Importation, Local Transmission, and Model Selection in Estimating the Transmissibility of COVID-19: The Outbreak in Shaanxi Province of China as a Case Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Data
2.2. Models
2.2.1. Richards Growth Model
2.2.2. Renewal Equation Model
2.2.3. SEDAR Transmission Model
2.2.4. SEEDAR Transmission Model
2.2.5. SEEDDAAR Transmission Model
2.3. Inference Method by Calibration to Shaanxi Outbreak
MCMC Sampling
Θj (t) = Θj (t−1) otherwise (rejected).
3. Results
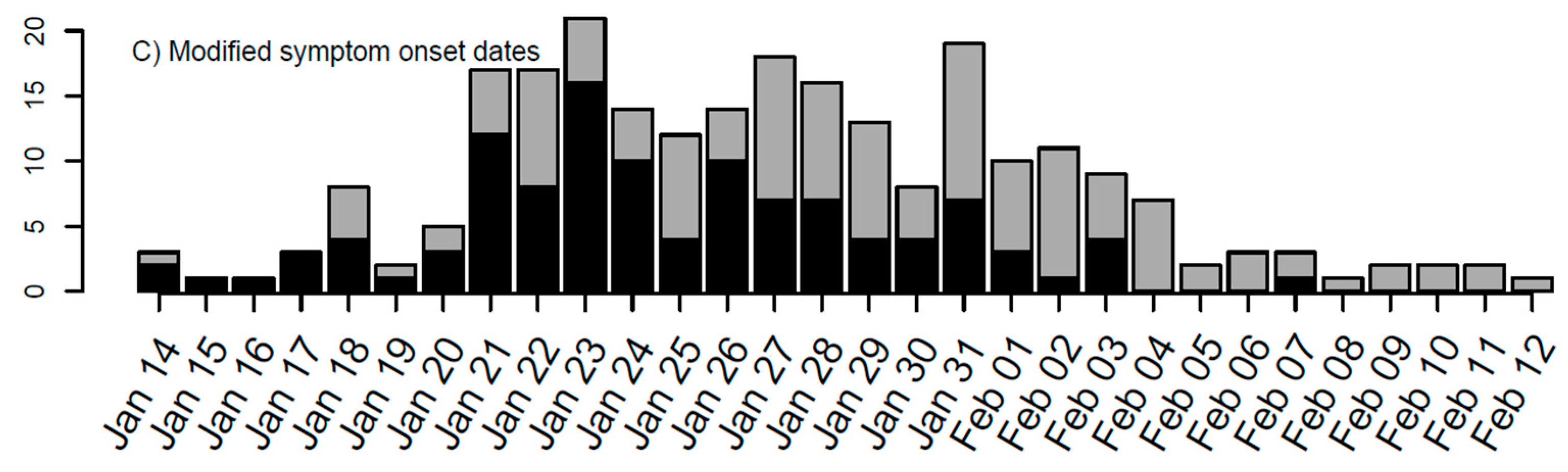
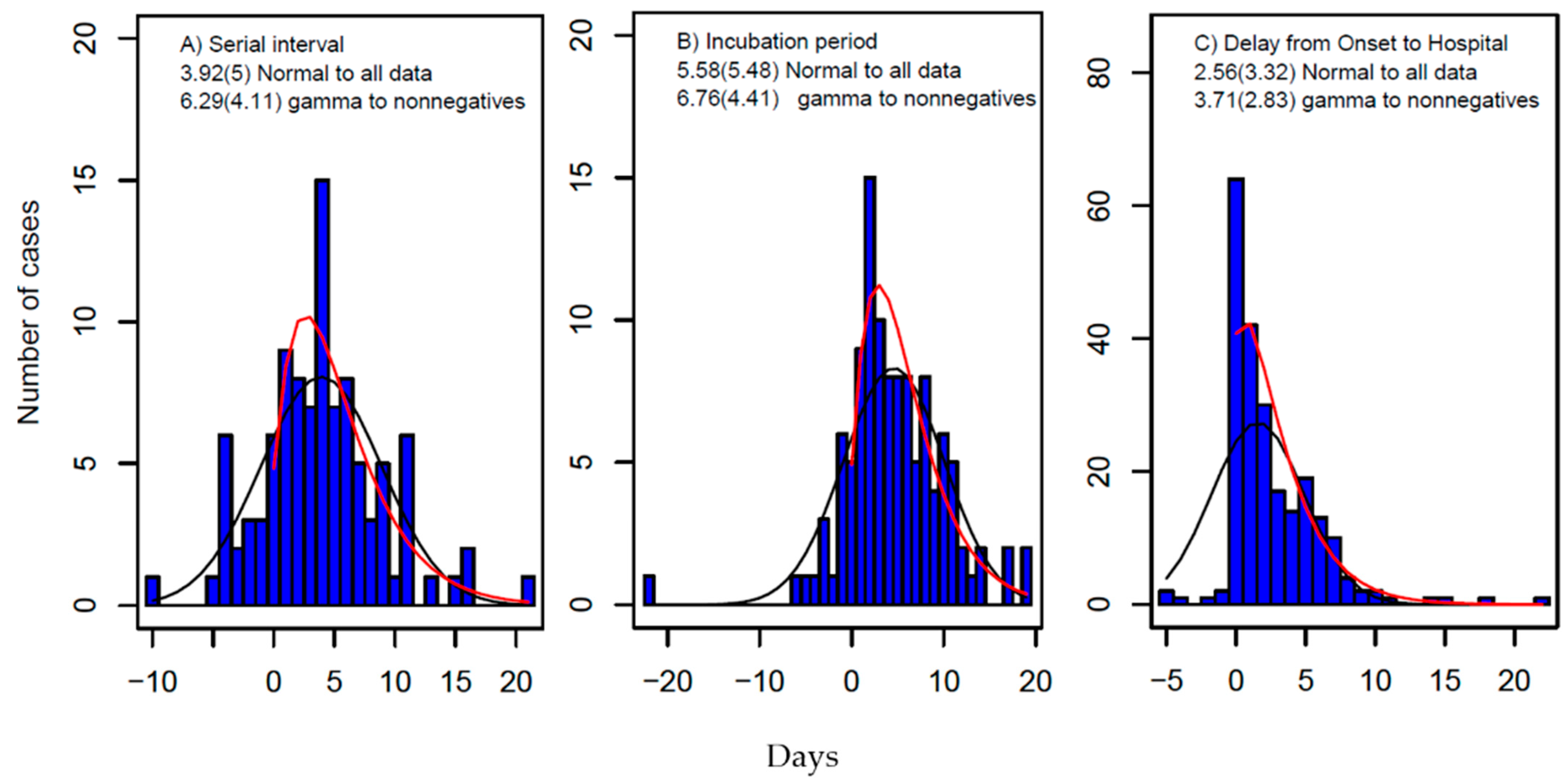
3.1. Estimates of SI and Incubation Period from Line List Data
3.2. Estimate of R0 in Shaanxi Outbreak
3.2.1. Richards Growth Model
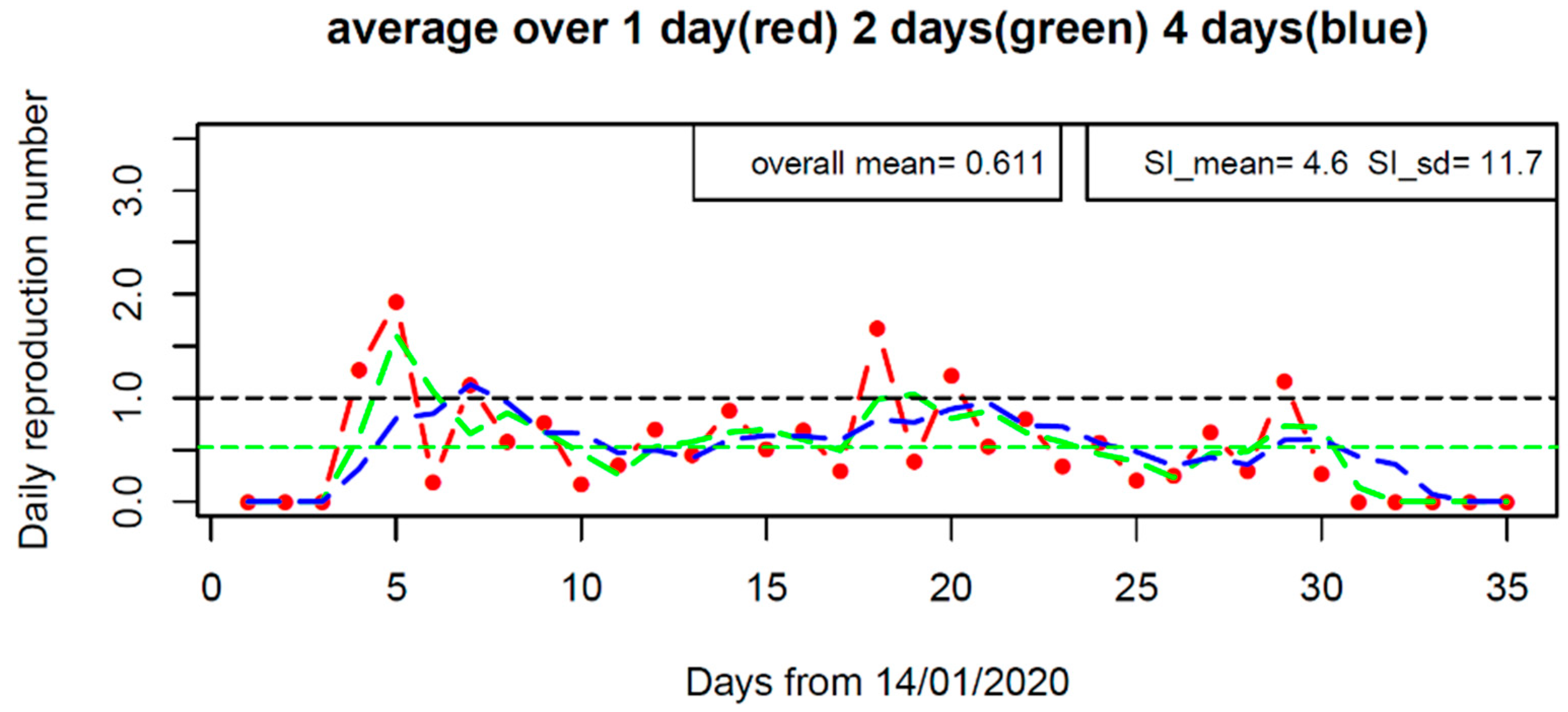
3.2.2. Renewal Equation Model
3.2.3. SEDAR Model
3.2.4. SEEDAR Model
3.2.5. SEEDDAAR Model
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Worldometer Coronavirus. Available online: https://www.worldometers.info/coronavirus/ (accessed on 19 September 2021).
- World Health Organization. Available online: https://www.who.int/publications/m/item/weekly-epidemiological-update-on-COVID-19 (accessed on 31 August 2021).
