Identification of Statewide Hotspots for Respiratory Disease Targets Using Wastewater Monitoring Data
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Location Inclusion Criteria and Sample Collection
2.2. Sample Concentration and Quantification of SARS-CoV-2, InfA, and InfB
2.3. Supplemental Data Collection and Preparation
- (i)
- Wastewater and Clinical Data
- (ii)
- Demographic and Infrastructure Data
2.4. Data Analyses
- (i)
- SARS-CoV-2 and Influenza Sewershed Ranks and Comparisons
- (ii)
- Hotspot Cluster and Outlier Analyses
- (iii)
- Pairwise Spearman Correlations
- (iv)
- Cross-Correlation and Forward Stepwise Multiple Regression
3. Results
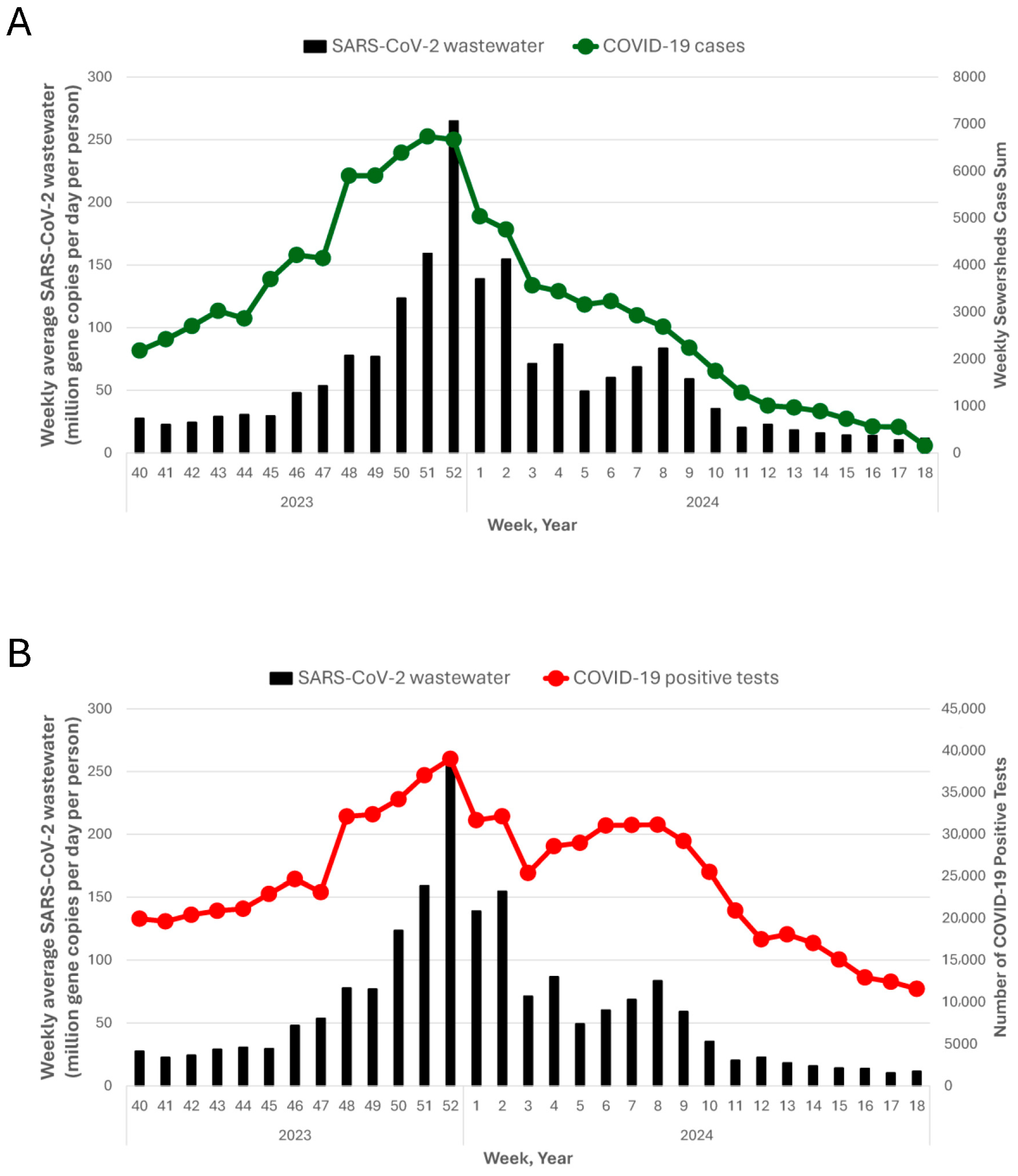
3.1. Comparison of Statewide Wastewater Concentrations to Clinical Indicators
3.2. Locations That Were First to Detect InfA and InfB in Wastewater
3.3. Locations Reaching High Prevalence of SARS-CoV-2, InfA, and InfB Earlier Using 80th Percentile Concentration Threshold
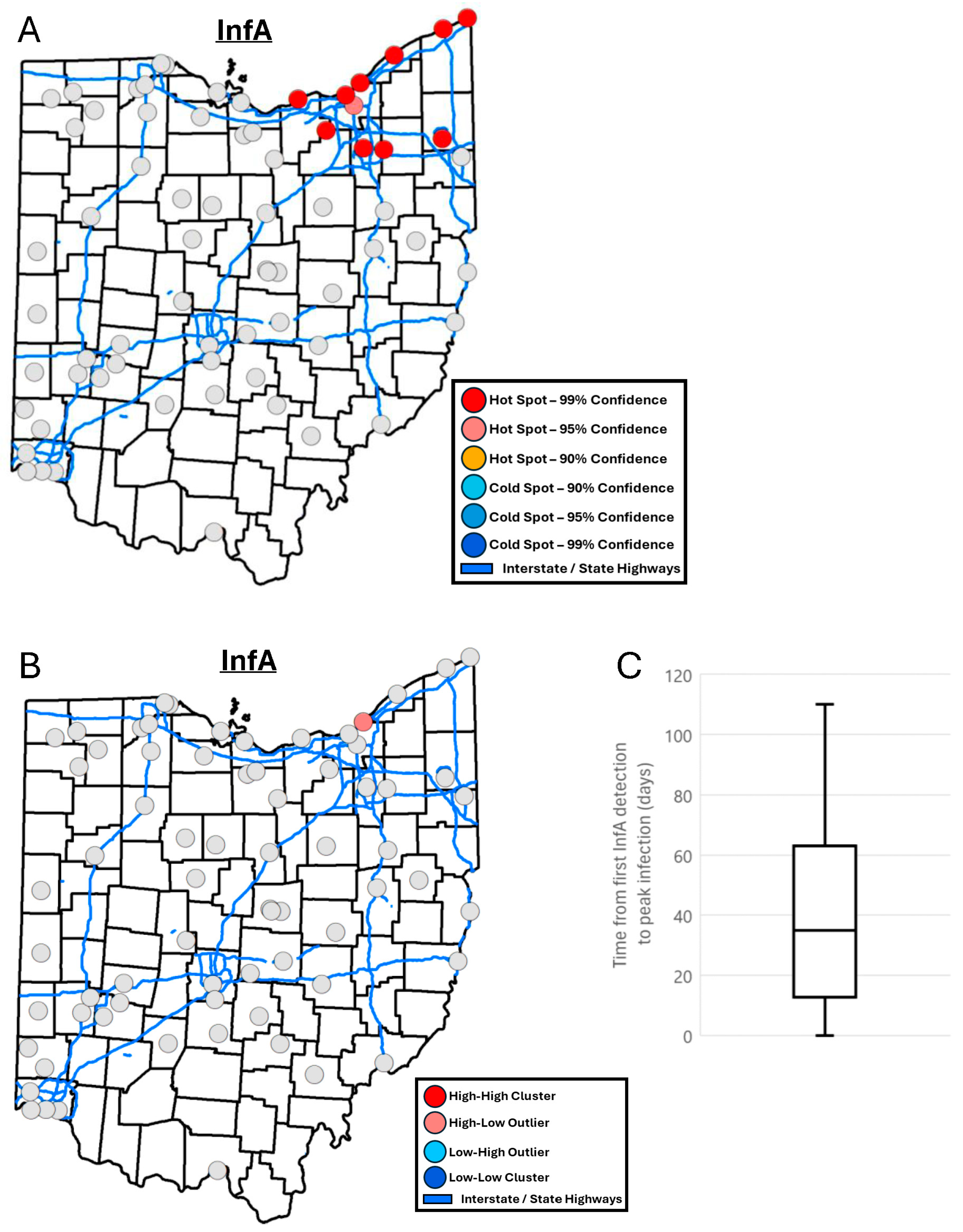
3.4. Locations Identified as Seasonal Hotspots and with Earliest Concentration Peaks of SARS-CoV-2, InfA, and InfB
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| InfA | Influenza A |
| InfB | Influenza B |
| SVI | Social Vulnerability Index |
| WWTP | Wastewater treatment plant |
| OWMN | Ohio Wastewater Monitoring Network |
