A Statistical Examination of Distinct Characteristics Influencing the Performance of Vector-Borne Epidemiological Agent-Based Simulation Models
Abstract
:1. Introduction
2. Materials and Methods
| Overview | Purpose | The model developed in this study, which represents a simplified abstraction of reality, was designed to investigate and highlight the importance of three different factors, namely, data quality, human behavior, and multi-strain, multi-vector, when developing epidemiological models. Rather than developing a detailed representation of an epidemic to predict the outcomes of future epidemics or to understand past epidemics, the goal was to call the attention of the academic community to the importance of investigating the aforementioned factors more in depth when developing epidemiological models. |
| State variables and scales | Agents: (i) humans, (ii) mosquitoes, and (iii) the environment. In this simplified abstraction, individuals and mosquitoes were randomly distributed within the environment. The epidemiological parameters, such as latent rate and recovery rate, were considered independent of age, gender, time, or any other parameters. Detailed information about the parameters used in each model is provided in Section 2.1. | |
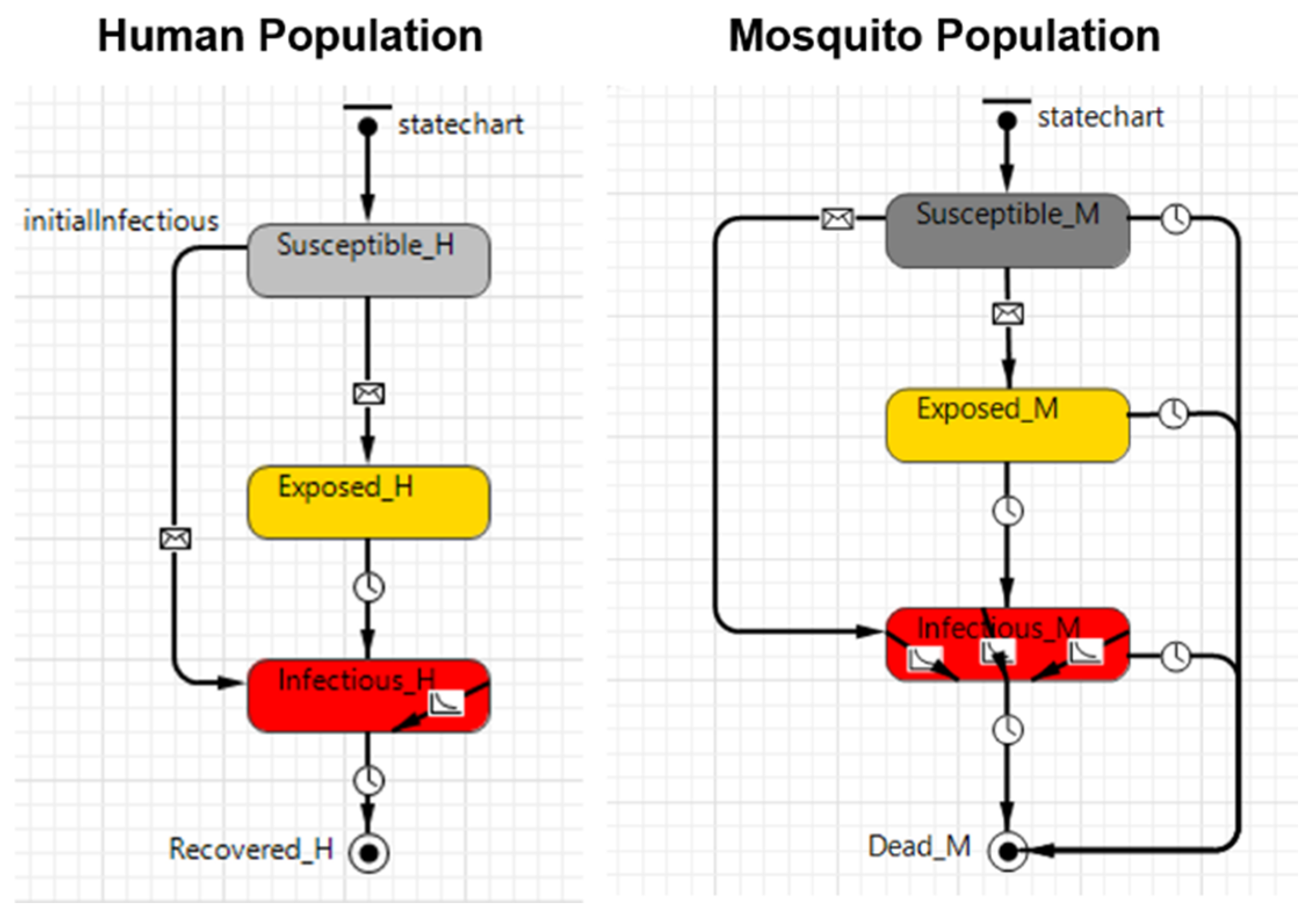
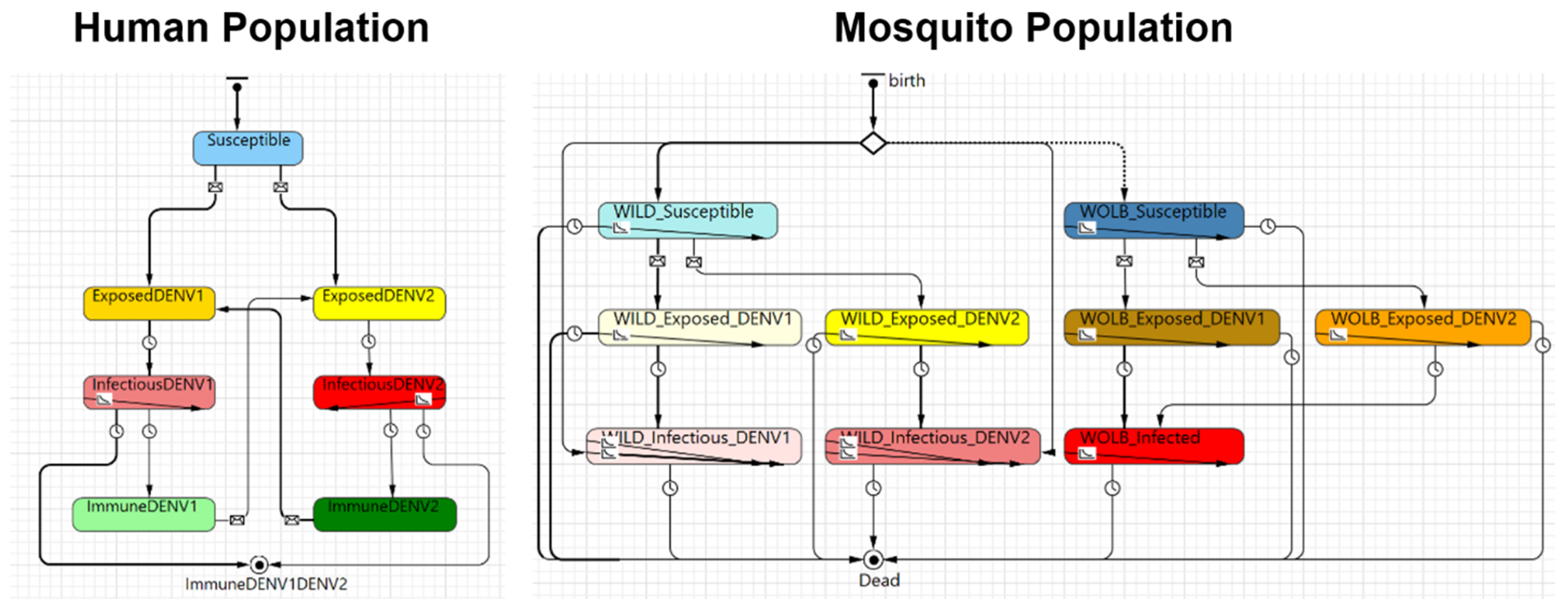
| Process overview and scheduling | In vector-borne diseases, there are generally three types of agents: (i) the pathogen, which may be a virus or bacteria; (ii) the vector, in this work a mosquito that can be either Aedes aegypti or Aedes albopictus; and, (iii) the final host, a human in this case. The model was built based on the traditional susceptible, exposed, infectious, and recovered (SEIR) and susceptible, exposed, and infectious (SEI) compartmental models. The SEIR model was used to represent humans, while the SEI model was used to represent the vector. The baseline conceptual model represented by the SEIR-SEI compartmental model is shown in Figure 1, and Table 2 provides the definition of the symbols presented in Figure 1. The life cycle of the pathogen can be described in four stages: (1) The pathogen is transmitted from an infectious mosquito (Mi) to a susceptible host (Hs) when the mosquito feeds on human blood. (2) The pathogen infects the exposed host (He), who still has no ability to transmit the disease to another mosquito. After the latent period, the pathogen reaches sufficiently high densities in the blood of the infectious host (Hi), who is now able to infect another susceptible mosquito (Ms). (3) Whenever a susceptible mosquito feeds on an infectious host, the susceptible mosquito inoculates the pathogen and becomes an exposed mosquito (Me). Similar to the host, the mosquito is not able to immediately transmit the disease to other susceptible hosts. (4) After the latent period, the pathogen develops in the mosquito to the point that the pathogen becomes present in the salivary glands of the mosquito, who becomes an infectious mosquito (Mi) and can now transmit the disease by biting a susceptible host. After the recovery period, the host is considered to be recovered (Hr) and immune to the pathogen (this is not true for all mosquito-borne diseases but it is true for some of them, such as dengue with respect to the same virus strain and chikungunya). An infectious mosquito never recovers from the disease and will stay infectious until its death. Three different models were developed in this work to meet the proposed goal. Each model works slightly differently and they are described in detail in Section 2.1.1, Section 2.1.2, Section 2.1.3 and Section 2.1.4. The description provided above refers to the baseline model (Model A). Figure 2, Figure 3 and Figure 4 shows the simulation model snapshot of the human population module and the mosquito population module of each one of these modules. The transition between each state is based on the epidemiological parameters as shown in Figure 1 and described in Table 2 and Table 3. The data were collected at the end of each day on the basis of an event. | |
| Design concepts | Design concepts | Stochasticity was considered in all models through the infectious disease parameters used as input. These data are provided in Table 4, Table 5, Table 6 and Table 7. Adaptation was considered in Model C when humans changed their behavior in response to the total number of infected individuals, either instantaneously or after a specific amount of time. This was considered at an individual and a population level. In summary, human behavior was considered to change in four different situations, as described in Section 2.1.3. |
| Details | Initialization | Humans and mosquitoes were uniformly randomly distributed in the continuous space. The human and mosquito population size and the initial number of infectious humans and mosquitoes are provided in Table 4, Table 5, Table 6 and Table 7. |
| Input | The input data used in the model were defined within the model and they are provided in Table 4, Table 5, Table 6 and Table 7. | |
| Sub-models | This consists of the “skeleton” of the model, as well as its description, which are provided in Figure 1, Figure 2, Figure 3 and Figure 4 and Section 2. |
| # | Symbol | Definition |
|---|---|---|
| 1 | Th | Human population size |
| 2 | Tm | Mosquito population size |
| 3 | βh | Infectivity rate from mosquito to human |
