Analyzing COVID-19 Medical Papers Using Artificial Intelligence: Insights for Researchers and Medical Professionals
Abstract
:1. Introduction
2. Materials and Methods
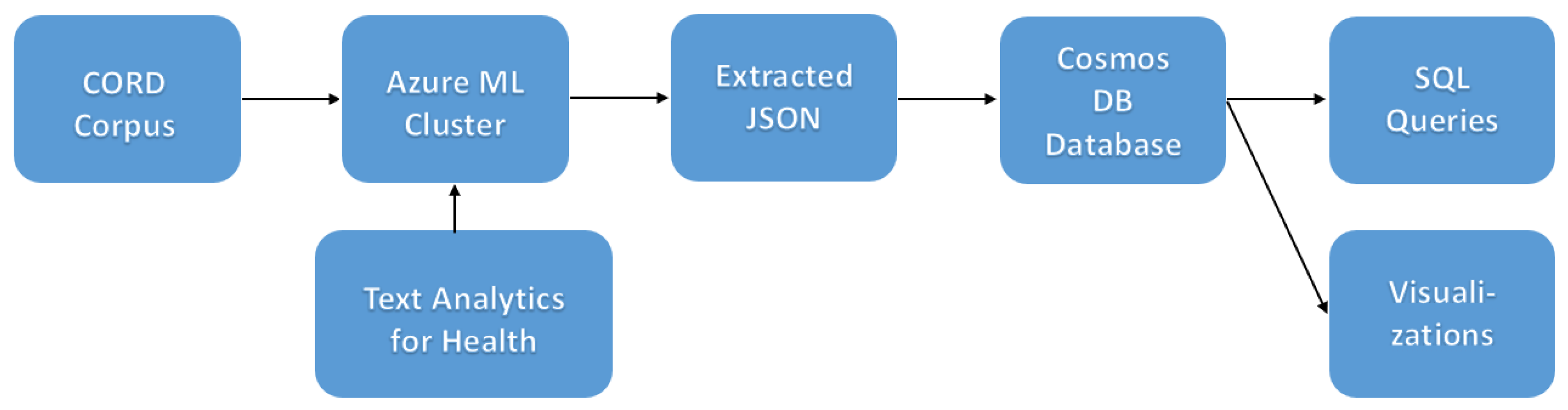
2.1. COVID-19 Scientific Papers and CORD Dataset
- Metadata file (“Metadata.csv”) contains the most important information for all publications in one place. Each paper in this table has an identifier “cord_uid”. This identifier is not completely unique throughout the table, but can mostly be used to identify the paper. Other information in metadata.csv includes:
- ○
- Title of publication,
- ○
- Journal,
- ○
- Authors,
- ○
- Abstract,
- ○
- Data of publication,
- ○
- DOI.
- Full-text papers in the “document_parses” directory, in the form of structured text in JSON format.
- Pre-built document embeddings that map cord_uid-s to float vectors that reflect the overall semantics of the paper.
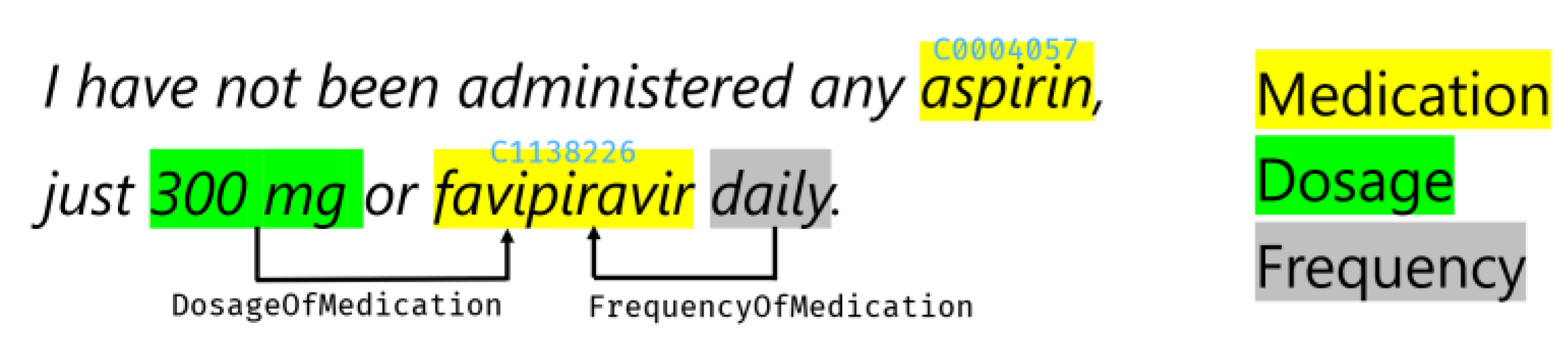
2.2. Text Analytics
- Perform basic NER on medical and near-medical terms and return the list of entities;
- Do entity mapping to standard medical ontologies, such as Unified Medical Language System (UMLS) [25];
- Extract relations between entities inside the text, such as «TimeOfCondition», etc.;
- Detect negation, which indicates that an entity was used in a negative context, for example, “COVID-19 diagnosis did not occur”.