| LOQ | Limit of quantification |
| ODHL | Ohio Department of Health Public Health Laboratory |
| BCoV | Bovine coronavirus |
| MGCPD | Million gene copies per person per day |
| IOP | Innovate Ohio Platform |
| ODRS | Ohio Disease Reporting System |
| NREVSS | National Respiratory and Enteric Virus Surveillance System |
| DHS | Department of Homeland Security |
| HIFLD | Homeland Infrastructure Foundation-Level Data |
| ATSDR | Agency for Toxic Substances and Disease Registry |
| CDC | Centers for Disease Control and Prevention |
| ODOT | Ohio Department of Transportation |
| SAS | Statistical Analysis Software |
References
- Hart, O.E.; Halden, R.U. Computational analysis of SARS-CoV-2/COVID-19 surveillance by wastewater-based epidemiology locally and globally: Feasibility, economy, opportunities and challenges. Sci. Total Environ. 2020, 730, 138875. [Google Scholar] [CrossRef] [PubMed]
- Adams, C.; Bias, M.; Welsh, R.M.; Webb, J.; Reese, H.; Delgado, S.; Person, J.; West, R.; Shin, S.; Kirby, A. The national wastewater surveillance system (NWSS): From inception to widespread coverage, 2020–2022, United States. Sci. Total Environ. 2024, 924, 171566. [Google Scholar] [CrossRef] [PubMed]
