Adapting the Parameters of RBF Networks Using Grammatical Evolution
Abstract
:1. Introduction
- The element represents the input pattern from the dataset describing the problem. For the rest of this paper, the notation d will be used to represent the number of elements in .
- The parameter k denotes the number of weights used to train the RBF network, and the associated vector of weights is denoted as .
- The vectors stand for the centers of the model.
- The value represents the value of the network for the given pattern .
- They have a simpler structure than other models used in machine learning, such as multilayer perceptron neural networks (MLPs) [11], since they have only one processing layer and therefore have faster training techniques, as well as faster response times.
- They can be used to efficiently approximate any continuous function [12].
- The first phase of the procedure seeks to locate a range of values for the parameters while also reducing the error of the network on the training dataset.
- The rules grammatical evolution uses in the first phase are simple and can be generalized to any dataset for data classification or fitting.
- The determination of the value interval is conducted in such a way that it is faster and more efficient to train the parameters with an optimization method during the second phase.
- After identifying a promising value interval from the first phase, any global optimization method can be used on that value interval to effectively minimize the network training error.
2. Method Description
2.1. Grammatical Evolution
- N is a set of the non-terminal symbols. A series of production rules is associated with every non-terminal symbol. The application of these production rules produces series of terminal symbols.
- T stands for the set of terminal symbols.
- S denotes the start symbol of the grammar and .
- P defines the set of production rules. These are rules that follow the following notations: or .
- Denote with V the next element form of the current chromosome.
- The next production rule is calculated as: Rule = V mod R. The number R stands for the total number of production rules for the non-terminal symbol that is currently under processing.
- A series of vectors that stand for the centers of the model.
- For every Gaussian unit, an additional parameter is required.
- The output weight vector .
| Algorithm 1 The BNF grammar used in the proposed method to produce intervals for the RBF parameters. By using this grammar in the first phase of the current work, the optimal interval of values for the parameters can be identified. |
S::=<expr> (0) <expr> ::= (<xlist> , <digit>,<digit>) (0) |<expr>,<expr> (1) <xlist>::=x1 (0) | x2 (1) ......... | xn (n) <digit> ::= 0 (0) | 1 (1) |
- For each center , there are d variables. As a consequence, every center requires parameters.
- Every Gaussian unit requires an additional parameter: , which means k more parameters.
- The weight vector used in the output has k parameters.
- The variable for which its original interval will be partitioned, for example, .
- An integer number with values 0 and 1 at the left margin of the interval. If this value is 1, then the left margin of the corresponding variable’s value field will be divided by two; otherwise, no change will be made.
- An integer number with values 0 and 1 at the right end of the range of values of the variable. If this value is 1, then the right end of the corresponding variable’s value field will be divided by two; otherwise, no change will be made.
2.2. The First Phase of the Proposed Algorithm
| Algorithm 2 The k-means algorithm. |
|
| Algorithm 3 The proposed algorithm used to locate the vectors |
|
- 1.
- Define as the number of chromosomes that will participate in the the grammatical evolution procedure.
- 2.
- Define as k the number of processing nodes of the used RBF model.
- 3.
- Define as the number of allowed generations.
- 4.
- Define as the used selection rate, with .
- 5.
- Define as the used mutation rate, with .
- 6.
- Define as the total number of RBF networks that will be created randomly in every fitness calculation.
- 7.
- Initialize chromosomes as sets of random numbers.
- 8.
- Set as the fitness of the best chromosome. The fitness function of any provided chromosome g is considered an interval
- 9.
- Set iter = 0.
- 10.
- For do
- (a)
- Produce the partition program using the grammar of Figure 1 for the chromosome i.
- (b)
- Produce the bounds for the partition program .
- (c)
- Set
- (d)
- For do
- i.
- Create randomly a set of parameters
- ii.
- Calculate the error
- iii.
- If , then
- iv.
- If , then
- (e)
- EndFor
- (f)
- Set the fitness
- 11.
- EndFor
- 12.
- Perform the procedure of selection. Initially, the chromosomes of the population are sorted according to their fitness values. Since the fitness values are intervals, the operator is defined asAs a consequence, the fitness value is considered smaller than if . The first chromosomes with smaller fitness values are copied without changes to the next generation of the algorithm. The rest of the chromosomes are replaced by chromosomes created in the crossover procedure.
- 13.
- Perform the crossover procedure. The crossover procedure will create new chromosomes. For every pair of created offspring, two parents are selected from the current population using the tournament selection method. These parent will produce the offspring and using the one-point crossover method shown in Figure 1.
- 14.
- Perform the mutation procedure. In this process, a random number is drawn for every element of each chromosome. The corresponding element is changed randomly if .
- 15.
- Set iter = iter + 1
- 16.
- If , go to step 10.