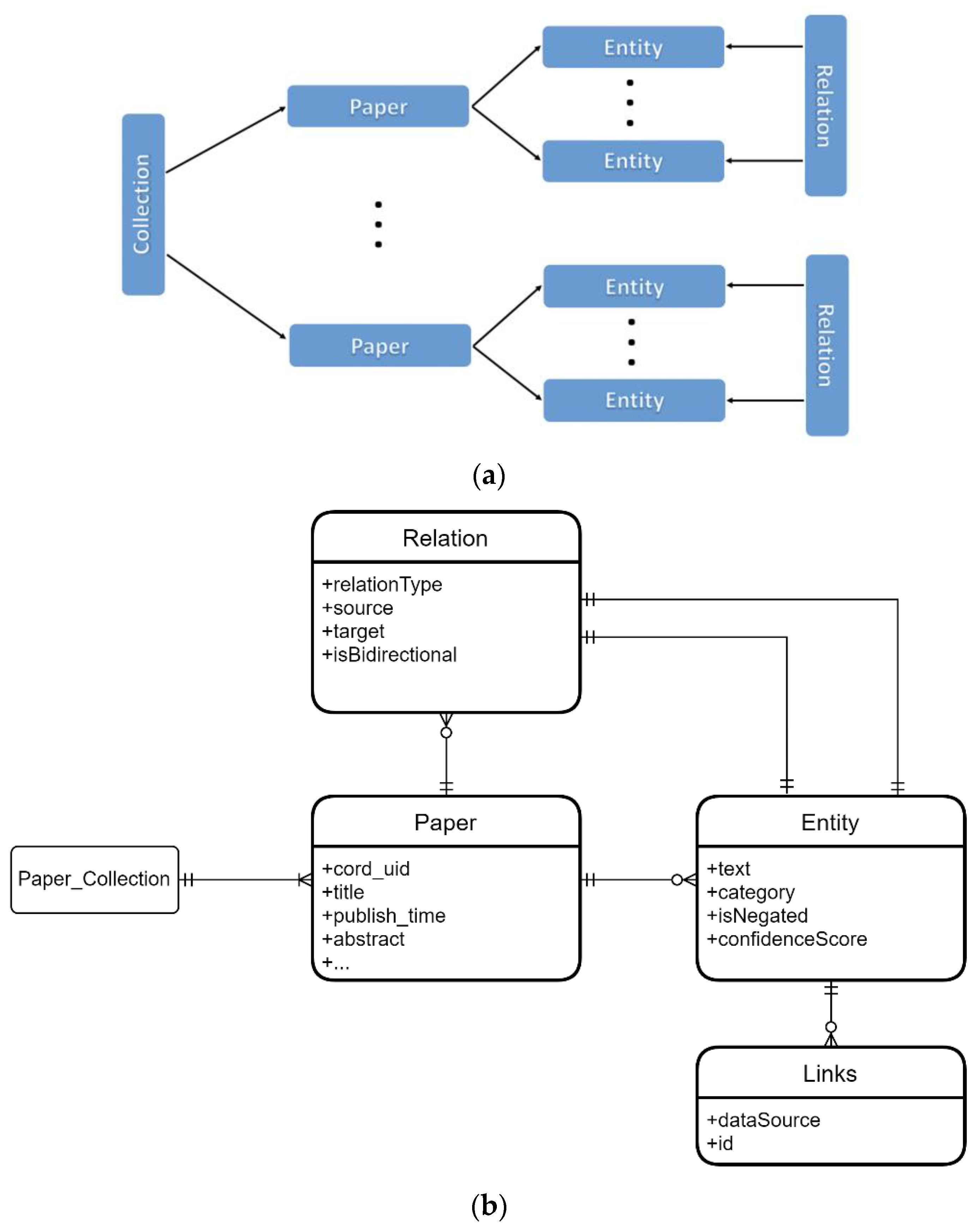
2.3. Entity/Relation Data Storage

2.4. Unified Medical Language System (UMLS)

2.5. Entity and Relation Types


3. Results and Discussion
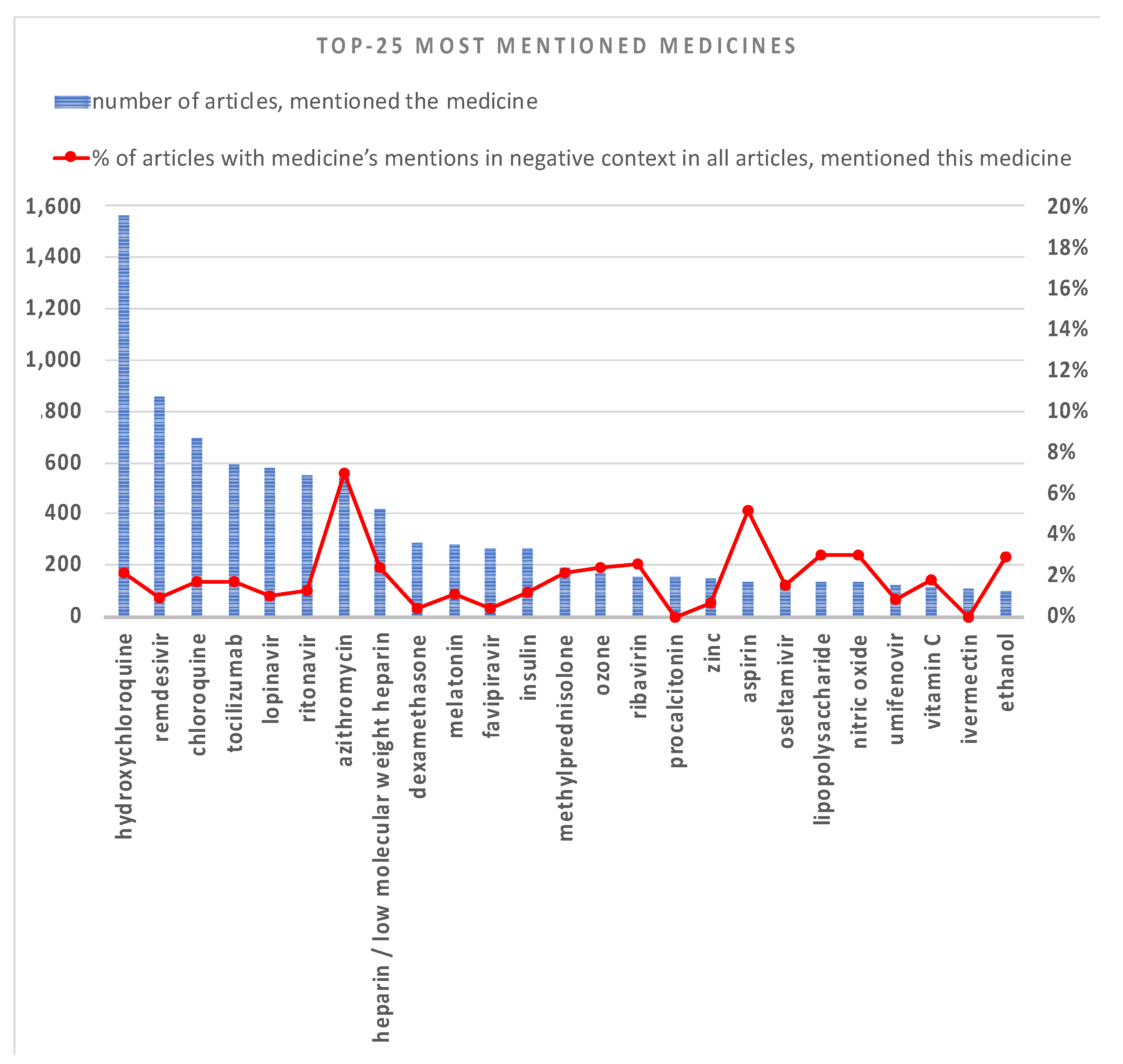
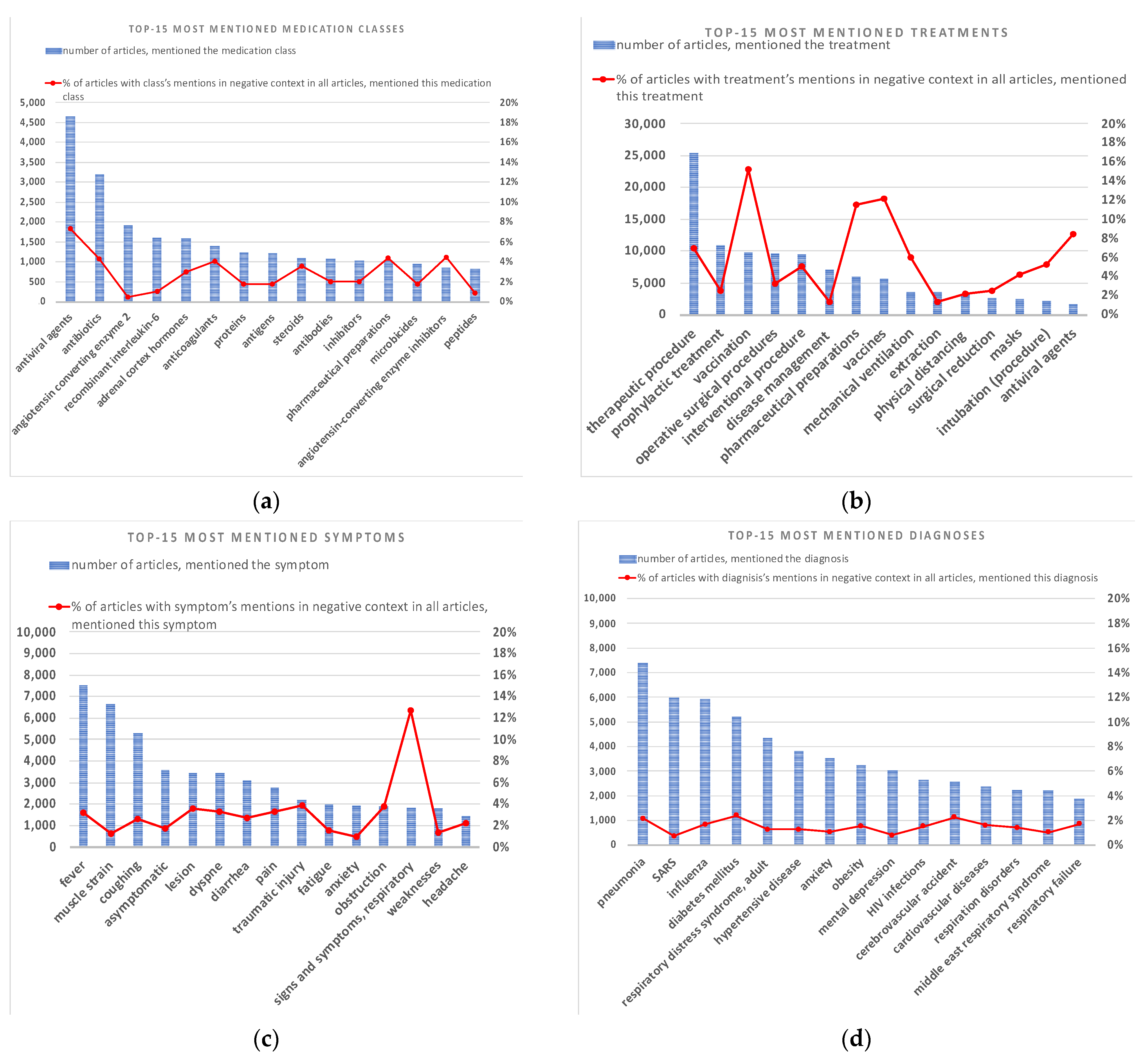
3.1. TOP Mentioned Entities
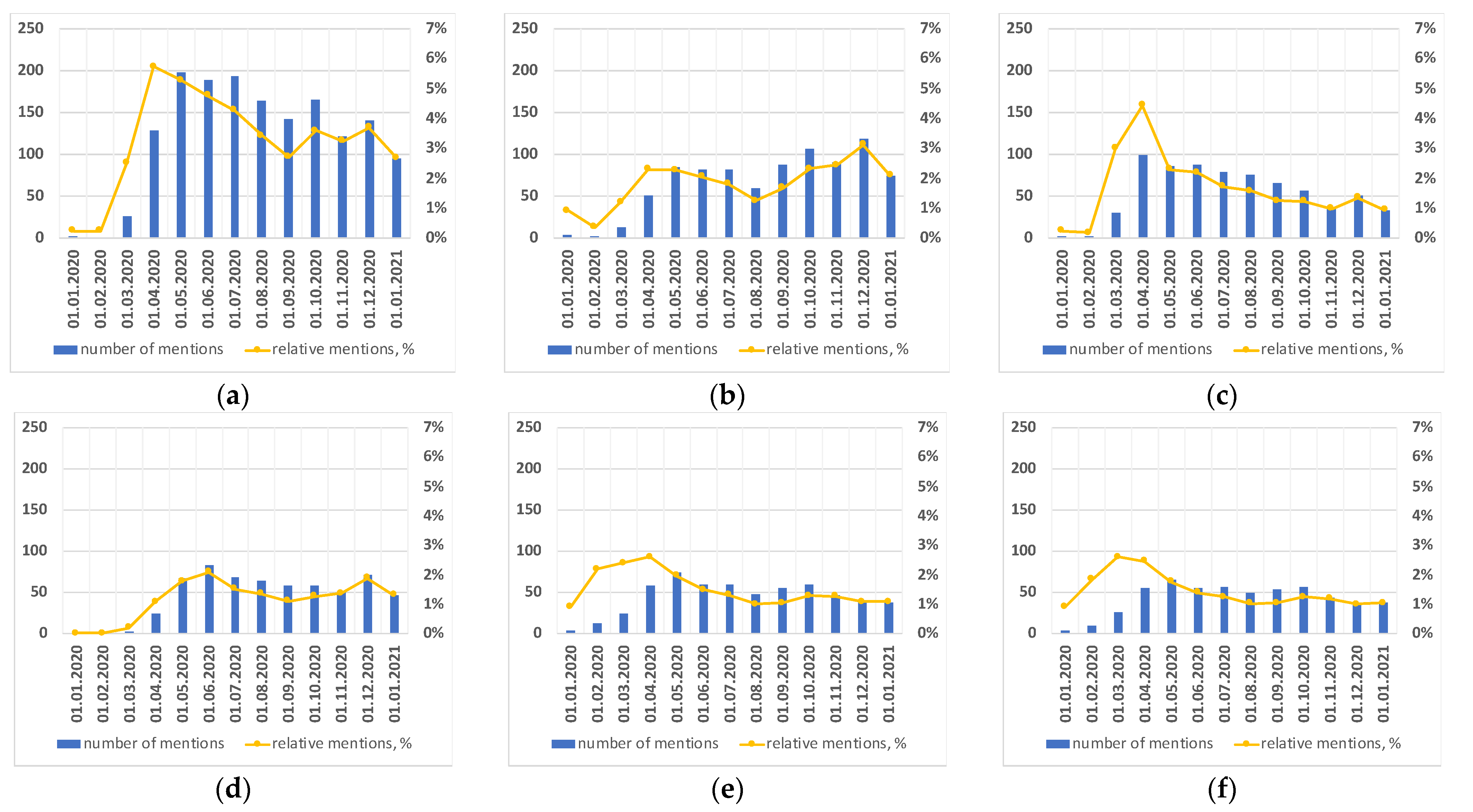
3.2. Treatment Strategy over Time
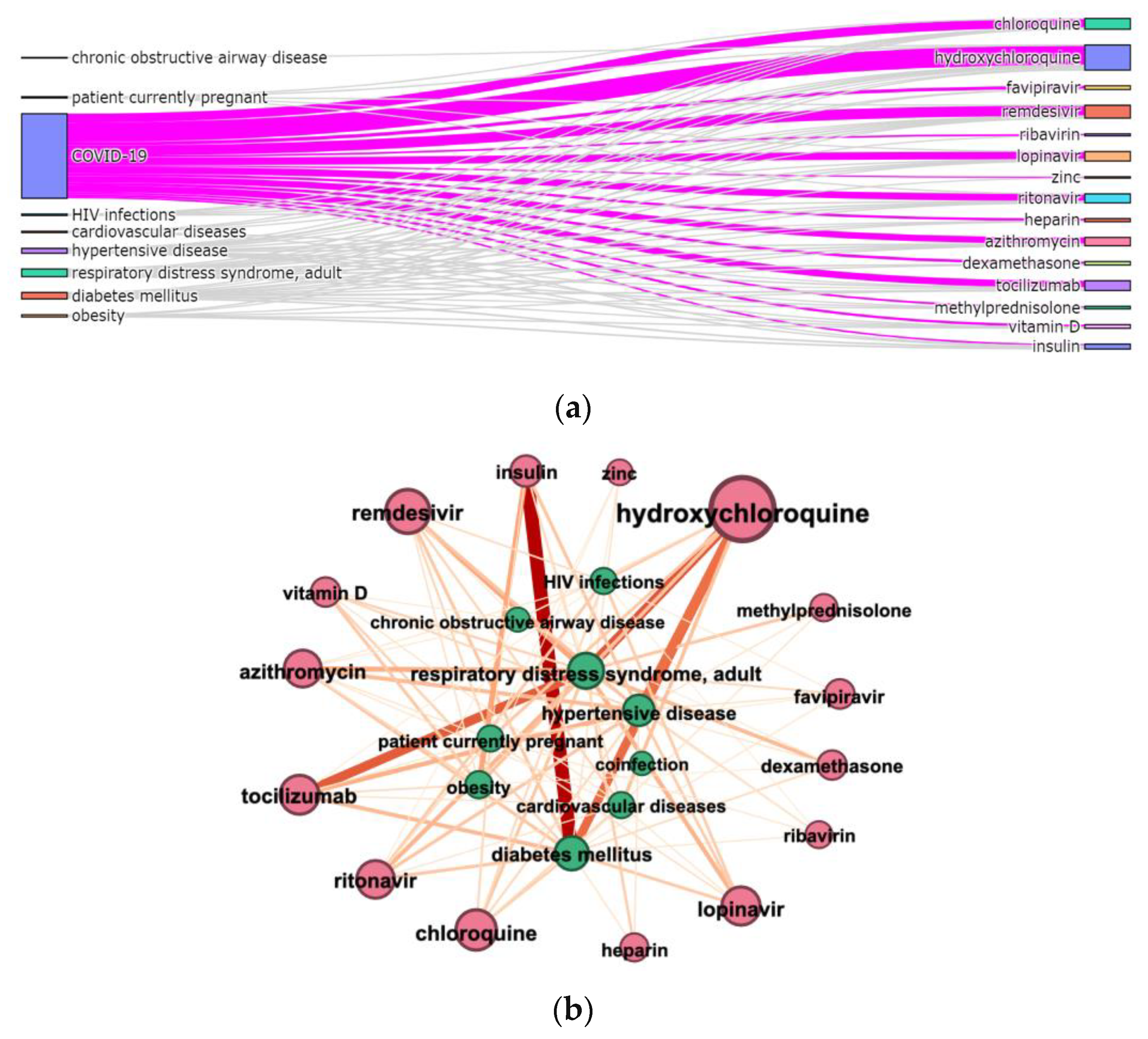
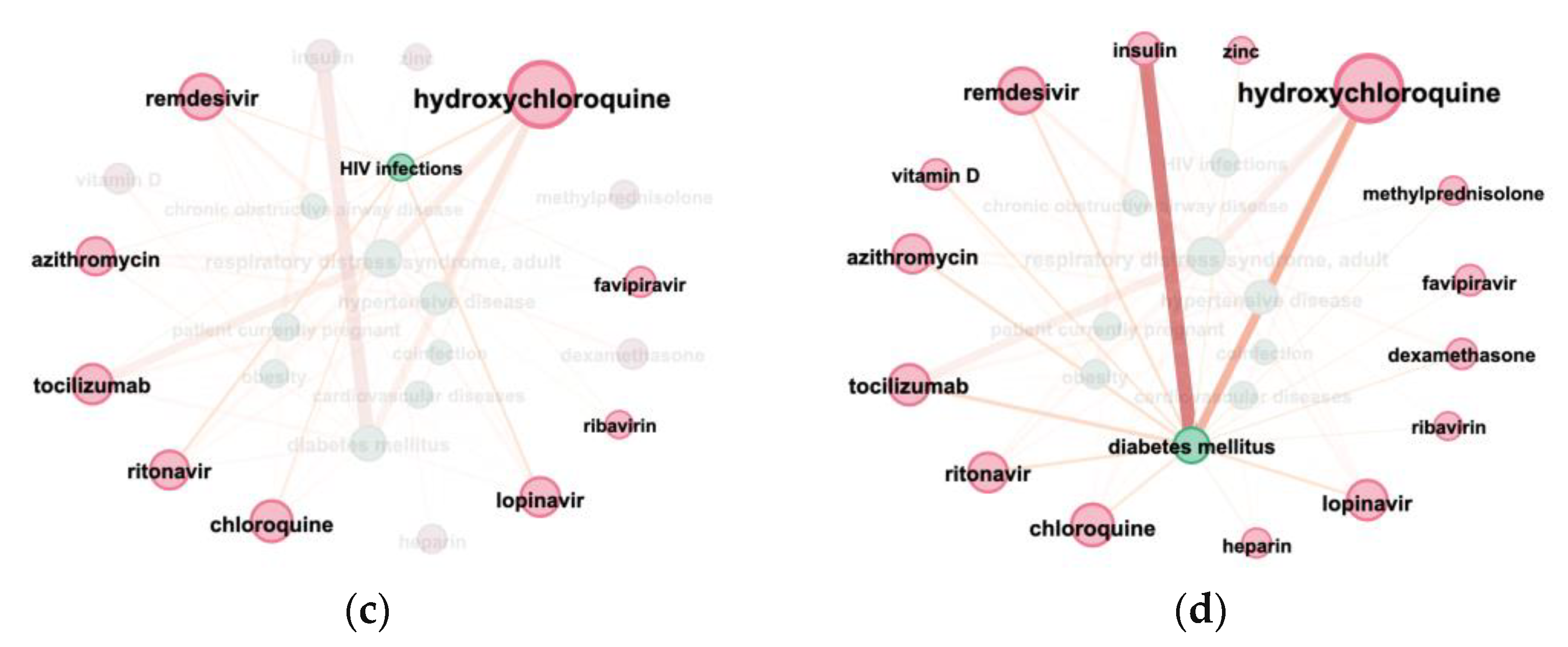
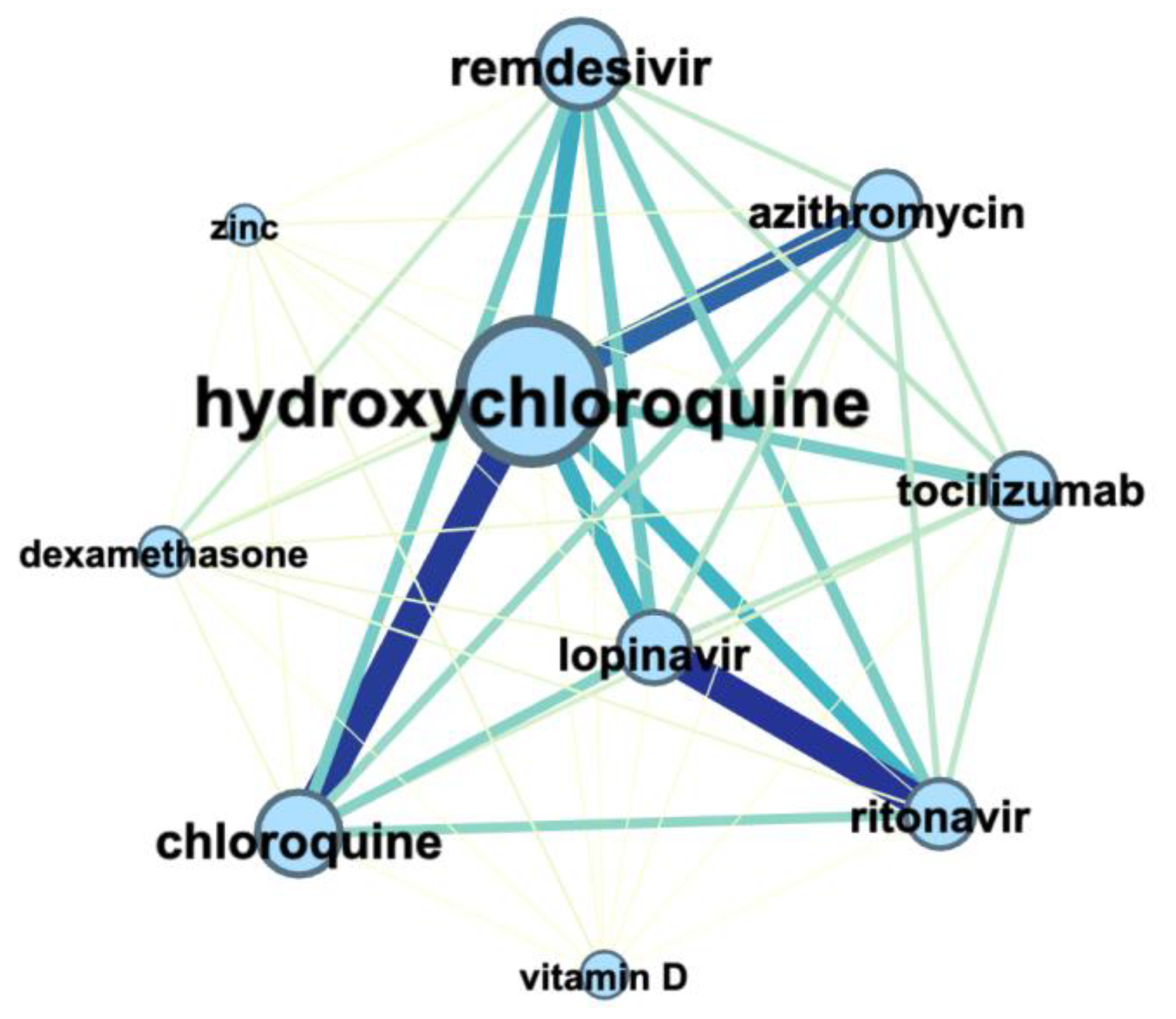
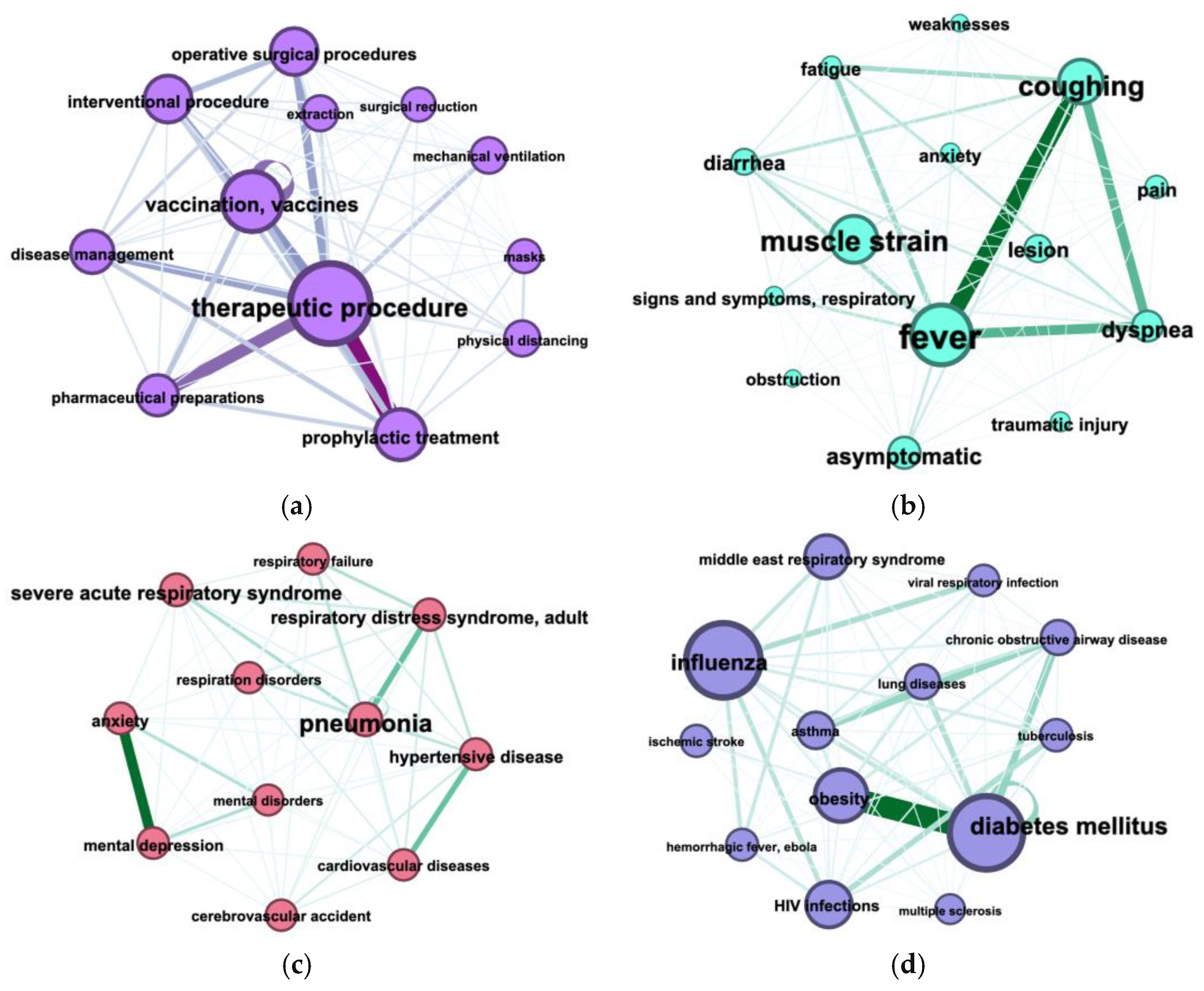
3.3. Terms Co-Occurrence
- Sankey diagram allowed us to investigate relations between two types of terms, e.g., diagnosis and treatment (Figure 7a);
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AI | artificial intelligence |
| BERT | Bidirectional Encoder Representations from Transformers |
| BioBERT | Bidirectional Encoder Representations from Transformers for Biomedical Text Mining |
| CORD | COVID-19 Open Research Dataset |
| COVID-19 | COronaVIrus Disease 2019 |
| CPU | central processing unit |
| DB | database |
| DBMS | database management system |
| DOI | digital object identifier |
| FHIR | fast healthcare interoperability resources |
| ID | identifier |
| LBD | literature-based discovery |
| ML | machine learning |