- D’SOuza, N.; Porter, A.M.; Rose, J.B.; Dreelin, E.; Peters, S.E.; Nowlin, P.J.; Carbonell, S.; Cissell, K.; Wang, Y.; Flood, M.T.; et al. Public health use and lessons learned from a statewide SARS-CoV-2 wastewater monitoring program (MiNET). Heliyon 2024, 10, e35790. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Neyra, M.; Hill, D.T.; Bennett, L.J.; Dunham, C.N.; Larsen, D.A. Establishing a statewide wastewater surveillance system in response to the covid-19 pandemic: A reliable model for continuous and emerging public health threats. J. Public Health Manag. Pract. 2023, 29, 854–862. [Google Scholar] [CrossRef]
- Bohrerova, Z.; Brinkman, N.E.; Chakravarti, R.; Chattopadhyay, S.; Faith, S.A.; Garland, J.; Herrin, J.M.; Hull, N.; Jahne, M.; Kang, D.-W.; et al. Ohio Coronavirus Wastewater Monitoring Network: Implementation of statewide monitoring for protecting public health. J. Public Health Manag. Pract. 2023, 29, 845–853. [Google Scholar] [CrossRef]
- Hrudey, S.E.; Bischel, H.N.; Charrois, J.; Chik, A.H.S.; Conant, B.; Delatolla, R.; Dorner, S.; Graber, T.E.; Hubert, C.; Isaac-Renton, J.; et al. Wastewater Surveillance for SARS-CoV-2 RNA in Canada. Facets 2022, 7, 1493–1597. [Google Scholar] [CrossRef]
- U.S. EPA (U.S. Environmental Protection Agency). A Compendium of U.S. Wastewater Surveillance to Support COVID-19 Public Health Response 2021. Available online: https://www.epa.gov/sustainable-water-infrastructure/compendium-us-wastewater-surveillance-support-covid-19-public (accessed on 16 September 2024).
