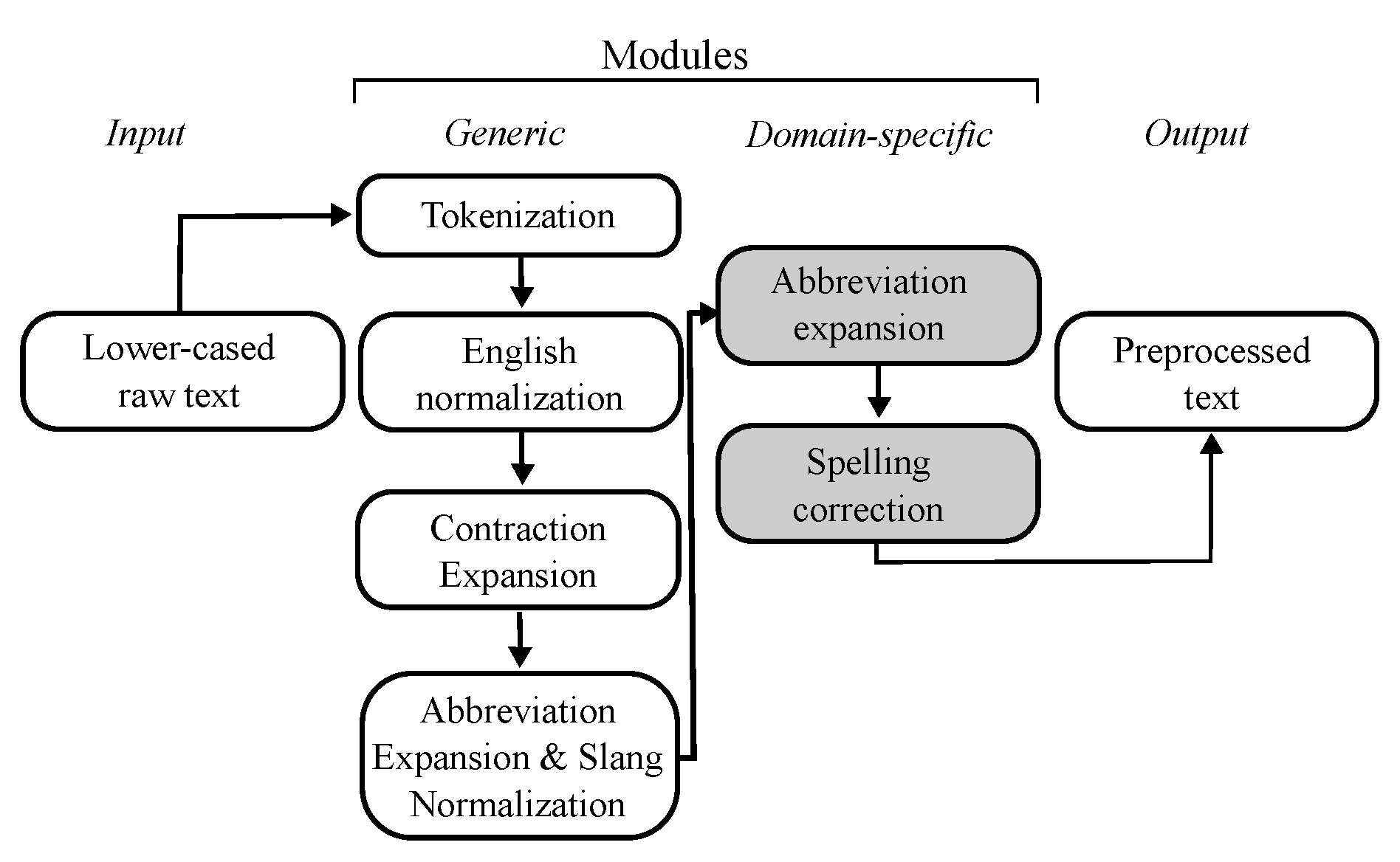
Figure 1.
Sequential processing pipeline.
Figure 1.
Sequential processing pipeline.
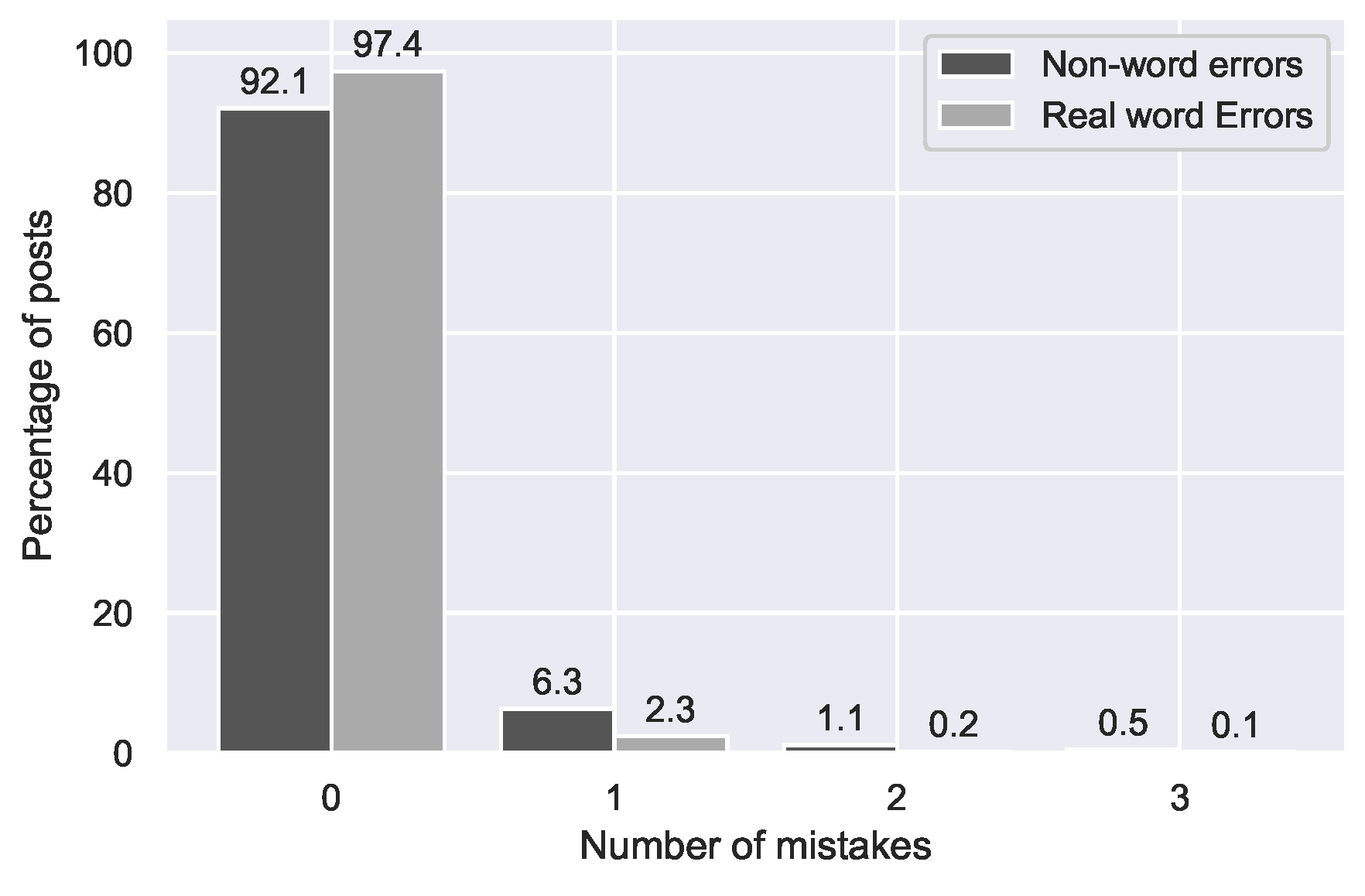
Figure 2.
