Potential Animal Reservoir of Mycobacterium ulcerans: A Systematic Review
Abstract
1. Introduction
2. Materials and Methods
3. Results
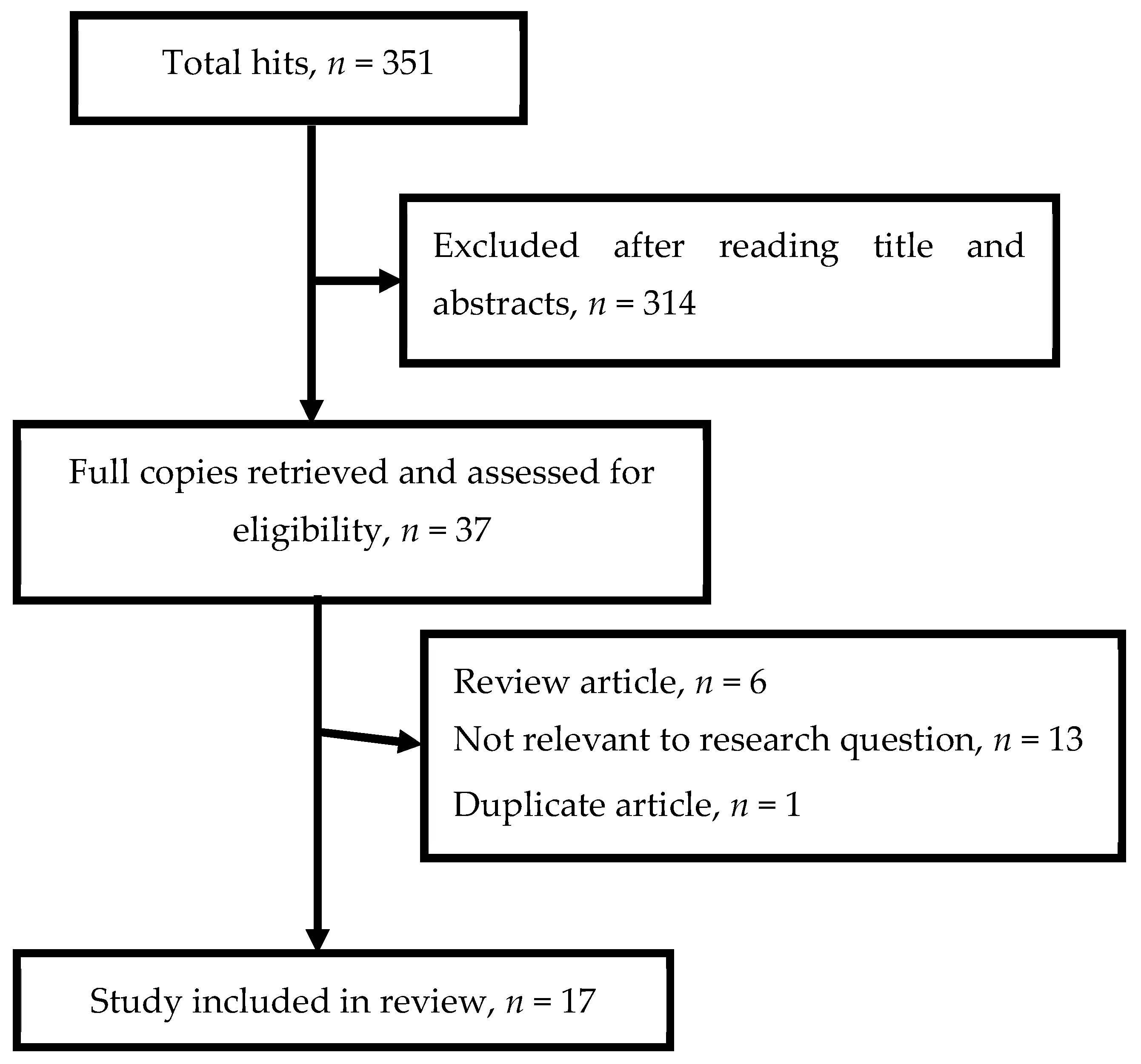
3.1. Results of the Literature Search and Method of Inclusion
3.2. Basic Characteristics of Selected Studies
4. Discussion on Possible Reservoirs and Vectors of Mycobacterium ulcerans by Country
4.1. Australia
4.2. Africa
4.3. Other Countries
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Alsop, D.G. The Bairnsdale ulcer. Aust. N. Z. J. Surg. 1972, 41, 317–319. [Google Scholar] [CrossRef] [PubMed]
- MacCallum, P.T.J.C.; Tolhurst, J.C.; Buckle, G.; Sissons, H.A. A new mycobacterial infection in man. J. Pathol. Bacteriol. 1948, 60, 93–122. [Google Scholar] [CrossRef] [PubMed]
- Clancey, J.; Dodge, R.; Lunn, H.F. Study of a Mycobacterium causing skin ulceration in Uganda. Ann. Soc. Belg. Med. Trop. 1920, 42, 585–590. [Google Scholar]
- Johnson, P.D.; Azuolas, J.; Lavender, C.J.; Wishart, E.; Stinear, T.P.; Hayman, J.A.; Brown, L.; Jenkin, G.A.; Fyfe, J.A. Mycobacterium ulcerans in mosquitoes captured during outbreak of Buruli ulcer, southeastern Australia. Emerg. Infect. Dis. 2007, 13, 1653–1660. [Google Scholar] [CrossRef] [PubMed]
- Carson, C.; Lavender, C.J.; Handasyde, K.A.; O’Brien, C.R.; Hewitt, N.; Johnson, P.D.; Fyfe, J.A. Potential wildlife sentinels for monitoring the endemic spread of human Buruli ulcer in south-east Australia. PLoS Negl. Trop. Dis. 2014, 8, e2668. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Neglected Tropical Diseases. 2018. Available online: http://www.who.int/neglected_diseases/diseases/en/ (accessed on 4 October 2017).
- World Health Organization. Distribution of Buruli Ulcer, Worldwide 2014; WHO: Geneva, Switzerland, 2014. [Google Scholar]
- Steffen, C.M.; Smith, M.; McBride, W.J. Mycobacterium ulcerans infection in North Queensland: The ‘Daintree ulcer’. ANZ J. Surg. 2010, 80, 732–736. [Google Scholar] [CrossRef] [PubMed]
- Röltgen, K.; Pluschke, G.; Johnson, P.D.; Fyfe, J. Mycobacterium ulcerans DNA in bandicoot excreta in Buruli ulcer-endemic area, northern Queensland, Australia. Emerg. Infect. Dis. 2017, 23, 2042–2045. [Google Scholar] [CrossRef] [PubMed]
- Francis, G.; Whitby, M.; Woods, M. Mycobacterium ulcerans infection: A rediscovered focus in the Capricorn Coast region of central Queensland. Med. J. Aust. 2006, 185, 179–180. [Google Scholar] [PubMed]
- Radford, A.J. Mycobacterium ulcerans in Australia. Aust. N. Z. J. Med. 1975, 5, 162–169. [Google Scholar] [CrossRef] [PubMed]