- Parkins, M.D.; Lee, B.E.; Acosta, N.; Bautista, M.; Hubert, C.R.; Hrudey, S.E.; Frankowski, K.; Pang, X.L. Wastewater-based surveillance as a tool for public health action: SARS-CoV-2 and beyond. Clin. Microbiol. Rev. 2024, 37, e00103-22. [Google Scholar] [CrossRef]
- Dos Santos, M.C.; Silva, A.C.C.; dos Reis Teixeira, C.; Prazeres, F.P.M.; Dos Santos, R.F.; de Araújo Rolo, C.; de Souza Santos, E.; da Fonseca, M.S.; Valente, C.O.; Hodel, K.V.S.; et al. Wastewater surveillance for viral pathogens: A tool for public health. Heliyon 2024, 10, e33873. [Google Scholar] [CrossRef]
- Levy, J.I.; Andersen, K.G.; Knight, R.; Karthikeyan, S. Wastewater surveillance for public health. Science 2023, 379, 26–27. [Google Scholar] [CrossRef]
- Charu, V.; Zeger, S.; Gog, J.; Bjørnstad, O.N.; Kissler, S.; Simonsen, L.; Grenfell, B.T.; Viboud, C. Human mobility and the spatial transmission of influenza in the United States. PLoS Comput. Biol. 2017, 13, e1005382. [Google Scholar] [CrossRef]
- de Souza, C.D.F.; Machado, M.F.; da Silva Junior, A.G.; Nunes, B.E.B.R.; do Carmo, R.F. Airports, highways and COVID-19: An analysis of spatial dynamics in Brazil. J. Transp. Health 2021, 21, 101067. [Google Scholar] [CrossRef]
- Urban, R.C.; Nakada, L.Y.K. GIS-based spatial modelling of COVID-19 death incidence in São Paulo, Brazil. Environ. Urban. 2021, 33, 229–238. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Jiang, B.; Yang, Y.; Chen, L.; Liu, X.; Wu, X.; Chen, B.; Webster, C.; Sullivan, W.C.; Larsen, L.; Wang, J.; et al. Green spaces, especially nearby forest, may reduce the SARS-CoV-2 infection rate: A nationwide study in the United States. Landsc. Urban. Plan. 2022, 228, 104583. [Google Scholar] [CrossRef] [PubMed]
- Boterman, W. Population density and SARS-CoV-2 pandemic: Comparing the geography of different waves in the Netherlands. Urban Stud. 2023, 60, 1377–1402. [Google Scholar] [CrossRef] [PubMed]
- da Costa, A.C.C.; Codeço, C.T.; Krainski, E.T.; Gomes, M.F.d.C.; Nobre, A.A. Spatiotemporal diffusion of influenza A (H1N1): Starting point and risk factors. PLoS ONE. 2018, 13, e0202832. [Google Scholar] [CrossRef]
- Holden, T.M.; Richardson, R.A.K.; Arevalo, P.; Duffus, W.A.; Runge, M.; Whitney, E.; Wise, L.; Ezike, N.O.; Patrick, S.; Cobey, S.; et al. Geographic and demographic heterogeneity of SARS-CoV-2 diagnostic testing in Illinois, USA, March to December 2020. BMC Public Health 2021, 21, 1105. [Google Scholar] [CrossRef]
- Baldwin, W.M.; Dayton, R.D.; Bivins, A.W.; Scott, R.S.; Yurochko, A.D.; Vanchiere, J.A.; Davis, T.; Arnold, C.L.; Asuncion, J.E.; Bhuiyan, M.A.; et al. Highly socially vulnerable communities exhibit disproportionately increased viral loads as measured in community wastewater. Environ. Res. 2023, 222, 115351. [Google Scholar] [CrossRef]
- Mota, C.R.; Bressani-Ribeiro, T.; Araújo, J.C.; Leal, C.D.; Leroy-Freitas, D.; Machado, E.C.; Espinosa, M.F.; Fernandes, L.; Leão, T.L.; Chamhum-Silva, L.; et al. Assessing spatial distribution of COVID-19 prevalence in Brazil using decentralised sewage monitoring. Water Res. 2021, 202, 117388. [Google Scholar] [CrossRef]
- Nelson, B. What poo tells us: Wastewater surveillance comes of age amid covid, monkeypox, and polio. BMJ 2022, 378, o1869. [Google Scholar] [CrossRef] [PubMed]
