Comparison of the Hybrid Capture II Method with a PCR-Based Screening Method Using a Carboxyfluorescein-Labeled Primer for Detecting Human Papillomavirus in Cervicovaginal Liquid-Based Cytology
Abstract
:1. Introduction
2. Methods and Materials
2.1. Samples
2.2. HPV Detection by PCR
2.2.1. DNA Extraction
2.2.2. PCR
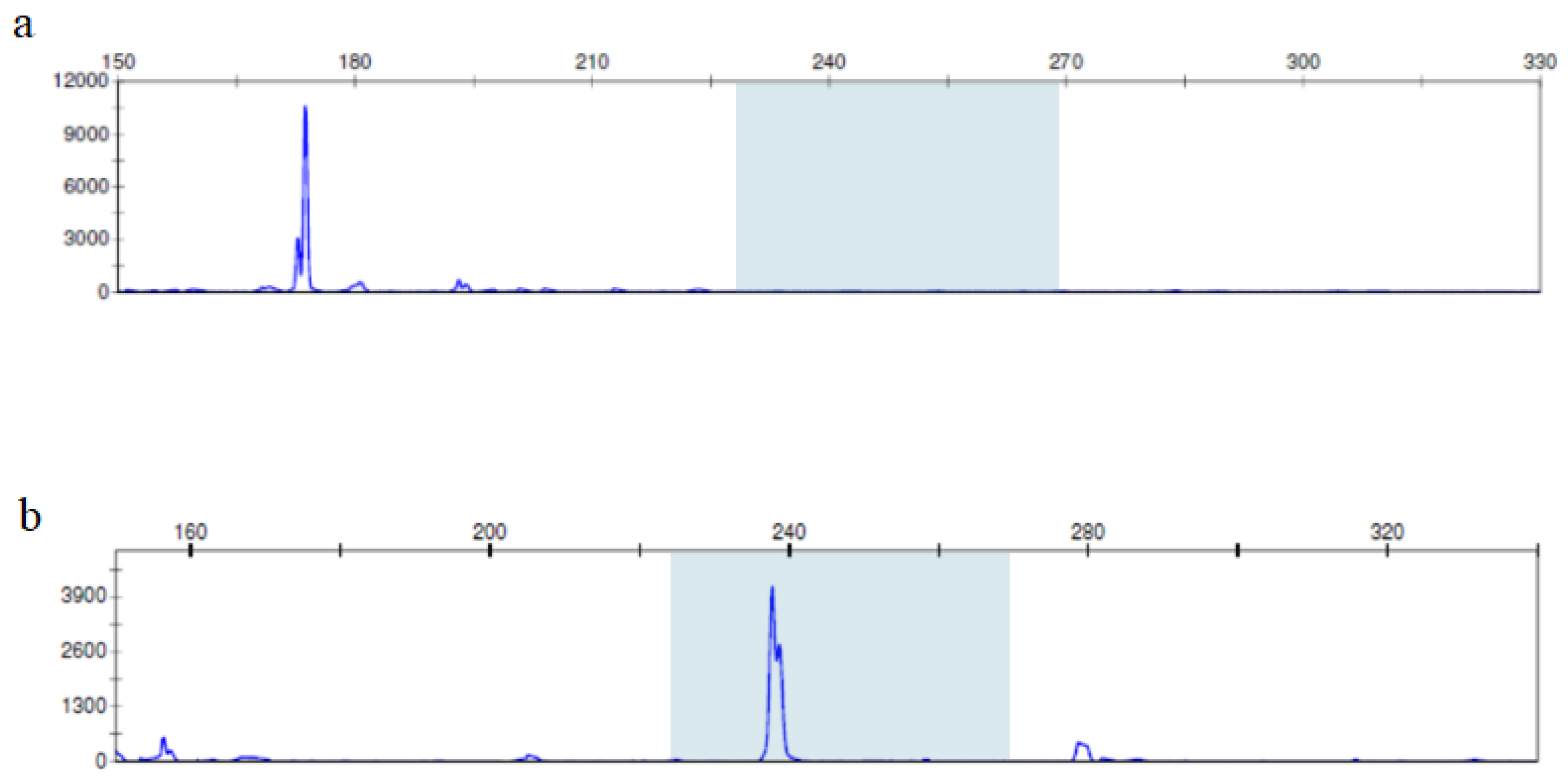
2.2.3. Fragment Analysis
2.3. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Muñoz, N.; Bosch, F.X.; De Sanjose, S.; Herrero, R.; Castellsagué, X.; Shah, K.V.; Snijders, P.J.; Meijer, C.J.L.M. Epidemiologic classification of human papillomavirus types associated with cervical cancer. N. Engl. J. Med. 2003, 348, 518–527. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bosch, F.X.; Manos, M.M.; Muñoz, N.; Sherman, M.; Jansen, A.M.; Peto, J.; Schiffman, M.H.; Moreno, V.; Kurman, R.; Shan, K.V.; et al. Prevalence of human papillomavirus in cervical cancer: A worldwide perspective. International biological study on cervical cancer (IBSCC) Study Group. J. Natl. Cancer Inst. 1995, 87, 796–802. [Google Scholar] [CrossRef]
- Khan, M.J.; Castle, P.E.; Lorincz, A.T.; Wacholder, S.; Sherman, M.; Scott, D.R.; Rush, B.B.; Glass, A.G.; Schiffman, M. The elevated 10-year risk of cervical precancer and cancer in women with human papillomavirus (HPV) type 16 or 18 and the possible utility of type-specific HPV testing in clinical practice. J. Natl. Cancer Inst. 2005, 97, 1072–1079. [Google Scholar] [CrossRef]
- Castle, P.E.; Hunt, W.C.; Langsfeld, E.; Wheeler, C.M. Three-year risk of cervical precancer and cancer after the detection of low-risk human papillomavirus genotypes targeted by a commercial test. Obs. Gynecol. 2014, 123, 49–56. [Google Scholar] [CrossRef] [Green Version]
- Giuliano, A.R.; Tortolero-Luna, G.; Ferrer, E.; Burchell, A.N.; de Sanjose, S.; Kjaer, S.K.; Muñoz, N.; Schiffman, M.; Bosch, F.X. Epidemiology of human papillomavirus infection in men, cancers other than cervical and benign conditions. Vaccine 2008, 26, K17–K28. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wright, T.C.J.; Stoler, M.H.; Sharma, A.; Zhang, G.; Behrens, C.; Wright, T.L. Evaluation of HPV-16 and HPV-18 genotyping for the triage of women with high-risk HPV+ cytology-negative results. Am. J. Clin. Pathol. 2011, 136, 578–586. [Google Scholar] [CrossRef] [PubMed]
- Thomsen, L.T.; Frederiksen, K.; Munk, C.; Junge, J.; Iftner, T.; Kjaer, S.K. Long-term risk of cervical intraepithelial neoplasia grade 3 or worse according to high-risk human papillomavirus genotype and semi-quantitative viral load among 33,288 women with normal cervical cytology. Int. J. Cancer 2015, 137, 193–203. [Google Scholar] [CrossRef] [PubMed]
- Clifford, G.M.; Smith, J.S.; Plummer, M.; Muñoz, N.; Franceschi, S. Human papillomavirus types in invasive cervical cancer worldwide: A meta-analysis. Br. J. Cancer 2003, 88, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Schiffman, M.; Castle, P.E.; Jeronimo, J.; Rodriguez, A.C.; Wacholder, S. Human papillomavirus and cervical cancer. Lancet 2007, 370, 890–907. [Google Scholar] [CrossRef]