| 4 | βm | Infectivity rate from human to mosquito |
| 5 | λh | Human latent rate |
| 6 | λm | Mosquito latent rate |
| 7 | γ | Human recovery rate |
| 8 | μ | Mosquito mortality rate |
| 9 | Hs | Number of susceptible humans |
| 10 | He | Number of exposed humans |
| 11 | Hi | Number of infectious humans |
| 12 | Hr | Number of recovered and immune humans |
| 13 | Ms | Number of susceptible mosquitoes |
| 14 | Me | Number of exposed mosquitoes |
| 15 | Mi | Number of infectious mosquitoes |


2.1. Experiment Design
| # | Agent | Parameter | If the Parameter is Used in the Model: | |||
|---|---|---|---|---|---|---|
| A | B | C | D | |||
| 1 | Human | Human population size | X | X | X | X |
| 2 | Human | Daily human to mosquito infect rate for DENV1 | X | X | X | X |
| 3 | Human | Daily human latent rate for DENV1 | X | X | X | X |
| 4 | Human | Daily human recovery rate for DENV1 | X | X | X | X |
| 5 | Mosquito | Mosquito population size | X | X | X | X |
| 6 | Mosquito | Daily wild mosquito to human infect rate for DENV1 | X | X | X | X |
| 7 | Mosquito | Daily wild mosquito latent rate for DENV1 | X | X | X | X |
| 8 | Mosquito | Daily wild mosquito mortality/birth rate | X | X | X | X |
| 9 | Human | Daily human to mosquito infect rate for DENV2 | X | |||
| 10 | Human | Daily human latent rate for DENV2 | X | |||
| 11 | Human | Daily human recovery rate for DENV2 | X | |||
| 12 | Mosquito | Daily wild mosquito to human infect rate for DENV2 | X | |||
| 13 | Mosquito | Daily wild mosquito latent rate for DENV2 | X | |||
| 14 | Mosquito | Daily Wolbachia-carrier mosquito latent rate for DENV1 | X | |||
| 15 | Mosquito | Daily Wolbachia-carrier mosquito latent rate for DENV2 | X | |||
| 16 | Mosquito | Daily Wolbachia-carrier mosquito mortality/birth rate | X | |||
| 17 | Mosquito | Mosquito type (wild or Wolbachia-carrier) | X | |||
| 18 | Mosquito | Proportion of wild and Wolbachia-carrier mosquitoes | X | |||
| 19 | Human | Initial number of DENV1 infectious humans | X | X | X | X |
| 20 | Human | Initial number of DENV2 infectious humans | X | |||
| 21 | Mosquito | Initial number of wild DENV1 infectious mosquitoes | X | X | X | X |
| 22 | Mosquito | Initial number of wild DENV2 infectious mosquitoes | X | |||
| 23 | Human | Exposed to DENV1 (0 or 1/no or yes) | X | |||
| 24 | Human | Exposed to DENV2 (0 or 1/no or yes) | X | |||
| 25 | Human—Population | Include same behavior for the whole population [0—False, 1—True] | X | |||
| 26 | Human—Individual | Include behavior for individual [0—False, 1—True] | X | |||
| 27 | Human | Percent of infectious individuals to trigger cautious behavior [%] | X | |||
| 28 | Human—Population | Population cautious factor [%] | X | |||
| 29 | Human—Individual | Individual cautious factor [%] | X | |||
| 30 | Human | Include time to switch behavior [0—False, 1—True] | X | |||
| 31 | Human—Population | Population time to switch behavior [days] | X | |||
| 32 | Human—Individual | Individual time to switch behavior [days] | X | |||
2.1.1. Model A—Baseline (Single-Strain, Single-Vector Dengue Spread Model)
| # | Input Data Description | Scenario #01 | Scenario #02 |
|---|---|---|---|
| 1 | Mosquito population size | 200 | 2000 |
| 2 | Initial number of infectious mosquitoes | 10 | 100 |
| 3 | Mosquito daily latent rate | Uniform (0.083, 0.125) | |
| 4 | Mosquito daily mortality rate | Uniform (0.033, 0.05) | |
| 5 | Daily infectivity rate from mosquito to human | Uniform (0.1, 0.18) | |
| 6 | Human population size | 100 | 1000 |
| 7 | Initial number of infectious humans | 0 | |
| 8 | Human daily latent rate | Uniform (0.1,0.25) | |
| 9 | Human daily recovery rate | Uniform (0.143, 0.45) | |
| 10 | Daily infectivity rate from human to mosquito | Uniform (0.36, 0.44) | |
2.1.2. Model B (Data Quality)
| # | Input Data Description | Parameter Level | ||||
|---|---|---|---|---|---|---|
| B1 | B2 | C1 | C2 | C3 | ||
| 1 | Mosquito population size | 200 | 2000 | 100 | 500 | 5000 |
| 2 | Initial number of infectious mosquitoes | 10 | 100 | 25 | 50 | 500 |
| 3 | Mosquito daily latent rate | Uniform (0.083, 0.125) | Uniform (0.05, 0.500) | Uniform (0.125, 0.500) | Uniform (0.05, 0.083) | |
| 4 | Mosquito daily mortality rate | Uniform (0.033, 0.05) | Uniform (0.020, 0.07) | Uniform (0.05, 0.07) | Uniform (0.020, 0.033) | |
| 5 | Daily infectivity rate from mosquito to human | Uniform (0.1, 0.18) | Uniform (0.07, 0.22) | Uniform (0.18, 0.22) | Uniform (0.07, 0.1) | |
| 6 | Human population size | 100 | 1000 | 500 | 2000 | 5000 |
| 7 | Initial number of infectious humans | 0 | 10 | 50 | 100 | |
| 8 | Human daily latent rate | Uniform (0.1,0.25) | Uniform (0.05, 0.500) | Uniform (0.25, 0.500) | Uniform (0.05, 0.1) | |
| 9 | Human daily recovery rate | Uniform (0.143, 0.450) | Uniform (0.070, 0.600) | Uniform (0.450, 0.600) | Uniform (0.070, 0.143) | |
| 10 | Daily infectivity rate from human to mosquito | Uniform (0.36, 0.44) | Uniform (0.20, 0.80) | Uniform (0.44,0.80) | Uniform (0.20,0.36) | |
- Mosquito population size: variation in mosquito population size may represent either the adoption of control strategies (e.g., insecticide use), the elimination of mosquito breeding sites (e.g., cleaning pots with standing water), climatic variation (e.g., increased rainfall and temperature that favor the reproduction of mosquitoes), or errors in estimating the mosquito population through techniques such as mosquito trap.