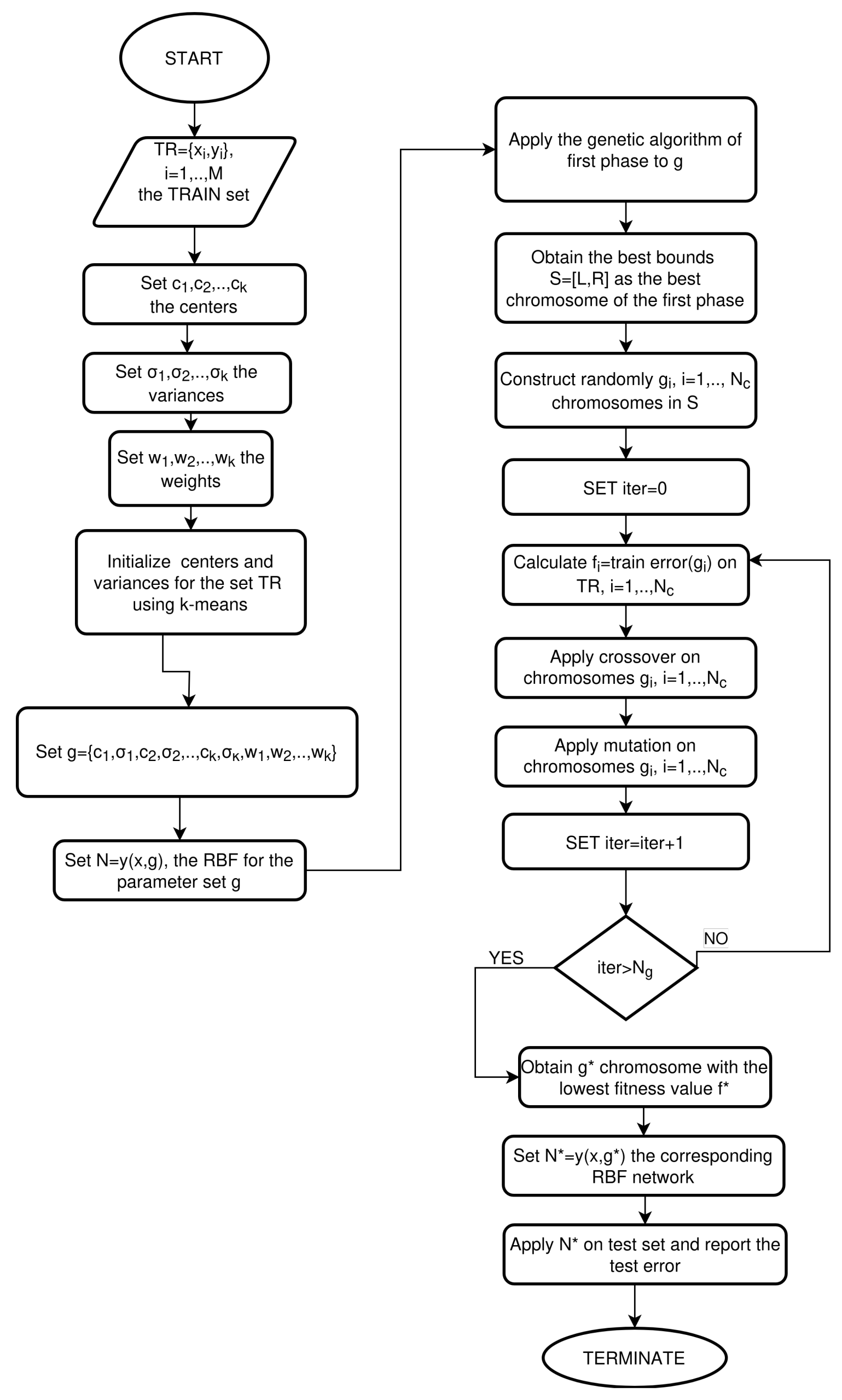
2.3. The Second Phase of the Proposed Algorithm
- Initialization Step
- (a)
- Define as the number of chromosomes.
- (b)
- Define as the total number of generations.
- (c)
- Define as k the number of processing nodes of the used RBF model.
- (d)
- Define as the best-located interval of the first stage of the algorithm of Section 2.2.
- (e)
- Produce random chromosomes in S.
- (f)
- Define as the used selection rate, with .
- (g)
- Define as the used mutation rate, with .
- (h)
- Set iter = 0.
- Fitness calculation step
- (a)
- For , do
- i.
- Compute the fitness of each chromosome as
- (b)
- EndFor
- Genetic operations step
- (a)
- Selection procedure. Initially, the population is sorted according to the fitness values. The first chromosomes with the lowest fitness values remain intact. The rest of the chromosomes are replaced by offspring that will be produced during the crossover procedure.
- (b)
- Crossover procedure: For every two new offspring , there are two parents that are selected from the current population with the selection procedure of tournament selection. The offspring are produced through the following process:The value is a random number, where [74].
- (c)
- Perform the mutation procedure. In this process, a random number is drawn for every element of each chromosome. The corresponding element is changed randomly if .
- Termination Check Step
- (a)
- Set
- (b)
- If , go to step 2.
3. Experiments
3.1. Experimental Datasets
- The UCI dataset repository, https://archive.ics.uci.edu/ml/index.php (accessed on 5 December 2023);
- The Keel repository, https://sci2s.ugr.es/keel/datasets.php (accessed on 5 December 2023) [75];
- The Statlib URL http://lib.stat.cmu.edu/datasets/ (accessed on 5 December 2023).
3.2. Experimental Results
- The column NEAT (NeuroEvolution of Augmenting Topologies) [115] denotes the application of the NEAT method for neural network training.
- The RBF-KMEANS column denotes the original two-phase training method for RBF networks.
- The column GENRBF stands for the RBF training method introduced in [116].
- The column PROPOSED stands for the results obtained using the proposed method.
- In the experimental tables, an additional row was added with the title AVERAGE. This row contains the average classification or regression error for all datasets.
4. Conclusions
- The proposed method could be applied to other variants of artificial neural networks.
- Intelligent learning techniques could be used in place of the k-means technique to initialize the neural network parameters.
- Techniques could be used to dynamically determine the number of necessary parameters for the neural network. For the time being, the number of parameters is considered constant, but this has the consequence of resulting in over-training phenomena being observed in various datasets.
- Crossover and mutation techniques that focus more on the existing interval construction technique for model parameters could be implemented.
- Efficient termination techniques for genetic algorithms could be used to obtain the most efficient termination of techniques without wasting computing time on unnecessary iterations.
