Precision Medicine in Pediatric Cancer: Current Applications and Future Prospects
Abstract
:1. Introduction
2. Pediatric Cancer Genome
3. Precision Applications in Childhood Cancer
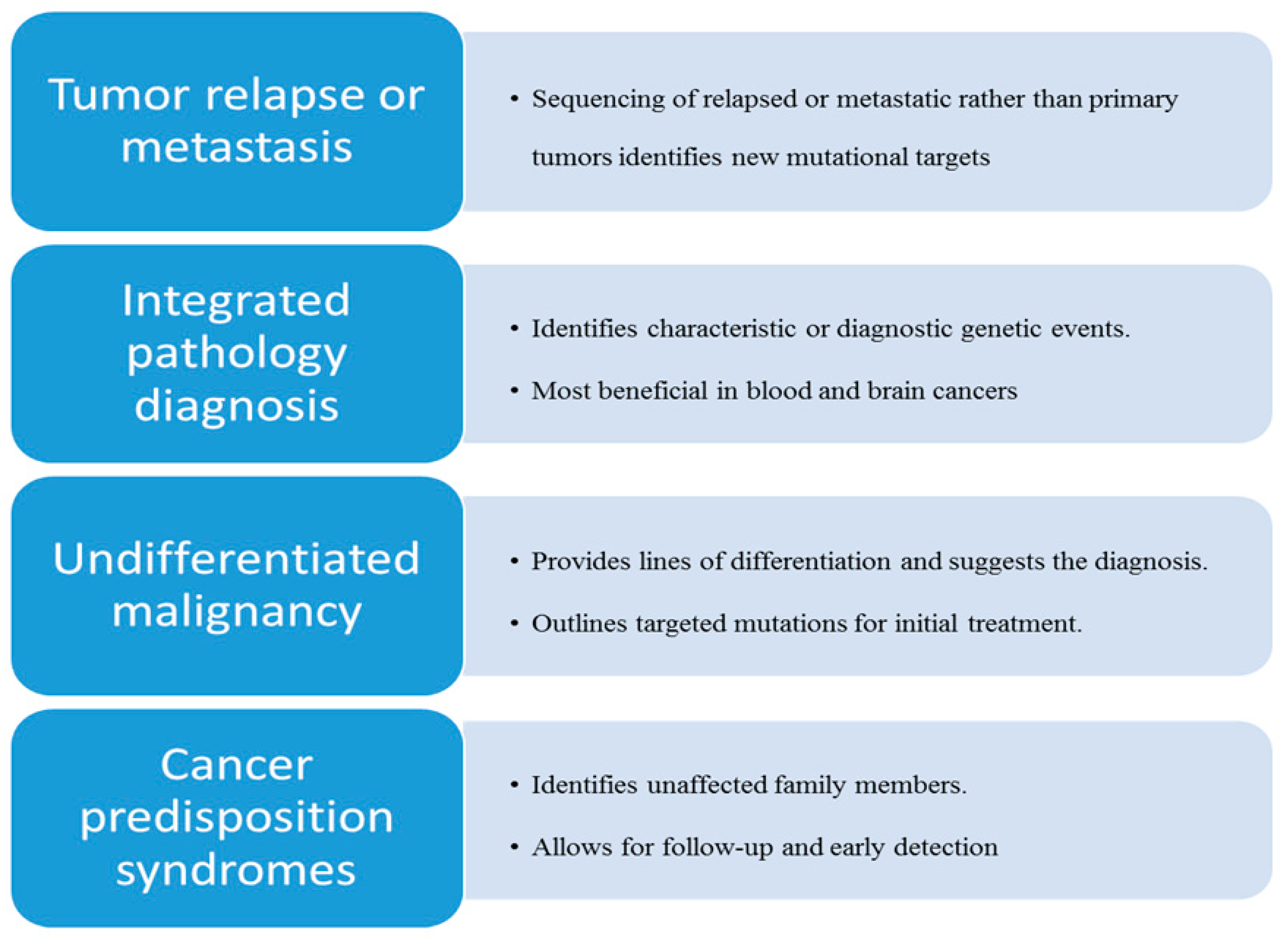
3.1. Tumor Recurrence or Metastasis during Therapy
3.2. Integrated Morphologic Molecular Diagnosis
3.3. Targeted Therapy of Undifferentiated Malignancies
3.4. Cancer Predisposition Syndromes (CPS)
4. Future Prospects
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Kalia, M. Personalized oncology: Recent advances and future challenges. Metabolism 2013, 62 (Suppl. 1), S11–S14. [Google Scholar] [CrossRef] [PubMed]
- Garraway, L.A.; Verweij, J.; Ballman, K.V. Precision oncology: An overview. J. Clin. Oncol. 2013, 31, 1803–1805. [Google Scholar] [CrossRef] [PubMed]
- Mendelsohn, J. Personalizing oncology: Perspectives and prospects. J. Clin. Oncol. 2013, 31, 1904–1911. [Google Scholar] [CrossRef] [PubMed]
- Seibel, N.L.; Janeway, K.; Allen, C.E.; Chi, S.N.; Cho, Y.J.; Glade Bender, J.L.; Kim, A.; Laetsch, T.W.; Irwin, M.S.; Takebe, N.; et al. Pediatric oncology enters an era of precision medicine. Curr. Probl. Cancer 2017, 41, 194–200. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, A.; Abedalthagafi, M. Cancer diagnostics: The journey from histomorphology to molecular profiling. Oncotarget 2016, 7, 58696–58708. [Google Scholar] [CrossRef] [Green Version]
- Blumenthal, G.M.; Mansfield, E.; Pazdur, R. Next-generation sequencing in oncology in the era of precision medicine. JAMA Oncol. 2016, 2, 13–14. [Google Scholar] [CrossRef]
- Horak, P.; Fröhling, S.; Glimm, H. Integrating next-generation sequencing into clinical oncology: Strategies, promises and pitfalls. ESMO Open 2016, 1, e000094. [Google Scholar] [CrossRef]
- Van Dijk, E.L.; Auger, H.; Jaszczyszyn, Y.; Thermes, C. Ten years of next-generation sequencing technology. Trends Genet. 2014, 30, 418–426. [Google Scholar] [CrossRef]
- Schwaederle, M.; Daniels, G.A.; Piccioni, D.E.; Fanta, P.T.; Schwab, R.B.; Shimabukuro, K.A.; Parker, B.A.; Kurzrock, R. On the road to precision cancer medicine: Analysis of genomic biomarker actionability in 439 Patients. Mol. Cancer Ther. 2015, 14, 1488–1494. [Google Scholar] [CrossRef]
- Tsui, P.C.; Lee, Y.F.; Liu, Z.W.Y.; Piao, W.; Chiang, A.K.S.; Lui, V.W.Y. An update on genomic-guided therapies for pediatric solid tumors. Future Oncol. 2017, 13, 1345–1358. [Google Scholar] [CrossRef] [Green Version]