Distribution of non-word and real word errors across posts in the GIST forum.
Figure 2.
Distribution of non-word and real word errors across posts in the GIST forum.
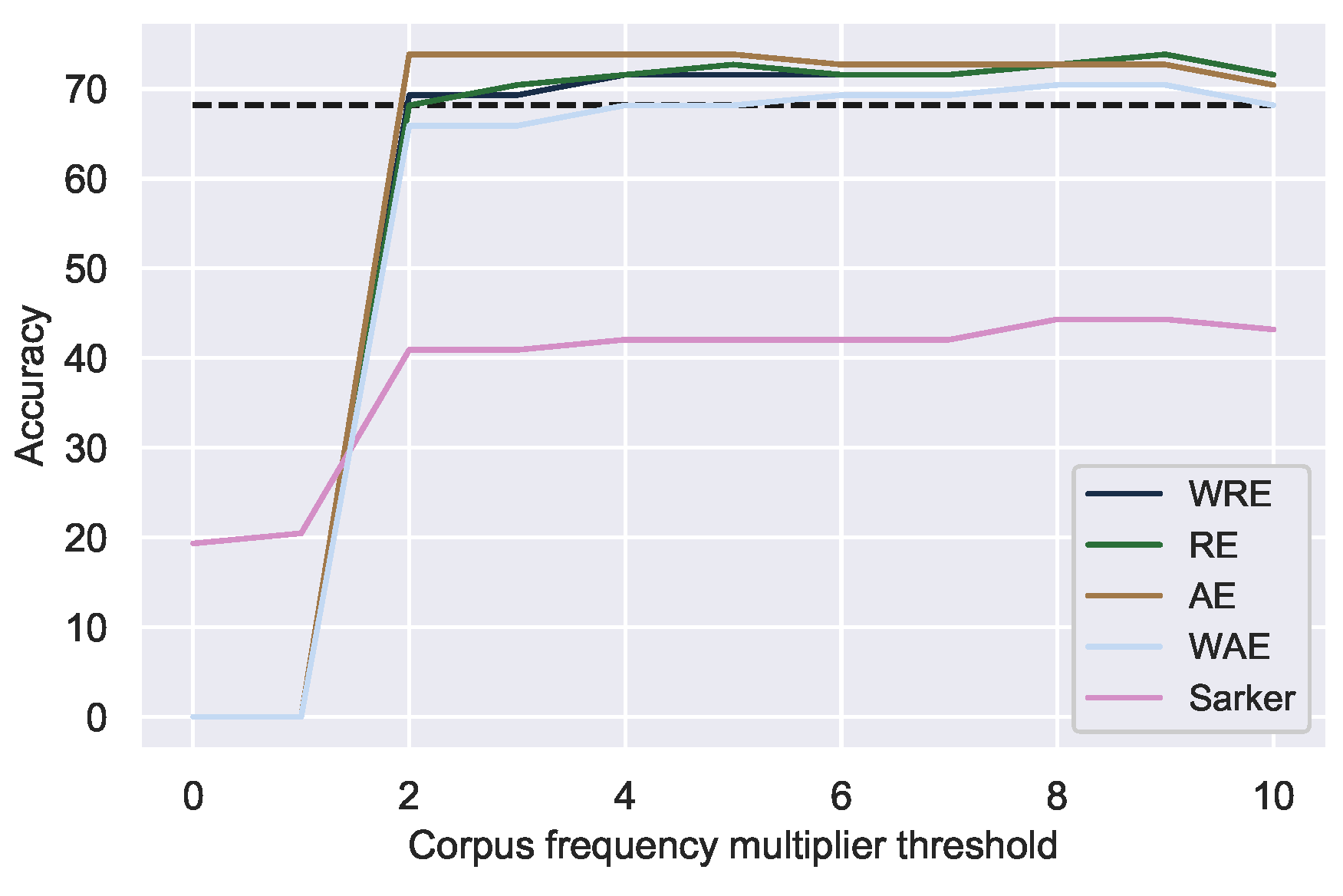
Figure 3.
Correction accuracy of unique mistakes using correction candidates from the data at various minimum relative corpus frequency thresholds. Dotted line indicates the best correction accuracy using dictionary-derived candidates.
Figure 3.
Correction accuracy of unique mistakes using correction candidates from the data at various minimum relative corpus frequency thresholds. Dotted line indicates the best correction accuracy using dictionary-derived candidates.
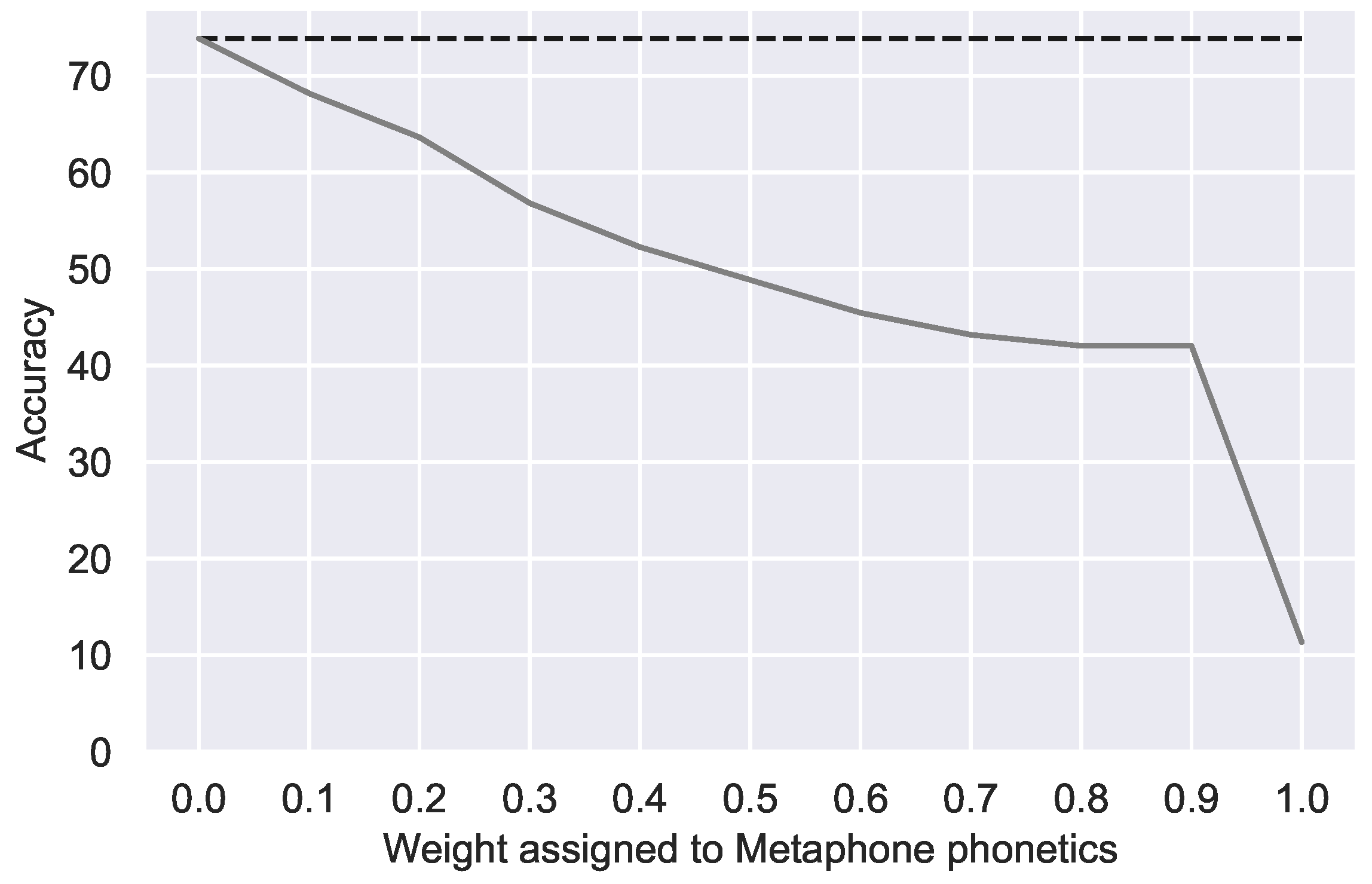
Figure 4.