- Haak, L.; Delic, B.; Li, L.; Guarin, T.; Mazurowski, L.; Dastjerdi, N.G.; Dewan, A.; Pagilla, K. Spatial and temporal variability and data bias in wastewater surveillance of SARS-CoV-2 in a sewer system. Sci. Total Environ. 2022, 805, 150390. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Lancaster, E.; Byrd, K.; Ai, Y.; Lee, J. Socioeconomic status correlations with confirmed COVID-19 cases and SARS-CoV-2 wastewater concentrations in small-medium sized communities. Environ. Res. 2022, 215, 114290. [Google Scholar] [CrossRef]
- Liu, J.; Suzuki, S. Real-Time Detection of Flu Season Onset: A Novel Approach to Flu Surveillance. Int. J. Environ. Res. Public Health 2022, 19, 3681. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Bloom-Feshbach, K.; Alonso, W.J.; Charu, V.; Tamerius, J.; Simonsen, L.; Miller, M.A.; Viboud, C. Latitudinal variations in seasonal activity of influenza and respiratory syncytial virus (RSV): A global comparative review. PLoS ONE 2013, 8, e54445. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Hoogeveen, M.J.; Hoogeveen, E.K. Comparable seasonal pattern for COVID-19 and flu-like illnesses. One Health 2021, 13, 100277. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ayala, A.; Villalobos Dintrans, P.; Elorrieta, F.; Castillo, C.; Vargas, C.; Maddaleno, M. Identification of COVID-19 waves: Considerations for research and policy. Int. J. Environ. Res. Public Health 2021, 18, 11058. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Gog, J.R.; Ballesteros, S.; Viboud, C.; Simonsen, L.; Bjornstad, O.N.; Shaman, J.; Chao, D.L.; Khan, F.; Grenfell, B.T.; Ferguson, N.M. Spatial transmission of 2009 pandemic influenza in the US. PLoS Comput. Biol. 2014, 10, e1003635. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Gong, S.; Huang, S.; Huo, X.; Wang, W. Geographical characteristics and influencing factors of the influenza epidemic in Hubei, China, from 2009 to 2019. PLoS ONE 2023, 18, e0280617. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ito, G.; Takazono, T.; Hosogaya, N.; Iwanaga, N.; Miyazawa, S.; Fujita, S.; Watanabe, H.; Mukae, H. Impact of meteorological and demographic factors on the influenza epidemic in Japan: A large observational database study. Sci. Rep. 2023, 13, 13000. [Google Scholar] [CrossRef]
- Kar, A.; Huyen, T.K.L.; Miller, H.J. What Is Essential Travel? Socioeconomic Differences in Travel Demand in Columbus, Ohio, during the COVID-19 Lockdown. Ann. Am. Assoc. Geogr. 2022, 112, 1023–1046. [Google Scholar] [CrossRef]
- Czaja, C.A.; Cockburn, M.G.; Colborn, K.; Miller, L.; Thomas, D.S.; Herlihy, R.K.; Alden, N.; Simões, E.A. Evaluation of rates of laboratory-confirmed influenza hospitalization in rural and urban census tracts over eight influenza seasons. Prev. Med. 2020, 139, 106184. [Google Scholar] [CrossRef]
- Jung, S.W.; Kim, Y.J.; Han, S.B.; Lee, K.-Y.; Kang, J.H. Differences in the age distribution of influenza B virus infection according to influenza B virus lineages in the Korean population. Postgrad. Med. 2021, 133, 82–88. [Google Scholar] [CrossRef] [PubMed]
