Induction of a Th17 Phenotype in Human Skin—A Mimic of Dermal Inflammatory Diseases
Abstract
:1. Introduction
2. Materials and Equipment
2.1. General Materials
- Serological pipets, various sizes (ThermoFisher Scientific, Waltham, MA, USA; Cat. nos.: 13-678-11C, 13-678-11D, 13-678-11E)
- Barrier pipet tips, various sizes (ThermoFisher Scientific, Waltham, MA, USA; Cat. nos.: 02-717-158, 02-717-161, 02-717-165, 02-717-166)
2.2. Materials for ex vivo Skin Culture
- Dulbecco’s modified Eagle’s medium (DMEM; MediaTech, Oceanside, CA, USA; Cat. no.: 10-013)
- Antibiotic/antimycotic (penicillin, streptomycin, and amphotericin B; ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 15-240-062)
- Sterile disposable 3 mm-diameter biopsy punches (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 12-460-406)
- Non-pyrogenic polystyrene multi-well plates (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 08-757-214)
- Cell culture dishes, 10 cm ((ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 07-000-386)
- Single-edge steel razor blades, individually wrapped (VWR, Radnor, PA, USA; Cat. no.: 55411-050)
2.3. Materials for Th17-Skewing Cocktail
- Anti-CD3 antibody (Abcam, Cambridge, MA, USA; Cat. no.: ab8090)
- Anti-CD28 antibody (R & D Systems, Minneapolis, MN, USA; Cat. no.: MAB342)
- Recombinant human IL-23 (R & D Systems, Minneapolis, MN, USA; Cat. no.: 1290-IL)
- Recombinant human IL-1β (Peprotech, Rocky Hill, NJ, USA; Cat. no.: 200-01B)
- Recombinant human IFNγ (R & D Systems, Minneapolis, MN, USA; Cat. no.: 285-IF)
- Recombinant human IL-4 (R & D Systems, Minneapolis, MN, USA; Cat. no.: 204-IL)
- Recombinant human IL-6 (R & D Systems, Minneapolis, MN, USA; Cat. no.: 206-IL)
- Recombinant human IL-21 (R & D Systems, Minneapolis, MN, USA; Cat. no.: 8879-IL)
- Recombinant human TGFβ1 (R & D Systems, Minneapolis, MN, USA; Cat. no.: 240-B)
- Sterile PBS (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 17-516F)
- Sterile RNAse-free water (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 10-977-015)
2.4. Materials for Endpoint Analyses
- TRIzolTM (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 15-596-018)
- Hard tissue homogenizing 2 mL reinforced bead tubes, nuclease-free (Omni International, Kennesaw, GA, USA; Cat. no.: 19-628)
- Sterile RNAse-free water (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 10-977-015)
- SuperScriptTM IV First-Strand Synthesis system (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 18091050)
- TaqManTM Gene Expression Master Mix (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 4369016)
- MicroAmpTM Fast Optical 96-well reaction PCR plates (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 43-469-07)
- IL-17A primers (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: Hs00174383_m1)
- IL-17F primers (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: Hs00369400_m1)
- IL-22 primers (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: Hs01574154_m1)
- IL-26 primers (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: Hs00218189_m1)
- IL-10 primers (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: Hs00961622_m1)
- PPIB primers (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: Hs00168719_m1)
- Human IL-17A Coated ELISA kit (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: BMS2017)
2.5. Equipment
- Biological Safety Cabinet (Labconco Corporation, Kansas City, MO, USA; Cat. no.: 362090412242)
- Pipet-Aid® (Drummond, Broomall, PA, USA; Cat. no.: 4-000-100)
- Pipettes capable of measuring 1 μL–1000 μL (Gilson, Middleton, WI, USA; Cat. no.: F167370)
- Stainless steel sharp-pointed scissors (ThermoFisher Scientific, Waltham, MA, USA; Cat. 08-940)
- Stainless steel fine point forceps (ThermoFisher Scientific, Waltham, MA, USA; Cat. 22-327379)
- Incubator (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 13-998-211)
- −80°C Ultra-Low Temperature Upright Freezer (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 907)
- Micro Centrifuge (VWR, Radnor, PA, USA; Cat. no.: 2405-37)
- BeadRuptor24 (Omni International, Kennesaw, GA, USA; Cat. no.: 19-2141A)
- NanoDrop Lite spectrophotometer (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: ND-LITE)
- C1000 TouchTM Thermal Cycler (Bio-Rad Laboratories, Hercules, CA, USA; Cat. no.: 1851148)
- StepOneTM Real-Time PCR System (ThermoFisher Scientific, Waltham, MA, USA; Cat. no.: 4376357)
- Synergy H1 microplate reader (BioTek Instruments, Inc., Winooski, VT, USA; Cat. no.: H1M)
3. Procedure
- Prior to treating skin, ensure that all reagent stocks have been prepared and treatment components have been resuspended in the vehicle recommended by the manufacturer (most require PBS). The suggested stock concentrations should range between 500×–1000× whenever possible.