- McOrist, S.; Jerrett, I.V.; Anderson, M.; Hayman, J. Cutaneous and respiratory tract infection with Mycobacterium ulcerans in two koalas (Phascolarctos cinereus). J. Wildl. Dis. 1985, 21, 171–173. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.J.; McOrist, S.; Bilney, R. Epidemiology of Mycobacterium ulcerans infection in koalas (Phascolarctos cinereus) on Raymond Island, southeastern Australia. J. Wildl. Dis. 1987, 23, 386–390. [Google Scholar] [CrossRef] [PubMed]
- Fyfe, J.A.; Lavender, C.J.; Handasyde, K.A.; Legione, A.R.; O’Brien, C.R.; Stinear, T.P.; Pidot, S.J.; Seemann, T.; Benbow, M.E.; Wallace, J.R.; et al. A major role for mammals in the ecology of Mycobacterium ulcerans. PLoS Negl. Trop. Dis. 2010, 4, e791. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, C.R.; Handasyde, K.A.; Hibble, J.; Lavender, C.J.; Legione, A.R.; McCowan, C.; Globan, M.; Mitchell, A.T.; McCracken, H.E.; Johnson, P.D.; et al. Clinical, microbiological and pathological findings of Mycobacterium ulcerans infection in three Australian possum species. PLoS Negl. Trop. Dis. 2014, 8, e2666. [Google Scholar] [CrossRef] [PubMed]
- van Zyl, A.; Daniel, J.; Wayne, J.; McCowan, C.; Malik, R.; Jelfs, P.; Lavender, C.J.; Fyfe, J.A. Mycobacterium ulcerans infections in two horses in south-eastern Australia. Aust. Vet. J. 2010, 88, 101–106. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, C.; Kuseff, G.; McMillan, E.; McCowan, C.; Lavender, C.; Globan, M.; Jerrett, I.; Oppedisano, F.; Johnson, P.; Fyfe, J. Mycobacterium ulcerans infection in two alpacas. Aust. Vet. J. 2013, 91, 296–300. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, C.R.; McMillan, E.; Harris, O.; O’Brien, D.P.; Lavender, C.J.; Globan, M.; Legione, A.R.; Fyfe, J.A. Localised Mycobacterium ulcerans infection in four dogs. Aust. Vet. J. 2011, 89, 506–510. [Google Scholar] [CrossRef] [PubMed]
- Elsner, L.; Wayne, J.; O’Brien, C.R.; McCowan, C.; Malik, R.; Hayman, J.A.; Globan, M.; Lavender, C.J.; Fyfe, J.A. Localised Mycobacterium ulcerans infection in a cat in Australia. J. Feline Med. Surg. 2008, 10, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; Prisma Group. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. Int. J. Surg. 2010, 8, 336–341. [Google Scholar]
- PROSPERO Registered Study Protocol. Available online: https://www.crd.york.ac.uk/prospero/display_record.php?RecordID=85484 (accessed on 12 January 2018).
- Fyfe, J.A.; Lavender, C.J.; Johnson, P.D.; Globan, M.; Sievers, A.; Azuolas, J.; Stinear, T.P. Development and application of two multiplex real-time PCR assays for the detection of Mycobacterium ulcerans in clinical and environmental samples. Appl. Environ. Microbiol. 2007, 73, 4733–4740. [Google Scholar] [CrossRef] [PubMed]
- Tobias, N.J.; Ammisah, N.A.; Ahortor, E.K.; Wallace, J.R.; Ablordey, A.; Stinear, T.P. Snapshot faecal survey of domestic animals in rural Ghana for Mycobacterium ulcerans. PeerJ 2016, 4, e2065. [Google Scholar] [CrossRef] [PubMed]