Correction accuracy with additional weighted double Metaphone phonetic similarity. Dotted line indicates the best accuracy with relative edit distance alone.
Figure 4.
Correction accuracy with additional weighted double Metaphone phonetic similarity. Dotted line indicates the best accuracy with relative edit distance alone.
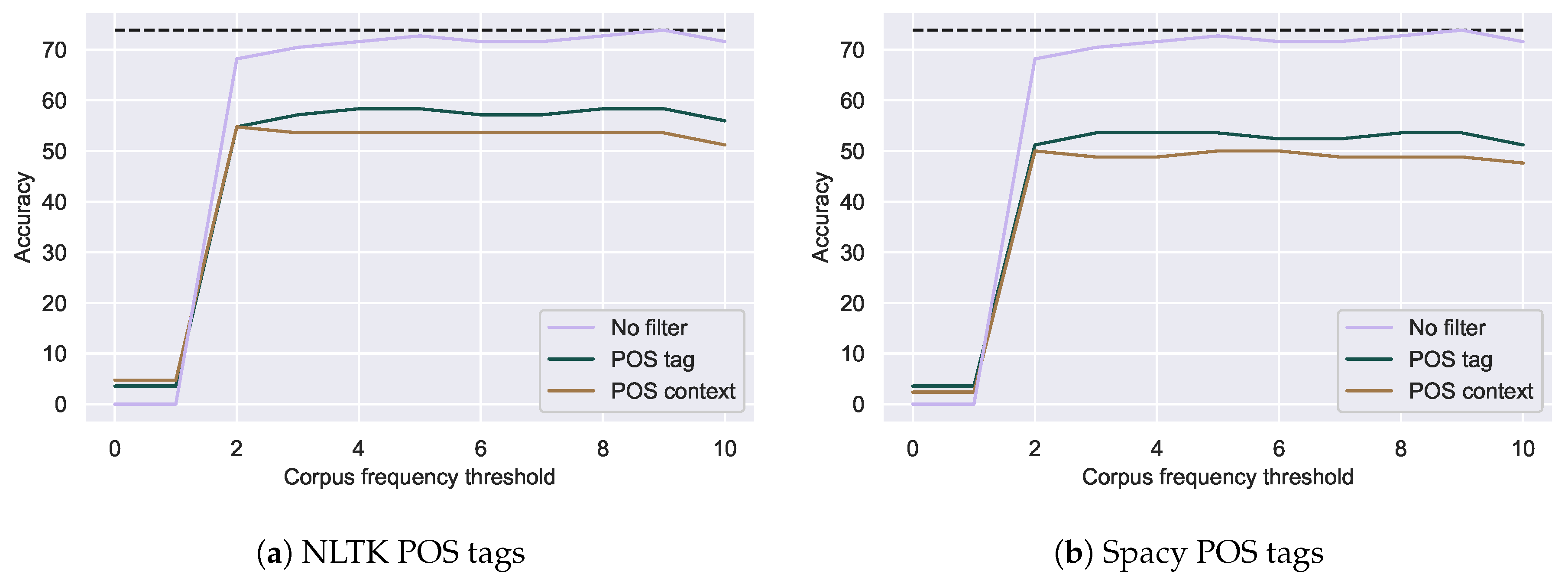
Figure 5.
Correction accuracy of spelling mistakes with additional POS tag filters using NLTK (a) or
Spacy (b). Dotted lines indicate the best accuracy with relative edit distance alone
Figure 5.
Correction accuracy of spelling mistakes with additional POS tag filters using NLTK (a) or
Spacy (b). Dotted lines indicate the best accuracy with relative edit distance alone
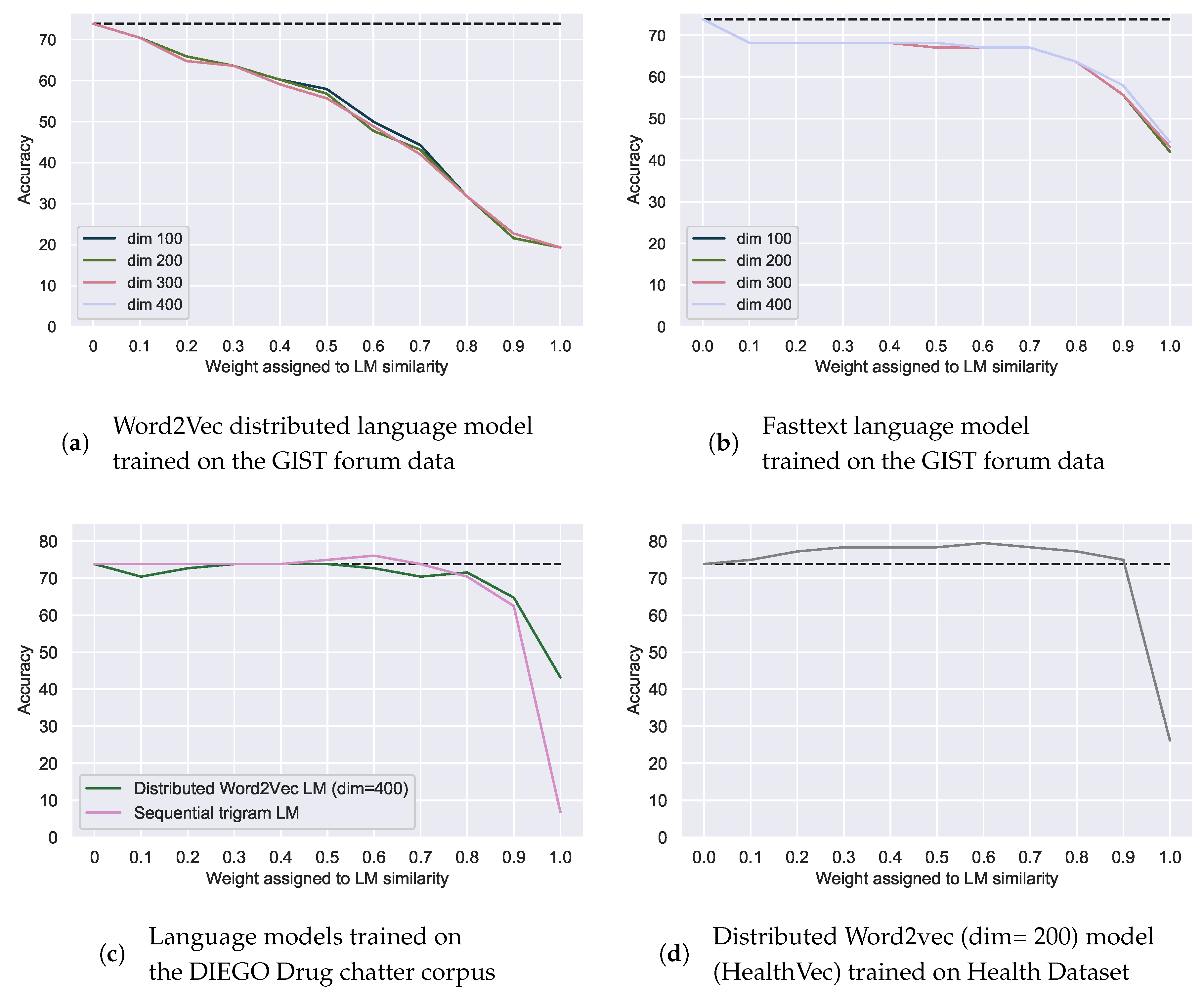
Figure 6.