- Initial number of infectious mosquitoes: this parameter was varied to represent regions in which the disease is imported by travelers who bring infectious mosquitoes to the area and regions where the disease is endemic.
- Mosquito daily latent rate, mosquito daily mortality rate, human daily latent rate, and human daily recovery rate: considered low and high rates, on the basis of the values found in the literature, as well as low and high variation. The variations of these rates represent the existence of several types of virus that can reproduce in mosquitoes and humans more slowly or quickly and, consequently, also affect the human recovery rate; the genetic and immune variation of humans and mosquitoes; the use of medical treatment that affects the recovery rate of individuals; and climatic variations and use of control measures, such as the use of screens in windows and insecticides, which may alter the mortality rate of mosquitoes.
- Daily infectivity rate from mosquito to human: the variations of this rate are due to reasons similar to mosquito daily latent rate, such as the existence of different types of virus, and genetic and immune variation of mosquitoes.
- Human population size: the size was varied to represent different neighborhoods or sizes of cities.
- Initial number of infectious humans: this parameter was varied for reasons similar to mosquito population size and initial number of infectious mosquitoes. The scenarios in Table 5 with initial number of infectious humans equal to 0 are equivalent to an epidemic-free population where a new epidemic is normally carried by a mosquito brought from an epidemic area. Besides representing an epidemic-free society and a society in which the disease is endemic, the variation may also represent large events, such as big sporting events, music events, or refugee entry into a region, which can lead to several cases imported at a single time.
- Daily infectivity rate from human to mosquito: the variations of this rate were for reasons similar to mosquito daily latent rate and daily infectivity rate from mosquito to human, such as the existence of different types of virus, genetic and immune variation of humans, and use of medical treatment.
2.1.3. Model C (Coupled Human Behavior and Dengue Spread Model)
| # | Input Data Description | Input Data Value | |||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | Mosquito population size | 200 | 2000 | ||||||
| 2 | Initial number of infectious mosquitoes | 10 | 100 | ||||||
| 3 | Mosquito daily latent rate | Uniform (0.083, 0.125) | |||||||
| 4 | Mosquito daily mortality rate | Uniform (0.033, 0.05) | |||||||
| 5 | Daily infectivity rate from mosquito to human | Uniform (0.1, 0.18) | |||||||
| 6 | Human population size | 100 | 1000 | ||||||
| 7 | Initial number of infectious humans | 0 | |||||||
| 8 | Human daily latent rate | Uniform (0.1,0.25) | |||||||
| 9 | Human daily recovery rate | Uniform (0.143, 0.45) | |||||||
| 10 | Daily infectivity rate from human to mosquito | Uniform (0.36, 0.44) | |||||||
| 11 | Percent of infectious individuals to trigger cautious behavior [%] | 5 | 15 | 5 | 15 | 5 | 15 | 5 | 15 |
| 12 | Cautious factor [%] * | Uniform (0.6, 0.9) | Uniform (0.8, 0.9) | Uniform (0.6, 0.9) | Uniform (0.8, 0.9) | ||||
| 13 | Time to switch behavior [days] | Uniform (1,3) | Uniform (1,7) | Uniform (1,3) | Uniform (1,7) | ||||
2.1.4. Model D (Multi-Strain, Multi-Vector Dengue Spread Model)
| # | Input Data Description | Input Data Value | |||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | Multi-strain only | Mosquito population size | Wild | 100 | 200 | 800 | 1000 | 2000 | 8000 |
| Wolbachia-carrier | 0 | ||||||||