| NER | named entity recognition |
| NLP | natural language processing |
| NoSQL | not only SQL |
| ODQA | open-domain question answering |
| OSI | open systems interconnection |
| PubMed | a search engine for the MEDLINE and some other databases of references and abstracts on life sciences and biomedical topics |
| PubMedBERT | domain-specific language model pretrained for biomedical texts |
| REST | representational state transfer |
| SARS-CoV-2 | Severe Acute Respiratory Syndrome-related Coronavirus 2 |
| SciBERT | Scientific BERT Trained on Semantic Scholar Data |
| SDK | software development kit |
| SeVeN | semantic vector networks |
| SemMedDB | semantic MEDLINE database |
| SemRep | semantic repository |
| SPECTER | scientific paper embeddings using citation-informed transformers |
| SQL | structured query language |
| TBED | text-based emotion detection |
| UID | user identifier |
| UMLS | unified medical language system |
References
- Minaee, S.; Kalchbrenner, N.; Cambria, E.; Nikzad, N.; Chenaghlu, M.; Gao, J. Deep Learning--based Text Classification: A Comprehensive Review. ACM Comput. Surv. 2021, 54, 40. [Google Scholar] [CrossRef]
- Ligthart, A.; Catal, C.; Tekinerdogan, B. Systematic reviews in sentiment analysis: A tertiary study. Artif. Intell. Rev. 2021, 54, 4997–5053. [Google Scholar] [CrossRef]