- O’Halloran, A.C.; Holstein, R.; Cummings, C.; Kirley, P.D.; Alden, N.B.; Yousey-Hindes, K.; Anderson, E.J.; Ryan, P.; Kim, S.; Lynfield, R.; et al. Rates of Influenza-Associated Hospitalization, Intensive Care Unit Admission, and In-Hospital Death by Race and Ethnicity in the United States From 2009 to 2019. JAMA Netw. Open. 2021, 4, e2121880. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ma, P.; Tang, X.; Zhang, L.; Wang, X.; Wang, W.; Zhang, X.; Wang, S.; Zhou, N. Influenza A and B outbreaks differed in their associations with climate conditions in Shenzhen, China. Int. J. Biometeorol. 2022, 66, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Zhu, A.; Li, Z.; Zhang, H. Spatial timing of circulating seasonal influenza A and B viruses in China from 2014 to 2018. Sci. Rep. 2023, 13, 7149. [Google Scholar] [CrossRef] [PubMed]
- Rosca, E.C.; Heneghan, C.; Spencer, E.A.; Brassey, J.; Plüddemann, A.; Onakpoya, I.J.; Evans, D.H.; Conly, J.M.; Jefferson, T. Transmission of SARS-CoV-2 associated with aircraft travel: A systematic review. J. Travel. Med. 2021, 28, taab133. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Khanh, N.C.; Thai, P.Q.; Quach, H.L.; Thi, N.H.; Dinh, P.C.; Duong, T.N.; Mai, L.T.Q.; Nghia, N.D.; Tu, T.A.; Quang, N.; et al. Transmission of SARS-CoV 2 During Long-Haul Flight. Emerg. Infect. Dis. 2020, 26, 2617–2624. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ngeh, S.; Vogt, F.; Sikazwe, C.T.; Levy, A.; Pingault, N.M.; Smith, D.W.; Effler, P.V. Travel-associated SARS-CoV-2 transmission documented with whole genome sequencing following a long-haul international flight. J. Travel. Med. 2022, 29, taac057. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Gartland, N.; Fishwick, D.; Coleman, A.; Davies, K.; Hartwig, A.; Johnson, S.; van Tongeren, M. Transmission and control of SARS-CoV-2 on ground public transport: A rapid review of the literature up to May 2021. J. Transp. Health 2022, 26, 101356. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Pilipenco, A.; Forinová, M.; Mašková, H.; Hönig, V.; Palus, M.; Lynn, N.S., Jr.; Víšová, I.; Vrabcová, M.; Houska, M.; Anthi, J.; et al. Negligible risk of surface transmission of SARS-CoV-2 in public transportation. J. Travel Med. 2023, 30, taad065. [Google Scholar] [CrossRef]
- Ellingjord-Dale, M.; Kalleberg, K.T.; Istre, M.S.; Nygaard, A.B.; Brunvoll, S.H.; Eggesbø, L.M.; Dahl, J.A.; Kjetland, E.F.; Ursin, G.; Søraas, A. The use of public transport and contraction of SARS-CoV-2 in a large prospective cohort in Norway. BMC Infect. Dis. 2022, 22, 252. [Google Scholar] [CrossRef]
- Bontempi, E.; Coccia, M.; Vergalli, S.; Zanoletti, A. Can commercial trade represent the main indicator of the COVID-19 diffusion due to human-to-human interactions? A comparative analysis between Italy, France, and Spain. Environ. Res. 2021, 201, 111529. [Google Scholar] [CrossRef]
- Grubesic, T.H.; Nelson, J.R.; Wallace, D.; Eason, J.; Towers, S.; Walker, J. Geodemographic insights on the COVID-19 pandemic in the State of Wisconsin and the role of risky facilities. GeoJournal 2022, 87, 4311–4333. [Google Scholar] [CrossRef]
- Blumenberg, E.; Siddiq, F. Commute distance and jobs-housing fit. Transportation 2023, 50, 869–891. [Google Scholar] [CrossRef]




