Institut Pasteur Dakar Mobile Lab: Part of the Solution to Tackle COVID Pandemic in Senegal, a Model to Be Exploited
Abstract
1. Introduction
2. Materials and Methods
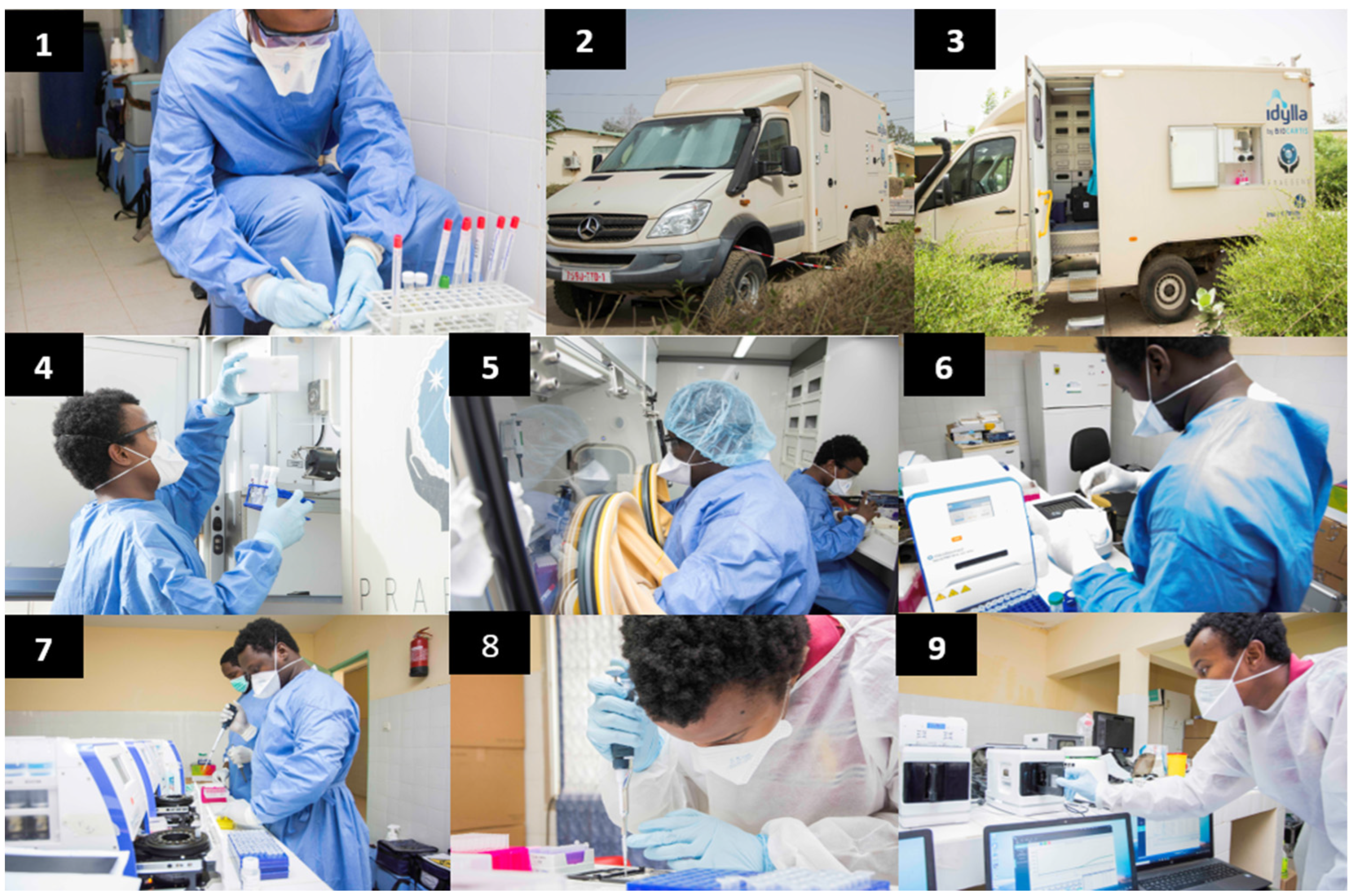
2.1. Organization of the Laboratory
2.2. Sample Collection
2.3. Sample Handling and RNA Extraction
2.4. RT-PCR Diagnostic Assays
2.5. Sequences Generation and Analysis
2.6. Data Management and Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.-M.; Wang, W.; Song, Z.-G.; Hu, Y.; Tao, Z.-W.; Tian, J.-H.; Pei, Y.-Y.; et al. A New Coronavirus Associated with Human Respiratory Disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef] [PubMed]
- WHO Coronavirus (COVID-19) Dashboard. Available online: https://covid19.who.int (accessed on 12 May 2022).