- Techniques that are based on parallel programming could be used to increase the speed of the method.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mjahed, M. The use of clustering techniques for the classification of high energy physics data. Nucl. Instrum. Methods Phys. Res. Sect. A Accel. Spectrometers Detect. Assoc. Equip. 2006, 559, 199–202. [Google Scholar] [CrossRef]
- Andrews, M.; Paulini, M.; Gleyzer, S.; Poczos, B. End-to-End Event Classification of High-Energy Physics Data. J. Phys. Conf. Ser. 2018, 1085, 042022. [Google Scholar] [CrossRef]
- He, P.; Xu, C.J.; Liang, Y.Z.; Fang, K.T. Improving the classification accuracy in chemistry via boosting technique. Chemom. Intell. Lab. Syst. 2004, 70, 39–46. [Google Scholar] [CrossRef]
- Aguiar, J.A.; Gong, M.L.; Tasdizen, T. Crystallographic prediction from diffraction and chemistry data for higher throughput classification using machine learning. Comput. Mater. Sci. 2020, 173, 109409. [Google Scholar]
- Kaastra, I.; Boyd, M. Designing a neural network for forecasting financial and economic time series. Neurocomputing 1996, 10, 215–236. [Google Scholar] [CrossRef]
- Hafezi, R.; Shahrabi, J.; Hadavandi, E. A bat-neural network multi-agent system (BNNMAS) for stock price prediction: Case study of DAX stock price. Appl. Soft Comput. 2015, 29, 196–210. [Google Scholar] [CrossRef]
- Yadav, S.S.; Jadhav, S.M. Deep convolutional neural network based medical image classification for disease diagnosis. J. Big Data 2019, 6, 113. [Google Scholar] [CrossRef]
- Qing, L.; Linhong, W.; Xuehai, D. A Novel Neural Network-Based Method for Medical Text Classification. Future Internet 2019, 11, 255. [Google Scholar] [CrossRef]
- Park, J.; Sandberg, I.W. Universal Approximation Using Radial-Basis-Function Networks. Neural Comput. 1991, 3, 246–257. [Google Scholar] [CrossRef]
- Montazer, G.A.; Giveki, D.; Karami, M.; Rastegar, H. Radial basis function neural networks: A review. Comput. Rev. J. 2018, 1, 52–74. [Google Scholar]
- Abiodun, O.I.; Jantan, A.; Omolara, A.E.; Dada, K.V.; Mohamed, N.A.; Arshad, H. State-of-the-art in artificial neural network applications: A survey. Heliyon 2018, 4, e00938. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Fang, S.C.; Nuttle, H.L.W. Relaxed conditions for radial-basis function networks to be universal approximators. Neural Netw. 2003, 16, 1019–1028. [Google Scholar] [CrossRef] [PubMed]
- Teng, P. Machine-learning quantum mechanics: Solving quantum mechanics problems using radial basis function networks. Phys. Rev. E 2018, 98, 033305. [Google Scholar] [CrossRef]
- Jovanović, R.; Sretenovic, A. Ensemble of radial basis neural networks with K-means clustering for heating energy consumption prediction. FME Trans. 2017, 45, 51–57. [Google Scholar] [CrossRef]
- Gorbachenko, V.I.; Zhukov, M.V. Solving boundary value problems of mathematical physics using radial basis function networks. Comput. Math. Math. Phys. 2017, 57, 145–155. [Google Scholar] [CrossRef]
- Määttä, J.; Bazaliy, V.; Kimari, J.; Djurabekova, F.; Nordlund, K.; Roos, T. Gradient-based training and pruning of radial basis function networks with an application in materials physics. Neural Netw. 2021, 133, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Nam, M.-D.; Thanh, T.-C. Numerical solution of differential equations using multiquadric radial basis function networks. Neural Netw. 2001, 14, 185–199. [Google Scholar]
- Mai-Duy, N. Solving high order ordinary differential equations with radial basis function networks. Int. J. Numer. Meth. Eng. 2005, 62, 824–852. [Google Scholar] [CrossRef]
- Sarra, S.A. Adaptive radial basis function methods for time dependent partial differential equations. Appl. Numer. 2005, 54, 79–94. [Google Scholar] [CrossRef]
- Lian, R.-J. Adaptive Self-Organizing Fuzzy Sliding-Mode Radial Basis-Function Neural-Network Controller for Robotic Systems. IEEE Trans. Ind. Electron. 2014, 61, 1493–1503. [Google Scholar] [CrossRef]
- Vijay, M.; Jena, D. Backstepping terminal sliding mode control of robot manipulator using radial basis functional neural networks. Comput. Electr. Eng. 2018, 67, 690–707. [Google Scholar] [CrossRef]
- Er, M.J.; Wu, S.; Lu, J.; Toh, H.L. Face recognition with radial basis function (RBF) neural networks. IEEE Trans. Neural Netw. 2002, 13, 697–710. [Google Scholar] [PubMed]
- Laoudias, C.; Kemppi, P.; Panayiotou, C.G. Localization Using Radial Basis Function Networks and Signal Strength Fingerprints in WLAN. In Proceedings of the GLOBECOM 2009—2009 IEEE Global Telecommunications Conference, Honolulu, HI, USA, 30 November–4 December 2009; pp. 1–6. [Google Scholar]
- Azarbad, M.; Hakimi, S.; Ebrahimzadeh, A. Automatic recognition of digital communication signal. Int. J. Energy Inf. Commun. 2012, 3, 21–33. [Google Scholar]
- Yu, D.L.; Gomm, J.B.; Williams, D. Sensor fault diagnosis in a chemical process via RBF neural networks. Control Eng. Pract. 1999, 7, 49–55. [Google Scholar] [CrossRef]