| Population Served | Population Density | County Traffic Counts | Number of Hospitals | Number of Hospital Beds | Number of Urgent Care Centers | Number of Nursing Homes/Assisted Living Facilities | Nocioeconomic Status SVI | Household Characteristics SVI | Minority and Ethnicity Status SVI | Housing Type and Transportation SVI | Overall SVI | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Population served | 0.34 * | 0.74 * | 0.92 * | 0.84 * | 0.87 * | 0.94 * | 0.20 | 0.09 | 0.6 * | −0.18 | 0.15 | |
| Population density | 0.34 * | 0.4 * | 0.36 * | 0.45 * | 0.30 | 0.18 | 0.48 * | 0.13 | 0.49 * | 0.11 | 0.4 * | |
| County traffic counts | 0.74 * | 0.4 * | 0.74 * | 0.72 * | 0.67 * | 0.69 * | 0.12 | −0.03 | 0.67 * | −0.34 * | 0.03 | |
| Number of hospitals | 0.92 * | 0.36 * | 0.74 * | 0.93 * | 0.7 * | 0.88 * | 0.23 | 0.13 | 0.67 * | −0.19 | 0.20 | |
| Number of hospital beds | 0.84 * | 0.45 * | 0.72 * | 0.93 * | 0.59 * | 0.77 * | 0.27 * | 0.13 | 0.66 * | −0.14 | 0.24 | |
| Number of urgent care centers | 0.87 * | 0.30 | 0.67 * | 0.7 * | 0.59 * | 0.82 * | 0.03 | 0.06 | 0.49 * | −0.21 | 0.02 | |
| Number of nursing homes/assisted living facilities | 0.94 * | 0.18 | 0.69 * | 0.88 * | 0.77 * | 0.82 * | 0.12 | 0.06 | 0.53 * | −0.24 | 0.07 | |
| Socioeconomic status SVI | 0.20 | 0.48 * | 0.12 | 0.23 | 0.27 | 0.03 | 0.12 | 0.52 * | 0.42 * | 0.5 * | 0.93 * | |
| Household characteristics SVI | 0.09 | 0.13 | −0.03 | 0.13 | 0.13 | 0.06 | 0.06 | 0.52 * | 0.16 | 0.35 * | 0.72 * | |
| Minority and ethnicity status SVI | 0.6 * | 0.49 * | 0.67 * | 0.67 * | 0.66 * | 0.49 * | 0.53 * | 0.42 * | 0.16 | −0.13 | 0.37 * | |
| Housing type and transportation SVI | −0.18 | 0.11 | −0.34 | −0.19 | −0.14 | −0.21 | −0.24 | 0.5 * | 0.35 * | −0.13 | 0.69 * | |
| Overall SVI | 0.15 | 0.4 * | 0.03 | 0.20 | 0.24 | 0.02 | 0.07 | 0.93 * | 0.72 * | 0.37 * | 0.69 * |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Servello, D.; Chalasani, P.; Leasure, E.; LeMaster, K.D.; Kellar, J.; Stiverson, J.; White, M.; Bohrerova, Z. Identification of Statewide Hotspots for Respiratory Disease Targets Using Wastewater Monitoring Data. Trop. Med. Infect. Dis. 2025, 10, 241. https://doi.org/10.3390/tropicalmed10090241
Servello D, Chalasani P, Leasure E, LeMaster KD, Kellar J, Stiverson J, White M, Bohrerova Z. Identification of Statewide Hotspots for Respiratory Disease Targets Using Wastewater Monitoring Data. Tropical Medicine and Infectious Disease. 2025; 10(9):241. https://doi.org/10.3390/tropicalmed10090241
Chicago/Turabian StyleServello, Dustin, Purnima Chalasani, Erica Leasure, Krysta Danielle LeMaster, Justin Kellar, Jill Stiverson, Michelle White, and Zuzana Bohrerova. 2025. "Identification of Statewide Hotspots for Respiratory Disease Targets Using Wastewater Monitoring Data" Tropical Medicine and Infectious Disease 10, no. 9: 241. https://doi.org/10.3390/tropicalmed10090241
APA StyleServello, D., Chalasani, P., Leasure, E., LeMaster, K. D., Kellar, J., Stiverson, J., White, M., & Bohrerova, Z. (2025). Identification of Statewide Hotspots for Respiratory Disease Targets Using Wastewater Monitoring Data. Tropical Medicine and Infectious Disease, 10(9), 241. https://doi.org/10.3390/tropicalmed10090241