- Hale, T.; Angrist, N.; Goldszmidt, R.; Kira, B.; Petherick, A.; Phillips, T.; Webster, S.; Cameron-Blake, E.; Hallas, L.; Majumdar, S.; et al. A global panel database of pandemic policies (Oxford COVID-19 Government Response Tracker). Nat. Hum. Behav. 2021, 5, 529–538. [Google Scholar] [CrossRef] [PubMed]
- Ross, R. The Prevention of Malaria; John Murray: London, UK, 1911. [Google Scholar]
- Anderson, R.M.; May, R.M. Infectious Diseases of Humans: Dynamics and Control; Oxford University Press: Oxford, UK, 1991. [Google Scholar]
- Heesterbeek, J.A.P.; Diekmann, O. Mathematical Epidemiology of Infectious Diseases: Model Building, Analysis and Interpretation; John Wiley & Sons: Hoboken, NJ, USA, 2000. [Google Scholar]
- Keeling, M.J.; Rohani, P. Modelling Infectious Diseases in Human and Animals; Princeton University Press: Princeton, NJ, USA, 2007. [Google Scholar]
- Hao, X.; Cheng, S.; Wu, D.; Wu, T.; Lin, X.; Wang, C. Reconstruction of the full transmission dynamics of COVID-19 in Wuhan. Nature 2020, 584, 420–424. [Google Scholar] [CrossRef] [PubMed]
- Kucharski, A.J.; Russell, T.W.; Diamond, C.; Liu, Y.; Edmunds, J.; Funk, S.; Eggo, R.M.; Sun, F.; Jit, M.; Munday, J.D.; et al. Early dynamics of transmission and control of COVID-19: A mathematical modelling study. Lancet Infect. Dis. 2020, 20, 553–558. [Google Scholar] [CrossRef]
- Lai, S.; Ruktanonchai, N.W.; Zhou, L.; Proper, O.; Luo, W.; Floyd, J.R.; Wesolowski, A.; Santillana, M.; Zhang, C.; Du, X.; et al. Effect of non-pharmaceutical interventions for containing the COVID-19 outbreak in China. Nature 2020, 585, 410–413. [Google Scholar] [CrossRef]
- Wu, J.T.; Leung, K.; Leung, G.M. Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: A modelling study. Lancet 2020, 395, 689–697. [Google Scholar] [CrossRef]
- Zhang, J.; Litvinova, M.; Wang, W.; Wang, Y.; Deng, X.; Chen, X.; Li, M.; Zheng, W.; Yi, L.; Chen, X.; et al. Evolving epidemiology and transmission dynamics of coronavirus disease 2019 outside Hubei province, China: A descriptive and modelling study. Lancet Infect. Dis. 2020, 20, 793–802. [Google Scholar] [CrossRef]
- Adekunle, A.I.; Adegboye, O.A.; Gayawan, E.; McBryde, E.S. Is Nigeria really on top of COVID-19? Message from effective reproduction number. Epidemiol. Infect. 2020, 148, e166. [Google Scholar] [CrossRef]
- Heffernan, J.M.; Smith, R.J.; Wahl, L.M. Perspectives on the basic reproductive ratio. J. R. Soc. Interface 2005, 2, 281–293. [Google Scholar] [CrossRef]
- Vynnycky, E.; White, R.G. An Introduction to Infectious Disease Modelling; Oxford University Press: Oxford, UK, 2010. [Google Scholar]
- Biggerstaff, M.; Cowling, B.J.; Cucunubá, Z.M.; Dinh, L.; Ferguson, N.M.; Gao, H.; Hill, V.; Imai, N.; Johansson, M.A.; Kada, S.; et al. Early Insights from Statistical and Mathematical Modeling of Key Epidemiologic Parameters of COVID-19. Emerg. Infect. Dis. 2020, 26, e201074. [Google Scholar] [CrossRef]
- Bai, Y.; Liu, K.; Chen, Z.; Chen, B.; Shao, Z. Early transmission dynamics of novel coronavirus pneumonia Epidemic in Shaanxi Province. Chin. J. Nosocomiol. 2020, 30, 834–838. [Google Scholar]