- Shankar, V.; Wright, G.B.; Fogelson, A.L.; Kirby, R.M. A radial basis function (RBF) finite difference method for the simulation of reaction–diffusion equations on stationary platelets within the augmented forcing method. Int. J. Numer. Meth. Fluids 2014, 75, 1–22. [Google Scholar] [CrossRef]
- Shen, W.; Guo, X.; Wu, C.; Wu, D. Forecasting stock indices using radial basis function neural networks optimized by artificial fish swarm algorithm. Knowl.-Based Syst. 2011, 24, 378–385. [Google Scholar] [CrossRef]
- Momoh, J.A.; Reddy, S.S. Combined Economic and Emission Dispatch using Radial Basis Function. In Proceedings of the 2014 IEEE PES General Meeting | Conference & Exposition, National Harbor, MD, USA, 27–31 July 2014; pp. 1–5. [Google Scholar]
- Sohrabi, P.; Shokri, B.J.; Dehghani, H. Predicting coal price using time series methods and combination of radial basis function (RBF) neural network with time series. Miner. Econ. 2021, 36, 207–216. [Google Scholar] [CrossRef]
- Ravale, U.; Marathe, N.; Padiya, P. Feature Selection Based Hybrid Anomaly Intrusion Detection System Using K Means and RBF Kernel Function. Procedia Comput. Sci. 2015, 45, 428–435. [Google Scholar] [CrossRef]
- Lopez-Martin, M.; Sanchez-Esguevillas, A.; Arribas, J.I.; Carro, B. Network Intrusion Detection Based on Extended RBF Neural Network With Offline Reinforcement Learning. IEEE Access 2021, 9, 153153–153170. [Google Scholar] [CrossRef]
- Kuncheva, L.I. Initializing of an RBF network by a genetic algorithm. Neurocomputing 1997, 14, 273–288. [Google Scholar] [CrossRef]
- Ros, F.; Pintore, M.; Deman, A.; Chrétien, J.R. Automatical initialization of RBF neural networks. Chemom. Intell. Lab. Syst. 2007, 87, 26–32. [Google Scholar] [CrossRef]
- Wang, D.; Zeng, X.J.; Keane, J.A. A clustering algorithm for radial basis function neural network initialization. Neurocomputing 2012, 77, 144–155. [Google Scholar] [CrossRef]
- Benoudjit, N.; Verleysen, M. On the Kernel Widths in Radial-Basis Function Networks. Neural Process. Lett. 2003, 18, 139–154. [Google Scholar] [CrossRef]
- Neruda, R.; Kudova, P. Learning methods for radial basis function networks. Future Gener. Comput. Syst. 2005, 21, 1131–1142. [Google Scholar] [CrossRef]
- Ricci, E.; Perfetti, R. Improved pruning strategy for radial basis function networks with dynamic decay adjustment. Neurocomputing 2006, 69, 1728–1732. [Google Scholar] [CrossRef]
- Huang, G.-B.; Saratchandran, P.; Sundararajan, N. A generalized growing and pruning RBF (GGAP-RBF) neural network for function approximation. IEEE Trans. Neural Netw. 2005, 16, 57–67. [Google Scholar] [CrossRef] [PubMed]
- Bortman, M.; Aladjem, M. A Growing and Pruning Method for Radial Basis Function Networks. IEEE Trans. Neural Netw. 2009, 20, 1039–1045. [Google Scholar] [CrossRef]
- Yokota, R.; Barba, L.A.; Knepley, M.G. PetRBF—A parallel O(N) algorithm for radial basis function interpolation with Gaussians. Comput. Methods Appl. Mech. Eng. 2010, 199, 1793–1804. [Google Scholar] [CrossRef]
- Lu, C.; Ma, N.; Wang, Z. Fault detection for hydraulic pump based on chaotic parallel RBF network. EURASIP J. Adv. Signal Process. 2011, 2011, 49. [Google Scholar] [CrossRef]
- Iranmehr, A.; Masnadi-Shirazi, H.; Vasconcelos, N. Cost-sensitive support vector machines. Neurocomputing 2019, 343, 50–64. [Google Scholar] [CrossRef]
- Cervantes, J.; Lamont, F.G.; Mazahua, L.R.; Lopez, A. A comprehensive survey on support vector machine classification: Applications, challenges and trends. Neurocomputing 2020, 408, 189–215. [Google Scholar] [CrossRef]
- Kotsiantis, S.B. Decision trees: A recent overview. Artif. Intell. Rev. 2013, 39, 261–283. [Google Scholar] [CrossRef]
- Bertsimas, D.; Dunn, J. Optimal classification trees. Mach. Learn. 2017, 106, 1039–1082. [Google Scholar] [CrossRef]
- Wang, Y.; Yao, H.; Zhao, S. Auto-encoder based dimensionality reduction. Neurocomputing 2016, 184, 232–242. [Google Scholar] [CrossRef]
- Agarwal, V.; Bhanot, S. Radial basis function neural network-based face recognition using firefly algorithm. Neural Comput. Applic. 2018, 30, 2643–2660. [Google Scholar] [CrossRef]
- Jiang, S.; Lu, C.; Zhang, S.; Lu, X.; Tsai, S.B.; Wang, C.K.; Gao, Y.; Shi, Y.; Lee, C.H. Prediction of Ecological Pressure on Resource-Based Cities Based on an RBF Neural Network Optimized by an Improved ABC Algorithm. IEEE Access 2019, 7, 47423–47436. [Google Scholar] [CrossRef]
- Khan, I.U.; Aslam, N.; Alshehri, R.; Alzahrani, S.; Alghamdi, M.; Almalki, A.; Balabeed, M. Cervical Cancer Diagnosis Model Using Extreme Gradient Boosting and Bioinspired Firefly Optimization. Sci. Program. 2021, 2021, 5540024. [Google Scholar] [CrossRef]
- Gyamfi, K.S.; Brusey, J.; Gaura, E. Differential radial basis function network for sequence modelling. Expert Syst. Appl. 2022, 189, 115982. [Google Scholar] [CrossRef]
- Li, X.Q.; Song, L.K.; Choy, Y.S.; Bai, G.C. Multivariate ensembles-based hierarchical linkage strategy for system reliability evaluation of aeroengine cooling blades. Aerosp. Sci. Technol. 2023, 138, 108325. [Google Scholar] [CrossRef]