- 1.1.
- Aliquot into small or single-use volumes to minimize freeze-thaw cycles and store at −80 °C until the day of the assay.
- 1.2.
- Time for Completion: ~1 h, including labeling of aliquot tubes.
- Defat and clean skin with ethanol and PBS, as previously described [21].
- 2.1.
- Briefly, place skin dermal-side down atop a layer of aluminum foil on the working surface in a biological safety cabinet and clean blood/debris off of dermis using 70% ethanol.
- 2.2.
- Separate the dermal and adipose layers using scissors and razor blade/scalpel.
- 2.2.1.
- Discard skin portions containing stretch marks, birth marks, and other imperfections, as well as all adipose tissue.
- 2.2.2.
- Using scissors, cut skin into pieces sized ~1–2 inches.
- 2.3.
- Assemble five conical tubes containing ~45 mL 70% ethanol and three conical tubes containing ~45 mL sterile PBS each to clean skin.
- 2.3.1.
- Vigorously dip a piece of skin into the first tube of ethanol 5–6 times, then repeat dipping likewise into each of the next four ethanol tubes, followed by each of the PBS tubes, in a successive manner to clean the skin.
- 2.3.2.
- Place the cleaned skin dermal-side down in a 10 cm dish.
- 2.3.3.
- Repeat for all pieces of skin, arranging the skin pieces in a non-overlapping manner into 10 cm dish(es).
- 2.4.
- Add serum-free DMEM with 2× antibiotic/antimycotic to cover the bottom of the dish, leaving the top of the skin exposed to air.
- 2.5.
- Incubate at 37 °C/5% CO2 until ready for use.
- 2.6.
- Time for Completion: 30–60 min including setup and cleanup.
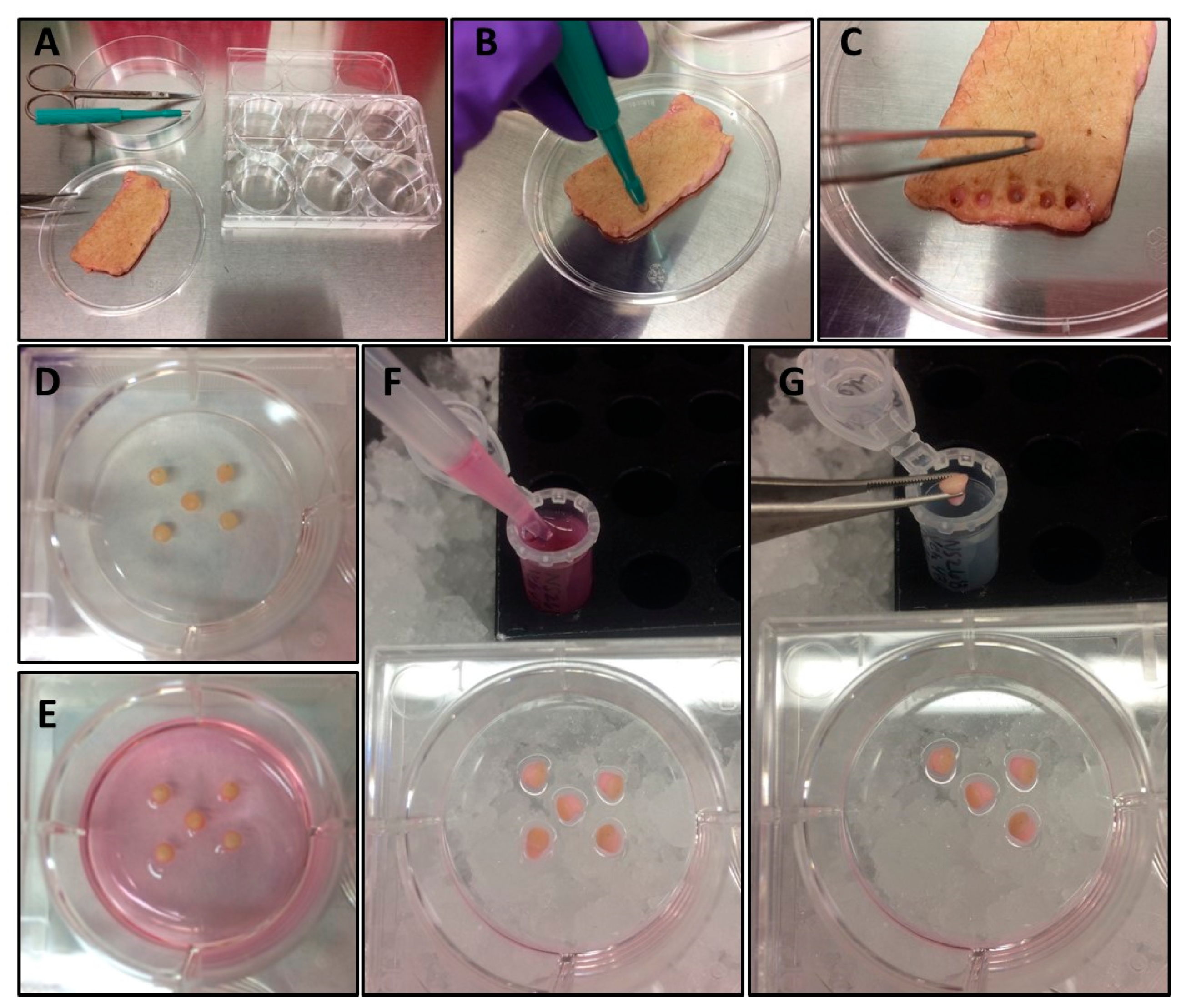
- Using a 3 mm punch biopsy, punch and distribute 3–5 biopsies from cleaned, defatted skin into each well of six-well tissue culture plate to be used for the experiment (Figure 1A–D).
- 3.1.
- Place each biopsy with the dermal side contacting the plastic and the epidermal side facing up (Figure 1D).
- 3.2.
- Time for Completion: Time is dependent on the size of the experiment. Allot ~20 min per 6-well plate.
