Machine learning has been defined as “the extraction of knowledge from data” [
7]. Within healthcare, various algorithms already outperform doctors and radiologists [
10]. However, [
11] addressed a number of ethical considerations and, although this study establishes a place for ML in healthcare, it also acknowledges some of the limitations that are present. Nevertheless, the discussion of these challenges lacks depth regarding causes and/or solutions. More recently, [
12] extended this discussion, articulating the significance of unconscious bias, overreliance, and interpretability. Ultimately, ML solutions must be developed with respect to these limitations, as acknowledged within the evaluation of the findings presented in this study.
2.1.1. Deep Learning
Deep learning (DL) algorithms learn feature representations with multiple levels of abstraction [
13]. Addressing this technology from the perspective of healthcare, the application ranges from the diagnosis of Alzheimer’s to the prognosis of COVID-19 [
14]. Many solutions adopt pre-defined networks and, in some cases, pre-trained weights to encourage faster convergence. Ref. [
15] observed that tuning pre-trained weights can be very effective, allowing the network to adapt to the classification problem. This highlights the potential for utilising a pre-trained network for the task of LC classification. However, it does not acknowledge the potential limitations of this method. Ref. [
16] articulated that their method was more effective when trained from scratch as opposed to fine-tuning VGG16 weights, as the gap between natural and pathological images was too large. This has been considered during the development of this study and, thus, pre-trained weights were only frozen where the distribution between training datasets were equal.
Ref. [
17] stated that pre-processing methods improve upon the accuracy of healthcare predictions. This implies that pre-processing methods will improve LC classification. However, their study only investigated the effect of this on Type II diabetes and, thus, the assumption cannot be made that this will apply to LC. Providing a more suitable analysis, [
18] proposed a DL neural network which utilised pre-processing steps to reduce the noise and dimensionality of the data, which subsequently improved the results. The critical comparison of these studies underlines the general consensus that applying considered pre-processing steps serves for a better feature representation and, thus, a more accurate model.
Ultimately, [
19] established a specific set of pre-processing steps for the task of malignant nodule detection in CT imaging. This method was recognised by many scholars due to the exceptional results which it produced. However, for some of these steps, minimal justification was provided, specifically a number of arbitrary values which determine the inclusive nature of the applied mask. As a result, these steps may lack repeatability for alternative LC datasets whereby the distribution differs from that of the data used by [
19]. Despite this, [
20] reinforces this implementation, albeit on the same dataset. These steps have been used within the data pipeline of the proposed solution presented in
Section 3. However, the implementation has been discussed with respect to the lack of justification provided by [
19].
Following the natural progression of data manipulation, [
19] also implemented a number of data augmentation techniques specific to malignant nodule detection in LC. Included within these steps was rotational and horizontal flip augmentation, thus, preserving the distribution of the data and improving model generalisation. Evidence presented by [
21] confirms that over 60% of prior medical studies implemented basic augmentation techniques, thus, validating the decision to include the techniques proposed by [
19] within the developed solution.
When screening at-risk patients for LC, CT is considered as one of the key methods [
22]. As a result, convolution neural networks (CNNs) have been extensively researched in regard to the feature extraction of medical images [
23]. In 2018, [
24] achieved a sensitivity and specificity of 0.87 and 0.991, respectively, using a deep convolution neural network to detect LC nodules in CT scan images from the KDSB17 dataset. Acknowledged as a limitation, their method downsized the input images to 128 × 128 from 512 × 512 due to hardware constraints, which could have led to the loss of important features [
24]. However, [
25] extended this approach by implementing transfer learning with AlexNet, a pre-trained variation of a CNN, achieving an accuracy of 96%. Despite this, the authors did not address what data they used, which negatively affects the validity and repeatability of this study. However, aligning with their predominant use, this study will adopt a CNN architecture for the feature extraction of CT scan images.