- Tian, R.B.; Niamké, S.; Tissot-Dupont, H.; Drancourt, M. Detection of Mycobacterium ulcerans DNA in the environment, Ivory Coast. PLoS ONE 2016, 11, e0151567. [Google Scholar] [CrossRef] [PubMed]
- Willson, S.J.; Kaufman, M.G.; Merritt, R.W.; Williamson, H.R.; Malakauskas, D.M.; Benbow, M.E. Fish and amphibians as potential reservoirs of Mycobacterium ulcerans, the causative agent of Buruli ulcer disease. Infect. Ecol. Epidemiol. 2013, 3, 19946. [Google Scholar] [CrossRef] [PubMed]
- Sakaguchi, K.; Iima, H.; Hirayama, K.; Okamoto, M.; Matsuda, K.; Miyasho, T.; Kasamatsu, M.; Hasegawa, K.; Taniyama, H. Mycobacterium ulcerans infection in an Indian flap-shelled turtle (Lissemys punctata punctata). J. Vet. Med. Sci. 2011, 73, 1217–1220. [Google Scholar] [CrossRef] [PubMed]
- Durnez, L.; Suykerbuyk, P.; Nicolas, V.; Barriere, P.; Verheyen, E.; Johnson, C.R.; Leirs, H.; Portaels, F. Terrestrial small mammals as reservoirs of Mycobacterium ulcerans in Benin. Appl. Environ. Microbiol. 2010, 76, 4574–4577. [Google Scholar] [CrossRef] [PubMed]
- Appleyard, G.D.; Clark, E.G. Histologic and genotypic characterization of a novel Mycobacterium species found in three cats. J. Clin. Microbiol. 2002, 40, 2425–2430. [Google Scholar] [CrossRef] [PubMed]
- Heckert, R.A.; Elankumaran, S.; Milani, A.; Baya, A. Detection of a new Mycobacterium species in wild striped bass in the Chesapeake Bay. J. Clin. Microbiol. 2001, 39, 710–715. [Google Scholar] [CrossRef] [PubMed]
- Ross, B.C.; Marino, L.; Oppedisano, F.; Edwards, R.; Robins-Browne, R.M.; Johnson, P.D. Development of a PCR assay for rapid diagnosis of Mycobacterium ulcerans infection. J. Clin. Microbiol. 1997, 35, 1696–1700. [Google Scholar] [PubMed]
- Portaels, F.; Hibble, J. Mycobacterium ulcerans in wild animals. Rev. Sci. Technol. 2001, 20, 252–264. [Google Scholar] [CrossRef]
| Author and Year | Sample and Sample Size | Collection Year, Location and Setting | Detection Method, Result or M. ulcerans Positive Signal |
|---|---|---|---|
| Roltgen, Pluschke, Johnson, & Fyfe, 2017 [9] | 102 environmental samples: 55 from soil/vegetation; 35 from insects or small insects pool and 12 from animal excreta | September 2013 Northern Queensland, Australia | RT-PCR IS 2404 positive: 1 soil specimen: 2 bandicoot faeces, one individual mosquito and 1 pool of 2 mosquitoes IS 2606 and KR (ketoreductase) positive: 2 bandicoot faeces and pool of two mosquitoes |
| Tobias et al., 2016 [23] | 180 faecal specimens from dominant domestic animals (ovine, porcine, avian, reptiles, canine) | September 2013 4 BU-endemic and one non-endemic villages of Ghana, West Africa | RT-PCR IS 2404 positive: 2/86 ovine; 1/69 avian: 1/16 reptiles IS 2606 and KR: all negative |
