1. Introduction
Of the 45 million breeding ewes in Australia, over 250,000 are artificially inseminated each year [
1]. The increasingly widespread use of artificial insemination (AI) is driven primarily by the advantages it offers over natural breeding, including the ease in use of alternative genetics, shortened lambing period, and increased number of ewes inseminated to one ejaculate. Successful conception in these assisted reproductive programs is crucial for profitable sheep farming. Low conception rates increase generation interval and reduce the number of lambs born and genetic gain, thereby having a negative impact on farm profitability. Moreover, large-scale assisted reproductive programs are time consuming, labour intensive and expensive; poor conception outcomes in these AI programs can also significantly impact farm profitability.
Reproductive success, meaning a ewe found to be pregnant following ultrasound scanning particularly after artificial insemination (AI), is dependent on a number of contributing factors on both the ewe and ram sides. While there has been considerable emphasis on characterising ewe side contributions to conception success (e.g., age, parity, etc.), relatively little emphasis has been placed on characterising ram side contributors. Literature suggests that failures during capacitation, fertilisation, or embryogenesis can all be of seminal origin [
2]. Moreover, specifically in the case of sheep, conception outcomes can vary considerably between sires, with semen from some sires yielding consistently high conception rates in contrast to other sires that yield either consistently poor or highly variable conception outcomes. Since a single ejaculate from a ram has the ability to inseminate hundreds of ewes through AI programs [
1], it is crucial to understand the mechanisms underlying these ram side factors that contribute to successful conception. Establishment and standardisation of accurate in vitro measures of the reproductive success of a ram are vital both prior to cryopreservation and use in artificial breeding programs.
Historically, the motility and morphology of semen ejaculates are assessed visually via a microscope [
3] prior to AI, in order to avoid the use of poor quality semen in AI programs that could lead to poor conception outcomes. While this suggests a relationship between semen quality parameters and conception outcomes, the magnitude of influence that semen quality parameters have on conception outcomes has been difficult to characterise. This is primarily because visual assessment bias between technicians has been reported to vary between 10–80% [
4], which means that semen quality determined via visual assessment can be highly subjective. However, in recent decades, Computer Assisted Semen Analysis (CASA) has become possible, which enables objective, repeatable assessment of semen quality parameters. This creates an opportunity to not only objectively characterise high and low quality semen samples, but also investigate the determinants of semen quality.
One of the key factors with respect to semen quality and seminal factors that contribute to conception outcomes relates to the influence of spermatozoal RNA and the physiological role of RNA in spermatozoal cells. There are several hypotheses in the literature which range from spermatozoal RNA influencing fertilisation [
5], promoting embryonic development [
6], and offspring phenotype [
7]. Accordingly, several studies in both animals (e.g., cattle [
8], pigs [
9], horses [
10]); and humans [
11] have attempted to characterise the spermatozoal transcriptome profiles. A previous study in pigs has also shown that spermatozoal transcript profiles can vary between high and low quality ejaculates and between breeds [
9]. If similar differences exist between breeds and between high and low quality ejaculates in sheep, then this could potentially be an important determinant of successful conception and farm profitability. However, only a few studies have attempted to characterise the ovine spermatozoal transcriptome, with previous research focused on expression of protein hormone adiponectin on motility parameters [
12], influence of pentose phosphate pathway enzymes on ram semen capacitation [
13], and expression of sperm transcriptome following heat stress [
14]. Therefore, the objective of the current study is to characterise the transcriptome of three sheep breeds common to Australia and determine whether spermatozoal transcript profiles vary between breeds and between ejaculates of varying quality.
4. Discussion
Spermatozoal cells contain a large repertoire of RNA transcripts that are transferred to the ovum during fertilisation. However, the physiological role of spermatozoal RNA, particularly in relation to fertility and embryonic development, remains largely unknown. Therefore, the key objectives of this study were to characterise the ovine spermatozoal transcriptome and to determine whether transcriptomic profiles varied between breeds and between semen ejaculates of varying quality.