Correction accuracy of spelling mistakes with additional weighted language model (LM) similarity using language models of the data itself (a,b) or trained on external datasets: the DIEGO Drug Chatter corpus (c) and the Health Dataset (d). Weight of the LM similarity is the inverse of the weight of the relative edit distance. Dotted line indicates the best accuracy with relative edit distance alone.
Figure 6.
Correction accuracy of spelling mistakes with additional weighted language model (LM) similarity using language models of the data itself (a,b) or trained on external datasets: the DIEGO Drug Chatter corpus (c) and the Health Dataset (d). Weight of the LM similarity is the inverse of the weight of the relative edit distance. Dotted line indicates the best accuracy with relative edit distance alone.
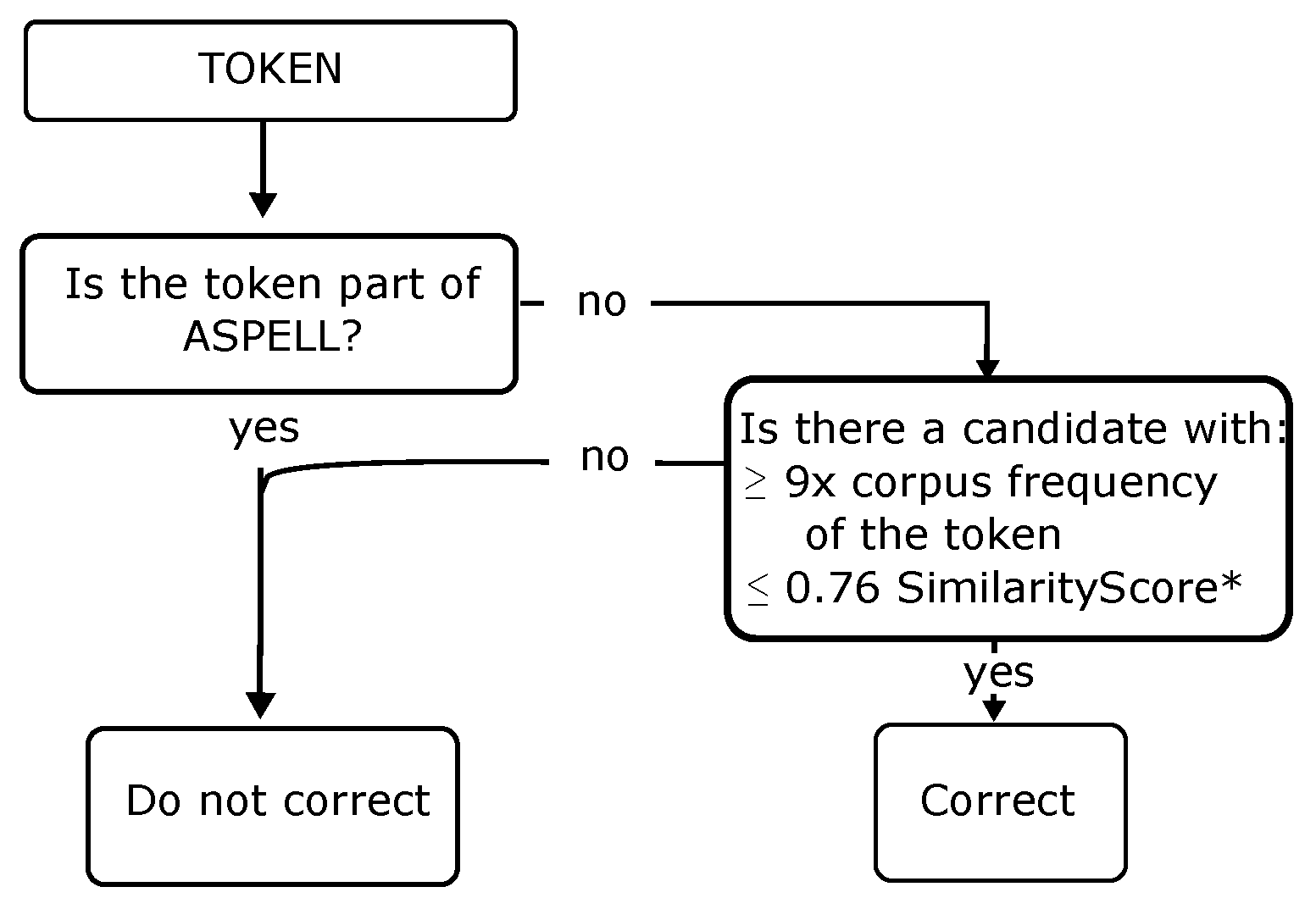
Figure 7.
Decision process. * SimilarityScore = 0.6 × model similarity + 0.4 × RE.
Figure 7.
Decision process. * SimilarityScore = 0.6 × model similarity + 0.4 × RE.