- Lowy, D.R.; Solomon, D.; Hildesheim, A.; Schiller, J.T.; Schiffman, M. Human papillomavirus infection and the primary and secondary prevention of cervical cancer. Cancer 2008, 113, 1980–1993. [Google Scholar] [CrossRef]
- Ferlay, J.; Shin, H.R.; Bray, F.; Forman, D.; Mathers, C.; Parkin, D.M. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int. J. Cancer 2010, 127, 2893–2917. [Google Scholar] [CrossRef] [PubMed]
- Mayrand, M.H.; Duarte-Franco, E.; Rodrigues, I.; Walter, S.D.; Hanley, J.; Ferenczy, A.; Ratnam, S.; Coutlée, F.; Franco, E. Human papillomavirus DNA versus Papanicolaou screening tests for cervical cancer. N. Engl. J. Med. 2007, 357, 1579–1588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cuzick, J.; Clavel, C.; Petry, K.U.; Meijer, C.J.; Hoyer, H.; Ratnam, S.; Szarewski, A.; Birembaut, P.; Kulasingam, S.; Sasieni, P.; et al. Overview of the European and North American studies on HPV testing in primary cervical cancer screening. Int. J. Cancer 2006, 119, 1095–1101. [Google Scholar] [CrossRef] [PubMed]
- Wright, T.C., Jr.; Schiffman, M.; Solomon, D.; Cox, J.T.; Garcia, F.; Goldie, S.; Hatch, K.; Noller, K.L.; Roach, N.; Runowicz, C.; et al. Interim guidance for the use of human papillomavirus DNA testing as an adjunct to cervical cytology for screening. Obs. Gynecol. 2004, 103, 304–309. [Google Scholar] [CrossRef] [PubMed]
- Basu, P.; Mittal, S.; Bhadra Vale, D.; Chami Kharaji, Y. Secondary prevention of cervical cancer. Best. Pract. Res. Clin. Obs. Gynaecol. 2018, 47, 73–85. [Google Scholar] [CrossRef]
- Castle, P.E.; Stoler, M.H.; Wright, T.C., Jr.; Sharma, A.; Wright, T.L.; Behrens, C.M. Performance of carcinogenic human papillomavirus (HPV) testing and HPV16 or HPV18 genotyping for cervical cancer screening of women aged 25 years and older: A subanalysis of the ATHENA study. Lancet Oncol. 2011, 12, 880–890. [Google Scholar] [CrossRef]
- Stoler, M.H.; Wright, T.C., Jr.; Sharma, A.; Apple, R.; Gutekunst, K.; Wright, T.L. High-risk human papillomavirus testing in women with ASC-US cytology: Results from the ATHENA HPV study. Am. J. Clin. Pathol. 2011, 135, 468–475. [Google Scholar] [CrossRef] [Green Version]
- Barroeta, J.E.; Adhikari-Guragain, D.; Grotkowski, C.E. Cervical cancer screening in the era of HPV vaccination: A review of shifting paradigms in cytopathology. Diagn. Cytopathol. 2017, 45, 903–914. [Google Scholar] [CrossRef]
- Clavel, C.; Masure, M.; Levert, M.; Putaud, I.; Mangeonjean, C.; Lorenzato, M.; Nazeyrollas, P.; Gabriel, R.; Quereux, C.; Birembaut, P. Human papillomavirus detection by the hybrid capture II assay: A reliable test to select women with normal cervical smears at risk for developing cervical lesions. Diagn. Mol. Pathol. 2000, 9, 145–150. [Google Scholar] [CrossRef]