A total of 754 differentially expressed genes (DEGs) were identified in this study when comparing spermatozoal transcript profiles of three common sheep breeds, and the top 10 DEGs identified in each breed contrast were subjected to a literature review. Genes of interest identified in the Merino vs. Dohne contrast include
5′-Aminolevulinate Synthase 1 (ALAS1),
Capping Protein Regulator and Myosin 1 Linked (
CARMIL1), and
Mevalonate Kinase (
MVK) (
Table 4). The gene
ALAS1 is known to regulate circadian networks in cattle, which could play a crucial role in the regulation of reproduction in seasonally breeding species like sheep [
31]. A bovine genome wide association study (GWAS) identified
CARMIL1 to be significantly associated with fertility [
37]. Finally,
MVK has been found to be significantly associated with regulation of cholesterol synthesis [
69], which in turn, is thought to play a significant role in spermatogenesis [
70].
In the Merino vs. Poll Dorset contrast,
Solute Carrier Family 35 Member A5 (
SLC35A5) and
Integral Membrane Protein 2C (
ITM2C) were identified as key DEGs, both of which have been reported for their association with conception and embryonic development [
40,
44] (
Table 4). Specifically, in cattle, a GWAS found
SLC35A5 to be associated with fertility [
40]; while the precise physiological role of
ITM2C is not yet known, expression of
ITM2C is significantly enriched in the epididymis and vas deferens in both humans and mice during sexual maturation [
46]. Lastly, key DEGs identified in the Dohne vs. Poll Dorset contrast included
DNA Polymerase Kappa (
POLK)
, which is developmentally regulated in the testis of human and mice, and is hypothesised to play a significant role in [
55]; and