- 3.3.
- Allow skin punches to adhere approximately 15 min. before adding treatment media.
- While waiting for punches to adhere, prepare treatments as follows:
- 4.1.
- Prepare cocktail and vehicle master mixes per Table 2 in serum-free DMEM +1% antibiotic/antimycotic.
- 4.1.1.
- Cocktail formulations (i.e., CT4, CT8, and CT9) are named according to the number of constituent components, as follows: CT9 contains 1 μg/mL each of anti-CD3, anti-CD28, IL-4, and IFNγ, 10 ng/mL each of IL-23, IL-1β, IL-6, and IL-21, and 1 ng/mL TGFβ; CT8 contains all of the CT9 components, except IL-1β; and, CT4 only contains anti-CD3, anti-CD28, IL-23, and IL-1β.
- 4.2.
- Whether treating a single well or more than one well with the same formulation, it is recommended to make a master mix, combining all of the components that are required in a conical tube first.
- 4.3.
- Mix the tubes containing the cocktail component formulations, and then distribute treatments into well(s) containing the skin punches in a dropwise fashion to avoid dislodging the skin punches from the plastic (Figure 1E).
- 4.3.1.
- For six-well tissue culture plates, a volume of 1 mL–2 mL is recommended, depending on the height of the skin punch.
- 4.3.2.
- Add sufficient media to cover the dermal layer of skin, leaving the epidermal layer exposed to air.
- 4.3.2.1.
- When viewed from the side of the well, the dermal layer should be completely submerged in media, leaving the topmost epidermal portion exposed to the air above the surface of the media.
- 4.4.
- Time for completion: ~15 min.
- Incubate at 37 °C/5% CO2 for 48 h.
- Harvest skin punches and supernatants into Eppendorf tubes and freeze at −80°C until analysis (Figure 1F–G).
- 6.1.
- Label three sets of Eppendorf tubes, one set to store skin punches and two sets for supernatants.
- 6.2.
- Using a P1000 pipette, transfer media from each treatment well into a corresponding pre-labeled Eppendorf tube (Figure 1F).
- 6.2.1.