- Coronavirus Disease 2019 (COVID-19)—Diagnosis Approach | BMJ Best Practice US. Available online: https://bestpractice.bmj.com/topics/en-us/3000168/diagnosis-approach (accessed on 3 December 2021).
- Dia, N.; Lakh, N.A.; Diagne, M.M.; Mbaye, K.D.; Taieb, F.; Fall, N.M.; Barry, M.A.; Ka, D.; Fall, A.; Diallo, V.M.P.C.; et al. COVID-19 Outbreak, Senegal, 2020. Emerg. Infect. Dis. 2020, 26, 2771–2773. [Google Scholar] [CrossRef] [PubMed]
- Fall, C.; Cappuyns, A.; Faye, O.; Pauwels, S.; Fall, G.; Dia, N.; Diagne, M.M.; Diagne, C.T.; Niang, M.; Mbengue, A.; et al. Field Evaluation of a Mobile Biosafety Laboratory in Senegal to Strengthen Rapid Disease Outbreak Response and Monitoring. Afr. J. Lab. Med. 2020, 9, 1041. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, H.; Mathieu, E.; Rodés-Guirao, L.; Appel, C.; Giattino, C.; Ortiz-Ospina, E.; Hasell, J.; Macdonald, B.; Beltekian, D.; Roser, M. Coronavirus Pandemic (COVID-19). Our World in Data 2020. Available online: https://ourworldindata.org/covid-vaccinations?country=JPN~USA (accessed on 26 November 2021).
- Dieng, I.; Diarra, M.; Diagne, M.M.; Faye, M.; Dior Ndione, M.H.; Ba, Y.; Diop, M.; Ndiaye, E.H.; Marinho de Andrade Zanotto, P.; Diop, B.; et al. Field Deployment of a Mobile Biosafety Laboratory Reveals the Co-Circulation of Dengue Viruses Serotype 1 and Serotype 2 in Louga City, Senegal, 2017. J. Trop. Med. 2021, 2021, e8817987. [Google Scholar] [CrossRef] [PubMed]
- Singhal, T. A Review of Coronavirus Disease-2019 (COVID-19). Indian J. Pediatr. 2020, 87, 281–286. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.; Hu, N.; Lou, J.; Chen, K.; Kang, X.; Xiang, Z.; Chen, H.; Wang, D.; Liu, N.; Liu, D.; et al. Characteristics of COVID-19 Infection in Beijing. J. Infect. 2020, 80, 401–406. [Google Scholar] [CrossRef] [PubMed]
- Lawal, Y. Africa’s Low COVID-19 Mortality Rate: A Paradox? Int. J. Infect. Dis. 2021, 102, 118–122. [Google Scholar] [CrossRef]
- Diop, B.Z.; Ngom, M.; Pougué Biyong, C.; Pougué Biyong, J.N. The Relatively Young and Rural Population May Limit the Spread and Severity of COVID-19 in Africa: A Modelling Study. BMJ Glob. Health 2020, 5, e002699. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, P.; Curtis, N. Why Is COVID-19 Less Severe in Children? A Review of the Proposed Mechanisms Underlying the Age-Related Difference in Severity of SARS-CoV-2 Infections. Arch. Dis. Child. 2021, 106, 429–439. [Google Scholar] [CrossRef] [PubMed]