| 2 | Initial number of wild infectious mosquitoes [%] | DENV1 | 2.5 | 2.5 | 2.5 | 2.5 | 2.5 | 2.5 | |
| DENV2 | 2.5 | 2.5 | 2.5 | 2.5 | 2.5 | 2.5 | |||
| 1 | Multi-vector only | Mosquito population size | Wild | 50 | 100 | 400 | 500 | 1000 | 4000 |
| Wolbachia-carrier | 50 | 100 | 400 | 500 | 1000 | 4000 | |||
| 2 | Initial number of wild infectious mosquitoes [%] | DENV1 | 5 | 10 | 40 | 50 | 100 | 400 | |
| DENV2 | 0 | ||||||||
| 1 | Multi-strain-multi-vector | Mosquito population size | Wild | 50 | 100 | 400 | 500 | 1000 | 4000 |
| Wolbachia-carrier | 50 | 100 | 400 | 500 | 1000 | 4000 | |||
| 2 | Initial number of wild infectious mosquitoes [%] | DENV1 | 2.5 | 2.5 | 2.5 | 2.5 | 2.5 | 2.5 | |
| DENV2 | 2.5 | 2.5 | 2.5 | 2.5 | 2.5 | 2.5 | |||
| 3 | Mosquito daily latent rate | Uniform (0.083, 0.125) | |||||||
| 4 | Mosquito daily mortality rate | Uniform (0.033, 0.05) | |||||||
| 5 | Daily infectivity rate from mosquito to human | Uniform (0.1, 0.18) | |||||||
| 6 | Human population size | 100 | 1000 | ||||||
| 7 | Initial number of infectious humans | 0 | |||||||
| 8 | Human daily latent rate | Uniform (0.1,0.25) | |||||||
| 9 | Human daily recovery rate | Uniform (0.143, 0.45) | |||||||
| 10 | Daily infectivity rate from human to mosquito | Uniform (0.36, 0.44) | |||||||
3. Results
3.1. Baseline
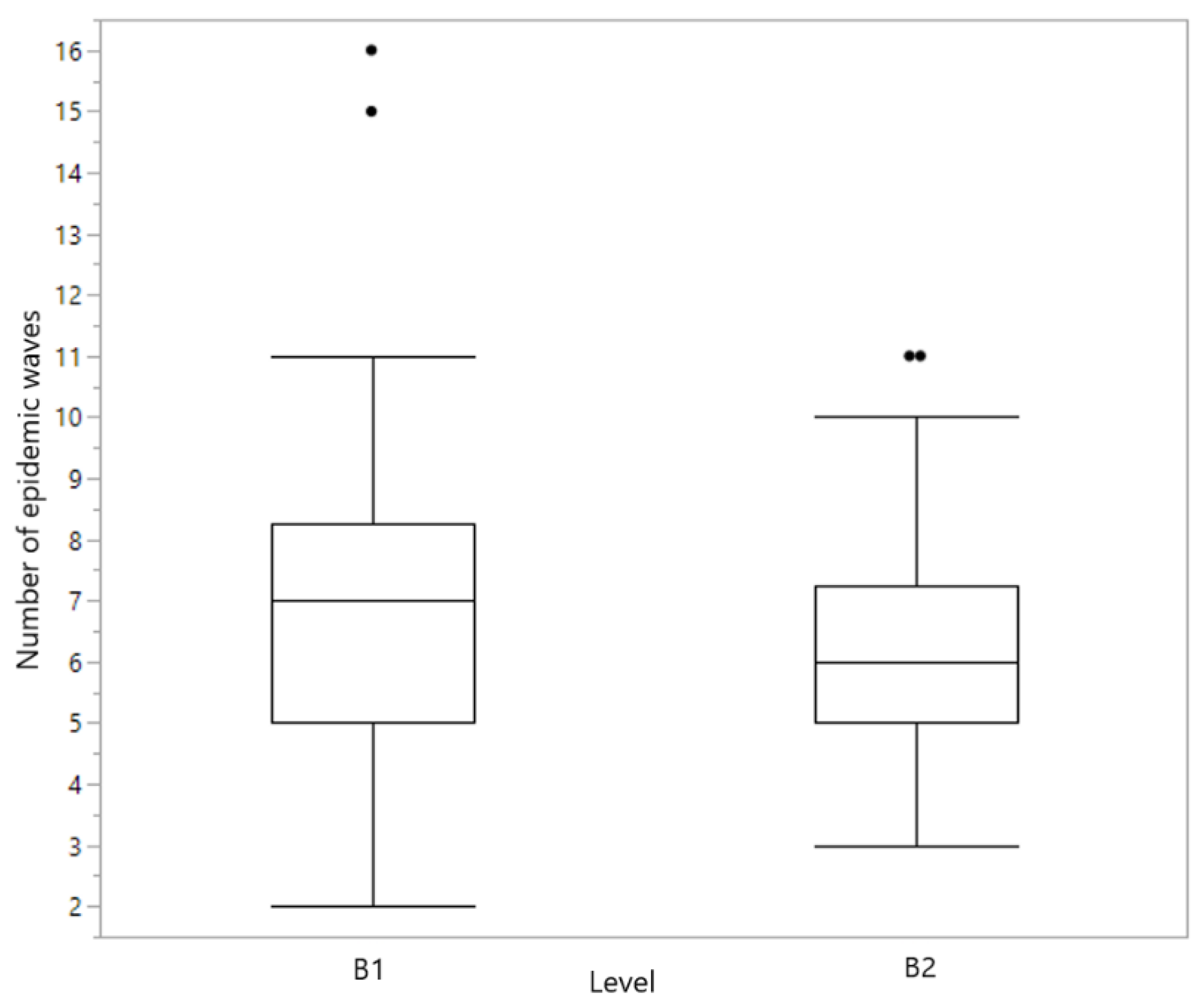
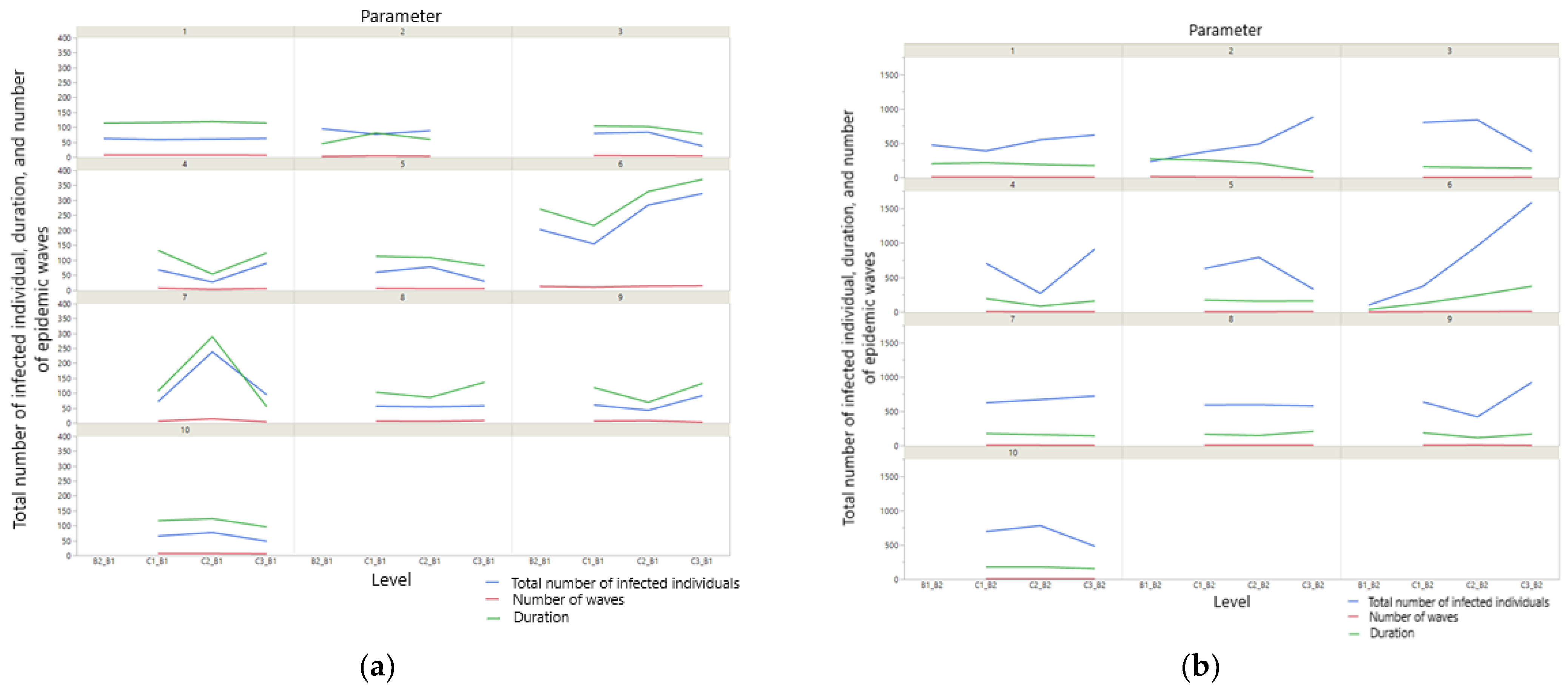
3.2. Impact of Data Quality
- Mosquito population size:
- 2.
- Initial number of infectious mosquitoes:
- 3.
- Mosquito daily latent rate:
- 4.
- Mosquito daily mortality rate:
- 5.
- Daily infectivity rate from mosquito to human:
- 6.
- Human population size:
- 7.
- Initial number of infectious humans:
- 8.
- Human daily latent rate:
- 9.
- Human daily recovery rate:
- 10.
- Daily infectivity from human to mosquito:
3.3. Impact of Human Behavior
3.4. Impact of Multi-Vector and Multi-Strain
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Santos, C.F. Reflections about the impact of the SARS-COV-2/COVID-19 pandemic on mental health. Braz. J. Psychiatry 2020, 42, 329. [Google Scholar] [CrossRef] [Green Version]
- Landry, M.D.; Geddes, L.; Moseman, A.P.; Lefler, J.P.; Raman, S.R.; van Wijchen, J. Early reflection on the global impact of COVID19, and implications for physiotherapy. Physiotherapy 2020, 107, A1–A3. [Google Scholar] [CrossRef]
- World Health Organization. A Year without Precedent: WHO’s COVID-19 Response. Available online: https://www.who.int/news-room/spotlight/a-year-without-precedent-who-s-covid-19-response (accessed on 18 January 2021).
- Venkatramanan, S.; Lewis, B.; Chen, J.; Higdon, D.; Vullikanti, A.; Marathe, M. Using data-driven agent-based models for forecasting emerging infectious diseases. Epidemics 2018, 22, 43–49. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Health in 2015: From MDGs, Millennium Development Goals to SDGs, Sustainable Development Goals. Available online: https://apps.who.int/iris/bitstream/handle/10665/200009/9789241565110_eng.pdf?sequence=1&isAllowed=y (accessed on 17 November 2020).
- Lamb, A.; Paul, M.J.; Dredze, M. Investigating Twitter as a Source for Studying Behavioral Responses to Epidemics. In Proceedings of the AAAI Fall Symposium: Information Retrieval and Knowledge Discovery in Biomedical Text, Arlington, VA, USA, 2–4 November 2012; Citeseer: Princeton, NJ, USA, 2012. [Google Scholar]
- Gozzi, N.; Perrotta, D.; Paolotti, D.; Perra, N. Towards a data-driven characterization of behavioral changes induced by the seasonal flu. PLoS Comput. Biol. 2020, 16, e1007879. [Google Scholar] [CrossRef]
- World Health Organization. Sierra Leone: A Traditional Healer and a Funeral. Available online: https://www.who.int/csr/disease/ebola/ebola-6-months/sierra-leone/en/ (accessed on 18 January 2021).