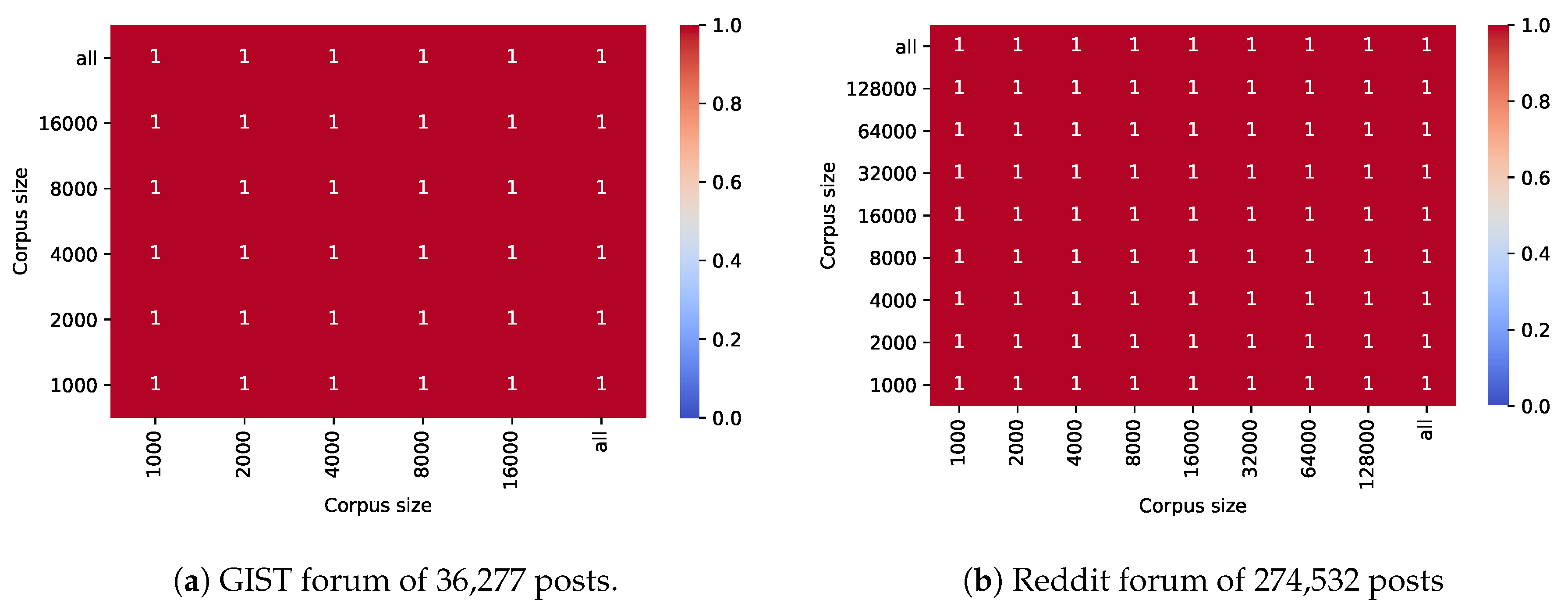
Figure 8.
Stability of error detection in 1000 posts with varying corpus size for the GIST forum (a) and the Reddit forum on cancer (b).
Figure 8.
Stability of error detection in 1000 posts with varying corpus size for the GIST forum (a) and the Reddit forum on cancer (b).
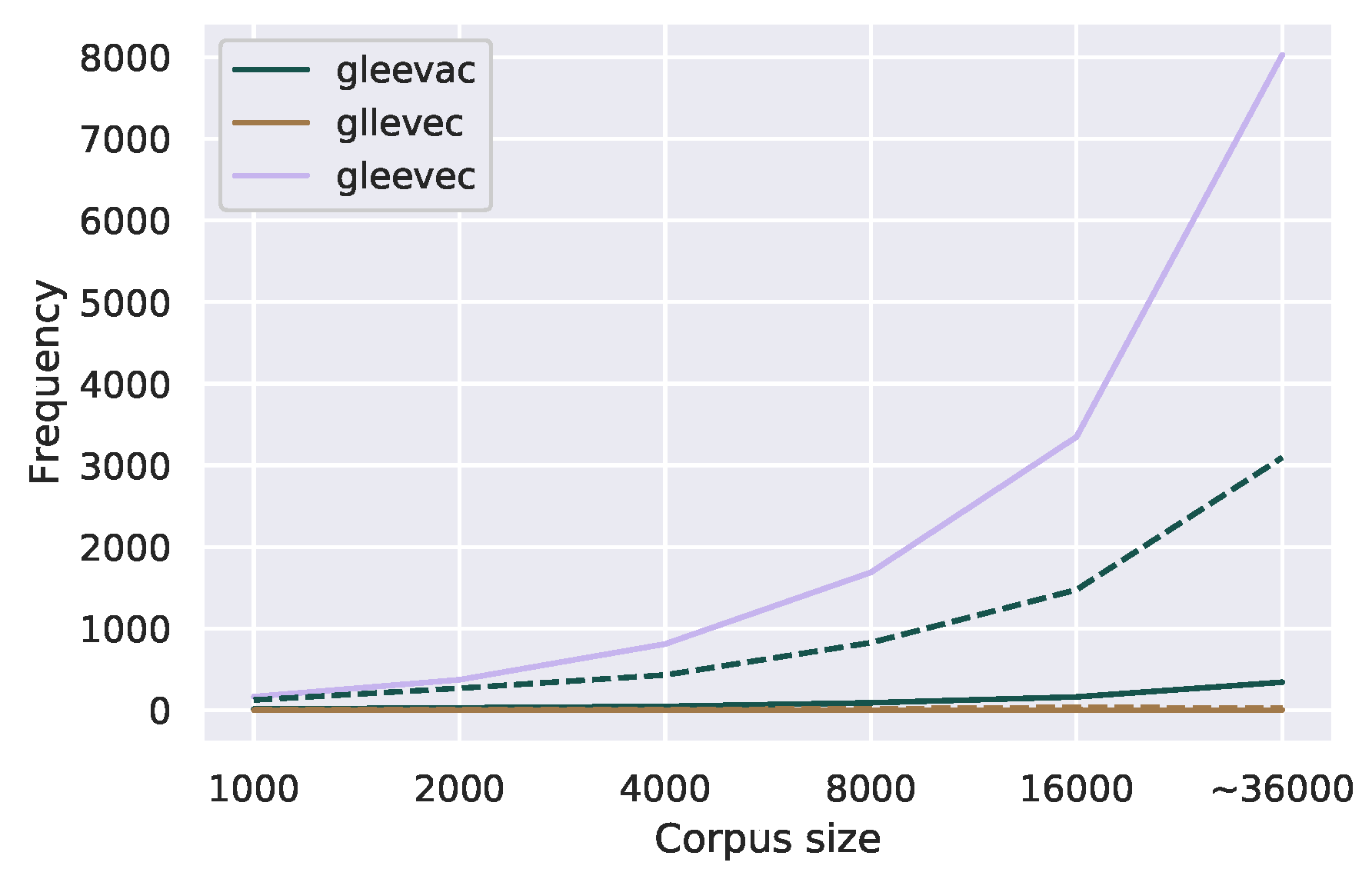
Figure 9.
Corpus frequency of one uncommon and one common misspelling of the medication Gleevec in the GIST forum with increasing corpus size. The dotted line indicates the corpus frequency threshold for correction candidates for each misspelling.
Figure 9.
Corpus frequency of one uncommon and one common misspelling of the medication Gleevec in the GIST forum with increasing corpus size. The dotted line indicates the corpus frequency threshold for correction candidates for each misspelling.