- Wu, W.; Bai, R.; Li, D.; Feng, A.; Xu, A.; Lü, J. Preliminary prediction of the epidemic trend of 2019 novel coronavirus (2019-nCoV) pneumonia in Guangdong province. J. Jinan Univ. 2020, 41, 1–6. [Google Scholar]
- D’Arienzo, M.; Coniglio, A. Assessment of the SARS-CoV_2 basic reproduction number, R0, based on the early phase of COVID-19 outbreak in Italy. Biosaf. Health 2020, 2, 57–59. [Google Scholar] [CrossRef]
- Cheng, Q.; Liu, Z.; Cheng, G.; Huang, J. Heterogeneity and effectiveness analysis of COVID-19 prevention and control in major cities in China through time-varying reproduction number estimation. Sci. Rep. 2020, 10, 21953. [Google Scholar] [CrossRef]
- Han, K.; Jia, W.; Cai, W.; Wang, S.; Song, Y.; Yang, S.; Li, J.; Kou, F.; Liu, M.; He, Y.; et al. Estimation of real-time basic reproduction number and epidemic status of 20-19 novel coronavirus disease (COVID-19) in first-tier cities. Acad. J. Chin. PLA Med. Sch. 2020, 4, 421–426. [Google Scholar]
- Song, P.; Chen, C.; Lou, Y.; Jiang, H.; Li, W.; Zhu, L. Assessing effectiveness of integrated strategies for preventing and controlling the outbreak of COVID-19 and predicting impact of opening exit channels to leave Hubei Province. Chin. J. Appl. Prob. Stat. 2020, 36, 321–330. [Google Scholar]
- Yuan, J.; Li, M.; Lv, G.; Lu, Z.K. Monitoring transmissibility and mortality of COVID-19 in Europe. Int. J. Infect Dis. 2020, 95, 311–315. [Google Scholar] [CrossRef]
- Yang, L.; Wang, C.; Shang, H.; Zhang, X.; Zhang, L.; Wang, K. Epidemiological parameter estimation and characteristics of the novel coronavirus (COVID-19) transmission in Shaanxi Province. J. Pub. Health Prev. Med. 2021, 32, 195–199. [Google Scholar]
- Birrell, P.J.; Zhang, X.-S.; Pebody, R.G.; Gay, N.J.; De Angelis, D. Reconstructing a spatially heterogeneous epidemic: Characterising the geographic spread of 2009 A/H1N1pdm infection in England. Sci. Rep. 2016, 6, 29004. [Google Scholar] [CrossRef] [PubMed]
- Chong, K.C.; Cheng, W.; Zhao, S.; Ling, F.; Mohammad, K.N.; Wang, M.; Zee, B.C.; Wei, L.; Xiong, X.; Liu, H.; et al. Transmissibility of coronavirus disease 2019 in Chinese cities with different dynamics of imported cases. PeerJ 2020, 8, e10350. [Google Scholar] [CrossRef]
- Yuan, H.Y.; Blakemore, C. The impact of multiple non-pharmaceutical interventions on controlling COVID-19 outbreak without lockdown in Hong Kong: A modelling study. Lancet Reg. Health West. Pac. 2021, 20, 100343. [Google Scholar] [CrossRef]
- Yuan, H.Y.; Blakemore, C. The impact of contact tracing and testing on controlling COVID-19 outbreak without lockdown in Hong Kong: An observational study. Lancet Reg. Health West. Pac. 2022, 20, 100374. [Google Scholar] [CrossRef] [PubMed]
- Bernal, J.L.; Panagiotopoulos, N.; Byers, C.; Vilaplana, T.G.; Boddington, N.; Zhang, X.-S.; Charlett, A.; Elgohari, S.; Coughlan, L.; Whillock, R.; et al. Transmission dynamics of COVID-19 in household and community settings in the United Kingdom, January to March 2020. Eurosurveillance 2022, 27, 2001551. [Google Scholar]