Mannosidase Alpha Class 1A Member 1 (MAN1A1), which has been reported to be associated with weight at 6 months in sheep in a GWAS [
57] (
Table 4).
There have been limited investigations aimed at characterising the spermatozoal transcriptomes of different breeds in livestock species. However, a previous study focused on pigs did report breed-specific differences in spermatozoal transcript profiles, which is similar to the findings in this study [
9]. Specifically, this study reported that Durocs differed significantly when compared to the Yorkshire and Landrace breeds, and ultimately suggested that the conception, growth, and offspring phenotype may be significantly influenced by spermatozoal transcripts of the pigs [
9]. Overall, results from both the current study and the previous porcine study indicate that breed-specific differences in spermatozoal transcriptome profiles exist, and the physiological relevance of these differences, particularly with respect to animal production, warrants further investigation.
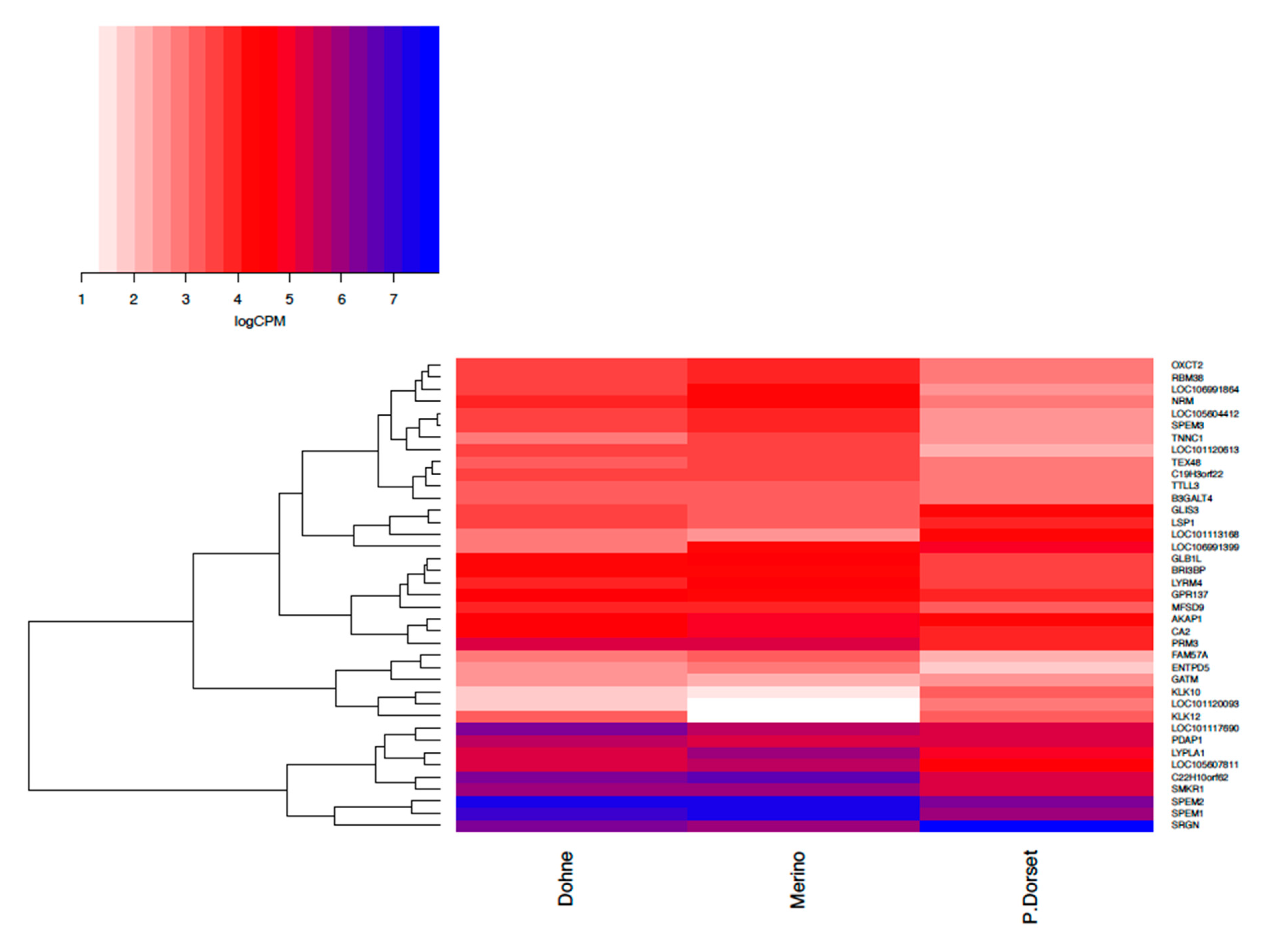
When contrasting ejaculates that were determined to be relatively high and low quality, the key DEGs identified included
SPEM Family Member 2 (
SPEM2),
3-Oxoacid CoA-Transferase 2 (
OXCT2), and
Family with Sequence Similarity 57 Member A (
FAM57A). In a GWAS in pigs,
SPEM2 was found to be associated with embryonic development and number of piglets born alive [
60]. Georgiadis et al. [
65] used
OXCT2 as a post-fertilisation and early embryonic marker using quantitative polymerase chain reaction (qPCR) when investigating high quality RNA in human semen. Similarly,
FAM57A has been shown to be upregulated in meiosis in mice, and is thought to play a crucial role in spermatogenesis [
67].