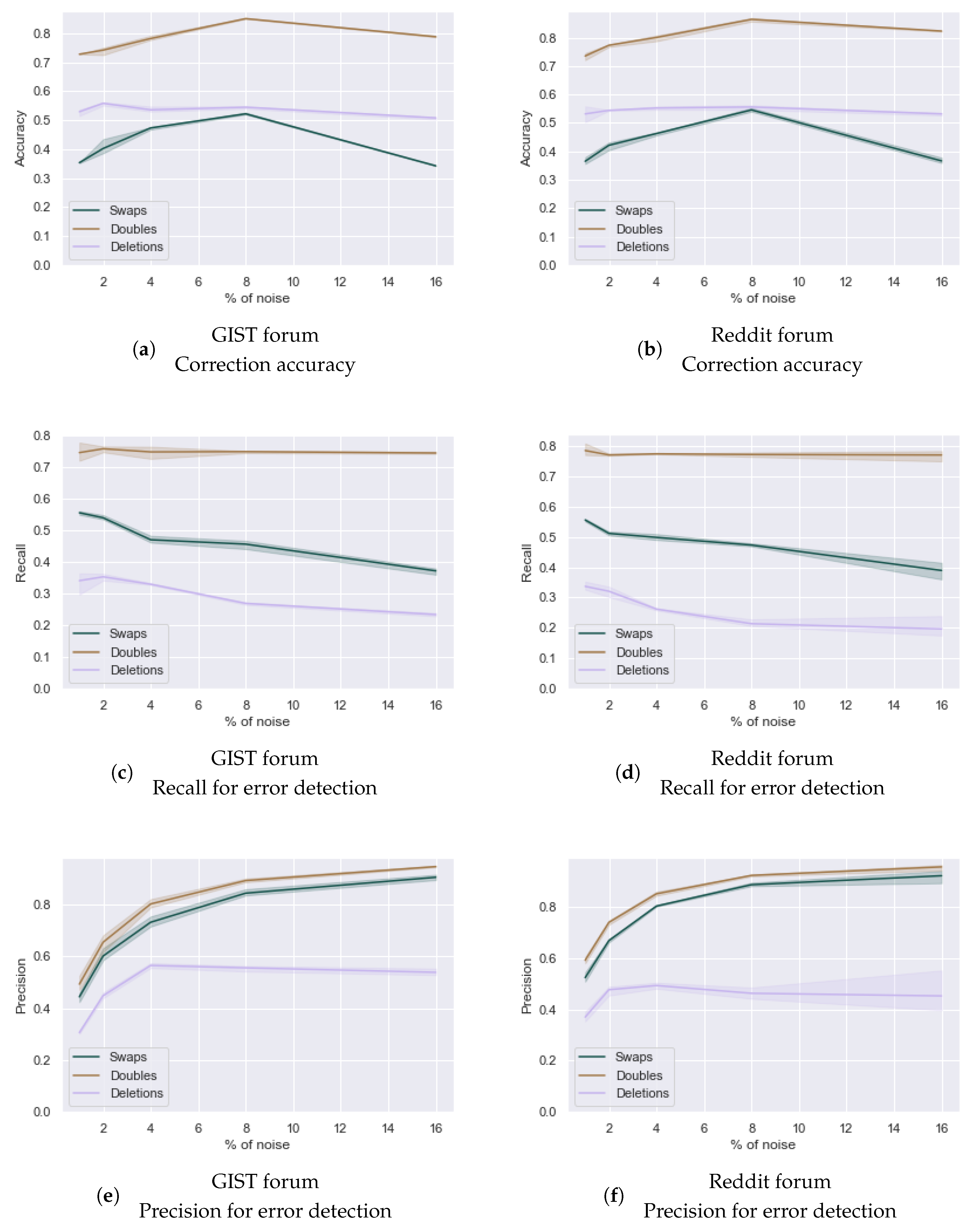
Figure 10.
Impact of degree of noisiness of the data (1%, 2%, 4%, 8% and 16% noise) on the detection (c–f) and correction accuracy (a,b) of three types of spelling errors (deletions of a single letter, doubling of a single letter and swaps of adjacent letters) in two cancer-related forums. The lines indicate the mean result while the band indicates the variance in results over three runs.
Figure 10.
Impact of degree of noisiness of the data (1%, 2%, 4%, 8% and 16% noise) on the detection (c–f) and correction accuracy (a,b) of three types of spelling errors (deletions of a single letter, doubling of a single letter and swaps of adjacent letters) in two cancer-related forums. The lines indicate the mean result while the band indicates the variance in results over three runs.
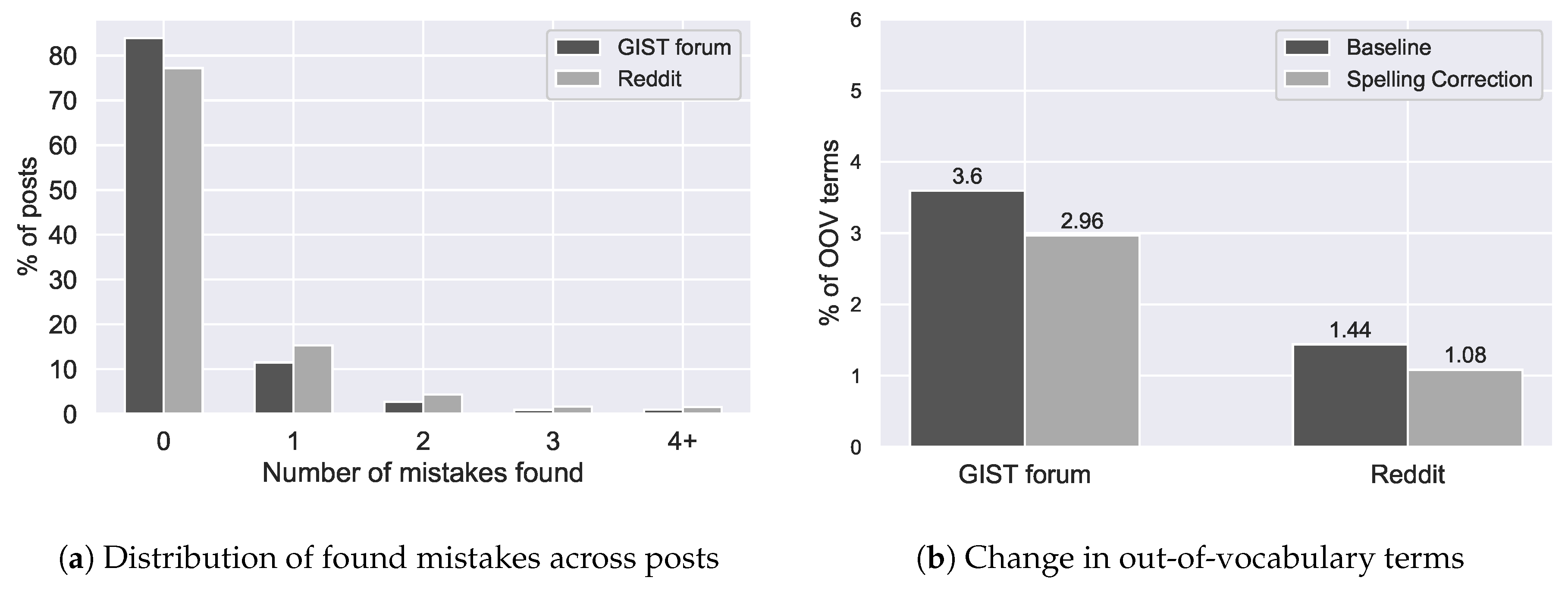
Figure 11.
The impact of spelling correction on the percentage of out-of-vocabulary terms in two cancer forums (b) and the distribution of the spelling mistakes that were found (a).
Figure 11.
The impact of spelling correction on the percentage of out-of-vocabulary terms in two cancer forums (b) and the distribution of the spelling mistakes that were found (a).