- Gröbner, S.N.; Worst, B.C.; Weischenfeldt, J.; Buchhalter, I.; Kleinheinz, K.; Rudneva, V.A.; Johann, P.D.; Balasubramanian, G.P.; Segura-Wang, M.; Brabetz, S.; et al. The landscape of genomic alterations across childhood cancers. Nature 2018, 555, 321–327. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, X.; Liu, Y.; Liu, Y.; Alexandrov, L.B.; Edmonson, M.N.; Gawad, C.; Zhou, X.; Li, Y.; Rusch, M.C.; Easton, J.; et al. Pan-cancer genome and transcriptome analyses of 1699 paediatric leukaemias and solid tumours. Nature 2018, 555, 371–376. [Google Scholar] [CrossRef] [PubMed]
- Rahal, Z.; Abdulhai, F.; Kadara, H.; Saab, R. Genomics of adult and pediatric solid tumors. Am. J. Cancer Res. 2018, 8, 1356–1386. [Google Scholar] [PubMed]
- Worst, B.C.; van Tilburg, C.M.; Balasubramanian, G.P.; Fiesel, P.; Witt, R.; Freitag, A.; Boudalil, M.; Previti, C.; Wolf, S.; Schmidt, S.; et al. Next-generation personalised medicine for high-risk paediatric cancer patients—The INFORM pilot study. Eur. J. Cancer 2016, 65, 91–101. [Google Scholar] [CrossRef] [PubMed]
- PDQ Pediatric Treatment Editorial Board. Childhood Cancer Genomics (PDQ®): Health Professional Version. In PDQ Cancer Information Summaries [Internet]; National Cancer Institute: Bethesda, MD, USA, 2018; pp. 66–90. [Google Scholar]
- Capper, D.; Stichel, D.; Sahm, F.; Jones, D.T.W.; Schrimpf, D.; Sill, M.; Schmid, S.; Hovestadt, V.; Reuss, D.E.; Koelsche, C.; et al. Practical implementation of DNA methylation and copy-number-based CNS tumor diagnostics: The Heidelberg experience. Acta Neuropathol. 2018, 136, 181–210. [Google Scholar] [CrossRef] [PubMed]
- Kou, T.; Kanai, M.; Matsumoto, S.; Okuno, Y.; Muto, M. The possibility of clinical sequencing in the management of cancer. Jpn. J. Clin. Oncol. 2016, 46, 399–406. [Google Scholar] [CrossRef] [Green Version]
- Maese, L.; Schiffman, J.D. The evidence for expanded genetic testing for pediatric patients with cancer. Future Oncol. 2018, 14, 187–190. [Google Scholar] [CrossRef]
- Beltran, H.; Eng, K.; Mosquera, J.M.; Sigaras, A.; Romanel, A.; Rennert, H.; Kossai, M.; Pauli, C.; Faltas, B.; Fontugne, J.; et al. Whole-exome sequencing of metastatic cancer and biomarkers of treatment response. JAMA Oncol. 2015, 1, 466–474. [Google Scholar] [CrossRef]
- Hirshfield, K.M.; Tolkunov, D.; Zhong, H.; Ali, S.M.; Stein, M.N.; Murphy, S.; Vig, H.; Vazquez, A.; Glod, J.; Moss, R.A.; et al. Clinical actionability of comprehensive genomic profiling for management of rare or refractory cancers. Oncologist 2016, 21, 1315–1325. [Google Scholar] [CrossRef]
- Mody, R.J.; Wu, Y.M.; Lonigro, R.J.; Cao, X.; Roychowdhury, S.; Vats, P.; Frank, K.M.; Prensner, J.R.; Asangani, I.; Palanisamy, N.; et al. Integrative clinical sequencing in the management of refractory or relapsed cancer in youth. JAMA 2015, 314, 913–925. [Google Scholar] [CrossRef]