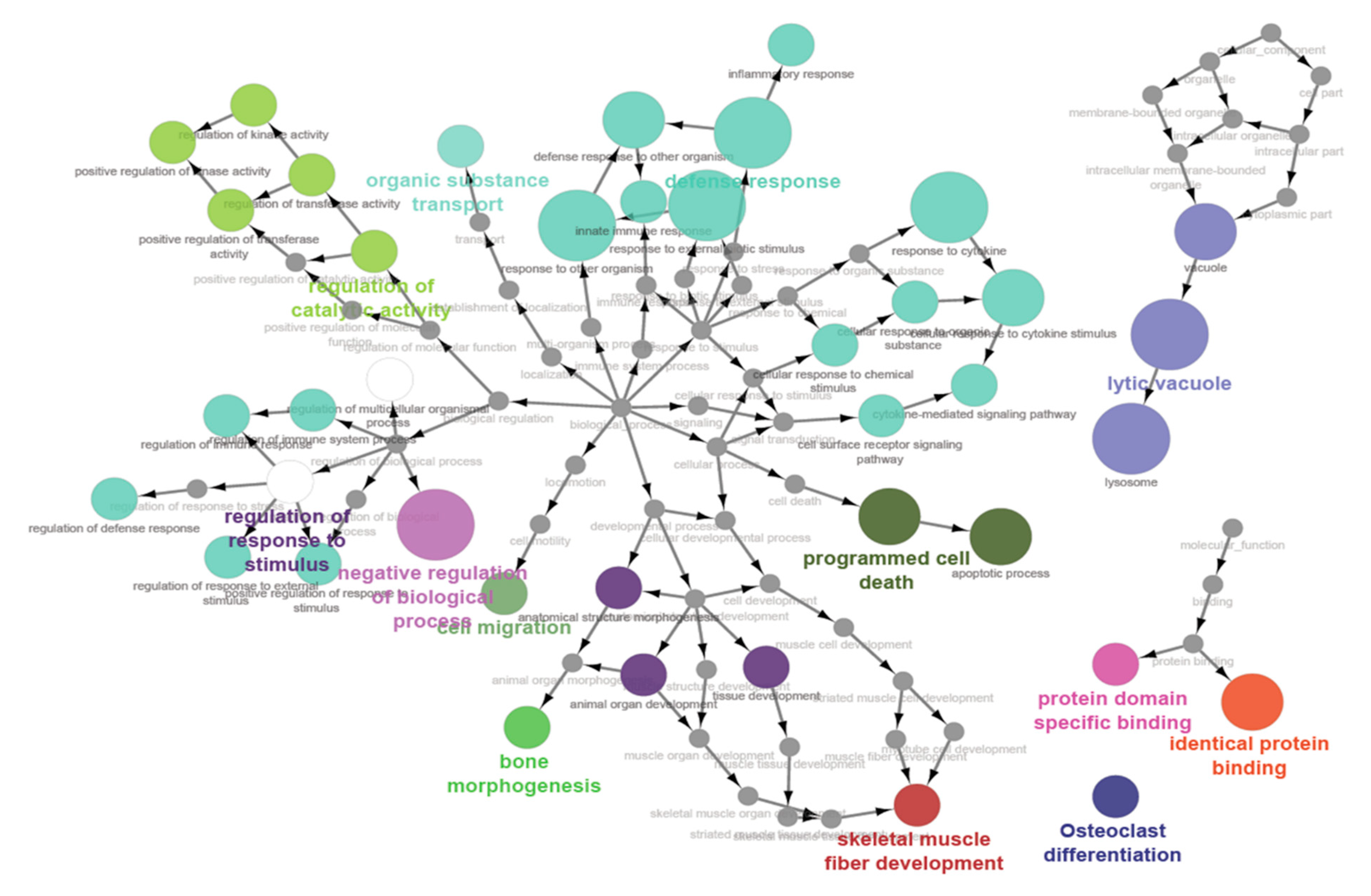
Results from gene set enrichment analyses further identified several biological pathways that accord with the results observed in differential gene expression analyses. The top pathways found to be significantly enriched in sampling groups included cell migration, regulation of molecular function, bone morphogenesis, organic substance transport, apoptotic processes, skeletal muscle fibre development, and animal organ development. It is noteworthy that a previous study in horses, which aimed to contrast the spermatozoal transcriptomes of stallions with differing conception rates, reported significantly enriched biological pathways similar to those identified in this study, including cellular function and maintenance, cellular growth and proliferation, cellular movement, cell death and survival, tissue morphology, and organ development [
10]. Given that a lot of these pathways would be relevant to embryonic development, these results indicate that spermatozoal transcripts could play a role in embryonic development and thereby contribute to either successful conception, or possibly the maintenance of pregnancy in animals.