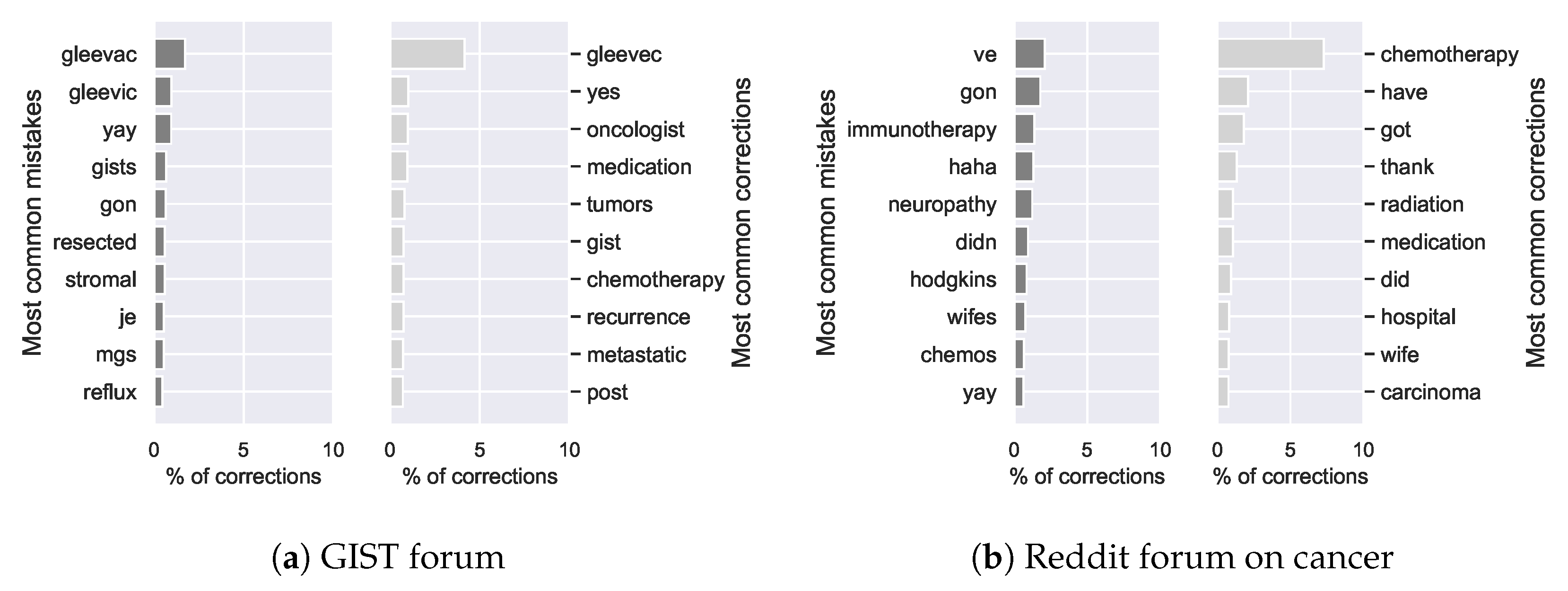
Figure 12.
Most frequent mistakes and corrections on the GIST forum (a) and the Reddit forum on cancer (b).
Figure 12.
Most frequent mistakes and corrections on the GIST forum (a) and the Reddit forum on cancer (b).
Table 1.
Raw data without punctuation. IQR, Inter-quartile range.
Table 1.
Raw data without punctuation. IQR, Inter-quartile range.
| | GIST Forum | Reddit Forum |
|---|
| # Tokens | 1,255,741 | 4,520,074 |
| # Posts | 36,277 | 274,532 |
| Median post length (IQR) | 20 (35) | 11 (18) |
Table 2.
Annotated data for spelling detection experiments. * Excluding punctuation, numbers and proper nouns.
Table 2.
Annotated data for spelling detection experiments. * Excluding punctuation, numbers and proper nouns.
| | Mistakes (%) | Total Word Count * |
|---|
| Training set | 57 (9.1%) | 627 |
| Test set | 45 (0.42%) | 10,760 |
Table 3.
Six classification datasets of health-related Twitter data. * SMM4H, Social Media Mining for Health Applications workshop.
Table 3.
Six classification datasets of health-related Twitter data. * SMM4H, Social Media Mining for Health Applications workshop.
| Dataset | Task | Size | Positive Class |
|---|
| Task 1 SMM4H 2019 * | Presence adverse drug reaction | 16,141 | 8.7% |
| Task 4 SMM4H 2019 * Flu vaccine | Personal health mention of flu vaccination | 6738 | 28.3% |
| Flu Vaccination Tweets [44] | Relevance to topic flu vaccination | 3798 | 26.4% |
| Twitter Health [45] | Relevance to health | 2598 | 40.1% |
| Task4 SMM4H 2019 * Flu infection | Personal health mention of having flu | 1034 | 54.4% |
| Zika Conspiracy Tweets [46] | Contains pseudo-scientific information | 588 | 25.9% |
Table 4.
Error distribution in 1000 GIST posts.
Table 4.
Error distribution in 1000 GIST posts.
| Error Type | Non-Word | Incorrect Splits | Incorrect Concatenations | Real Word |
|---|
| Amount | 109 | 17 | 24 | 30 |
| Non-Medical/Medical | 57/52 | 25/5 | 14/3 | 18/6 |
| Percentage of tokens | 0.32% | 0.05% | 0.07% | 0.09% |
| Example mistake | gleevac | gall bladder | sideeffects | scant |
| Example correction | gleevec | gallbladder | side effects | scan |
Table 5.
Correction accuracy using a specialized vocabulary. AE, absolute edit distance; RE, relative edit distance; WAE, weighted absolute edit distance; WRE, weighted relative edit distance. * Only the best corpus frequency threshold is reported.
Table 5.
Correction accuracy using a specialized vocabulary. AE, absolute edit distance; RE, relative edit distance; WAE, weighted absolute edit distance; WRE, weighted relative edit distance. * Only the best corpus frequency threshold is reported.
| Source of Correction Candidates | Ceiling | AE | RE | WAE | WRE | Sarker | TISC |
|---|
| Specialized vocabulary | 92.0% | 58.0% | 64.7% | 63.3% | 68.2% | 19.3% | 14.8% |
| GIST forum text * | 97.6% | 73.9% | 73.9% | 70.4% | 72.7% | 44.3% | - |
Table 6.
Mean computation time over 5 runs.
Table 6.
Mean computation time over 5 runs.
| AE | RE | WAE | WRE | Sarker |
|---|
| 13.36 ms | 14.04 ms | 29.45 ms | 32.00 ms | 904.33 ms |
Table 7.
Corrections by different methods with candidates from a specialized vocabulary. * Gleevec and Sutent are important medications for GIST patients.
Table 7.
Corrections by different methods with candidates from a specialized vocabulary. * Gleevec and Sutent are important medications for GIST patients.
| Mistake | Correction | AE | RE | WAE | WRE | Sarker | TISC |
|---|
| gleevac | gleevec * | gleevec | gleevec | gleevec | gleevec | colonic | gleevac |
| stomack | stomach | stomach | stomach | smack | stomach | smack | smack |
| ovari | ovary | ovary | ovary | ovary | ovary | ova | atari |
| sutant | sutent * | mutant | mutant | sutent | sutent | mutant | dunant |
| mestastis | metastasis | miscasts | metastasis | metastasis | metastasis | miscasts | mestastis |
Table 8.