- Østrup, O.; Nysom, K.; Scheie, D.; Schmidt, A.Y.; Mathiasen, R.; Hjalgrim, L.L.; Olsen, T.E.; Skjøth-Rasmussen, J.; Henriksen, B.M.; Nielsen, F.C.; et al. Importance of comprehensive molecular profiling for clinical outcome in children with recurrent cancer. Front. Pediatr. 2018, 6, 114. [Google Scholar] [CrossRef] [PubMed]
- Weidenbusch, B.; Richter, G.H.S.; Kesper, M.S.; Guggemoos, M.; Gall, K.; Prexler, C.; Kazantsev, I.; Sipol, A.; Lindner, L.; Nathrath, M.; et al. Transcriptome based individualized therapy of refractory pediatric sarcomas: Feasibility, tolerability and efficacy. Oncotarget 2018, 9, 20747–20760. [Google Scholar] [CrossRef] [PubMed]
- Greenawalt, D.M.; Liang, W.S.; Saif, S.; Johnson, J.; Todorov, P.; Dulak, A.; Enriquez, D.; Halperin, R.; Ahmed, A.; Saveliev, V.; et al. Comparative analysis of primary versus relapse/refractory DLBCL identifies shifts in mutation spectrum. Oncotarget 2017, 8, 99237–99244. [Google Scholar] [CrossRef] [PubMed]
- Groisberg, R.; Roszik, J.; Conley, A.; Patel, S.R.; Subbiah, V. The role of next-generation sequencing in sarcomas: Evolution from light microscope to molecular microscope. Curr. Oncol. Rep. 2017, 19, 78. [Google Scholar] [CrossRef] [PubMed]
- Klein, M.E.; Parvez, M.M.; Shin, J.G. Clinical implementation of pharmacogenomics for personalized precision medicine: Barriers and solutions. J. Pharm. Sci. 2017, 106, 2368–2379. [Google Scholar] [CrossRef] [PubMed]
- Deverka, PA.; Dreyfus, J.C. Clinical integration of next generation sequencing: Coverage and reimbursement challenges. J. Law Med. Ethics 2014, 42 (Suppl. 1), 22–41. [Google Scholar] [CrossRef]
- Bavle, A.A.; Lin, F.Y.; Parsons, D.W. Applications of genomic sequencing in pediatric CNS tumors. Oncology (Williston Park) 2016, 30, 411–423. [Google Scholar]
- Kline, C.N.; Joseph, N.M.; Grenert, J.P.; van Ziffle, J.; Talevich, E.; Onodera, C.; Aboian, M.; Cha, S.; Raleigh, D.R.; Braunstein, S.; et al. Targeted next-generation sequencing of pediatric neuro-oncology patients improves diagnosis, identifies pathogenic germline mutations, and directs targeted therapy. NeuroOncol. 2017, 19, 699–709. [Google Scholar] [CrossRef]
- Oberg, J.A.; Glade Bender, J.L.; Sulis, M.L.; Pendrick, D.; Sireci, A.N.; Hsiao, S.J.; Turk, A.T.; Dela Cruz, F.S.; Hibshoosh, H.; Remotti, H.; et al. Implementation of next generation sequencing into pediatric hematology-oncology practice: Moving beyond actionable alterations. Genome Med. 2016, 8, 133. [Google Scholar] [CrossRef]