- vom Steeg, L.G.; Klein, S.L. SeXX matters in infectious disease pathogenesis. PLoS Pathog. 2016, 12, e1005374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tolhurst, R.; De Koning, K.; Price, J.; Kemp, J.; Theobald, S.; Squire, S.B. The challenge of infectious disease: Time to take gender into account. J. Health Manag. 2002, 4, 135–151. [Google Scholar] [CrossRef]
- Stemple, L.; Karegeya, P.; Gruskin, S. Human rights, gender, and infectious disease: From HIV/AIDS to Ebola. Hum. Rights Q. 2016, 38, 993–1021. [Google Scholar] [CrossRef]
- Zhou, M.; Zhang, N.; Zhang, M.; Ma, G. Culture, eating behavior, and infectious disease control and prevention. J. Ethn. Foods 2020, 7, 1–7. [Google Scholar]
- Evans, J.C.; Silk, M.J.; Boogert, N.J.; Hodgson, D.J. Infected or informed? Social structure and the simultaneous transmission of information and infectious disease. Oikos 2020, 129, 1271–1288. [Google Scholar] [CrossRef]
- Agaba, G.O.; Kyrychko, Y.N.; Blyuss, K.B. Mathematical model for the impact of awareness on the dynamics of infectious diseases. Math. Biosci. 2017, 286, 22–30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Greenhalgh, D.; Rana, S.; Samanta, S.; Sardar, T.; Bhattacharya, S.; Chattopadhyay, J. Awareness programs control infectious disease–multiple delay induced mathematical model. Appl. Math. Comput. 2015, 251, 539–563. [Google Scholar] [CrossRef] [Green Version]
- Lipsitch, M.; Santillana, M. Enhancing Situational Awareness to Prevent Infectious Disease Outbreaks from Becoming Catastrophic. In Global Catastrophic Biological Risks; Inglesby, T.V., Adalja, A.A., Eds.; Springer International Publishing: Cham, Switzerland, 2019; pp. 59–74. [Google Scholar]
- Huaman, M.A.; Araujo-Castillo, R.V.; Soto, G.; Neyra, J.M.; Quispe, J.A.; Fernandez, M.F.; Mundaca, C.C.; Blazes, D.L. Impact of two interventions on timeliness and data quality of an electronic disease surveillance system in a resource limited setting (Peru): A prospective evaluation. BMC Med. Inform. Decis. Mak. 2009, 9, 16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, J.C.; Sampson, L.A. High rates of incarceration as a social force associated with community rates of sexually transmitted infection. J. Infect. Dis. 2005, 191, S55–S60. [Google Scholar] [CrossRef] [Green Version]
- Marathe, M.V.; Ramakrishnan, N. Recent advances in computational epidemiology. IEEE Intell. Syst. 2013, 28, 96–101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- World Health Organization. Medicines in Health Systems: Advancing Access, Affordability and Appropriate Use. Available online: https://apps.who.int/iris/bitstream/handle/10665/179197/9789241507622_eng.pdf?sequence=1&isAllowed=y (accessed on 15 November 2020).
- Centers for Disease Control and Prevention. Epidemiology: Transmission of the Dengue Virus. Available online: https://www.cdc.gov/dengue/epidemiology/index.html (accessed on 9 September 2017).
- European Centre for Disease Prevention and Control. How is Zika Virus Transmitted? Available online: https://ecdc.europa.eu/en/publications-data/how-zika-virus-transmitted (accessed on 10 September 2017).
- Pan American Health Organization. Tool for the Diagnosis and Care of Patients with Suspected Arboviral Diseases. Available online: https://iris.paho.org/handle/10665.2/33895 (accessed on 12 November 2017).
- Aguiar, M.; Stollenwerk, N.; Halstead, S.B. The risks behind Dengvaxia recommendation. Lancet Infect. Dis. 2016, 16, 882–883. [Google Scholar] [CrossRef]
- Ferguson, N.M.; Rodríguez-Barraquer, I.; Dorigatti, I.; Mier-y-Teran-Romero, L.; Laydon, D.J.; Cummings, D.A.T. Benefits and risks of the Sanofi-Pasteur dengue vaccine: Modeling optimal deployment. Science 2016, 353, 1033–1036. [Google Scholar] [CrossRef] [Green Version]
- Grimm, V.; Berger, U.; Bastiansen, F.; Eliassen, S.; Ginot, V.; Giske, J.; Goss-Custard, J.; Grand, T.; Heinz, S.K.; Huse, G.; et al. A standard protocol for describing individual-based and agent-based models. Ecol. Model. 2006, 198, 115–126. [Google Scholar] [CrossRef]
- World Health Organization. Dengue: Guidelines for Diagnosis, Treatment, Prevention and Control. Available online: https://apps.who.int/iris/bitstream/handle/10665/44188/9789241547871_eng.pdf?sequence=1&isAllowed=y (accessed on 15 November 2020).
- World Health Organization. Dengue Control. Available online: http://www.who.int/denguecontrol/human/en/ (accessed on 20 November 2017).
- Araújo, H.R.C.D. Caracterização Morfológica dos Hemócitos do Aedes Aegypti e do Aedes Albopictus e a Resposta Imune dos Hemócitos do Aedes Aegypti Após a Infecção pelo Dengue Virus. Ph.D. Thesis, Fundação Oswaldo Cruz, Centro de Pesquisas René Rachou, Belo Horizonte/MG, Brazil, 2011. [Google Scholar]
- Yakob, L.; Clements, A.C. A mathematical model of chikungunya dynamics and control: The major epidemic on Reunion Island. PLoS ONE 2013, 8, e57448. [Google Scholar] [CrossRef] [PubMed]
- Ross, R.; Thomson, D. Some enumerative studies on malarial fever. Proc. R. Soc. Lond. 1910, 83, 159–173. [Google Scholar] [CrossRef]
- Kermack, W.O.; McKendrick, A.G. A contribution to the mathematical theory of epidemics. Proc. R. Soc. Lond. A Math. Phys. Eng. Sci. 1927, 115, 700–721. [Google Scholar]
- Dumont, Y.; Chiroleu, F.; Domerg, C. On a temporal model for the Chikungunya disease: Modeling, theory and numerics. Math. Biosci. 2008, 213, 80–91. [Google Scholar] [CrossRef] [PubMed]