Corrections by different methods with data-driven candidates. AE, absolute edit distance; RE, relative edit distance; WAE, weighted absolute edit distance; WRE, weighted relative edit distance.
Table 8.
Corrections by different methods with data-driven candidates. AE, absolute edit distance; RE, relative edit distance; WAE, weighted absolute edit distance; WRE, weighted relative edit distance.
| Mistake | Correction | AE | RE | WAE | WRE | Sarker |
|---|
| gleevac | gleevec | gleevec | gleevec | gleevec | gleevec | gleevec |
| stomack | stomach | stomach | stomach | stomach | stomach | stuck |
| ovari | ovary | ovary | ovary | ovary | ovary | ovarian |
| sutant | sutent | sutent | sutent | sutent | sutent | mutant |
| mestastis | metastasis | metastis | metastis | metastis | metastis | metastis |
Table 9.
Effect of lemmatization of the errors (LemmatizedInput), their corrections (LemmatizedOutput) and correction candidates (LemmatizedCandidates) on spelling correction accuracy using RE ( = 9).
Table 9.
Effect of lemmatization of the errors (LemmatizedInput), their corrections (LemmatizedOutput) and correction candidates (LemmatizedCandidates) on spelling correction accuracy using RE ( = 9).
| NoLemmatization | LemmatizedInput | + LemmatizedOutput | + LemmatizedCandidates |
|---|
| 73.6% | 73.6% | 64.7% | 67.0% |
Table 10.
Changes in corrections when HealthVec was added (weight = 0.6) to the relative edit distance (weight = 0.4) with = 9. LM = language model.
Table 10.
Changes in corrections when HealthVec was added (weight = 0.6) to the relative edit distance (weight = 0.4) with = 9. LM = language model.
| | Error | Correct Word | Correction |
|---|
| Without LM | With LM |
|---|
| Improved | alse | else | false | else |
| lm | im | am | im |
| esle | else | resolve | else |
| explane | explain | explained | explain |
| ovarie | ovary | ovary | ovaries |
| surgerys | surgeries | surgeries | surgery |
| Missed | surgerys | surgery | surgery | surgury |
Table 11.
Results for mistake detection methods on the test set.
Table 11.
Results for mistake detection methods on the test set.
| Method | Mistakes Found | Recall | Precision | F0.5 | F1 |
|---|
| CELEX | 395 | 1.0 | 0.11 | 0.13 | 0.20 |
| Aspell dictionary | 163 | 1.0 | 0.26 | 0.31 | 0.42 |
| TISC | 270 | 0.74 | 0.12 | 0.14 | 0.21 |
| Microsoft word | 395 | 0.88 | 0.10 | 0.12 | 0.18 |
| Our method (RE = 0.76) | 90 | 0.91 | 0.46 | 0.51 | 0.61 |
| Our method (RE = 0.76) + ConcatCorrection | 92 | 0.96 | 0.47 | 0.52 | 0.63 |
Table 12.
Examples of false positives and negatives of our error detection method.
Table 12.
Examples of false positives and negatives of our error detection method.
| | Mistakes (their Corrections with our Method) |
|---|
| False positives | intolerances (intolerant) | resected (removed) | reflux (really) | condroma (syndrome) |
| False negatives | istological (histological) | vechile (vehicle) | | |
Table 13.
Manual error analysis of 50 most frequent OOV terms after spelling detection.
Table 13.
Manual error analysis of 50 most frequent OOV terms after spelling detection.
| | GIST | Example | Reddit | Example |
|---|
| Real word | 33 | unpredictable, internet | 42 | misdiagnosed, website |
| Spelling mistake | 5 | side-effects, wildtype, copay, listserve, listserve | 2 | side-effects, inpatient |
| Abbreviation | 2 | mos, wk | 3 | aka |
| Slang | 6 | scanxiety, gister | 1 | rad |
| Drug name | 2 | stivarga, mastinib | 1 | ativan |
| Not English | 2 | que, moi | - | |
| TOTAL | 50 | | 50 | |
Table 14.
Mean classification accuracy before and after normalization for six health-related classification tasks. Only the results for the best performing classifier per dataset are reported. indicates p < 0.005; * indicates p < 0.01; † indicates absolute change.
Table 14.
Mean classification accuracy before and after normalization for six health-related classification tasks. Only the results for the best performing classifier per dataset are reported. indicates p < 0.005; * indicates p < 0.01; † indicates absolute change.
| | | F1 | Recall | Precision |
|---|
| Dataset | Words Altered | Pre | Post | † | Pre | Post | † | Pre | Post | † |
| Task1 SMM4H 2019 | 1.53% | 0.410 | 0.410 | −0.0007 | 0.373 | 0.387 | +0.014 | 0.470 | 0.445 | −0.025 |
| Task4 SMM4H 2019 Flu Vaccination | 0.50% | 0.780 | 0.787 | +0.006 | 0.834 | 0.843 | +0.008 | 0.733 | 0.738 | +0.005 |
| Flu Vaccination Tweets | 0.50% | 0.939 | 0.941 | +0.002 | 0.935 | 0.939 | +0.004 | 0.943 | 0.943 | +0.0004 |
| Twitter Health | 0.71% | 0.702 | 0.718 | +0.016 * | 0.657 | 0.685 | +0.028 * | 0.756 | 0.755 | −0.0009 |
| Task4 SMM4H 2019 Flu Infection | 0.57% | 0.784 | 0.800 | +0.012 ** | 0.842 | 0.854 | +0.013 | 0.735 | 0.754 | +0.019 ** |
| Zika Conspiracy | 0.36% | 0.822 | 0.817 | −0.005 | 0.817 | 0.829 | +0.012 | 0.835 | 0.814 | −0.021 |
Table 15.
Results for unconstrained systems of ACL W-NUT 2015.
Table 15.
Results for unconstrained systems of ACL W-NUT 2015.
| | F1 | Precision | Recall |
|---|
| MoNoise [16] | 0.864 | 0.934 | 0.803 |
| Sarker’s method [5] | 0.836 | 0.880 | 0.796 |
| IHS_RD [50] | 0.827 | 0.847 | 0.808 |
| USZEGED [51] | 0.805 | 0.861 | 0.756 |
| BEKLI [52] | 0.757 | 0.774 | 0.742 |
| LYSGROUP [53] | 0.531 | 0.459 | 0.630 |
| Our method | 0.743 | 0.734 | 0.753 |
Table 16.
Manual analysis of 100 most frequent errors in W-NUT. * also considered non-word mistakes.
Table 16.
Manual analysis of 100 most frequent errors in W-NUT. * also considered non-word mistakes.
| Type of Error | Frequency | Example | Our Correction | W-NUT Annotation |
|---|
| Should not have been altered | 46 | info, kinda | information, kind of | info, kinda |
Abbreviation not or
incorrectly expanded | 19 | smh | smh | shaking my head |
| Uncorrected slang | 14 | esp | esp | especially |
| Missed concatenation error * | 6 | incase | incase | in case |
| Missed apostrophe * | 5 | youre | youre | you’re |
| Wrong correction | 4 | u | your | your |
| Missed split mistake * | 3 | i g g y | i g g y | iggy |
| Missed non-word spelling mistake | 2 | limites | limites | limits |
| American English | 1 | realise | realize | realise |
| TOTAL | 100 | | | |