- Zhang, X.S.; Vynnycky, E.; Charlett, A.; De Angelis, D.; Chen, Z.J.; Liu, W. Transmission dynamics and control measures of COVID-19 outbreak in China: A modelling study. Sci. Rep. 2021, 11, 2652. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Zeng, Z.; Wang, K.; Wong, S.-S.; Liang, W.; Zanin, M.; Liu, P.; Cao, X.; Gao, Z.; Mai, Z.; et al. Modified SEIR and AI prediction of the epidemics trend of COVID-19 in China under public health interventions. J. Torac. Dis. 2020, 12, 165–174. [Google Scholar] [CrossRef] [PubMed]
- Flaxman, S.; Mishra, S.; Gandy, A.; Unwin, H.J.T.; Mellan, T.A.; Coupland, H.; Whittaker, C.; Zhu, H.; Berah, T.; Eaton, J.W.; et al. Estimating the effects of non-pharmaceutical interventions on COVID-19 in Europe. Nature 2020, 584, 257–261. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.Y.; Lei, B.; Mallick, B. Estimation of COVID-19 spread curves integrating global data and borrowing information. PLoS ONE 2020, 15, e0236860. [Google Scholar] [CrossRef] [PubMed]
- Dahal, S.; Luo, R.; Subedi, R.K.; Dhimal, M.; Chowell, G. Transmission Dynamics and Short-Term Forecasts of COVID-19: Nepal 2020/2021. Epidemiologia 2021, 2, 639–659. [Google Scholar] [CrossRef]
- Swaraj, A.; Verma, K.; Kaur, A.; Singh, G.; Kumar, A.; Melo, L.; Sale, D. Implementation of stacking based ARIMA model for prediction of COVID-19 cases in India. J. Biomed. Inform. 2021, 121, 103887. [Google Scholar] [CrossRef]
- Shaanxi Provincial Health Committee. Available online: http://sxwjw.shaanxi.gov.cn/sy/wjyw/index_66.html (accessed on 18 October 2020).
- Richards, F. A flexible growth function for empirical use. J. Exp. Bot. 1959, 10, 290–300. [Google Scholar] [CrossRef]
- Nelder, J.A. 182. note: An alternative form of a generalized logistic equation. Biometrics 1962, 18, 614–616. [Google Scholar] [CrossRef]
- Hsieh, Y.-H. 2015 Middle East Respiratory Syndrome Coronavirus (MERS-CoV) nosocomial outbreak in South Korea: Insights from modeling. PeerJ 2015, 3, e1505. [Google Scholar] [CrossRef]
- Wallinga, J.; Lipsitch, M. How generation intervals shape the relationship between growth rates and reproductive number. Proc. Biol. Sci. 2007, 27, 599–604. [Google Scholar] [CrossRef]
- Nishiura, H.; Chowell, G.; Safan, M.; Castillo-Chavez, C. Pros and cons of estimating the reproduction number from early epidemic growth rate of influenza A (H1N1) 2009. Theor. Biol. Med. Model. 2010, 7, 1. [Google Scholar] [CrossRef]
- Fraser, C. Estimating Individual and Household Reproduction Numbers in an Emerging Epidemic. PLoS ONE 2007, 2, e758. [Google Scholar] [CrossRef]
- Cori, A.; Ferguson, N.M.; Fraser, C.; Cauchemez, S. A new Framework and software to estimate time varying reproduction numbers during epidemics. Am. J. Epidemiol. 2013, 178, 1505–1512. [Google Scholar] [CrossRef] [PubMed]
- Roberts, M.G.; Nishiura, H. Early Estimation of the Reproduction Number in the Presence of Imported Cases: Pandemic Influenza H1N1-2009 in New Zealand. PLoS ONE 2011, 6, e17835. [Google Scholar] [CrossRef]