- Mecenas, P.; Bastos, R.T.d.R.M.; Vallinoto, A.C.R.; Normando, D. Effects of Temperature and Humidity on the Spread of COVID-19: A Systematic Review. PLoS ONE 2020, 15, e0238339. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Tang, K.; Feng, K.; Li, X.; Lv, W.; Chen, K.; Wang, F. Impact of Temperature and Relative Humidity on the Transmission of COVID-19: A Modeling Study in China and the United States. BMJ Open 2021, 11, e043863. [Google Scholar] [CrossRef] [PubMed]
- Motayo, B.O.; Oluwasemowo, O.O.; Olusola, B.A.; Akinduti, P.A.; Arege, O.T.; Obafemi, Y.D.; Faneye, A.O.; Isibor, P.O.; Aworunse, O.S.; Oranusi, S.U. Evolution and Genetic Diversity of SARS-CoV-2 in Africa Using Whole Genome Sequences. Int. J. Infect. Dis. 2021, 103, 282–287. [Google Scholar] [CrossRef] [PubMed]
- Ngoi, J.M.; Quashie, P.K.; Morang’a, C.M.; Bonney, J.H.; Amuzu, D.S.; Kumordjie, S.; Asante, I.A.; Bonney, E.Y.; Eshun, M.; Boatemaa, L.; et al. Genomic Analysis of SARS-CoV-2 Reveals Local Viral Evolution in Ghana. Exp. Biol. Med. 2021, 246, 960–970. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, E.; Giovanetti, M.; Tegally, H.; San, J.E.; Lessells, R.; Cuadros, D.; Martin, D.P.; Rasmussen, D.A.; Zekri, A.-R.N.; Sangare, A.K.; et al. A Year of Genomic Surveillance Reveals How the SARS-CoV-2 Pandemic Unfolded in Africa. Science 2021, 374, 423–431. [Google Scholar] [CrossRef] [PubMed]
- Lai, A.; Bergna, A.; Caucci, S.; Clementi, N.; Vicenti, I.; Dragoni, F.; Cattelan, A.M.; Menzo, S.; Pan, A.; Callegaro, A.; et al. Molecular Tracing of SARS-CoV-2 in Italy in the First Three Months of the Epidemic. Viruses 2020, 12, 798. [Google Scholar] [CrossRef] [PubMed]
- MHSA. Daily Report; Number 09; Minister of Health and Social Action: Dakar, Senegal, 2020.
- Hoang, V.-T.; Colson, P.; Levasseur, A.; Delerce, J.; Lagier, J.-C.; Parola, P.; Million, M.; Fournier, P.-E.; Raoult, D.; Gautret, P. Clinical Outcomes in Patients Infected with Different SARS-CoV-2 Variants at One Hospital during Three Phases of the COVID-19 Epidemic in Marseille, France. Infect. Genet. Evol. 2021, 95, 105092. [Google Scholar] [CrossRef] [PubMed]
- Acharya, A.; Pandey, K.; Thurman, M.; Klug, E.; Trivedi, J.; Sharma, K.; Lorson, C.L.; Singh, K.; Byrareddy, S.N. Discovery and Evaluation of Entry Inhibitors for SARS-CoV-2 and Its Emerging Variants. J. Virol. 2021, 95, e01437-21. [Google Scholar] [CrossRef] [PubMed]

| All Patients | Sars-CoV-2 Pos. | Sars-CoV-2 Neg. | OR (95% IC) | p-Value | |
|---|---|---|---|---|---|
| Positivity no. (%) | 11,693 | 1240 (10.6) | 10,453 (89.4) | ||
| Gender no. (%) | |||||
| Male | 6981 (59.7) | 743 (59.9) | 6238 (59.8) | 1.0 (0.8–1.1) | |
| Female | 4712 (40.3) | 497 (40.0) | 4215 (40.3) | ||
| Age (years) no. (%) | |||||
| [0–5] | 631 (5.4) | 31 (2.5) | 600 (05.7) | 0.05 (0.0–0.07) | <0.0005 |