- Centrifuge at 5000 RPM for 5 min to pellet cellular debris and dead cells.
- 6.2.2.
- Transfer liquid portion (supernatant) to clean pre-labeled Eppendorf tube on ice. Discard the tube containing pellet.
- 6.3.
- Transfer skin punches to corresponding Eppendorf tube on ice (Figure 1G).
- 6.3.1.
- Skin punches from the same treatment well can be frozen into one Eppendorf tube. Stagger the punches along the inside wall of the tube to ease future removal once frozen.
- 6.4.
- Freeze tubes containing skin punches and centrifuged supernatants at −80 °C.
- 6.5.
- Time for completion: Time is dependent on the size of the experiment. Allot ~20 min per 6-well plate.
- RNA isolation, reverse transcription, and quantitative real-time polymerase chain reaction.
- 7.1.
- Homogenize one skin punch per treatment in 1 mL TRIzolTM using a Bead Ruptor 24 tissue homogenizer for 30 s at a speed of 6.95, followed by two minutes on dry ice, repeated for a total of 4–5 cycles.
- 7.1.1.
- Time for Completion: 20–30 min.
- 7.2.
- Transfer homogenate out of bead tubes and into clean Eppendorf tubes.
- 7.2.1.
- Time for Completion: 5 min.
- 7.3.
- Isolate RNA using the TRIzolTM separation method per the manufacturer’s instructions and resuspend in purified nuclease-free water.
- 7.3.1.
- Time for Completion: 3 h.
- 7.4.
- Quantify RNA and reverse-transcribe cDNA from 2 μg RNA per 20 μL cDNA reaction using the SuperScript® IV First-Strand Synthesis system and oligo (dT) primers.
- 7.4.1.
- Time for Completion: 1 h.
- 7.5.
- In a 10 μL final reaction volume including best-coverage TaqMan® Gene Expression primers, include 2 μL cDNA and plate in replicate for real-time PCR.
- 7.5.1.
- The baseline expression of target genes tends to be undetectable after 55 cycles in vehicle-treated samples when starting from 1 μg of RNA and 1 μL of cDNA. Using greater quantities of starting material specified in this protocol helps to ensure a calculable Ct value for vehicles, which aids in quantifiable data analysis.
- 7.5.2.
- Time for Completion: 3 h.
- 7.6.
- Determine gene expression using the delta delta Ct method [24], normalizing data to the housekeeping gene peptidylprolyl isomerase B (PPIB).
- 7.6.1.
- Relative fold change = 2−ΔΔCt where Ct = threshold cycle and ΔΔCt = ((Treatment gene of interest Ct value − Vehicle gene of interest Ct value) − (Treatment PPIB Ct value − Vehicle PPIB Ct value)).
- 7.6.2.
- Time for Completion: 1 h.
- IL-17A ELISA.
- 8.1.
- Measure the secreted IL-17A in supernatants diluted 1:1 using a commercially available sandwich ELISA kit with a dynamic detection range between 1.6 pg/mL–100 pg/mL per manufacturer’s instructions.
- 8.1.1.
CRITICAL STEP Thaw supernatants on ice.
- 8.2.
- Using a plate reader, read absorbance at 450 nm.
- 8.3.
- When calculating IL-17A secretion, be sure to multiply values by 2 to account for the dilution that was performed according to kit instructions.
- 8.3.1.
- Time for Completion: 6–7 h.