- Fujinaga, Y.; Shimada, M.; Okazawa, K.; Fukushima, M.; Kato, I.; Fujinaga, K. Simultaneous detection and typing of genital human papillomavirus DNA using the polymerase chain reaction. J. Gen. Virol. 1991, 72, 1039–1044. [Google Scholar] [CrossRef]
- Sasagawa, T.; Minemoto, Y.; Basha, W.; Yamazaki, H.; Nakamura, M.; Yoshimoto, H.; Sakaike, J.; Inoue, M. A new PCR-based assay amplifies the E6-E7 genes of most mucosal human papillomaviruses (HPV). Virus Res. 2000, 67, 127–139. [Google Scholar] [CrossRef]
- Tsiodras, S.; Georgoulakis, J.; Chranioti, A.; Voulgaris, Z.; Psyrri, A.; Tsivilika, A.; Panayiotides, J.; Karakitsos, P. Hybrid capture vs. PCR screening of cervical human papilloma virus infections. Cytological and histological associations in 1270 women. BMC Cancer 2010, 10, 53. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kulmala, S.M.; Syrjänen, S.; Shabalova, I.; Petrovichev, N.; Kozachenko, V.; Podistov, J.; Ivanchenko, O.; Zakharenko, S.; Nerovjna, R.; Kljukina, L.; et al. Human papillomavirus testing with the hybrid capture 2 assay and PCR as screening tools. J. Clin. Microbiol. 2004, 42, 2470–2475. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peyton, C.L.; Schiffman, M.; Lörincz, A.T.; Hunt, W.C.; Mielzynska, I.; Bratti, C.; Eaton, S.; Hildesheim, A.; Morera, L.A.; Rodriguez, A.C.; et al. Comparison of PCR- and hybrid capture-based human papillomavirus detection systems using multiple cervical specimen collection strategies. J. Clin. Microbiol. 1998, 36, 3248–3254. [Google Scholar] [CrossRef] [Green Version]
- Saini, R.; Shen, T.H.; Othman, N.H.; Santhanam, J.; Othman, N.; Tang, T.H. Evaluation of polymerase chain reaction (PCR) method and hybrid capture II (HCII) assay for the detection of human papillomavirus in cervical scrapings. Med. J. Malays. 2007, 62, 206–209. [Google Scholar]
- Kirschner, B.; Simonsen, K.; Junge, J. Comparison of conventional Papanicolaou smear and SurePath liquid-based cytology in the Copenhagen population screening programme for cervical cancer. Cytopathology 2006, 17, 187–194. [Google Scholar] [CrossRef]
- Roberts, J.M.; Thurloe, J.K. Comparative sensitivities of ThinPrep and Papanicolaou smear for adenocarcinoma in situ (AIS) and combined AIS/high-grade squamous intraepithelial lesion (HSIL): Comparison with HSIL. Cancer 2007, 111, 482–486. [Google Scholar] [CrossRef]
- Landis, J.R.; Koch, G.G. The measurement of observer agreement for categorical data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef] [Green Version]
- Yamazaki, H.; Sasagawa, T.; Basha, W.; Segawa, T.; Inoue, M. Hybrid capture-II and LCR-E7 PCR assays for HPV typing in cervical cytologic samples. Int. J. Cancer 2001, 94, 222–227. [Google Scholar] [CrossRef]
- de Freitas, A.C.; Gurgel, A.P.; Chagas, B.S.; Coimbra, E.C.; do Amaral, C.M. Susceptibility to cervical cancer: An overview. Gynecol. Oncol. 2012, 126, 304–311. [Google Scholar] [CrossRef]