- Nadeesha, P.; Dehmer, M.; Emmert-Streib, F. Named Entity Recognition and Relation Detection for Biomedical Information Extraction. Front. Cell Dev. Biol. 2020, 8, 673. [Google Scholar] [CrossRef]
- Widyassari, A.P.; Rustad, S.; Shidik, G.F.; Noersasongko, E.; Syukur, A.; Affandy, A. Review of automatic text summarization techniques & methods. J. King Saud Univ.—Comput. Inf. Sci. 2020, 1319–1578. [Google Scholar] [CrossRef]
- Mutabazi, E.; Ni, J.; Tang, G.; Cao, W. A Review on Medical Textual Question Answering Systems Based on Deep Learning Approaches. Appl. Sci. 2021, 11, 5456. [Google Scholar] [CrossRef]
- Zhu, P.; Li, X.; Li, J.; Zhao, H. Unsupervised Open-Domain Question Answering. arXiv 2021, arXiv:2108.13817. [Google Scholar]
- Wang, L.; Lo, K.; Chandrasekhar, Y.; Reas, R.; Yang, J.; Eide, D.; Funk, K.; Kinney, R.; Liu, Z. CORD-19: The Covid-19 Open Research Dataset. arXiv 2020, arXiv:2004.10706v2. [Google Scholar]
- Extance, A. How AI technology can tame the scientific literature. Nature 2018, 561, 273–274. [Google Scholar] [CrossRef] [PubMed]
- Bullock, J.; Luccioni, A.; Pham, K.H.; Lam, C.S.N.; Luengo-Oroz, M. Mapping the landscape of artificial intelligence applications against COVID-19. J. Artif. Inteill. Res. 2020, 69, 807–845. [Google Scholar] [CrossRef]