- MacQueen, J. Some methods for classification and analysis of multivariate observations. In Proceedings of the Fifth Berkeley Symposium on Mathematical Statistics and Probability, Los Angeles, CA, USA, 21 June 21–18 July 1967; pp. 281–297. [Google Scholar]
- O’Neill, M.; Ryan, C. Grammatical evolution. IEEE Trans. Evol. Comput. 2001, 5, 349–358. [Google Scholar] [CrossRef]
- Wang, H.Q.; Huang, D.S.; Wang, B. Optimisation of radial basis function classifiers using simulated annealing algorithm for cancer classification. Electron. Lett. 2005, 41, 630–632. [Google Scholar] [CrossRef]
- Fathi, V.; Montazer, G.A. An improvement in RBF learning algorithm based on PSO for real time applications. Neurocomputing 2013, 111, 169–176. [Google Scholar] [CrossRef]
- Goldberg, D. Genetic Algorithms in Search, Optimization and Machine Learning; Addison-Wesley Publishing Company: Reading, MA, USA, 1989. [Google Scholar]
- Michaelewicz, Z. Genetic Algorithms + Data Structures = Evolution Programs; Springer-Verlag: Berlin/Heidelberg, Germany, 1996. [Google Scholar]
- Grady, S.A.; Hussaini, M.Y.; Abdullah, M.M. Placement of wind turbines using genetic algorithms. Renew. Energy 2005, 30, 259–270. [Google Scholar] [CrossRef]
- Holland, J.H. Genetic algorithms. Sci. Am. 1992, 267, 66–73. [Google Scholar] [CrossRef]
- Stender, J. Parallel Genetic Algorithms: Theory & Applications; IOS Press: Amsterdam, The Netherlands, 1993. [Google Scholar]
- Backus, J.W. The Syntax and Semantics of the Proposed International Algebraic Language of the Zurich ACM-GAMM Conference. In Proceedings of the International Conference on Information Processing, UNESCO, Paris, France, 15–20 June 1959; pp. 125–132. [Google Scholar]
- Ryan, C.; Collins, J.; O’Neill, M. Grammatical evolution: Evolving programs for an arbitrary language. In Genetic Programming. EuroGP 1998; Banzhaf, W., Poli, R., Schoenauer, M., Fogarty, T.C., Eds.; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 1998; Volume 1391. [Google Scholar]
- O’Neill, M.; Ryan, M.C. Evolving Multi-line Compilable C Programs. In Genetic Programming; Poli, R., Nordin, P., Langdon, W.B., Fogarty, T.C., Eds.; EuroGP 1999. Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 1999; Volume 1598. [Google Scholar]
- Ryan, C.; O’Neill, M.; Collins, J.J. Grammatical evolution: Solving trigonometric identities. In Proceedings of Mendel; Technical University of Brno, Faculty of Mechanical Engineering: Brno, Czech Republic, 1998; Volume 98. [Google Scholar]
- Puente, A.O.; Alfonso, R.S.; Moreno, M.A. Automatic composition of music by means of grammatical evolution. In Proceedings of the APL ’02: 2002 Conference on APL: Array Processing Languages: Lore, Problems, and Applications, Madrid, Spain, 22–25 July 2002; pp. 148–155. [Google Scholar]
- De Campos, L.M.L.; de Oliveira, R.C.L.; Roisenberg, M. Optimization of neural networks through grammatical evolution and a genetic algorithm. Expert Syst. Appl. 2016, 56, 368–384. [Google Scholar] [CrossRef]
- Soltanian, K.; Ebnenasir, A.; Afsharchi, M. Modular Grammatical Evolution for the Generation of Artificial Neural Networks. Evol. Comput. 2022, 30, 291–327. [Google Scholar] [CrossRef] [PubMed]
- Dempsey, I.; Neill, M.O.; Brabazon, A. Constant creation in grammatical evolution. Int. J. Innov. Appl. 2007, 1, 23–38. [Google Scholar] [CrossRef]
- Galván-López, E.; Swafford, J.M.; O’Neill, M.; Brabazon, A. Evolving a Ms. PacMan Controller Using Grammatical Evolution. In Applications of Evolutionary Computation; EvoApplications 2010. Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2010; Volume 6024. [Google Scholar]
- Shaker, N.; Nicolau, M.; Yannakakis, G.N.; Togelius, J.; O’Neill, M. Evolving levels for Super Mario Bros using grammatical evolution. In Proceedings of the 2012 IEEE Conference on Computational Intelligence and Games (CIG), Granada, Spain, 11–14 September 2012; pp. 304–331. [Google Scholar]
- Martínez-Rodríguez, D.; Colmenar, J.M.; Hidalgo, J.I.; Micó, R.J.V.; Salcedo-Sanz, S. Particle swarm grammatical evolution for energy demand estimation. Energy Sci. Eng. 2020, 8, 1068–1079. [Google Scholar] [CrossRef]
- Sabar, N.R.; Ayob, M.; Kendall, G.; Qu, R. Grammatical Evolution Hyper-Heuristic for Combinatorial Optimization Problems. IEEE Trans. Evol. Comput. 2013, 17, 840–861. [Google Scholar] [CrossRef]
- Ryan, C.; Kshirsagar, M.; Vaidya, G.; Cunningham, A.; Sivaraman, R. Design of a cryptographically secure pseudo random number generator with grammatical evolution. Sci. Rep. 2022, 12, 8602. [Google Scholar] [CrossRef]
- Kaelo, P.; Ali, M.M. Integrated crossover rules in real coded genetic algorithms. Eur. J. Oper. Res. 2007, 176, 60–76. [Google Scholar] [CrossRef]
- Alcalá-Fdez, J.; Fernandez, A.; Luengo, J.; Derrac, J.; García, S.; Sánchez, L.; Herrera, F. KEEL Data-Mining Software Tool: Data Set Repository, Integration of Algorithms and Experimental Analysis Framework. J. Mult.-Valued Log. Soft Comput. 2011, 17, 255–287. [Google Scholar]