4. Expected Results
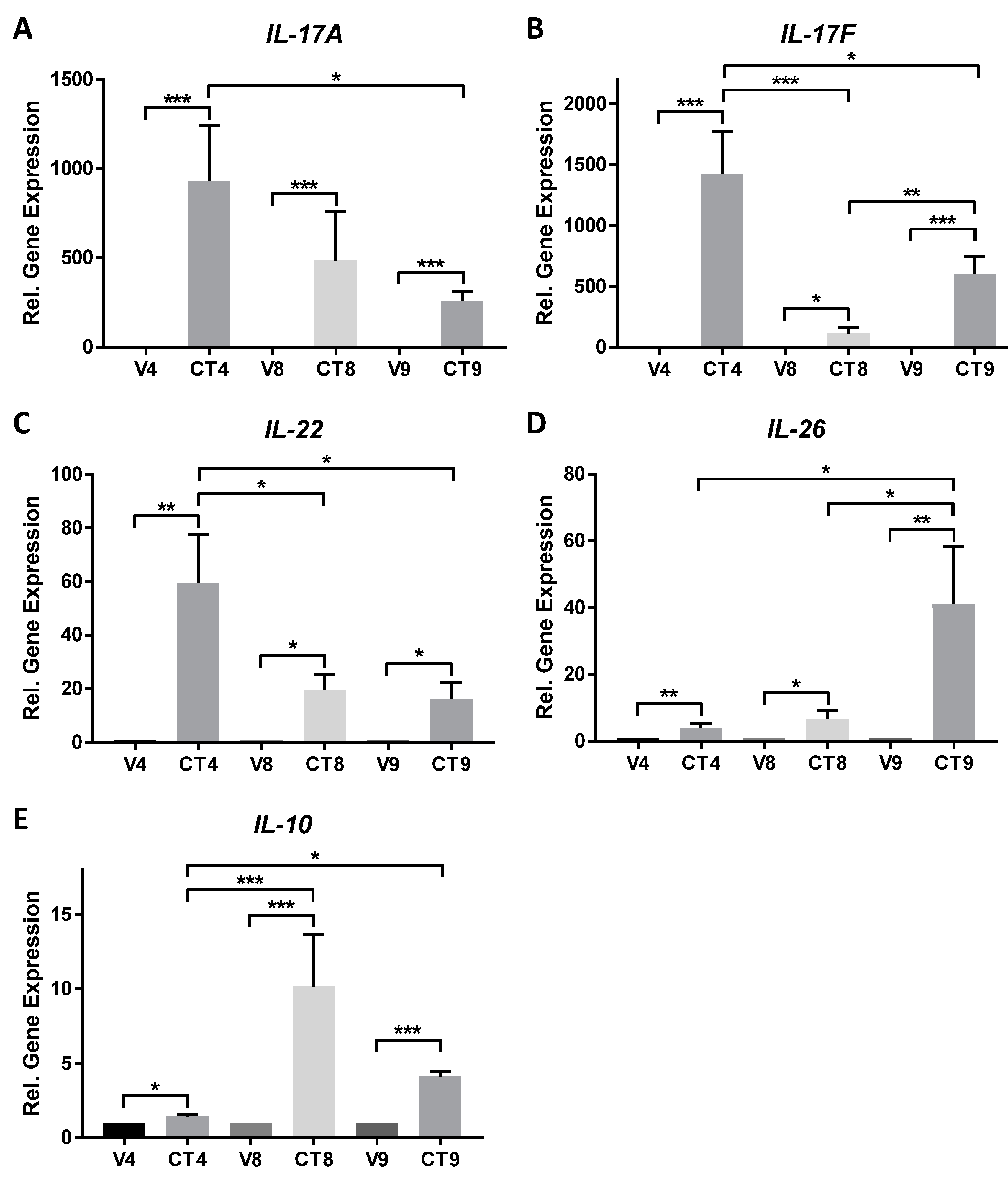
4.1. Regulation of Gene Expression by Different Cocktail Formulations
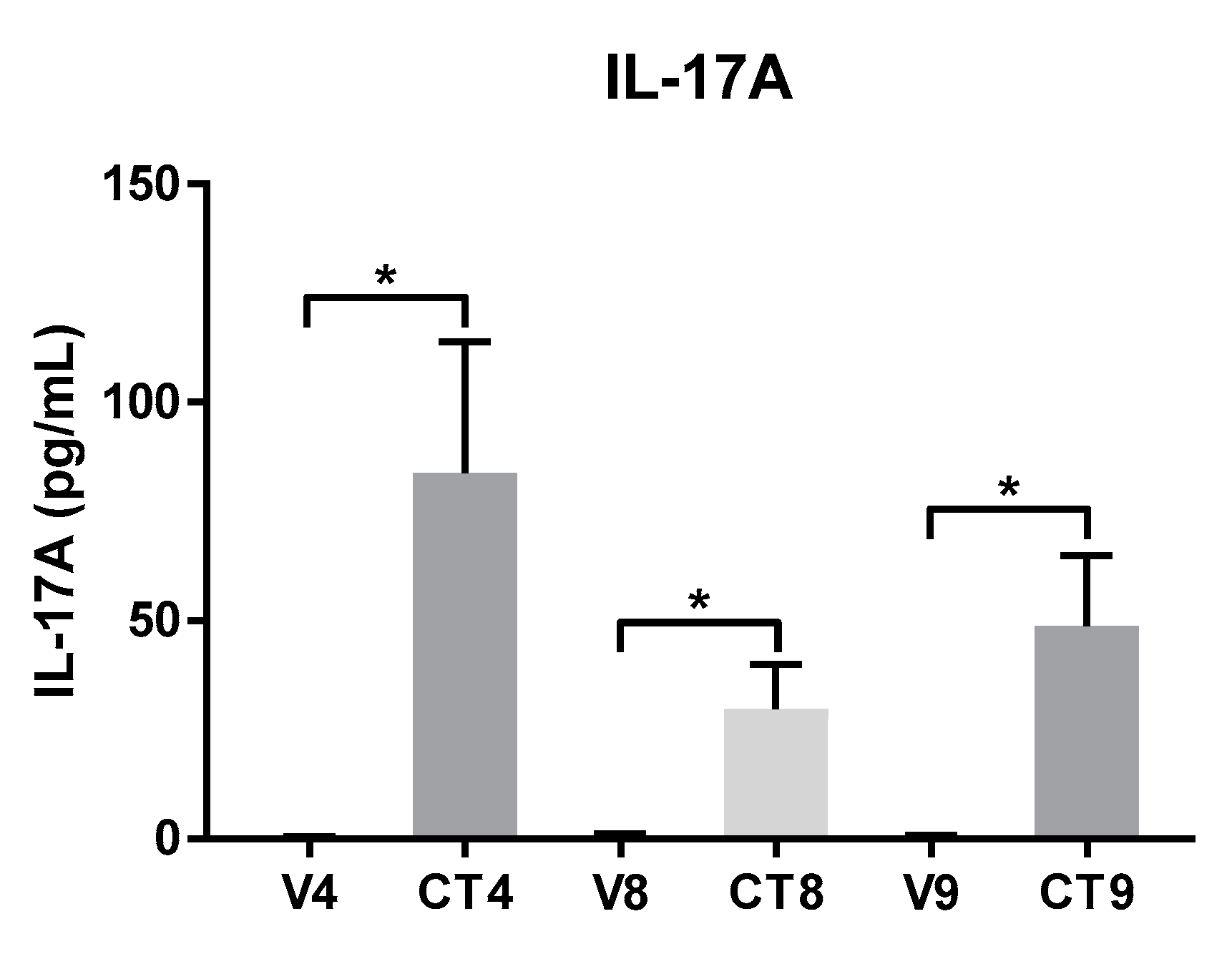
4.2. Regulation of Protein Levels of IL-17A by Different Cocktail Formulations
4.3. Interpretation/Discussion:
Author Contributions
Funding
Conflicts of Interest
References
- Bettelli, E.; Carrier, Y.; Gao, W.; Korn, T.; Strom, T.B.; Oukka, M.; Weiner, H.L.; Kuchroo, V.K. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature 2006, 441, 235–238. [Google Scholar] [CrossRef] [PubMed]
- Harrington, L.E.; Mangan, P.R.; Weaver, C.T. Expanding the effector CD4 T-cell repertoire: The Th17 lineage. Curr. Opin. Immunol. 2006, 18, 349–356. [Google Scholar] [CrossRef] [PubMed]
- Luckheeram, R.V.; Zhou, R.; Verma, A.D.; Xia, B. CD4(+)T cells: Differentiation and functions. Clin. Dev. Immunol. 2012, 2012, 925135. [Google Scholar] [CrossRef] [PubMed]
- Tzartos, J.S.; Friese, M.A.; Craner, M.J.; Palace, J.; Newcombe, J.; Esiri, M.M.; Fugger, L. Interleukin-17 production in central nervous system-infiltrating T cells and glial cells is associated with active disease in multiple sclerosis. Am. J. Pathol. 2008, 172, 146–155. [Google Scholar] [CrossRef] [PubMed]
- Yamagata, T.; Skepner, J.; Yang, J. Targeting Th17 effector cytokines for the treatment of autoimmune diseases. Arch. Immunol. Ther. Exp. 2015, 63, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Yang, J.; Xing, X.; Wan, L.; Li, M. Increased frequency of Th17 cells in systemic sclerosis is related to disease activity and collagen overproduction. Arthritis Res. Ther. 2014, 16, R4. [Google Scholar] [CrossRef] [PubMed]