- Meijer, C.J.; Berkhof, J.; Castle, P.E.; Hesselink, A.T.; Franco, E.L.; Ronco, G.; Arbyn, M.; Bosch, F.X.; Cuzick, J.; Dillner, J.; et al. Guidelines for human papillomavirus DNA test requirements for primary cervical cancer screening in women 30 years and older. Int. J. Cancer 2009, 124, 516–520. [Google Scholar] [CrossRef] [Green Version]
- Inoue, M.; Sakaguchi, J.; Sasagawa, T.; Tango, M. The evaluation of human papillomavirus DNA testing in primary screening for cervical lesions in a large Japanese population. Int. J. Gynecol. Cancer 2006, 16, 1007–1013. [Google Scholar] [CrossRef]
- Serrano, B.; Brotons, M.; Bosch, F.X.; Bruni, L. Epidemiology and burden of HPV-related disease. Best Pract. Res. Clin. Obs. Gynaecol. 2018, 47, 14–26. [Google Scholar] [CrossRef] [PubMed]
- Hwang, H.S.; Park, M.; Lee, S.Y.; Kwon, K.H.; Pang, M.G. Distribution and prevalence of human papillomavirus genotypes in routine pap smear of 2,470 korean women determined by DNA chip. Cancer Epidemiol. Biomark. Prev. 2004, 13, 2153–2156. [Google Scholar]
- An, H.J.; Cho, N.H.; Lee, S.Y.; Kim, I.H.; Lee, C.; Kim, S.J.; Mun, M.S.; Kim, S.H.; Jeong, J.K. Correlation of cervical carcinoma and precancerous lesions with human papillomavirus (HPV) genotypes detected with the HPV DNA chip microarray method. Cancer 2003, 97, 1672–1680. [Google Scholar] [CrossRef] [PubMed]
- Nishiwaki, M.; Yamamoto, T.; Tone, S.; Murai, T.; Ohkawara, T.; Matsunami, T.; Koizumi, M.; Takagi, Y.; Yamaguchi, J.; Kondo, N.; et al. Genotyping of human papillomaviruses by a novel one-step typing method with multiplex PCR and clinical applications. J. Clin. Microbiol. 2008, 46, 1161–1168. [Google Scholar] [CrossRef] [Green Version]
- Cox, J.T.; Lorincz, A.T.; Schiffman, M.H.; Sherman, M.E.; Cullen, A.; Kurman, R.J. Human papillomavirus testing by hybrid capture appears to be useful in triaging women with a cytologic diagnosis of atypical squamous cells of undetermined significance. Am. J. Obs. Gynecol. 1995, 172, 946–954. [Google Scholar] [CrossRef]
- Wong, A.K.; Chan, R.C.; Nichols, W.S.; Bose, S. Invader human papillomavirus (HPV) type 16 and 18 assays as adjuncts to HPV screening of cervical papanicolaou smears with atypical squamous cells of undetermined significance. Cancer 2009, 115, 823–832. [Google Scholar] [CrossRef] [PubMed]
- Safaeian, M.; Solomon, D.; Wacholder, S.; Schiffman, M.; Castle, P. Risk of precancer and follow-up management strategies for women with human papillomavirus-negative atypical squamous cells of undetermined significance. Obs. Gynecol. 2007, 109, 1325–1331. [Google Scholar] [CrossRef]
- Stern, P.L. Immune control of human papillomavirus (HPV) associated anogenital disease and potential for vaccination. J. Clin. Virol. 2005, 32, S72–S81. [Google Scholar] [CrossRef]