- Weiss, S.M.; Kulikowski, C.A. Computer Systems That Learn: Classification and Prediction Methods from Statistics, Neural Nets, Machine Learning, and Expert Systems; Morgan Kaufmann Publishers Inc.: Burlington, MA, USA, 1991. [Google Scholar]
- Quinlan, J.R. Simplifying Decision Trees. Int. Man-Mach. Stud. 1987, 27, 221–234. [Google Scholar] [CrossRef]
- Shultz, T.; Mareschal, D.; Schmidt, W. Modeling Cognitive Development on Balance Scale Phenomena. Mach. Learn. 1994, 16, 59–88. [Google Scholar] [CrossRef]
- Zhou, Z.H.; Jiang, Y. NeC4.5: Neural ensemble based C4.5. IEEE Trans. Knowl. Data Eng. 2004, 16, 770–773. [Google Scholar] [CrossRef]
- Setiono, R.; Leow, W.K. FERNN: An Algorithm for Fast Extraction of Rules from Neural Networks. Appl. Intell. 2000, 12, 15–25. [Google Scholar] [CrossRef]
- Demiroz, G.; Govenir, H.A.; Ilter, N. Learning Differential Diagnosis of Eryhemato-Squamous Diseases using Voting Feature Intervals. Artif. Intell. Med. 1998, 13, 147–165. [Google Scholar]
- Hayes-Roth, B.; Hayes-Roth, B.F. Concept learning and the recognition and classification of exemplars. J. Verbal Learn. Verbal Behav. 1977, 16, 321–338. [Google Scholar] [CrossRef]
- Kononenko, I.; Šimec, E.; Robnik-Šikonja, M. Overcoming the Myopia of Inductive Learning Algorithms with RELIEFF. Appl. Intell. 1997, 7, 39–55. [Google Scholar] [CrossRef]
- French, R.M.; Chater, N. Using noise to compute error surfaces in connectionist networks: A novel means of reducing catastrophic forgetting. Neural Comput. 2002, 14, 1755–1769. [Google Scholar] [CrossRef]
- Dy, J.G.; Brodley, C.E. Feature Selection for Unsupervised Learning. J. Mach. Learn. Res. 2004, 5, 845–889. [Google Scholar]
- Perantonis, S.J.; Virvilis, V. Input Feature Extraction for Multilayered Perceptrons Using Supervised Principal Component Analysis. Neural Process. Lett. 1999, 10, 243–252. [Google Scholar] [CrossRef]
- Garcke, J.; Griebel, M. Classification with sparse grids using simplicial basis functions. Intell. Data Anal. 2002, 6, 483–502. [Google Scholar] [CrossRef]
- Elter, M.; Schulz-Wendtland, R.; Wittenberg, T. The prediction of breast cancer biopsy outcomes using two CAD approaches that both emphasize an intelligible decision process. Med. Phys. 2007, 34, 4164–4172. [Google Scholar] [CrossRef] [PubMed]
- Little, M.A.; McSharry, P.E.; Hunter, E.J.; Spielman, J.; Ramig, L.O. Suitability of dysphonia measurements for telemonitoring of Parkinson’s disease. IEEE Trans. Biomed. Eng. 2009, 56, 1015. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.W.; Everhart, J.E.; Dickson, W.C.; Knowler, W.C.; Johannes, R.S. Using the ADAP learning algorithm to forecast the onset of diabetes mellitus. In Proceedings of the Symposium on Computer Applications and Medical Care, Washington, DC, USA, 6–9 November 1988; IEEE Computer Society Press: Washington, DC, USA, 1988; pp. 261–265. [Google Scholar]
- Lucas, D.D.; Klein, R.; Tannahill, J.; Ivanova, D.; Brandon, S.; Domyancic, D.; Zhang, Y. Failure analysis of parameter-induced simulation crashes in climate models. Geosci. Model Dev. 2013, 6, 1157–1171. [Google Scholar] [CrossRef]
- Gavrilis, D.; Tsoulos, I.G.; Dermatas, E. Selecting and constructing features using grammatical evolution. Pattern Recognit. Lett. 2008, 29, 1358–1365. [Google Scholar] [CrossRef]
- Giannakeas, N.; Tsipouras, M.G.; Tzallas, A.T.; Kyriakidi, K.; Tsianou, Z.E.; Manousou, P.; Hall, A.; Karvounis, E.C.; Tsianos, V.; Tsianos, E. A clustering based method for collagen proportional area extraction in liver biopsy images. In Proceedings of the Annual International Conference of the IEEE Engineering in Medicine and Biology Society, EMBS, Milan, Italy, 25–29 August 2015; pp. 3097–3100. [Google Scholar]
- Hastie, T.; Tibshirani, R. Non-parametric logistic and proportional odds regression. JRSS-C (Appl. Stat.) 1987, 36, 260–276. [Google Scholar] [CrossRef]
- Dash, M.; Liu, H.; Scheuermann, P.; Tan, K.L. Fast hierarchical clustering and its validation. Data Knowl. Eng. 2003, 44, 109–138. [Google Scholar] [CrossRef]
- Wolberg, W.H.; Mangasarian, O.L. Multisurface method of pattern separation for medical diagnosis applied to breast cytology. Proc. Natl. Acad. Sci. USA 1990, 87, 9193–9196. [Google Scholar] [CrossRef]
- Raymer, M.; Doom, T.E.; Kuhn, L.A.; Punch, W.F. Knowledge discovery in medical and biological datasets using a hybrid Bayes classifier/evolutionary algorithm. IEEE Trans. Syst. Man, And Cybern. Part B Cybern. 2003, 33, 802–813. [Google Scholar] [CrossRef] [PubMed]
- Zhong, P.; Fukushima, M. Regularized nonsmooth Newton method for multi-class support vector machines. Optim. Methods Softw. 2007, 22, 225–236. [Google Scholar] [CrossRef]
- Andrzejak, R.G.; Lehnertz, K.; Mormann, F.; Rieke, C.; David, P.; Elger, C.E. Indications of nonlinear deterministic and finite-dimensional structures in time series of brain electrical activity: Dependence on recording region and brain state. Phys. Rev. E 2001, 64, 1–8. [Google Scholar] [CrossRef]