- World Mosquito Program. Wolbachia. Available online: http://www.eliminatedengue.com/our-research/wolbachia (accessed on 10 October 2017).
- Murray, J.V.; Jansen, C.C.; De Barro, P. Risk Associated with the Release of Wolbachia-Infected Aedes aegypti Mosquitoes into the Environment in an Effort to Control Dengue. Front. Public Health 2016, 4, 1–12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scheidegger, A.P.G.; Banerjee, A. An agent-based model to investigate behavior impacts on vector-borne disease spread. In Proceedings of the 2017 Winter Simulation Conference (WSC), Las Vegas, NV, USA, 3–6 December 2017; IEEE: Piscataway, NJ, USA, 2017. [Google Scholar] [CrossRef]



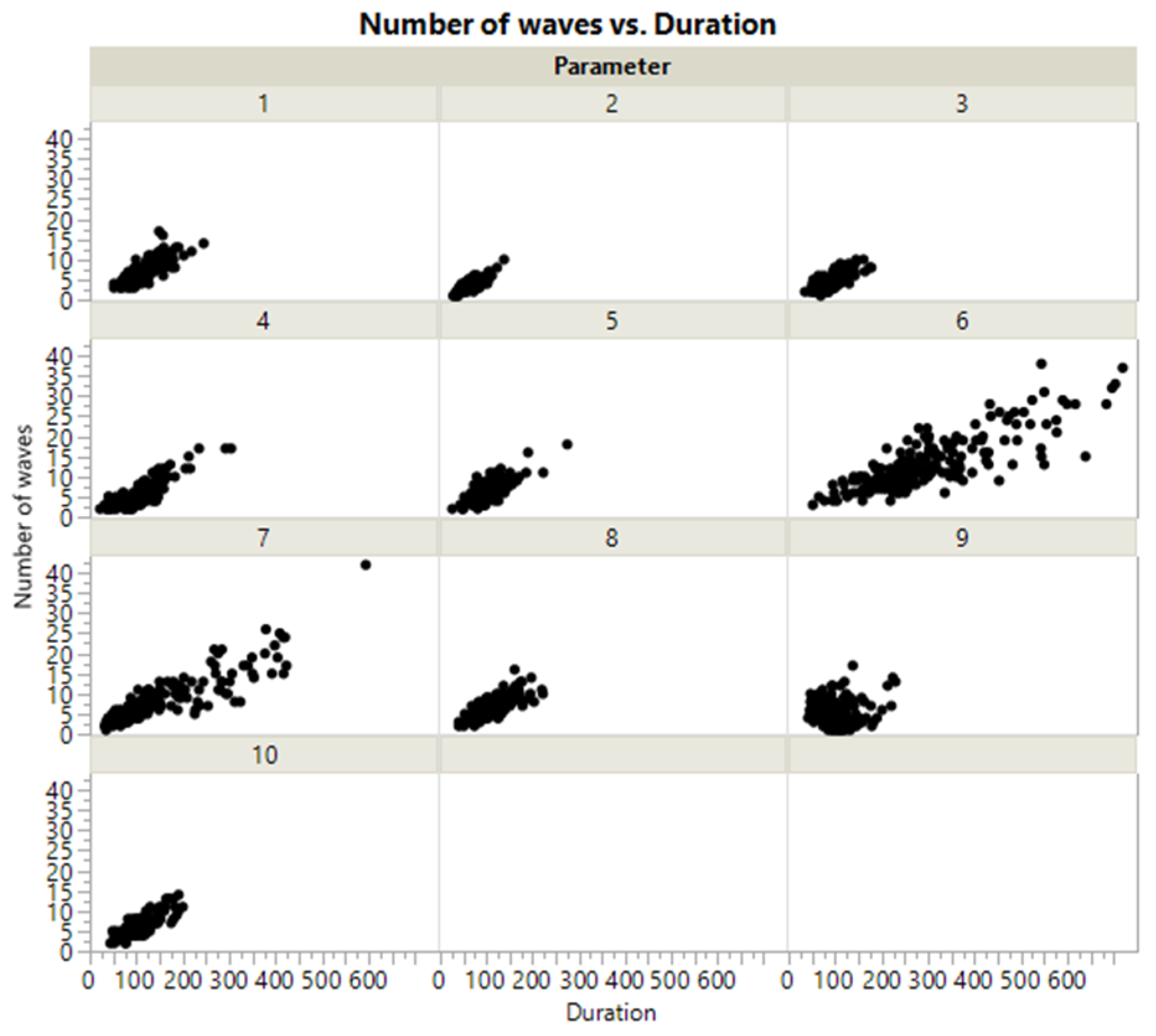
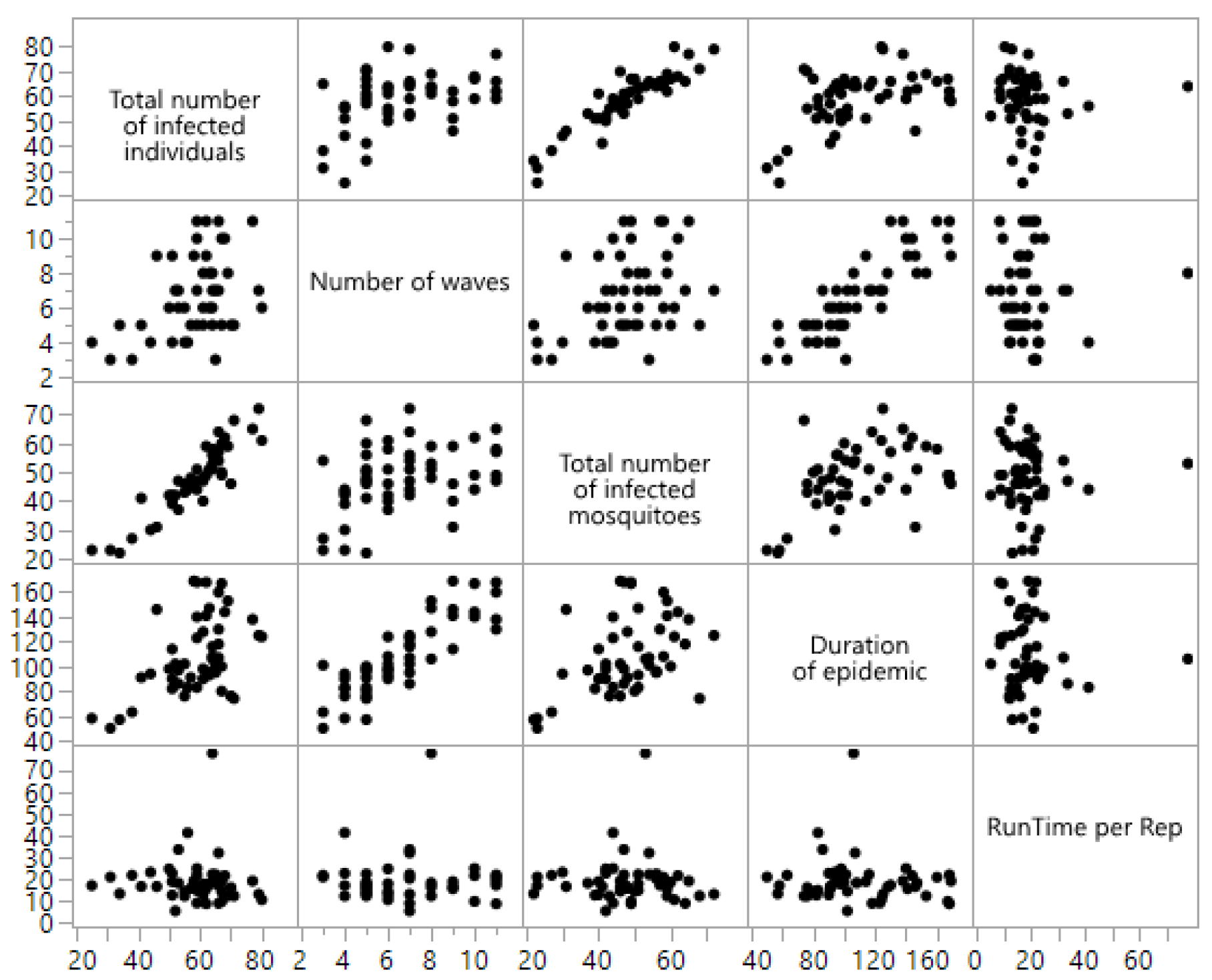
| Output Response | p-Value | 95% Confidence Interval | Conclusion |
|---|---|---|---|
| Total number of infected individuals | <0.0001 | [545.849, 564.511] | Statistically different |
| Duration of epidemic in days | <0.0001 | [55.313, 82.127] | Statistically different |
| Number of epidemic waves | 0.1429 | [−1.734, 0.254] | Not statistically different |
| Runtime per replication in seconds | <0.0001 | [59.794, 62.100] | Statistically different |
| # | Input Data Description | Total Number of Infected Individuals | Number of Epidemic Waves | Duration of the Epidemic in Days | Runtime Per Replication in Seconds | ||||
| B1 | B2 | B1 | B2 | B1 | B2 | B1 | B2 | ||
| Model p-Value | Model p-Value | Model p-Value | Model p-Value | Model p-Value | MODEL P-VALUE | Model p-Value | Model p-Value | ||
| <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | ||
| Input p-Value | Input p-Value | Input p-Value | Input p-Value | Input p-Value | Input p-Value | Input p-Value | Input p-Value | ||
| 1 | Mosquito population size | 0.9502 | <0.0001 | 0.9747 | <0.0001 | 0.9759 | <0.0001 | 0.0504 | <0.0001 |
| 2 | Initial number of infectious mosquitoes | 0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | 0.8953 | <0.0001 |
| 3 | Mosquito daily latent rate | <0.0001 | <0.0001 | 0.0008 | 0.0016 | 0.0249 | <0.0001 | 0.0714 | <0.0001 |
| 4 | Mosquito daily mortality rate | <0.0001 | <0.0001 | <0.0001 | 0.0003 | <0.0001 | <0.0001 | 0.0767 | <0.0001 |
| 5 | Daily infectivity rate from mosquito to human | <0.0001 | <0.0001 | 0.2034 | 0.0306 | 0.0159 | 0.0191 | 0.0799 | <0.0001 |
| 6 | Human population size | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 |
| 7 | Initial number of infectious humans | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 |
| 8 | Human daily latent rate | 0.9893 | 0.1435 | 0.0001 | 0.0046 | <0.0001 | <0.0001 | 0.0626 | <0.0001 |
| 9 | Human daily recovery rate | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | 0.0790 | <0.0001 |
| 10 | Daily infectivity rate from human to mosquito | 0.0117 | <0.0001 | 0.4050 | 0.3405 | 0.0891 | 0.0052 | 0.0829 | <0.0001 |
| Model Baseline Level | Input | Pair Level Being Tested for the Input | Total Number of Infected Individuals | Number of Epidemic Waves | Duration of the Epidemic in Days | Runtime Per Replication in Seconds | |
|---|---|---|---|---|---|---|---|
| Level 1 | Level 2 | p-Value | p-Value | p-Value | p-Value | ||
| B1 | 1 | B1 | B2 | N/A | N/A | N/A | N/A |
| B1 | C1 | N/A | N/A | N/A | N/A | ||
| B1 | C2 | N/A | N/A | N/A | N/A | ||
| B1 | C3 | N/A | N/A | N/A | N/A | ||
| B2 | C1 | N/A | N/A | N/A | N/A | ||
| B2 | C2 | N/A | N/A | N/A | N/A | ||
| B2 | C3 | N/A | N/A | N/A | N/A | ||
| C1 | C2 | N/A | N/A | N/A | N/A | ||
| C1 | C3 | N/A | N/A | N/A | N/A | ||
| C2 | C3 | N/A | N/A | N/A | N/A | ||
| B2 | 1 | B2 | B1 | <0.0001 | 0.0464 | 0.0758 | 0.0004 |
| B2 | C1 | <0.0001 | 0.3639 | 0.4405 | 0.9997 | ||
| B2 | C2 | <0.0001 | 0.2930 | 0.6252 | 0.0125 | ||
| B2 | C3 | <0.0001 | 0.0013 | 0.0075 | 0.0002 | ||
| B1 | C1 | <0.0001 | <0.0001 | 0.0002 | 0.0008 | ||
| B1 | C2 | 0.0003 | 0.9273 | 0.7690 | 0.8825 | ||
| B1 | C3 | 0.9707 | 0.8216 | 0.9387 | 0.9998 | ||
| C1 | C2 | <0.0001 | 0.0016 | 0.0171 | 0.0234 | ||
| C1 | C3 | <0.0001 | <0.0001 | <0.0001 | 0.0004 | ||
| C2 | C3 | <0.0001 | 0.3274 | 0.2960 | 0.7983 | ||
| B1 | 2 | B1 | B2 | 0.0001 | <0.0001 | <0.0001 | N/A |
| B1 | C1 | 0.1160 | 0.0016 | 0.0328 | N/A | ||
| B1 | C2 | 0.0023 | <0.0001 | <0.0001 | N/A | ||
| B1 | C3 | N/A | N/A | N/A | N/A | ||
| B2 | C1 | 0.1719 | 0.0051 | 0.0072 | N/A | ||
| B2 | C2 | 0.8837 | 0.3870 | 0.5617 | N/A | ||
| B2 | C3 | N/A | N/A | N/A | N/A | ||
| C1 | C2 | 0.5554 | 0.3070 | 0.2213 | N/A | ||
| C1 | C3 | N/A | N/A | N/A | N/A | ||
| C2 | C3 | N/A | N/A | N/A | N/A | ||
| B2 | 2 | B2 | B1 | <0.0001 | <0.0001 | <0.0001 | <0.0001 |
| B2 | C1 | <0.0001 | <0.0001 | 0.1921 | 0.1167 | ||
| B2 | C2 | <0.0001 | <0.0001 | <0.0001 | 0.0678 | ||
| B2 | C3 | <0.0001 | <0.0001 | <0.0001 | 0.9978 | ||
| B1 | C1 | <0.0001 | <0.0001 | <0.0001 | 0.0055 | ||
| B1 | C2 | <0.0001 | 0.9951 | 0.0076 | 0.0117 | ||
| B1 | C3 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| C1 | C2 | <0.0001 | <0.0001 | <0.0001 | 0.9995 | ||
| C1 | C3 | <0.0001 | <0.0001 | <0.0001 | 0.0525 | ||
| C2 | C3 | <0.0001 | <0.0001 | <0.0001 | 0.0280 | ||
| B1 | 3 | B1 | C1 | 0.0499 | 0.1076 | 0.9168 | N/A |
| B1 | C2 | 0.0149 | 0.0084 | 0.8261 | N/A | ||
| B1 | C3 | 0.1882 | 0.0008 | 0.0201 | N/A | ||
| C1 | C2 | 0.9752 | 0.7993 | 0.9966 | N/A | ||