- Won, K.H.; Lee, J.Y.; Cho, H.Y.; Suh, D.H.; No, J.H.; Kim, Y.B. Impact of age on the false negative rate of human papillomavirus DNA test in patients with atypical squamous cells of undetermined significance. Obs. Gynecol. Sci. 2015, 58, 117–123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beyazit, F.; Sılan, F.; Gencer, M.; Aydin, B.; Paksoy, B.; Unsal, M.A.; Ozdemir, O. The prevelance of human papillomavirus (HPV) genotypes detected by PCR in women with normal and abnormal cervico-vaginal cytology. Ginekol. Pol. 2018, 89, 62–67. [Google Scholar] [CrossRef] [PubMed]
| Age (Years) | n (%) | PCR+ (High-Risk) | PCR+ (Low-Risk) | HC-II+ |
|---|---|---|---|---|
| 20–29 | 4 (6.8%) | 2 (28.6%) | 0 | 3 (33.3%) |
| 30–39 | 22 (37.3%) | 3 (42.9%) | 1 (50.0%) | 4 (44.4%) |
| 40–49 | 15 (25.4%) | 2 (28.6%) | 0 | 2 (22.2%) |
| 50–59 | 11 (18.6%) | 0 | 0 | 0 |
| 60–69 | 5 (8.5%) | 0 | 1 (50.0%) | 0 |
| 70–79 | 1 (1.7%) | 0 | 0 | 0 |
| 80–89 | 1 (1.7%) | 0 | 0 | 0 |
| Total | 59 | 7 | 2 | 9 |
| Range | 22–80 | 24–47 | 30–69 | 24–47 |
| Mean | 44.4 | 35.1 | 49.5 | 34.6 |
| Median | 45 | 35 | 49.5 | 35 |
| HC-II | Total | |||
|---|---|---|---|---|
| + | − | |||
| PCR | + | 7 | 0 | 7 |
| (78%) | (0%) | |||
| − | 2 | 50 | 52 | |
| (22%) | (100%) | |||
| Total | 9 | 50 | 59 | |
| Author | Year | n | Details of HPV Subtypes |
|---|---|---|---|
| Tsiodras S et al. [22] | 2010 | 1270 | 6, 11, 13, 16, 18, 30, 31, 32, 33, 35, 39, 40, 43, 45, 51, 52, 54, 55, 56, 58, 59, 66 |
| Kulmala SM et al. [23] | 2004 | 1511 | 16, 18, 31, 33, 35, 39, 45, 51, 52, 54, 56, 58 |
| Peyton CL et al. [24] | 1998 | 208 | 16, 18, 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, 68 (+20 subtypes: detail omission) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Saiki, Y.; Gion, Y.; Nishikori, A.; Norimatsu, Y.; Sato, Y. Comparison of the Hybrid Capture II Method with a PCR-Based Screening Method Using a Carboxyfluorescein-Labeled Primer for Detecting Human Papillomavirus in Cervicovaginal Liquid-Based Cytology. J. Mol. Pathol. 2020, 1, 9-18. https://doi.org/10.3390/jmp1010003
Saiki Y, Gion Y, Nishikori A, Norimatsu Y, Sato Y. Comparison of the Hybrid Capture II Method with a PCR-Based Screening Method Using a Carboxyfluorescein-Labeled Primer for Detecting Human Papillomavirus in Cervicovaginal Liquid-Based Cytology. Journal of Molecular Pathology. 2020; 1(1):9-18. https://doi.org/10.3390/jmp1010003
Chicago/Turabian StyleSaiki, Yusuke, Yuka Gion, Asami Nishikori, Yoshiaki Norimatsu, and Yasuharu Sato. 2020. "Comparison of the Hybrid Capture II Method with a PCR-Based Screening Method Using a Carboxyfluorescein-Labeled Primer for Detecting Human Papillomavirus in Cervicovaginal Liquid-Based Cytology" Journal of Molecular Pathology 1, no. 1: 9-18. https://doi.org/10.3390/jmp1010003
APA StyleSaiki, Y., Gion, Y., Nishikori, A., Norimatsu, Y., & Sato, Y. (2020). Comparison of the Hybrid Capture II Method with a PCR-Based Screening Method Using a Carboxyfluorescein-Labeled Primer for Detecting Human Papillomavirus in Cervicovaginal Liquid-Based Cytology. Journal of Molecular Pathology, 1(1), 9-18. https://doi.org/10.3390/jmp1010003





