef] [PubMed]
- Wolbrette, D.L. Drugs that cause Torsades de pointes and increase the risk of sudden cardiac death. Curr. Cardiol. Rep. 2004, 6, 379–384. [Google Scholar] [CrossRef] [PubMed]
- Witt, O.; Deubzer, H.E.; Milde, T.; Oehme, I. HDAC family: What are the cancer relevant targets? Cancer Lett. 2009, 277, 8–21. [Google Scholar] [CrossRef] [PubMed]
- Shultz, M.D.; Cao, X.; Chen, C.H.; Cho, Y.S.; Davis, N.R.; Eckman, J.; Fan, J.; Fekete, A.; Firestone, B.; Flynn, J.; et al. Optimization of the in vitro cardiac safety of hydroxamate-based histone deacetylase inhibitors. J. Med. Chem. 2011, 54, 4752–4772. [Google Scholar] [CrossRef] [PubMed]
- Shi, M.Q.; Xu, Y.; Fu, X.; Pan, D.S.; Lu, X.P.; Xiao, Y.; Jiang, Y.Z. Advances in targeting histone deacetylase for treatment of solid tumors. J. Hematol. Oncol. 2024, 17, 37. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhao, Y.T.; Zhao, T.C. Histone deacetylases in modulating cardiac disease and their clinical translational and therapeutic implications. Exp. Biol. Med. 2021, 246, 213–225. [Google Scholar] [CrossRef] [PubMed]
- Bagchi, R.A.; Weeks, K.L. Histone deacetylases in cardiovascular and metabolic diseases. J. Mol. Cell. Cardiol. 2019, 130, 151–159. [Google Scholar] [CrossRef] [PubMed]




| Classification | HDAC Inhibitor | Chemical Structure | The Target |
|---|---|---|---|
| Short-chain fatty acid | Sodium Butyrate (NaB) |  | Pan inhibitor |
| Valproic acid (VPA) |  | Pan inhibitor | |
| Cyclic peptide | Apicidin |  | Pan inhibitor |
| Romidepsin |  | HDAC1/2/4/6 | |
| Hydroxamate | Trichostatin A (TSA) |  | Class I/II |
| Vorinostat (SAHA) |  | Pan inhibitor | |
| Belinostat |  | Pan inhibitor | |
| Panobinostat |  | Pan inhibitor | |
| Benzamide | Entinostat |  | ClassI |
| Mocetinostat |  | Pan inhibitor |
| Biological Function | Drug | Target Protein | Modification Site | The Acetylation Level | Reference |
|---|---|---|---|---|---|
| Inhibition of cardiac hypertrophy | TSA | HDAC6 | H3K9 | ↓ | [173] |
| VPA | HDACs | ↑ | [174] | ||
| Fibroblast Growth Factor 21 (FGF21) | SIRT1 | ↓ | [175] | ||
| Atherosclerosis | TSA | HDAC6 | H3K9 | [176] | |
| Protecting the heart from ischemia/reperfusion injury | Entinostat | HDAC1/2/3 | ↑ | [177] | |
| SAHA | HDAC1/2/3/6/7/11 | H3K14 | ↑ | [178] | |
| VPA | HDAC1/2 | H3K4 | ↓ | [179] | |
| Tubastatin A (TubA) | HDAC6 | ↓ | [180] | ||
| TSA | HDACs | ↑ | [181] | ||
| Sodium valproate | HDACs | ↑ | [182] | ||
| Remifentanil | HDAC3 | ↑ | [183] | ||
| TubA | HDAC6 | Prdx1 | ↑ | [150] | |
| Slowing myocardial conduction and increasing susceptibility to refractory arrhythmias | Romidepsin | HDAC1/2/4/6 | ↓ | [184] | |
| SAHA | HDACs | ↑ | [185] | ||
| Reduction in infarct size in rats with myocardial ischemia/reperfusion injury | Panobinostat | HDAC6 | ↑ | [186] | |
| Reduce myocardial infarction | TSA | HDAC6 | H3K9 | [187] | |
| Apoptosis and cardiac fibrosis | β-hydroxybutyrate (β-OHB) | HDAC2 | ↑ | [188] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, S.; Sun, Y.; Yu, W. HDACs and Their Inhibitors on Post-Translational Modifications: The Regulation of Cardiovascular Disease. Cells 2025, 14, 1116. https://doi.org/10.3390/cells14141116
Yang S, Sun Y, Yu W. HDACs and Their Inhibitors on Post-Translational Modifications: The Regulation of Cardiovascular Disease. Cells. 2025; 14(14):1116. https://doi.org/10.3390/cells14141116
Chicago/Turabian StyleYang, Siyi, Yidong Sun, and Wei Yu. 2025. "HDACs and Their Inhibitors on Post-Translational Modifications: The Regulation of Cardiovascular Disease" Cells 14, no. 14: 1116. https://doi.org/10.3390/cells14141116
APA StyleYang, S., Sun, Y., & Yu, W. (2025). HDACs and Their Inhibitors on Post-Translational Modifications: The Regulation of Cardiovascular Disease. Cells, 14(14), 1116. https://doi.org/10.3390/cells14141116