| Tian, Niamke, Tissot-Dupont, &Drancourt, 2016 [24] | 496 environmental samples: 100 from soil (endemic n = 50 and non-endemic n = 50); 200 from stagnant water (endemic n = 100 and non-endemic n = 100); 100 from plants (endemic n = 50 and non-endemic n = 50) and 96 animal faeces (Thryonomys swinderianus (agouti) stools) (endemic n = 48 and non-endemic n = 48) | June–October 2014 Ivory Coast, West Africa | RT-PCR 43 samples with at least one positive IS 2404 and KR Out of 43, only 10 positive for both IS2404 and KR, IS 2606 not performed: 7 water specimen; 2 T. swinderianus (agouti) faeces and one soil specimen |
| Carson et al., 2014 [5] | Fecal sample: 216 common ringtail possums and 6 common brushtail possums | Southeast Australia, State Victoria | RT-PCR targeting IS 2404, IS 2606 and KR 20 common ringtail possums and 4 common brushtail possums |
| O’Brien et al., 2014 [15] | 69 possums (ringtail and brushtail) trapped at Point Lonsdale: Faecal samples: 57; blood samples: 63; buccal swab: 67; urine sample: 16; pouch swab: 15; cloacal swab: 20 69 fecal samples from 15 mountain brushtail possums | 1998–2011 Victoria, Australia | RT-PCR targeting IS 2404, IS 2606 and KR Point Lonsdale: Positive: faecal sample: 12 (25%); blood sample: 0; buccal swab: 7 (16%); urine sample: 0; pouch swab: 3 (20%) Bellbird Creek: Positive: 4 mountain brushtail possums (27%) |
| C. O’Brien et al., 2013 [17] | Case report: two alpacas (Vicugna pacos) ulcerated tissue | Case 1: September 1997 Case 2: May 2011 Victoria, Australia | RT-PCR targeting IS 2404, IS 2606 and KR positive |
| Willson et al., 2013 [25] | 587 fish representing 13 genera and 17 species and 351 amphibians representing 10 genera: external swab | 2008–2009 Ghana, West Africa | RT-PCR targeting IS 2606 and KR not performed. Not confirmed |
| C. R. O’Brien et al., 2011 [18] | Case report: Case 1: 14 months old female kelpie Case 2: 3 years old female kelpie Case 3: 6 years old male whippet Case 4: 3 years old male koolie | 2011 Victoria, Australia | RT-PCR targeting IS 2404, IS 2606 and KR All 4 dogs positive for M. ulcerans |
| Sakaguchi et al., 2011 [26] | Case report; Indian flap-shelled turtle, Lissemys punctata punctata | Imported from India to aquarium in Japan | PCR assays targeting the rpoβ gene: unable to differentiate M. ulcerans from mycolactone-producing M. marinum (MPMM) |
| Fyfe et al., 2010 [14] | 589 fecal samples from ringtail possums and 250 samples from brushtail possums. Live trapping: 42 ringtail possums and 21 brushtail possums | 2007–2009 Victoria, Australia | RT-PCR targeting IS 2404, IS 2606 and KR M. ulcerans DNA detected in 43% of ringtail possum and 29% of brushtail possum faecal samples. 38% ringtail possum have M. ulcerans lesion and/or positive faeces Lower in brushtail possums: 1 with M. ulcerans lesion and/or positive faeces and 4 with no lesions and low M. ulcerans DNA in faeces. |