- O’Reilly, S.; Hügle, T.; Van Laar, J.M. T cells in systemic sclerosis: A reappraisal. Rheumatology 2012, 51, 1540–1549. [Google Scholar] [CrossRef] [PubMed]
- Huber, S.; Gagliani, N.; Esplugues, E.; O’Connor, W., Jr.; Huber, F.J.; Chaudhry, A.; Kamanaka, M.; Kobayashi, Y.; Booth, C.J.; Rudensky, A.Y.; et al. Th17 cells express interleukin-10 receptor and are controlled by Foxp3(-) and Foxp3+ regulatory CD4+ T cells in an interleukin-10-dependent manner. Immunity 2011, 34, 554–565. [Google Scholar] [CrossRef] [PubMed]
- Ivanov, I.I.; McKenzie, B.S.; Zhou, L.; Tadokoro, C.E.; Lepelley, A.; Lafaille, J.J.; Cua, D.J.; Littman, D.R. The orphan nuclear receptor RORgammat directs the differentiation program of proinflammatory IL-17+ T helper cells. Cell 2006, 126, 1121–1133. [Google Scholar] [CrossRef] [PubMed]
- Nalbant, A.; Eskier, D. Genes associated with T helper 17 cell differentiation and function. Front. Biosci. 2016, 8, 427–435. [Google Scholar] [CrossRef]
- Busman-Sahay, K.O.; Walrath, T.; Huber, S.; O’Connor, W. Cytokine crowdsourcing: Multicellular production of TH17-associated cytokines. J. Leukoc. Biol. 2015, 97, 499–510. [Google Scholar] [CrossRef] [PubMed]
- Miossec, P.; Kolls, J.K. Targeting IL-17 and TH17 cells in chronic inflammation. Nat. Rev. Drug Discov. 2012, 11, 763–776. [Google Scholar] [CrossRef] [PubMed]
- O’Connor, W., Jr.; Kamanaka, M.; Booth, C.J.; Town, T.; Nakae, S.; Iwakura, Y.; Kolls, J.K.; Flavell, R.A. A protective function for interleukin 17A in T cell–mediated intestinal inflammation. Nat. Immunol. 2009, 10, 603–609. [Google Scholar] [CrossRef] [PubMed]