- Roberts, K.; Alam, T.; Bedrick, S.; Demner-Fushman, D.; Lo, K.; Soboroff, I.; Voorhees, E.; Wang, L.L.; Hersh, W.R. TREC-Covid: Rationale and structure of an information retrieval shared task for covid-19. J. Am. Med. Inform. Assoc. 2020, 27, 1431–1436. [Google Scholar] [CrossRef] [PubMed]
- Tang, R.; Nogueira, R.; Zhang, E.; Gupta, N.; Cam, P.; Cho, K.; Lin, J. Rapidly bootstrapping a question answering dataset for COVID-19. arXiv 2020, arXiv:2004.11339. [Google Scholar]
- Wang, L.L.; Lo, K. Text mining approaches for dealing with the rapidly expanding literature on COVID-19. Brief. Bioinform. 2021, 22, 781–799. [Google Scholar] [CrossRef] [PubMed]
- Devlin, J.; Chang, M.-W.; Lee, K.; Toutanova, K. BERT: Pre-training of deep bidirectional transformers for language understanding. In Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, Vol. 1 (Long and Short Papers). Minneapolis, Minnesota: Association for Computational Linguistics; Association for Computational Linguistics: Minneapolis, Minnesota, 2019; pp. 4171–4186. [Google Scholar]