| Durnez et al., 2010 [27] | 565 small mammals: 326 rodents and 222 shrews | 2006 Benin, West Africa | RT-PCR: No M. ulcerans specific DNA detected |
| Van Zyl et al., 2010 [16] | 2 horses: Case report Case 1: 21-year-old quarterhorse-cross Case 2: 32-year-old standard bredgelding | Case 1: May 2006 Case 2: October 2006 Southeastern Australia | RT-PCR M. ulcerans specific DNA detected from both horses |
| Elsner et al., 2008 [19] | Cat: Case report 10-year-old castrated male domestic cat | 2006 Victoria, Australia | RT-PCR M. ulcerans specific DNA detected |
| Appleyard & Clark, 2002 [28] | Case report: three cats Case 1: An 8-year-old spayed female shorthair Case 2: 6-year-old spayed female shorthair Case 3: 11-year-old domestic longhair cat | 2002 North America | PCR Could not differentiate M. ulcerans from other Mycobacterium spp. (a new Mycobacterial spp. namely ‘Mycobacterium visibilis’ suggested) |
| Heckert, Elankumaran, Milani, &Baya, 2001 [29] | 60 wild striped bass: Swab from external ulcerative dermatitis and granulomatous-like lesions in the internal organs | 1997 Chesapeake Bay, USA | PCR No M. ulcerans specific DNA detected (a new mycobacterial spp. suggested) |
| Mitchell, McOrist, &Bilney, 1987 [13] | 36 male and 51 female adult koalas captured | 1980–1985 Raymond Island, southeastern Australia | Pathological and bacteriological examination 18 out of 87 captured koalas had skin wound 11 koalas were found positive for M. ulcerans |
| McOrist, Jerrett, Anderson, & Hayman, 1985 [12] | Case study: 2 koalas: one male and one female Ulcerated tissue | 1982 Raymond Island, southeastern Australia | Pathological and bacteriological examination Both koalas suggested positive for M. ulcerans |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Singh, A.; McBride, W.J.H.; Govan, B.; Pearson, M. Potential Animal Reservoir of Mycobacterium ulcerans: A Systematic Review. Trop. Med. Infect. Dis. 2018, 3, 56. https://doi.org/10.3390/tropicalmed3020056
Singh A, McBride WJH, Govan B, Pearson M. Potential Animal Reservoir of Mycobacterium ulcerans: A Systematic Review. Tropical Medicine and Infectious Disease. 2018; 3(2):56. https://doi.org/10.3390/tropicalmed3020056
Chicago/Turabian StyleSingh, Avishek, William John Hannan McBride, Brenda Govan, and Mark Pearson. 2018. "Potential Animal Reservoir of Mycobacterium ulcerans: A Systematic Review" Tropical Medicine and Infectious Disease 3, no. 2: 56. https://doi.org/10.3390/tropicalmed3020056
APA StyleSingh, A., McBride, W. J. H., Govan, B., & Pearson, M. (2018). Potential Animal Reservoir of Mycobacterium ulcerans: A Systematic Review. Tropical Medicine and Infectious Disease, 3(2), 56. https://doi.org/10.3390/tropicalmed3020056