2.1.2. Multimodal Learning
Modality has been defined as the way in which something happens or is experienced [
26]. Multiple modalities are inherently present within the realm of medicine [
27] and, thus, multimodal learning can be highly effective within DL and healthcare, improving the accuracy, sensitivity, and specificity of some classification problems [
28,
29]. This establishes that the utilisation of multimodality can result in robust and accurate predictions. As seen within the current diagnosis pathway, the method of diagnosis includes the type, size and location, and overall clinical status of the patient [
30]. However, scholars have commented upon the difficulty of exploiting
Supplementary Data as opposed to just complementary data within DL multimodal models [
26]. In contrast, [
31] identified that the utilisation of multiple modalities provides superior results regarding the effect of
Supplementary Data; a characteristic which is sought to be applied to the task of LC classification.
The abundance of fusion techniques has accelerated the growth of multimodal ML. Such techniques include joint and co-ordinated representations, as in [
26]. Ref. [
32] provided a critical comparison between these two techniques and presented equal arguments for both approaches. However, this balanced discussion is not reflected in the literature, as a consequence of the lack of ability to interpret more than two modalities within coordinated representations. In a more recent study, [
33] presented a multimodal architecture, projecting intermediate features into a joint space for classification. This architecture enriched the feature representations with unimodal models including a CNN and stacked denoising autoencoders. This outperformed other methods, but only for the task of Alzheimer’s disease (AD) classification. Thus, by taking inspiration from this approach, the model presented in this paper adopts similar steps.
In respect to LC, several modalities have been identified as good indicators, including CT/PET/MRI scans [
22], clinical/metabolomic biomarkers [
34], and volatile organic compounds [
35]. However, current screening trials, such as NLST, oversimplify LC risk prediction, reducing the cost efficacy due to the primary use of low-dose CT scans. It has been recognised that the pre-test probability can be improved if other clinical biomarkers, such as cancer history, history of other diseases, and asbestos exposure, are used [
36]. This provides evidence on the suitability of multimodal learning for LC classification. However, in contrast to AD, there is a lack of multimodal datasets suitable for the inclusion of these biomarkers. Ref. [
20] utilised the NLST and VLSP datasets and applied a co-learning approach, achieving an AUC (area under curve) of 0.91. However, it was articulated that their approach could be improved with additional CDEs, thus, providing an impetus for the implementation of a novel dataset.
The Lung Cancer Screening (LUCAS) dataset, published in 2020, provides 830 samples, including 76 CDEs and CT scans for each patient. This dataset was also presented alongside a benchmark ML architecture with an F1 and AUC score of 0.25 and 0.702, respectively [
37]. More recently, the SAMA model improved this to an F1 score of 0.341 with a standard deviation of 0.058 [
38]. The additional CDE data, despite the smaller sample size, provides a solution to the aforementioned limitation expressed by [
20]. However, the data presented by [
37] lacked clarity regarding the categorical nature of the CDEs. Despite the critical comparison against more established datasets, [
37] offers an alternative dataset to provide an incentive for the development of multimodal ML for LC classification.
While the aforementioned approach proposed by [
33] yielded good results for AD classification, the implementation for LC must be further validated. Ref. [
39] identified that simple concatenation of the output features of unimodal models may lack the depth needed to exploit intermodal interactions. This implies that the implementation presented by [
33] may fail to exploit the full potential of multiple modalities. In contrast, [
16] extracts intermediate features from a pre-trained CNN model. It was identified that extracting these multi-level features from various layers within a CNN provided a richer feature representation and higher AUC than those purely extracted from the last fully connected layer. This richer fusion technique was more effective at exploiting the complex multimodal associations within heterogenous data, and increased the accuracy from 83.6 to 91.1 [
16]. The method presented by [
16] provides an approach which can be applied to the task of LC classification, as demonstrated in the subsequent sections of this study.