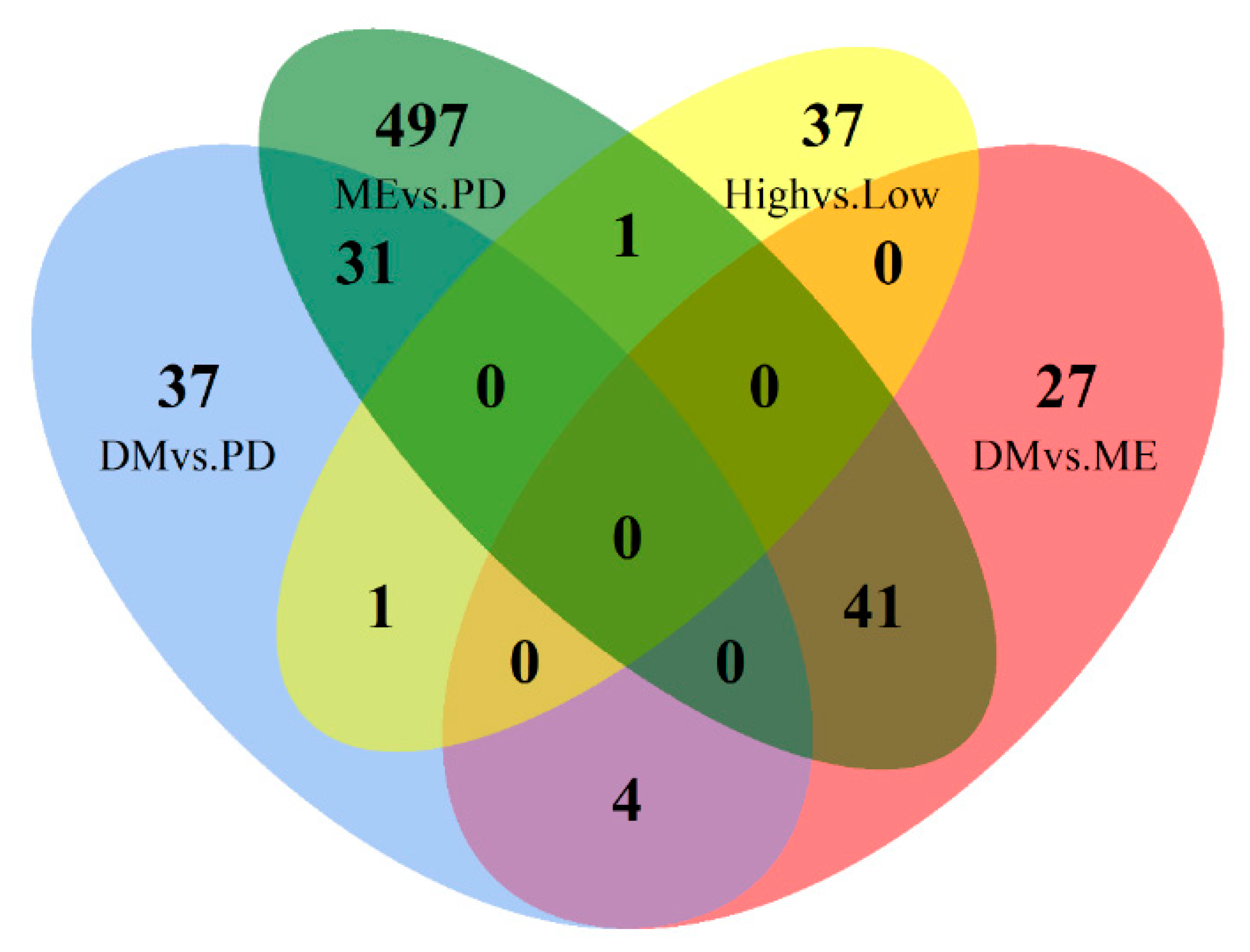
Comparison of the differentially expressed genes from one breed with the other two breeds indicated unique expression profiles. 41, 4, and 31 genes were found to be expressed at a significantly different level when compared to the other two breeds for Merino, Dohne, and Poll Dorset, respectively (
Table 5), suggesting that each gene has a unique expression profile in said breed.