- Beltagy, I.; Lo, K.; Cohan, A. SciBERT: A pretrained language model for scientific text. In Proceedings of the 2019 Conference on Empirical Methods in Natural Language Processing and the 9th International Joint Conference on Natural Language Processing (EMNLP-IJCNLP); Association for Computational Linguistics: Hong Kong, China, 2019; pp. 3615–3620. [Google Scholar]
- Lee, J.; Yoon, W.; Kim, S.; Kim, D.; Kim, S.; So, C.H.; Kang, J. BioBERT: A pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 2019, 36, 1234–1240. [Google Scholar] [CrossRef] [PubMed]
- Yuxian, G.; Robert Tinn, R.; Hao Cheng, H.; Lucas, M.; Usuyama, N.; Liu, X.; Naumann, T.; Gao, J.; Poon, H. Domain-Specific Language Model Pretraining for Biomedical Natural Language Processing. arXiv 2020, arXiv:abs/2007.15779. [Google Scholar]
- National Library of Medicine. Available online: https://pubmed.ncbi.nlm.nih.gov (accessed on 1 December 2021).
- COVID-19 Knowledge Graph. Available online: https://covidgraph.org/ (accessed on 1 December 2021).
- Ilievski, F.; Garijo, D.; Chalupsky, H.; Divvala, N.T.; Yao, Y.; Rogers, C.; Li, R.; Liu, J.; Singh, A.; Schwabe, D.; et al. KGTK: A toolkit for large knowledge graph manipulation and analysis. In Proceedings of the 19th International Semantic Web Conference, Athens, Greece, 2–6 November 2020. [Google Scholar]