| [6–14] | 1059 (9.1) | 53 (4.27) | 1006 (9.62) | 1.0 (0.6–1.6) | 0.933 |
| [15–50] | 7249 (62.0) | 754 (80.8) | 6495 (62.1) | 2.2 (1.5–3.3) | <0.0005 |
| 50+ | 2676 (22.9) | 396 (31.9) | 2280 (21.8) | 3.3 (2.3–4.9 | <0.0005 |
| Missing | 78 (0.7) | 6 (0.48) | 72 (0.68) | 1.0 (0.5–3.7) | 0.302 |
| Clinical signs no. (%) | |||||
| Fever | 2891 (24.7) | 572 (46.1) | 2319 (22.2) | 3.0 (2,6–3.3) | <0.0005 |
| Cough | 2231 (19.1) | 445 (35.9) | 1786 (17.1) | 2.7 (2.3–3.0) | <0.0005 |
| Sore throat | 899 (7.7) | 111 (08.9) | 0788 (07.5) | 1.2 (0.9–1,4) | 0.08 |
| Headache | 1773 (15.2) | 378 (30.4) | 1395 (13.3) | 2.8 (2,4–3.2) | <0.0005 |
| Myalgia | 400 (3.4) | 112 (09.0) | 288 (02.8) | 3.5 (2.7–4.3 | <0.0005 |
| Dyspnea | 547 (4.7) | 086 (06.9) | 461 (04.4) | 1.6 (1.2–2.0) | 0.0001 |
| Rhinorrhea | 612 (5.2) | 117 (09.4) | 495 (04.7) | 2.0 (1.6–2.5) | <0.0005 |
| Ageusia | 114 (1.0) | 037 (03.0) | 077 (0.7) | 4.1 (2.7–6.1) | <0.0005 |
| Anosmia | 229 (2.0) | 062 (05.0) | 167 (1.6) | 3.2 (2.4–4.3) | <0.0005 |
| Diarrhea | 077 (0.7) | 010 (00.8) | 067 (00.6) | 1.2 (0.6–2.4) | 0.2 |
| Vomiting | 080 (0.7) | 009 (00.7) | 071 (00.7) | 1.0 (0.4–2.14) | 0.4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fall, A.; Dieng, I.; Touré, C.T.; Mhamadi, M.; Sadio, B.D.; Ndione, M.H.D.; Diagne, M.M.; Ndiaye, M.; Barry, M.A.; Diaw, Y.; et al. Institut Pasteur Dakar Mobile Lab: Part of the Solution to Tackle COVID Pandemic in Senegal, a Model to Be Exploited. COVID 2022, 2, 1509-1517. https://doi.org/10.3390/covid2100108
Fall A, Dieng I, Touré CT, Mhamadi M, Sadio BD, Ndione MHD, Diagne MM, Ndiaye M, Barry MA, Diaw Y, et al. Institut Pasteur Dakar Mobile Lab: Part of the Solution to Tackle COVID Pandemic in Senegal, a Model to Be Exploited. COVID. 2022; 2(10):1509-1517. https://doi.org/10.3390/covid2100108
Chicago/Turabian StyleFall, Amary, Idrissa Dieng, Cheikh Talibouya Touré, Moufid Mhamadi, Bacary Djilocalisse Sadio, Marie Henriette Dior Ndione, Moussa Moise Diagne, Mignane Ndiaye, Mamadou Aliou Barry, Yague Diaw, and et al. 2022. "Institut Pasteur Dakar Mobile Lab: Part of the Solution to Tackle COVID Pandemic in Senegal, a Model to Be Exploited" COVID 2, no. 10: 1509-1517. https://doi.org/10.3390/covid2100108
APA StyleFall, A., Dieng, I., Touré, C. T., Mhamadi, M., Sadio, B. D., Ndione, M. H. D., Diagne, M. M., Ndiaye, M., Barry, M. A., Diaw, Y., Dieng, A. B., Diop, N. M., Sankhe, S., Diop, B., Ndiaye, M., Diallo, A., Diop, M., Dieng, M., Cappuyns, A., ... Faye, O. (2022). Institut Pasteur Dakar Mobile Lab: Part of the Solution to Tackle COVID Pandemic in Senegal, a Model to Be Exploited. COVID, 2(10), 1509-1517. https://doi.org/10.3390/covid2100108