- Koivisto, M.; Sood, K. Exact Bayesian Structure Discovery in Bayesian Networks. J. Mach. Learn. Res. 2004, 5, 549–573. [Google Scholar]
- Nash, W.J.; Sellers, T.L.; Talbot, S.R.; Cawthor, A.J.; Ford, W.B. The Population Biology of Abalone (_Haliotis_ Species) in Tasmania. I. Blacklip Abalone (_H. rubra_) from the North Coast and Islands of Bass Strait, Sea Fisheries Division; Technical Report No. 48; Marine Research Laboratories, Department of Primary Industries and Fisheries: Hobart, Australia, 1994. [Google Scholar]
- Brooks, T.F.; Pope, D.S.; Marcolini, A.M. Airfoil Self-Noise and Prediction; Technical Report, NASA RP-1218; NASA: Washington, DC, USA, 1989. [Google Scholar]
- Simonoff, J.S. Smooting Methods in Statistics; Springer: Berlin/Heidelberg, Germany, 1996. [Google Scholar]
- Cheng Yeh, I. Modeling of strength of high performance concrete using artificial neural networks. Cem. Concr. Res. 1998, 28, 1797–1808. [Google Scholar]
- Harrison, D.; Rubinfeld, D.L. Hedonic prices and the demand for clean ai. J. Environ. Econ. Manag. 1978, 5, 81–102. [Google Scholar] [CrossRef]
- Mackowiak, P.A.; Wasserman, S.S.; Levine, M.M. A critical appraisal of 98.6 degrees f, the upper limit of the normal body temperature, and other legacies of Carl Reinhold August Wunderlich. J. Am. Med. Assoc. 1992, 268, 1578–1580. [Google Scholar] [CrossRef]
- King, R.D.; Muggleton, S.; Lewis, R.; Sternberg, M.J.E. Drug design by machine learning: The use of inductive logic programming to model the structure-activity relationships of trimethoprim analogues binding to dihydrofolate reductase. Proc. Natl. Acad. Sci. USA 1992, 89, 11322–11326. [Google Scholar] [CrossRef]
- Sikora, M.; Wrobel, L. Application of rule induction algorithms for analysis of data collected by seismic hazard monitoring systems in coal mines. Arch. Min. Sci. 2010, 55, 91–114. [Google Scholar]
- Sanderson, C.; Curtin, R. Armadillo: A template-based C++ library for linear algebra. J. Open Source Softw. 2016, 1, 26. [Google Scholar] [CrossRef]
- Bishop, C. Neural Networks for Pattern Recognition; Oxford University Press: Oxford, UK, 1995. [Google Scholar]
- Cybenko, G. Approximation by superpositions of a sigmoidal function. Math. Control Signals Syst. 1989, 2, 303–314. [Google Scholar] [CrossRef]
- Riedmiller, M.; Braun, H. A Direct Adaptive Method for Faster Backpropagation Learning: The RPROP algorithm. In Proceedings of the International Conference on Neural Networks (ICNN’88), San Francisco, CA, USA, 28 March–1 April 1993; pp. 586–591. [Google Scholar]
- Kingma, D.P.; Ba, J.L. ADAM: A method for stochastic optimization. In Proceedings of the 3rd International Conference on Learning Representations (ICLR 2015), San Diego, CA, USA, 7–9 May 2015; pp. 1–15. [Google Scholar]
- Xue, Y.; Tong, Y.; Neri, F. An ensemble of differential evolution and Adam for training feed-forward neural networks. Inf. Sci. 2022, 608, 453–471. [Google Scholar] [CrossRef]
- Stanley, K.O.; Miikkulainen, R. Evolving Neural Networks through Augmenting Topologies. Evol. Comput. 2002, 10, 99–127. [Google Scholar] [CrossRef]
- Ding, S.; Xu, L.; Su, C.; Jin, F. An optimizing method of RBF neural network based on genetic algorithm. Neural Comput. Appl. 2012, 21, 333–336. [Google Scholar] [CrossRef]
- Gropp, W.; Lusk, E.; Doss, N.; Skjellum, A. A high-performance, portable implementation of the MPI message passing interface standard. Parallel Comput. 1996, 22, 789–828. [Google Scholar] [CrossRef]
- Chandra, R.; Dagum, L.; Kohr, D.; Maydan, D.; McDonald, J.; Menon, R. Parallel Programming in OpenMP; Morgan Kaufmann Publishers Inc.: Burlington, MA, USA, 2001. [Google Scholar]






| Expression | Chromosome | Operation |
|---|---|---|
| 9, 8, 6, 4, 15, 9, 16, 23, 8 | 9 mod 2 = 1 | |
| <expr>,<expr> | 8, 6, 4, 15, 9, 16, 23, 8 | 8 mod 2 = 0 |
| (<xlist>,<digit>,<digit>),<expr> | 6, 4, 15, 9, 16, 23, 8 | 6 mod 8 = 6 |
| (x7,<digit>,<digit>),<expr> | 4, 15, 9, 16, 23, 8 | 4 mod 2 = 0 |
| (x7,0,<digit>),<expr> | 15, 9, 16, 23, 8 | 15 mod 2 = 1 |
| (x7,0,1),<expr> | 9, 16, 23, 8 | 9 mod 2 = 1 |
| (x7,0,1),(<xlist>,<digit>,<digit>) | 16, 23, 8 | 16 mod 8 = 0 |
| (x7,0,1),(x1,<digit>,<digit>) | 23, 8 | 23 mod 2 = 1 |
| (x7,0,1),(x1,1,<digit>) | 8 | 8 mod 2 = 0 |
| (x7,0,1),(x1,1,0) |
| Dataset | Classes | Reference |
|---|---|---|
| APPENDICITIS | 2 | [76] |
| AUSTRALIAN | 2 | [77] |
| BALANCE | 3 | [78] |
| CLEVELAND | 5 | [79,80] |
| DERMATOLOGY | 6 | [81] |
| HAYES ROTH | 3 | [82] |
| HEART | 2 | [83] |
| HOUSEVOTES | 2 | [84] |
| IONOSPHERE | 2 | [85,86] |
| LIVERDISORDER | 2 | [87] |
| MAMMOGRAPHIC | 2 | [88] |
| PARKINSONS | 2 | [89] |
| PIMA | 2 | [90] |
| POPFAILURES | 2 | [91] |
| SPIRAL | 2 | [92] |
| REGIONS2 | 5 | [93] |
| SAHEART | 2 | [94] |
| SEGMENT | 7 | [95] |
| WDBC | 2 | [96] |
| WINE | 3 | [97,98] |
| Z_F_S | 3 | [99] |
| ZO_NF_S | 3 | [99] |
| ZONF_S | 2 | [99] |
| ZOO | 7 | [100] |
| Dataset | Reference |
|---|---|
| ABALONE | [101] |
| AIRFOIL | [102] |
| BASEBALL | STATLIB |
| BK | [103] |