- Griffin, J.T.; Garske, T.; Ghani, A.C.; Clarke, P.S. Joint estimation of the basic reproduction number and generation time parameters for infectious disease outbreaks. Biostatistics 2011, 12, 303–312. [Google Scholar] [CrossRef]
- Ferretti, L.; Ledda, A.; Wymant, C.; Zhao, L.; Ledda, V.; Abeler-Dörner, L.; Kendall, M.; Nurtay, A.; Cheng, H.-Y.; Ng, T.-C.; et al. The timing of COVID-19 transmission. medRxiv 2020. [Google Scholar] [CrossRef]
- Wearing, H.J.; Rohani, P.; Keeling, M.J. Appropriate models for the management of infectious diseases. PLoS Med. 2005, 2, e174. [Google Scholar] [CrossRef]
- Pasetto, D.; Lemaitre, J.C.; Bertuzzo, E.; Gatto, M.; Rinaldo, A. Range of reproduction number estimates for COVID-19 spread. Biochem. Biophys. Res. Commun. 2021, 538, 253–258. [Google Scholar] [CrossRef]
- Xiang, Y.; Jia, Y.; Chen, L.; Guo, L.; Shu, B.; Long, E. COVID-19 epidemic prediction and the impact of public health interventions: A review of COVID-19 epidemic models. Infect. Dis. Model. 2021, 6, 324–342. [Google Scholar] [CrossRef] [PubMed]
- Röst, G.; Bartha, F.A.; Bogya, N.; Boldog, P.; Dénes, A.; Ferenci, T.; Horváth, K.J.; Juhász, A.; Nagy, C.; Tekeli, T.; et al. Early phase of the COVID-19 outbreak in Hungary and post-lockdown scenarios. Viruses 2020, 12, 708. [Google Scholar] [CrossRef] [PubMed]
- Polver, M.; Previdi, F.; Mazzoleni, M.; Zucchi, A. A SIAT3 HE model of the COVID-19 pandemic in Bergamo, Italy. IFAC Pap. 2021, 54, 263–268. [Google Scholar] [CrossRef]
- Champredon, D.; Dushoff, J.; Earn, D.J.D. Equivalence of the Erlang-distributed SEIR epidemicmodel and the renewal equation. SIAM J. Appl. Math. 2018, 78, 3258–3278. [Google Scholar] [CrossRef]
- Bettencourt, L.M.A.; Ribeiro, R.M. Real time Bayesian estimation of the epidemic potential of emerging infectious diseases. PLoS ONE 2008, 3, e2185. [Google Scholar] [CrossRef]
- Cauchemez, S.; Carrat, F.; Viboud, C.; Valleron, A.J.; Boëlle, P.Y. A Bayesian MCMC approach to study transmission of influenza: Application to household longitudinal data. Stat. Med. 2004, 23, 3469–3487. [Google Scholar] [CrossRef]
- Burnham, K.P.; Anderson, D.R. Model Selection and Multimodel Inference: A Practical Information—Theoretic Approach, 2nd ed.; Springer: New York, NY, USA, 2002. [Google Scholar]
- Spiegelhalter, D.J.; Best, N.; Carlin, B.P.; van der Linde, A. Bayesian measures of model complexity and fit. J. Roy. Stat. Soc. B 2002, 64, 583–639. [Google Scholar] [CrossRef]
- Alene, M.; Yismaw, L.; Assemie, M.A.; Ketema, D.B.; Gietaneh, W.; Birhan, T.Y. Serial interval and incubation period of COVID-19: A systematic review and meta-analysis. BMC Infect. Dis. 2021, 21, 257. [Google Scholar] [CrossRef]
- Chen, Z.J.; Wei, T.; Li, H.D.; Feng, L.H.; Liu, H.N.; Li, N.; Gu, R.; Zhang, N.; Lu, W.; Zhang, X.-S. Renewal equation model for the COVID-19 in Yunnan. Chin. Med. Humanit. 2021, 7, 53–55. [Google Scholar]
- Liu, Y.; Gayle, A.A.; Wilder-Smith, A.; Rocklov, J. The reproductive number of COVID-19 is higher compared to SARS coronavirus. J. Travel Med. 2020, 27, taaa021. [Google Scholar] [CrossRef]