- Marks, L.J.; Oberg, J.A.; Pendrick, D.; Sireci, A.N.; Glasser, C.; Coval, C.; Zylber, R.J.; Chung, W.K.; Pang, J.; Turk, A.T.; et al. Precision medicine in children and young adults with hematologic malignancies and blood disorders: The Columbia University experience. Front. Pediatr. 2017, 5, 265. [Google Scholar] [CrossRef]
- Laetsch, T.W.; Roy, A.; Xu, L.; Black, J.O.; Coffin, C.M.; Chi, Y.Y.; Tian, J.; Spunt, S.L.; Hawkins, D.S.; Bridge, J.A.; et al. Undifferentiated sarcomas in children harbor clinically relevant oncogenic fusions and gene copy-number alterations: A report from the Children’s Oncology Group. Clin. Cancer Res. 2018, 24, 3888–3897. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Walsh, M.F.; Wu, G.; Edmonson, M.N.; Gruber, T.A.; Easton, J.; Hedges, D.; Ma, X.; Zhou, X.; Yergeau, D.A.; et al. Germline Mutations in Predisposition Genes in Pediatric Cancer. N. Engl. J. Med. 2015, 373, 2336–2346. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuhlen, M.; Taeubner, J.; Brozou, T.; Wieczorek, D.; Siebert, R.; Borkhardt, A. Family-based germline sequencing in children with cancer. Oncogene 2018. [Google Scholar] [CrossRef] [PubMed]
- Kuhlen, M.; Borkhardt, A. Trio sequencing in pediatric cancer and clinical implications. EMBO Mol. Med. 2018, 10, E8641. [Google Scholar] [CrossRef] [PubMed]
- Brozou, T.; Taeubner, J.; Velleuer, E.; Dugas, M.; Wieczorek, D.; Borkhardt, A.; Kuhlen, M. Genetic predisposition in children with cancer—Affected families’ acceptance of Trio-WES. Eur. J. Pediatr. 2018, 177, 53–60. [Google Scholar] [CrossRef] [PubMed]
- Coury, S.A.; Schneider, K.A.; Schienda, J.; Tan, W.H. Recognizing and managing children with a pediatric cancer predisposition syndrome: A Guide for the Pediatrician. Pediatr. Ann. 2018, 47, e204–e216. [Google Scholar] [CrossRef] [PubMed]
- Gallipoli, P.; Giotopoulos, G.; Huntly, B.J. Epigenetic regulators as promising therapeutic targets in acute myeloid leukemia. Ther. Adv. Hematol. 2015, 6, 103–119. [Google Scholar] [CrossRef] [Green Version]
- Ramos, K.N.; Ramos, I.N.; Zeng, Y.; Ramos, K.S. Genetics and epigenetics of pediatric leukemia in the era of precision medicine. F1000Res 2018, 7, F1000. [Google Scholar] [CrossRef]
- Bhattacharjee, S.; Nandi, S. Synthetic lethality in DNA repair network: A novel avenue in targeted cancer therapy and combination therapeutics. IUBMB Life 2017, 69, 929–937. [Google Scholar] [CrossRef]
- Chen, E.S. Targeting epigenetics using synthetic lethality in precision medicine. Cell. Mol. Life Sci. 2018, 75, 3381–3392. [Google Scholar] [CrossRef]