| C1 | C3 | <0.0001 | 0.4040 | 0.1111 | N/A | ||
| C2 | C3 | <0.0001 | 0.9169 | 0.1755 | N/A | ||
| B2 | 3 | B2 | C1 | <0.0001 | 0.0836 | 0.0402 | <0.0001 |
| B2 | C2 | <0.0001 | 0.0053 | 0.0003 | <0.0001 | ||
| B2 | C3 | <0.0001 | 0.9882 | <0.0001 | <0.0001 | ||
| C1 | C2 | 0.0506 | 0.7828 | 0.4941 | 0.9996 | ||
| C1 | C3 | <0.0001 | 0.1723 | 0.0456 | 0.0035 | ||
| C2 | C3 | <0.0001 | 0.0152 | 0.6288 | 0.0025 | ||
| B1 | 4 | B1 | C1 | 0.5157 | 0.8834 | 0.2250 | N/A |
| B1 | C2 | 0.0140 | <0.0001 | <0.0001 | N/A | ||
| B1 | C3 | 0.0011 | 0.6797 | 0.6794 | N/A | ||
| C1 | C2 | <0.0001 | <0.0001 | <0.0001 | N/A | ||
| C1 | C3 | 0.0837 | 0.2503 | 0.8584 | N/A | ||
| C2 | C3 | <0.0001 | 0.0006 | <0.0001 | N/A | ||
| B2 | 4 | B2 | C1 | <0.0001 | 0.9159 | 0.4339 | <0.0001 |
| B2 | C2 | <0.0001 | 0.0014 | <0.0001 | <0.0001 | ||
| B2 | C3 | <0.0001 | 0.0124 | 0.0700 | <0.0001 | ||
| C1 | C2 | <0.0001 | 0.0138 | <0.0001 | 0.0048 | ||
| C1 | C3 | <0.0001 | 0.0772 | 0.0005 | 0.5340 | ||
| C2 | C3 | <0.0001 | 0.9269 | <0.0001 | <0.0001 | ||
| B1 | 5 | B1 | C1 | 0.9636 | N/A | 0.9985 | N/A |
| B1 | C2 | 0.0703 | N/A | 0.9970 | N/A | ||
| B1 | C3 | 0.0319 | N/A | 0.0433 | N/A | ||
| C1 | C2 | 0.2040 | N/A | 0.9833 | N/A | ||
| C1 | C3 | 0.0072 | N/A | 0.0275 | N/A | ||
| C2 | C3 | <0.0001 | N/A | 0.0733 | N/A | ||
| B2 | 5 | B2 | C1 | 0.6224 | 0.5644 | 0.7787 | <0.0001 |
| B2 | C2 | <0.0001 | 0.0836 | 0.0292 | <0.0001 | ||
| B2 | C3 | <0.0001 | 0.9939 | 0.0755 | <0.0001 | ||
| C1 | C2 | <0.0001 | 0.7075 | 0.2652 | 0.6238 | ||
| C1 | C3 | <0.0001 | 0.4035 | 0.4608 | 0.0728 | ||
| C2 | C3 | <0.0001 | 0.0429 | 0.9846 | 0.0017 | ||
| B1 | 6 | B1 | B2 | <0.0001 | <0.0001 | <0.0001 | <0.0001 |
| B1 | C1 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| B1 | C2 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| B1 | C3 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| B2 | C1 | <0.0001 | 0.0040 | <0.0001 | <0.0001 | ||
| B2 | C2 | <0.0001 | 0.5257 | <0.0001 | <0.0001 | ||
| B2 | C3 | <0.0001 | 0.0061 | <0.0001 | <0.0001 | ||
| C1 | C2 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| C1 | C3 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| C2 | C3 | 0.0003 | 0.3451 | 0.0029 | <0.0001 | ||
| B2 | 6 | B2 | B1 | <0.0001 | <0.0001 | <0.0001 | <0.0001 |
| B2 | C1 | <0.0001 | <0.0001 | <0.0001 | 0.0989 | ||
| B2 | C2 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| B2 | C3 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| B1 | C1 | <0.0001 | 0.4023 | <0.0001 | <0.0001 | ||
| B1 | C2 | <0.0001 | 0.9912 | <0.0001 | <0.0001 | ||
| B1 | C3 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| C1 | C2 | <0.0001 | 0.1777 | <0.0001 | <0.0001 | ||
| C1 | C3 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| C2 | C3 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| B1 | 7 | B1 | C1 | 0.7241 | 0.9610 | 0.9407 | 0.0573 |
| B1 | C2 | <0.0001 | <0.0001 | <0.0001 | 0.0317 | ||
| B1 | C3 | <0.0001 | <0.0001 | <0.0001 | 0.0657 | ||
| C1 | C2 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| C1 | C3 | <0.0001 | <0.0001 | <0.0001 | 0.9999 | ||
| C2 | C3 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | ||
| B2 | 7 | B2 | C1 | <0.0001 | <0.0001 | <0.0001 | <0.0001 |
| B2 | C2 | 0.0037 | 0.2318 | 0.0897 | <0.0001 | ||
| B2 | C3 | 0.0083 | 0.6037 | 0.0651 | <0.0001 | ||
| C1 | C2 | <0.0001 | <0.0001 | <0.0001 | 0.3773 | ||
| C1 | C3 | <0.0001 | <0.0001 | <0.0001 | 0.0786 | ||
| C2 | C3 | <0.0001 | 0.0008 | <0.0001 | 0.8586 | ||
| B1 | 8 | B1 | C1 | N/A | 0.6430 | 0.8663 | N/A |
| B1 | C2 | N/A | 0.0297 | 0.0932 | N/A | ||
| B1 | C3 | N/A | 0.3702 | 0.1223 | N/A | ||
| C1 | C2 | N/A | 0.3870 | 0.4082 | N/A | ||
| C1 | C3 | N/A | 0.0273 | 0.0155 | N/A | ||
| C2 | C3 | N/A | <0.0001 | <0.0001 | N/A | ||
| B2 | 8 | B2 | C1 | N/A | 0.4620 | 0.3110 | <0.0001 |
| B2 | C2 | N/A | 0.3307 | 0.0008 | <0.0001 | ||
| B2 | C3 | N/A | 0.4422 | 0.0060 | <0.0001 | ||
| C1 | C2 | N/A | 0.9959 | 0.1515 | 0.9987 | ||
| C1 | C3 | N/A | 0.0168 | <0.0001 | 0.9998 | ||
| C2 | C3 | N/A | 0.0082 | <0.0001 | 0.9999 | ||
| B1 | 9 | B1 | C1 | 0.9598 | 0.8577 | 0.9324 | N/A |
| B1 | C2 | 0.4592 | 0.6978 | 0.0010 | N/A | ||
| B1 | C3 | 0.0007 | <0.0001 | 0.2521 | N/A | ||
| C1 | C2 | 0.2031 | 0.2372 | <0.0001 | N/A | ||
| C1 | C3 | 0.0043 | <0.0001 | 0.5968 | N/A | ||
| C2 | C3 | <0.0001 | <0.0001 | <0.0001 | N/A | ||
| B2 | 9 | B2 | C1 | 0.3465 | 0.4822 | 0.8990 | <0.0001 |
| B2 | C2 | <0.0001 | 0.0042 | <0.0001 | <0.0001 | ||
| B2 | C3 | <0.0001 | <0.0001 | 0.4452 | <0.0001 | ||
| C1 | C2 | <0.0001 | <0.0001 | <0.0001 | 0.2078 | ||
| C1 | C3 | <0.0001 | <0.0001 | 0.1284 | 0.3175 | ||
| C2 | C3 | <0.0001 | <0.0001 | <0.0001 | 0.0015 | ||
| B1 | 10 | B1 | C1 | 0.7481 | N/A | N/A | N/A |
| B1 | C2 | 0.0981 | N/A | N/A | N/A | ||
| B1 | C3 | 0.8307 | N/A | N/A | N/A | ||
| C1 | C2 | 0.5636 | N/A | N/A | N/A | ||
| C1 | C3 | 0.2495 | N/A | N/A | N/A | ||
| C2 | C3 | 0.0090 | N/A | N/A | N/A | ||
| B2 | 10 | B2 | C1 | <0.0001 | N/A | 0.9998 | <0.0001 |
| B2 | C2 | <0.0001 | N/A | 1.0000 | <0.0001 | ||
| B2 | C3 | <0.0001 | N/A | 0.0171 | <0.0001 | ||
| C1 | C2 | <0.0001 | N/A | 0.9998 | 0.8493 | ||
| C1 | C3 | <0.0001 | N/A | 0.0220 | 0.4090 | ||
| C2 | C3 | <0.0001 | N/A | 0.0173 | 0.0855 | ||
| Factor | Total Number of Infected Individuals | Number of Epidemic Waves | Duration of the Epidemic in Days | Runtime per Replication in Seconds | ||||