| BL | STATLIB |
| CONCRETE | [104] |
| DEE | KEEL |
| DIABETES | KEEL |
| FA | STATLIB |
| HOUSING | [105] |
| MB | [103] |
| MORTGAGE | KEEL |
| NT | [106] |
| PY | [107] |
| QUAKE | [108] |
| TREASURY | KEEL |
| WANKARA | KEEL |
| Parameter | Value |
|---|---|
| 200 | |
| 100 | |
| 50 | |
| F | 10.0 |
| B | 100.0 |
| k | 10 |
| 0.90 | |
| 0.05 |
| Dataset | Rprop | Adam | Neat | Rbf-Kmeans | Genrbf | Proposed |
|---|---|---|---|---|---|---|
| Appendicitis | 16.30% | 16.50% | 17.20% | 12.23% | 16.83% | 15.77% |
| Australian | 36.12% | 35.65% | 31.98% | 34.89% | 41.79% | 22.40% |
| Balance | 8.81% | 7.87% | 23.14% | 33.42% | 38.02% | 15.62% |
| Cleveland | 61.41% | 67.55% | 53.44% | 67.10% | 67.47% | 50.37% |
| Dermatology | 15.12% | 26.14% | 32.43% | 62.34% | 61.46% | 35.73% |
| Hayes Roth | 37.46% | 59.70% | 50.15% | 64.36% | 63.46% | 35.33% |
| Heart | 30.51% | 38.53% | 39.27% | 31.20% | 28.44% | 15.91% |
| HouseVotes | 6.04% | 7.48% | 10.89% | 6.13% | 11.99% | 3.33% |
| Ionosphere | 13.65% | 16.64% | 19.67% | 16.22% | 19.83% | 9.30% |
| Liverdisorder | 40.26% | 41.53% | 30.67% | 30.84% | 36.97% | 28.44% |
| Mammographic | 18.46% | 46.25% | 22.85% | 21.38% | 30.41% | 17.72% |
| Parkinsons | 22.28% | 24.06% | 18.56% | 17.41% | 33.81% | 14.53% |
| Pima | 34.27% | 34.85% | 34.51% | 25.78% | 27.83% | 23.33% |
| Popfailures | 4.81% | 5.18% | 7.05% | 7.04% | 7.08% | 4.68% |
| Regions2 | 27.53% | 29.85% | 33.23% | 38.29% | 39.98% | 25.18% |
| Saheart | 34.90% | 34.04% | 34.51% | 32.19% | 33.90% | 29.46% |
| Segment | 52.14% | 49.75% | 66.72% | 59.68% | 54.25% | 49.22% |
| Spiral | 46.59% | 48.90% | 50.22% | 44.87% | 50.02% | 23.58% |
| Wdbc | 21.57% | 35.35% | 12.88% | 7.27% | 8.82% | 5.20% |
| Wine | 30.73% | 29.40% | 25.43% | 31.41% | 31.47% | 5.63% |
| Z_F_S | 29.28% | 47.81% | 38.41% | 13.16% | 23.37% | 3.90% |
| ZO_NF_S | 6.43% | 47.43% | 43.75% | 9.02% | 22.18% | 3.99% |
| ZONF_S | 27.27% | 11.99% | 5.44% | 4.03% | 17.41% | 1.67% |
| ZOO | 15.47% | 14.13% | 20.27% | 21.93% | 33.50% | 9.33% |
| AVERAGE | 26.56% | 32.36% | 30.11% | 28.84% | 33.35% | 18.73% |
| Dataset | Rprop | Adam | Neat | Rbf-Kmeans | Genrbf | Proposed |
|---|---|---|---|---|---|---|
| ABALONE | 4.55 | 4.30 | 9.88 | 7.37 | 9.98 | 5.16 |
| AIRFOIL | 0.002 | 0.005 | 0.067 | 0.27 | 0.121 | 0.004 |
| BASEBALL | 92.05 | 77.90 | 100.39 | 93.02 | 98.91 | 81.26 |
| BK | 1.60 | 0.03 | 0.15 | 0.02 | 0.023 | 0.025 |
| BL | 4.38 | 0.28 | 0.05 | 0.013 | 0.005 | 0.0004 |
| CONCRETE | 0.009 | 0.078 | 0.081 | 0.011 | 0.015 | 0.006 |
| DEE | 0.608 | 0.630 | 1.512 | 0.17 | 0.25 | 0.16 |
| DIABETES | 1.11 | 3.03 | 4.25 | 0.49 | 2.92 | 1.74 |
| HOUSING | 74.38 | 80.20 | 56.49 | 57.68 | 95.69 | 21.11 |
| FA | 0.14 | 0.11 | 0.19 | 0.015 | 0.15 | 0.033 |
| MB | 0.55 | 0.06 | 0.061 | 2.16 | 0.41 | 0.19 |
| MORTGAGE | 9.19 | 9.24 | 14.11 | 1.45 | 1.92 | 0.014 |
| NT | 0.04 | 0.12 | 0.33 | 8.14 | 0.02 | 0.007 |
| PY | 0.039 | 0.09 | 0.075 | 0.012 | 0.029 | 0.019 |
| QUAKE | 0.041 | 0.06 | 0.298 | 0.07 | 0.79 | 0.034 |
| TREASURY | 10.88 | 11.16 | 15.52 | 2.02 | 1.89 | 0.098 |
| WANKARA | 0.0003 | 0.02 | 0.005 | 0.001 | 0.002 | 0.003 |
| AVERAGE | 11.71 | 11.02 | 11.97 | 10.17 | 12.54 | 6.46 |
| Dataset | |||
|---|---|---|---|
| Appendicitis | 15.57% | 16.60% | 15.77% |
| Australian | 24.29% | 23.94% | 22.40% |
| Balance | 17.22% | 15.39% | 15.62% |
| Cleveland | 52.09% | 51.65% | 50.37% |
| Dermatology | 37.23% | 36.81% | 35.73% |
| Hayes Roth | 35.72% | 32.31% | 35.33% |
| Heart | 16.32% | 15.54% | 15.91% |
| HouseVotes | 4.35% | 3.90% | 3.33% |
| Ionosphere | 12.50% | 11.44% | 9.30% |
| Liverdisorder | 28.08% | 28.19% | 28.44% |
| Mammographic | 17.49% | 17.15% | 17.72% |
| Parkinsons | 16.25% | 15.17% | 14.53% |
| Pima | 23.29% | 23.97% | 23.33% |
| Popfailures | 5.31% | 5.86% | 4.68% |
| Regions2 | 25.97% | 26.29% | 25.18% |
| Saheart | 28.52% | 28.59% | 29.46% |
| Segment | 44.95% | 48.77% | 49.22% |
| Spiral | 15.49% | 18.19% | 23.58% |
| Wdbc | 5.43% | 5.01% | 5.20% |
| Wine | 7.59% | 8.39% | 5.63% |
| Z_F_S | 4.37% | 4.26% | 3.90% |
| ZO_NF_S | 3.79% | 4.21% | 3.99% |
| ZONF_S | 2.34% | 2.26% | 1.67% |
| ZOO | 11.90% | 10.50% | 9.33% |
| AVERAGE | 19.03% | 18.93% | 18.73% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsoulos, I.G.; Tzallas, A.; Karvounis, E. Adapting the Parameters of RBF Networks Using Grammatical Evolution. AI 2023, 4, 1059-1078. https://doi.org/10.3390/ai4040054
Tsoulos IG, Tzallas A, Karvounis E. Adapting the Parameters of RBF Networks Using Grammatical Evolution. AI. 2023; 4(4):1059-1078. https://doi.org/10.3390/ai4040054
Chicago/Turabian StyleTsoulos, Ioannis G., Alexandros Tzallas, and Evangelos Karvounis. 2023. "Adapting the Parameters of RBF Networks Using Grammatical Evolution" AI 4, no. 4: 1059-1078. https://doi.org/10.3390/ai4040054
APA StyleTsoulos, I. G., Tzallas, A., & Karvounis, E. (2023). Adapting the Parameters of RBF Networks Using Grammatical Evolution. AI, 4(4), 1059-1078. https://doi.org/10.3390/ai4040054