- A Free and Open Knowledge Base. Available online: https://www.wikidata.org/ (accessed on 1 December 2021).
- Cohan, A.; Feldman, S.; Beltagy, I.; Downey, D.; Weld, D.S. Specter: Document-level representation learning using citation-informed transformers. In Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics, Online, 5–10 July 2020. [Google Scholar]
- Newman-Griffis, D.; Lai, A.M.; Fosler-Lussier, E. Jointly embedding entities and text with distant supervision. In Proceedings of the Third Workshop on Representation Learning for NLP, Association for Computational Linguistics, Melbourne, Australia, 20 July 2018; pp. 195–206. [Google Scholar]
- Espinosa-Anke, L.; Schockaert, S. SeVeN: Augmenting word embeddings with unsupervised relation vectors. In Proceedings of the 27th International Conference on Computational Linguistics, Santa Fe, NM, USA, 20 August 2018; pp. 2653–2665. [Google Scholar]
- Oniani, D.; Jiang, G.; Liu, H.; Shen, F. Constructing co-occurrence network embeddings to assist association extraction for COVID-19 and other coronavirus infectious diseases. J. Am. Med. Inform. Assoc. 2020, 27, 1259–1267. [Google Scholar] [CrossRef] [PubMed]
- Unified Medical Language System (UMLS). Available online: https://www.nlm.nih.gov/research/umls/index.html (accessed on 1 December 2021).