In total, 41 DEGs were identified in common, when contrasting Merinos with either Poll Dorsets or Dohnes. Of these, 9 have been found, in the literature, to significantly influence reproductive output and embryonic development. Specifically,
Mitogen-Activated Protein Kinase 3 (
MAPK3),
Mediator Complex Subunit 6 (
MED6),
WAS/WASL-Interacting Protein Family Member 2 (
GTL2)
, and
Spectrin Beta, Non-Erythocytic 2 (
SPTBN2) have been reported for associations with in vitro oocyte maturation [
71], stem cell development [
36], normal embryonic development [
72], and cytoskeletal development [
73], respectively. Similarly,
Mevalonate Kinase (
MVK) [
69] and
Chondroitin Sulfate N-Acetylgalactosaminyltransferase 1 (
CSGALNACT1) [
74] have been linked with regulation of cholesterol synthesis and copper metabolism respectively, both of which are thought to play crucial roles in spermatogenesis and reproductive performance. Finally,
Tensin 3 (
TNS3) is known to be expressed in the endometrium during parturition in pigs [
30], and
Obscurin Like Cyotoskeletal Adaptor 1 (
OBSL1) is significantly expressed in ovarian follicles in cattle [
75], which could indicate a physiological role in either conception or embryonic development.
Of the 41 DEGs identified in common when contrasting Merinos with either Poll Dorsets or Dohnes, nine of those genes have been associated with growth and development following a literature search. Previous studies show that genes
Muskelin (
MKLN1) [
76],
Carbohydrate Sulfotransferase 4 (
CHST4) [
77],
Family with Sequence Similarity 210 Member B (
FAM210B) [
44], and
Limb Development Membrane Protein 1 (
LMBR1) [
78] have been shown to be respectively associated with back fat accumulation in pigs, enriched in pathways associated with feed intake in cattle, meat quality in pigs, and normal limb and embryonic development in both chickens and pigs, suggesting roles in growth and development of offspring. The following common DEGs for Merino rams are associated with breed specific production performance traits:
Transforming Growth Factor Beta 2 (
TGFB2) [
79],
Keratin 4 (
KRT4) [
80], and
Involucrin (
IVL) [
81]. These have all been found in the literature to be expressed during skin tissue and wool follicle development. Similarly,
Intersectin 1 (
ITSN1) is directly associated with the polled trait in cattle [
82], suggesting it is significant in the Poll Merino ram phenotype, as this gene was upregulated for the Merino when contrasting Merino and Poll Dorset rams.
Four genes identified in common when contrasting Dohnes with either Merino or Poll Dorsets have been found in the literature to have various influences on growth and development:
Capping Protein Regulator and Myosin 1 Linker 1 (
CARMIL1)
, Fibronectin Leucine Rich Transmembrane Protein 2 (
FLRT2),
ATP Binding Cassette Subfamily A Member 4 (
ABCA4), and
Zinc Finger Protein 382 (
ZNF382). Cole et al. [
38] found that a SNP located close to
FLRT2 is associated with significant effects on embryonic development and live birth weight in cattle. Similarly, previous studies have highlighted the importance of
ABCA4, ZNF382, and
CARMIL1 in the development of neural tissue [
83], heart development, and function [
84], and the importance of ensuring motility of cells, such as lamellipodium, which is important for cell survival [
37].
Of the 31 DEGs found in common when contrasting Poll Dorset rams with either Merinos or Dohnes, 11 were found to be associated with growth and development, 3 with fertility and reproduction, and 8 with carcass performance.
Peptidase Domain Containing Associated With Muscle Regeneration 1 (
PAMR1) and
Sorbin and SH3 Domain Containing 2 (
SORBS2) have been found to respectively influence formation and development of skin [
49], and muscle tissue and disposition of marbling (IMF) in cattle [
85]. Puttabyatappa [
58] found
Chintase 3 Like 1 (CHI3L1) to be differentially expressed and play a role in sheep follicular function, suggesting an importance in embryonic development. Similarly,
Lymphocyte Antigen 6 Family Member E (
LY6E) [
50] has been discovered to be significantly expressed in biological pathways associated with embryonic development in cattle. The
Ring Finger Protein 151 (
RNF151) gene has been found to be exclusively expressed in the testis and plays a crucial role in the spermatogenesis, and as such is crucial for the regulation of male reproductive performance [
86,
87].
Previous studies emphasise the findings of this current study, with common DEGs between each sampling breed suggesting that the spermatozoal transcriptome plays an important role in conception and embryonic development.