- Shapira, A.; Livney, Y.D.; Broxterman, H.J.; Assaraf, Y.G. Nanomedicine for targeted cancer therapy: Towards the overcoming of drug resistance. Drug Resist. Updates 2011, 14, 150–163. [Google Scholar] [CrossRef] [PubMed]
- Bar-Zeev, M.; Livney, Y.D.; Assaraf, Y.G. Targeted nanomedicine for cancer therapeutics: Towards precision medicine overcoming drug resistance. Drug Resist. Updates 2017, 31, 15–30. [Google Scholar] [CrossRef] [PubMed]
- Santhosh, P.B.; Ulrih, N.P. Multifunctional superparamagnetic iron oxide nanoparticles: Promising tools in cancer theranostics. Cancer Lett. 2013, 336, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Mao, K.; Zhang, B.; Zhao, Y. Superparamagnetic iron oxide nanoparticles conjugated with folic acid for dual target-specific drug delivery and MRI in cancer theranostics. Mater. Sci. Eng. C Mater. Biol. Appl. 2017, 70, 763–771. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S.; Riddell, S.R. Engineering CAR-T cells: Design concepts. Trends Immunol. 2015, 36, 494–502. [Google Scholar] [CrossRef] [Green Version]
- Xia, A.L.; Wang, X.C.; Lu, Y.J.; Lu, X.J.; Sun, B. Chimeric-antigen receptor T (CAR-T) cell therapy for solid tumors: Challenges and opportunities. Oncotarget 2017, 8, 90521–90531. [Google Scholar] [CrossRef]
- Mody, R.J.; Prensner, J.R.; Everett, J.; Parsons, D.W.; Chinnaiyan, A.M. Precision medicine in pediatric oncology: Lessons learned and next steps. Pediatr. Blood Cancer 2017, 64, e26288. [Google Scholar] [CrossRef]
| Tumor | Structural Rearrangements | Significantly Mutated Genes |
|---|---|---|
| Ewing’s sarcoma | EWS-ETS | STAG2, CDKN2A, TP53 |
| Ewing’s-like round cell sarcoma | CIC-FOX4; CIC-DUX4; BCOR-CCNB3; EWS-POU5F1/PATZI | CIC, BCOR |
| Alveolar rhabdomyosarcoma | PAX3/PAX7-FOXO1; PAX3-NCOA1 | BCOR, PIK3CA, GAB1, PTEN, ARID1A, ROBO1, AKAP9, NEB, C15orf2, PTPRO, COL5A2, PXDNL, NLRC5, TTN |
| Fusion-negative rhabdomyosarcoma | None | FGFR4, RAS, AKT, PIK3CA, MYOD1, DICER1, CTNNB1, FBXW7, BCOR, TP53 |
| Osteosarcoma | None | TP53, MDM2, RB1, ATRX, DLG2, PTEN |
| Neuroblastoma | None | ALK, MYCN amplification, ATRX, TERT, PTPN11, ARID1A, ARID1B, NF1, RAS, BRAF, FGFR1 |
| Wilms tumor | None | WT1, CTNNB1, WTX, DICER1, DIS3L2, SIX1/2, MLLT1, TP53, FBXW7, MYCN, CTR9, REST |
| Malignant rhabdoid tumor | None | SMARCB1, SMARCA4 |
| Translocation renal cell carcinoma | Xp11 (TFE3) | TFE, SMARCC2, KDM5C, INO80D, CHD, MLL3 |
| Clear cell sarcoma of the kidney | YWHAE-NUTM2 | BCOR duplication |
| Synovial sarcoma | SS18-SSX1/2 | NGDN, RASAL3, KLHL34, MUM1L1, EP300 |
| Dermatofibrosarcoma protuberans | COL1A1-PDGFB | CARD10, PPP1R39, SAFB2, STARD9 |