| Low Baseline (B1) | High Baseline (B2) | Low Baseline (B1) | High Baseline (B2) | Low Baseline (B1) | High Baseline (B2) | Low Baseline (B1) | High Baseline (B2) | |
| Model p-value | Model p-value | Model p-value | Model p-value | Model p-value | Model p-value | Model p-value | Model p-value | |
| 0.0002 | 0.0837 | 0.0398 | 0.5439 | 0.0441 | 0.5621 | 0.9999 | <0.0001 | |
| Factor p-value | Factor p-value | Factor p-value | Factor p-value | Factor p-value | Factor p-value | Factor p-value | Factor p-value | |
| Individual behavior, no time to switch behavior, low percentage of infectious individuals to trigger behavior, low cautious factor | 0.3478 | N/A | 0.3251 | N/A | 0.3412 | N/A | N/A | 0.0954 |
| Individual behavior, no time to switch behavior, high percentage of infectious individuals to trigger behavior, low cautious factor | 0.0398 | N/A | 0.7511 | N/A | 0.6404 | N/A | N/A | 0.5011 |
| Individual behavior, no time to switch behavior, low percentage of infectious individuals to trigger behavior, large cautious factor | 0.0123 | N/A | 0.3251 | N/A | 0.2149 | N/A | N/A | 0.4266 |
| Individual behavior, no time to switch behavior, high percentage of infectious individuals to trigger behavior, large cautious factor | 0.0604 | N/A | 0.2421 | N/A | 0.5538 | N/A | N/A | 0.0541 |
| Individual behavior, low time switch behavior, low percentage of infectious individuals to trigger behavior | 0.7592 | N/A | 0.8468 | N/A | 0.5492 | N/A | N/A | <0.0001 |
| Individual behavior, low time switch behavior, high percentage of infectious individuals to trigger behavior | 0.5851 | N/A | 0.0114 | N/A | 0.2764 | N/A | N/A | 0.3715 |
| Individual behavior, large time switch behavior, low percentage of infectious individuals to trigger behavior | 0.6930 | N/A | 0.8582 | N/A | 0.0358 | N/A | N/A | <0.0001 |
| Individual behavior, large time switch behavior, high percentage of infectious individuals to trigger behavior | 0.3519 | N/A | 0.2956 | N/A | 0.5883 | N/A | N/A | 0.0326 |
| Population behavior, no time to switch behavior, low percentage of infectious individuals to trigger behavior, low cautious factor | 0.0005 | N/A | 0.3565 | N/A | 0.3951 | N/A | N/A | 0.5968 |
| Population behavior, no time to switch behavior, high percentage of infectious individuals to trigger behavior, low cautious factor | 0.0567 | N/A | 0.6591 | N/A | 0.0147 | N/A | N/A | <0.0001 |
| Population behavior, no time to switch behavior, low percentage of infectious individuals to trigger behavior, large cautious factor | 0.2410 | N/A | 0.0131 | N/A | 0.0287 | N/A | N/A | 0.0779 |
| Population behavior, no time to switch behavior, high percentage of infectious individuals to trigger behavior, large cautious factor | 0.5890 | N/A | 0.8099 | N/A | 0.5041 | N/A | N/A | 0.0873 |
| Population behavior, low time switch behavior, low percentage of infectious individuals to trigger behavior | 0.6930 | N/A | 0.6591 | N/A | 0.9993 | N/A | N/A | <0.0001 |
| Population behavior, low time switch behavior, high percentage of infectious individuals to trigger behavior | 0.0011 | N/A | 0.2127 | N/A | 0.6046 | N/A | N/A | <0.0001 |
| Population behavior, large time switch behavior, low percentage of infectious individuals to trigger behavior | 0.8657 | N/A | 0.9942 | N/A | 0.5766 | N/A | N/A | <0.0001 |
| Population behavior, large time switch behavior, high percentage of infectious individuals to trigger behavior | 0.6871 | N/A | 0.1352 | N/A | 0.2518 | N/A | N/A | 0.9207 |
| Factor | Total Number of Infected Individuals | Number of Epidemic Waves | Duration of the Epidemic in Days | Runtime Per Replication in Seconds | ||||
| Low Baseline (B1) | High Baseline (B2) | Low Baseline (B1) | High Baseline (B2) | Low Baseline (B1) | High Baseline (B2) | Low Baseline (B1) | High Baseline (B2) | |
| Model p-value | Model p-value | Model p-value | Model p-value | Model p-value | Model p-value | Model p-value | Model p-value | |
| <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | 0.0429 | |
| Factor p-value | Factor p-value | Factor p-value | Factor p-value | Factor p-value | Factor p-value | Factor p-value | Factor p-value | |
| Multi-strain model | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | 0.4606 |
| Multi-vector model | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | 0.2959 |
| Multi-strain, multi-vector model | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | <0.0001 | 0.0334 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Galvão Scheidegger, A.P.; dos Santos Maxir, H.; Banerjee, A. A Statistical Examination of Distinct Characteristics Influencing the Performance of Vector-Borne Epidemiological Agent-Based Simulation Models. Modelling 2021, 2, 166-196. https://doi.org/10.3390/modelling2020009
Galvão Scheidegger AP, dos Santos Maxir H, Banerjee A. A Statistical Examination of Distinct Characteristics Influencing the Performance of Vector-Borne Epidemiological Agent-Based Simulation Models. Modelling. 2021; 2(2):166-196. https://doi.org/10.3390/modelling2020009
Chicago/Turabian StyleGalvão Scheidegger, Anna Paula, Henrique dos Santos Maxir, and Amarnath Banerjee. 2021. "A Statistical Examination of Distinct Characteristics Influencing the Performance of Vector-Borne Epidemiological Agent-Based Simulation Models" Modelling 2, no. 2: 166-196. https://doi.org/10.3390/modelling2020009