- Achaiah, N.C.; Subbarajasetty, S.B.; Shetty, R.M. R0 and Re of COVID-19: Can We Predict When the Pandemic Outbreak will be Contained? Indian J. Crit. Care Med. 2020, 24, 1125. [Google Scholar] [CrossRef]




| Parameter | Richards Growth | Renewal Equation | SEDAR | SEEDAR | SEEDDAAR | ||||
|---|---|---|---|---|---|---|---|---|---|
| Prior | Posterior | Prior | Posterior | Prior | Posterior | Prior | Posterior | Posterior | |
| Growth rate (r) | [0,1.0] | 0.02 [0.012,0.032] | – | – | – | – | – | – | – |
| Final epidemic size (K) | [1,6600] | 3315 [56,6521] | – | – | – | – | – | – | – |
| Scaling exponent (ν) | [0.1,50] | 24.51 [0.72,48.81] | – | – | – | – | – | – | – |
| Mean of SI (SI_mean) | – | – | U [3.5,10.0] | 4.66 [3.53,7.18] | – | – | – | – | – |
| Standard deviation of SI (SI_sd) | – | – | U [3.0,15.0] | 11.73 [5.85,14.88] | – | – | – | – | – |
| Transmission coefficient (β) | – | – | – | – | U [.001,0.5] | 0.155 [0.117,0.186] | U [.001,0.5] | 0.066 [0.029,0.154] | 0.072 [0.032,0.180] |
| Latent period (L1) * | – | – | – | – | U [1.6,14.0] | 1.81 [1.61,2.82] | U [1.0,10.0] | 5.04 [1.25,9.65] | 5.25 [1.28,9.76] |
| Pre-symptomatic infectious period (L3) | – | – | – | – | – | – | U [1.0,10.0] | 1.45 [1.04,4.43] | 1.45 [1.04,4.43] |
| Infectious period (D1) of diseased infections * | – | – | – | – | U [3.5,25.0] | 3.75 [3.51,5.16] | U [1.5,15.0] | 4.78 [1.61,14.06] | 5.40 [1.68,13.97] |
| Dispersion parameter (η) | – | – | U [1.01,50.0] | 1.58 [1.06,2.86] | U [1.01,50.0] | 2.47 [1.56,4.431] | U [1.01,50.0] | 1.73 [1.08,3.26] | 1.71 [1.09,3.18] |
| R0♦ | – | 1.13 [1.08,1.21] | U [0.05,3.0] | 0.61 [0.54,0.68] | – | 0.59 [0.50,0.70] | – | 0.45 [0.30,0.76] | 0.53 [0.35,0.85] |
| DIC ♣ | – | 140.2 | – | 127.9 | – | 175.1 | – | 160.5 | 160.8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, X.-S.; Xiong, H.; Chen, Z.; Liu, W. Importation, Local Transmission, and Model Selection in Estimating the Transmissibility of COVID-19: The Outbreak in Shaanxi Province of China as a Case Study. Trop. Med. Infect. Dis. 2022, 7, 227. https://doi.org/10.3390/tropicalmed7090227
Zhang X-S, Xiong H, Chen Z, Liu W. Importation, Local Transmission, and Model Selection in Estimating the Transmissibility of COVID-19: The Outbreak in Shaanxi Province of China as a Case Study. Tropical Medicine and Infectious Disease. 2022; 7(9):227. https://doi.org/10.3390/tropicalmed7090227
Chicago/Turabian StyleZhang, Xu-Sheng, Huan Xiong, Zhengji Chen, and Wei Liu. 2022. "Importation, Local Transmission, and Model Selection in Estimating the Transmissibility of COVID-19: The Outbreak in Shaanxi Province of China as a Case Study" Tropical Medicine and Infectious Disease 7, no. 9: 227. https://doi.org/10.3390/tropicalmed7090227
APA StyleZhang, X.-S., Xiong, H., Chen, Z., & Liu, W. (2022). Importation, Local Transmission, and Model Selection in Estimating the Transmissibility of COVID-19: The Outbreak in Shaanxi Province of China as a Case Study. Tropical Medicine and Infectious Disease, 7(9), 227. https://doi.org/10.3390/tropicalmed7090227