| Cancer Predisposition Syndrome | Common Tumors | Mutated Genes | Dysregulated Pathways and Functions |
|---|---|---|---|
| Li Fraumeni syndrome | Leukemias, osteosarcoma, soft tissue sarcoma, adrenocortical, brain tumors | TP53, CHEK2 | Cell cycle, apoptosis |
| Familial adenomatous polyposis | Colonic polyps, osteomas, desmoid, thyroid and adrenal tumors | CTNNB1, MUTYH | WNT/β-catenin |
| Fanconi anemia | Leukemias, hepatic tumors, upper airways carcinomas | FANC family of genes, BRCA2 | DNA repair |
| Nevoid basal cell carcinoma | Basal cell carcinomas, jaw cysts | PTCH | Sonic hedgehog (SHH) |
| Neurofibromatosis | Peripheral nerve and nerve sheath tumors, brain gliomas, GIST tumors | NF1 | GTPase, RAS/MAPK inactivation |
| Von Hippel–Lindau | Renal carcinoma, hemangioblastoma, pheochromocytoma, pancreatic tumors | VHL | HIF degradation, RNA polymerase II regulation |
| Dicer1 syndrome | Pleuropulmonary blastoma | Dicer1 | miRNA synthesis |
| Xeroderma pigmentosum | Skin cancers | XP family of genes | DNA repair |
| Hereditary paraganglioma-pheochromocytoma | Paraganglioma, pheochromocytoma, renal cell carcinoma | SDH | Citric acid cycle, oxidative phosphorylation |
| Ataxia telangiectasia | Leukemia, lymphoma | ATM | DNA repair |
| Bloom syndrome | Leukemia, lymphoma | BLM | DNA repair |
| PTEN hamartoma | Breast, thyroid, GI polyps | PTEN | Tyrosine kinase de-phosphorylation |
| Hamartomatous polyposis (Peutz-Jeghers/juvenile polyposis) | Gastrointestinal polyps | LKB1/STK11; SMAD4; BMPR1A; ENG | DNA mismatch repair |
| Hereditary non-polyposis colorectal cancer | Colorectal cancer (very rare in children) | MLH1, MSH2, MSH6, or PMS2 | DNA mismatch repair |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ahmed, A.A.; Vundamati, D.S.; Farooqi, M.S.; Guest, E. Precision Medicine in Pediatric Cancer: Current Applications and Future Prospects. High-Throughput 2018, 7, 39. https://doi.org/10.3390/ht7040039
Ahmed AA, Vundamati DS, Farooqi MS, Guest E. Precision Medicine in Pediatric Cancer: Current Applications and Future Prospects. High-Throughput. 2018; 7(4):39. https://doi.org/10.3390/ht7040039
Chicago/Turabian StyleAhmed, Atif A., Divya S. Vundamati, Midhat S. Farooqi, and Erin Guest. 2018. "Precision Medicine in Pediatric Cancer: Current Applications and Future Prospects" High-Throughput 7, no. 4: 39. https://doi.org/10.3390/ht7040039
APA StyleAhmed, A. A., Vundamati, D. S., Farooqi, M. S., & Guest, E. (2018). Precision Medicine in Pediatric Cancer: Current Applications and Future Prospects. High-Throughput, 7(4), 39. https://doi.org/10.3390/ht7040039