- Introducing Text Analytics for Health. Available online: https://techcommunity.microsoft.com/t5/azure-ai/introducing-text-analytics-for-health/ba-p/1505152 (accessed on 1 December 2021).
- Azure Cosmos DB. Available online: https://azure.microsoft.com/en-us/services/cosmos-db/ (accessed on 1 December 2021).
- Python. Available online: https://www.python.org/ (accessed on 1 December 2021).
- The Open Graph Viz Platform. Available online: https://gephi.org (accessed on 1 December 2021).
- Azure Text Analytics Client Library for Python. Available online: https://github.com/Azure/azure-sdk-for-python/blob/main/sdk/textanalytics/azure-ai-textanalytics/README.md (accessed on 1 December 2021).
- Azure Machine Learning. Available online: https://azure.microsoft.com/en-us/services/machine-learning/ (accessed on 1 December 2021).
- COVID-19 Treatment Guidelines. Chloroquine or Hydroxychloroquine and/or Azithromycin. Available online: https://www.covid19treatmentguidelines.nih.gov/therapies/antiviral-therapy/chloroquine-or-hydroxychloroquine-and-or-azithromycin/ (accessed on 1 December 2021).
- COVID-19 Treatment Guidelines. Lopinavir/Ritonavir and Other HIV Protease Inhibitors. Available online: https://www.covid19treatmentguidelines.nih.gov/therapies/antiviral-therapy/lopinavir-ritonavir-and-other-hiv-protease-inhibitors/ (accessed on 1 December 2021).
- Hamidi Alamdari, D.; Bagheri Moghaddam, A.; Amini, S.; Alamdari, A.H.; Damsaz, M.; Yarahmadi, A. The Application of a Reduced Dye Used in Orthopedics as a Novel Treatment against Coronavirus (COVID-19): A Suggested Therapeutic Protocol. Arch. Bone Jt. Surg. 2020, 8 (Supp. Sl1), 291–294. [Google Scholar] [CrossRef] [PubMed]
- Pundir, H.; Joshi, T.; Joshi, T.; Sharma, P.; Mathpal, S.; Chandra, S.; Tamta, S. Using Chou’s 5-steps rule to study pharmacophore-based virtual screening of SARS-CoV-2 Mpro inhibitors. Mol. Divers. 2021, 25, 1731–1744. [Google Scholar] [CrossRef] [PubMed]
- Caruso, A.; Caccuri, F.; Bugatti, A.; Zani, A.; Vanoni, M.; Bonfanti, P.; Cazzaniga, M.E.; Perno, C.F.; Messa, C.; Alberghina, L. Methotrexate inhibits SARS-CoV-2 virus replication “in vitro”. J. Med. Virol. 2021, 93, 1780–1785. [Google Scholar] [CrossRef] [PubMed]
- COVID-19 Treatment Guidelines. Remdesivir. Available online: https://www.covid19treatmentguidelines.nih.gov/therapies/antiviral-therapy/remdesivir/ (accessed on 1 December 2021).

| Entity Type | Count | Relation Type | Count |
|---|---|---|---|
| AdministrativeEvent | 171,937 | Abbreviation | 713,475 |
| Age | 155,693 | DirectionOfBodyStructure | 12,675 |
| BodyStructure | 245,787 | DirectionOfCondition | 13,437 |
| CareEnvironment | 130,770 | DirectionOfExamination | 3391 |
| ConditionQualifier | 486,762 | DirectionOfTreatment | 4276 |
| Date | 42,153 | DosageOfMedication | 23,760 |
| Diagnosis | 2,477,847 | FormOfMedication | 6510 |
| Direction | 34,398 | FrequencyOfMedication | 4903 |
| Dosage | 35,550 | FrequencyOfTreatment | 6921 |
| ExaminationName | 2,226,245 | QualifierOfCondition | 459,129 |
| FamilyRelation | 80,454 | RelationOfExamination | 144,470 |
| Frequency | 21,274 | RouteOfMedication | 12,612 |
| Gender | 67,145 | TimeOfCondition | 145,256 |
| GeneOrProtein | 39,782 | TimeOfEvent | 27,825 |
| HealthcareProfession | 120,570 | TimeOfExamination | 101,553 |
| MeasurementUnit | 384,921 | TimeOfMedication | 14,981 |
| MeasurementValue | 1,030,936 | TimeOfTreatment | 59,917 |
| MedicationClass | 242,516 | UnitOfCondition | 41,130 |
| MedicationForm | 7697 | UnitOfExamination | 279,532 |
| MedicationName | 297,463 | ValueOfCondition | 58,324 |
| MedicationRoute | 16,988 | ValueOfExamination | 853,503 |
| RelationalOperator | 190,191 | ||
| SymptomOrSign | 1,117,712 | ||
| Time | 336,369 | ||
| TreatmentName | 1,314,612 | ||
| Variant | 4427 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Soshnikov, D.; Petrova, T.; Soshnikova, V.; Grunin, A. Analyzing COVID-19 Medical Papers Using Artificial Intelligence: Insights for Researchers and Medical Professionals. Big Data Cogn. Comput. 2022, 6, 4. https://doi.org/10.3390/bdcc6010004
Soshnikov D, Petrova T, Soshnikova V, Grunin A. Analyzing COVID-19 Medical Papers Using Artificial Intelligence: Insights for Researchers and Medical Professionals. Big Data and Cognitive Computing. 2022; 6(1):4. https://doi.org/10.3390/bdcc6010004
Chicago/Turabian StyleSoshnikov, Dmitry, Tatiana Petrova, Vickie Soshnikova, and Andrey Grunin. 2022. "Analyzing COVID-19 Medical Papers Using Artificial Intelligence: Insights for Researchers and Medical Professionals" Big Data and Cognitive Computing 6, no. 1: 4. https://doi.org/10.3390/bdcc6010004
APA StyleSoshnikov, D., Petrova, T., Soshnikova, V., & Grunin, A. (2022). Analyzing COVID-19 Medical Papers Using Artificial Intelligence: Insights for Researchers and Medical Professionals. Big Data and Cognitive Computing, 6(1), 4. https://doi.org/10.3390/bdcc6010004