Although the argument presented by [
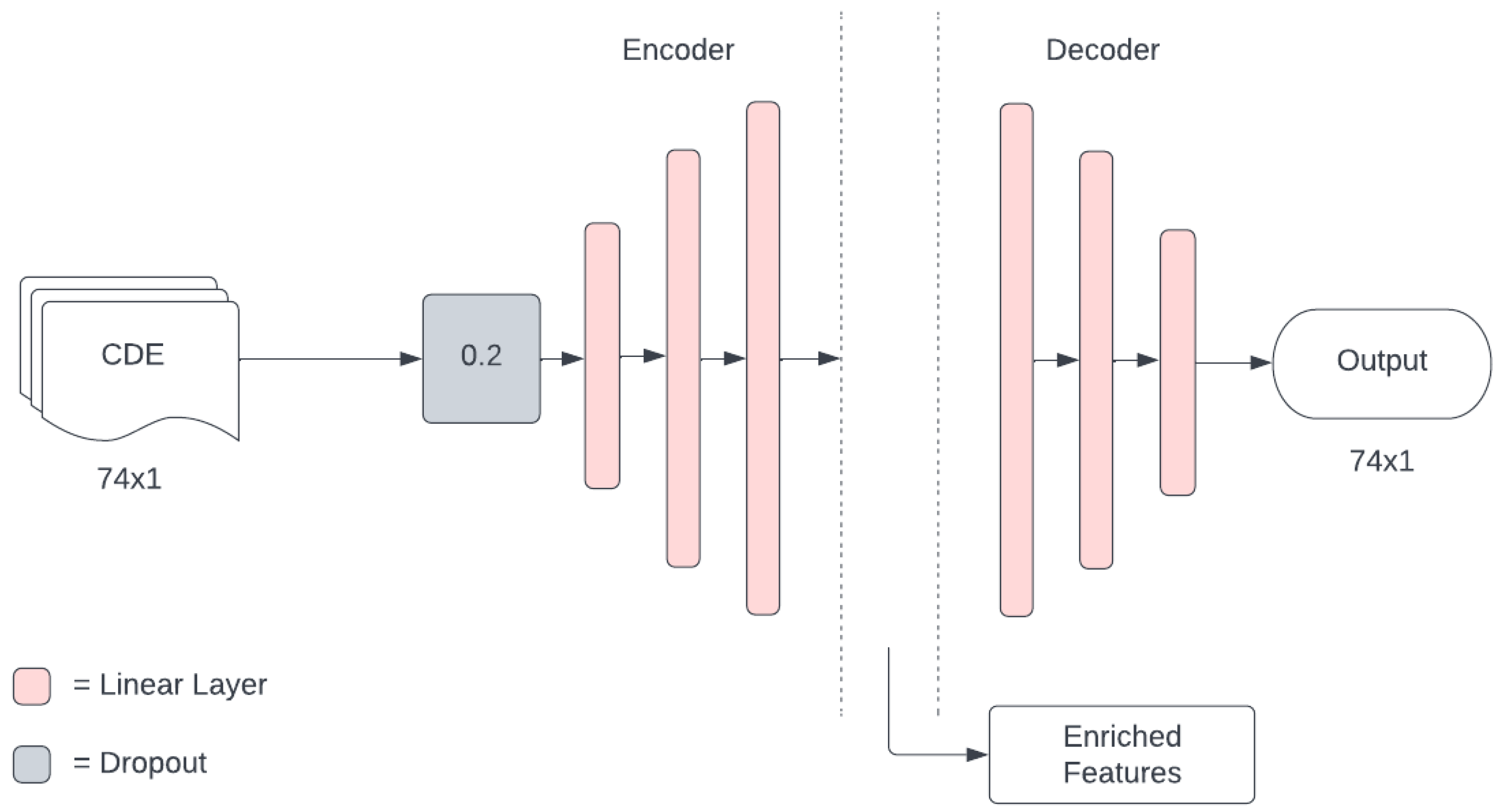
16] highlights the potential benefits of extracting a richer feature representation from the CT scan, there is a requirement to learn a good feature representation in every modality before information fusion. This establishes a need to increase the dimensionality of the CDEs. Ref. [
16] used a denoising autoencoder to achieve this, with an architecture in which the dimension of the encoded layer is greater than the dimension of the input layer. This further questions the validity of the approach used by [
33], as the down-sampling from successive layers within a CNN is also a process of information loss [
16]. Acknowledging the approaches of both [
16] and [
33], the solution presented in this study aimed to improve the feature representation of each modality, prior to fusion, for the task of LC classification.