- Sziksz, E.; Pap, D.; Lippai, R.; Béres, N.J.; Fekete, A.; Szabó, A.J.; Vannay, Á. Fibrosis related inflammatory mediators: Role of the IL-10 cytokine family. Mediat. Inflamm. 2015, 2015, 764641. [Google Scholar] [CrossRef] [PubMed]
- Couper, K.N.; Blount, D.G.; Riley, E.M. IL-10: The master regulator of Immunity to infection. J. Immunol. 2008, 180, 5771–5777. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, M.D.; Simpson, K.L.; Keelan, J.A. Paradoxical proinflammatory actions of interleukin-10 in human amnion: Potential roles in term and preterm labour. J. Clin. Endocrinol. Metab. 2004, 89, 4149–4152. [Google Scholar] [CrossRef] [PubMed]
- Muhl, H. Pro-inflammatory signaling by IL-10 and IL-22: Bad habit stirred up by interferons? Front. Immunol. 2013, 4, 18. [Google Scholar] [CrossRef] [PubMed]
- Rajkovic, I.; Dragicevic, A.; Vasilijic, S.; Bozic, B.; Dzopalic, T.; Tomic, S.; Majstorovic, I.; Vucevic, D.; Djokic, J.; Balint, B.; et al. Differences in T-helper polarizing capability between human monocyte-derived dendritic cells and monocyte-derived Langerhans’-like cells. Immunology 2011, 132, 217–225. [Google Scholar] [CrossRef] [PubMed]
- Vanderlocht, J.; Van Elssen, C.H.; Senden-Gijsbers, B.L.; Meek, B.; Cloosen, S.; Libon, C.; Bos, G.M.; Germeraad, W.T. Increased tumor-specific CD8+ T cell induction by dendritic cells matured with a clinical grade TLR-agonist in combination with IFN-gamma. Int. J. Immunopathol. Pharmacol. 2010, 23, 35–50. [Google Scholar] [CrossRef]
- Wilson, N.J.; Boniface, K.; Chan, J.R.; McKenzie, B.S.; Blumenschein, W.M.; Mattson, J.D.; Basham, B.; Smith, K.; Chen, T.; Morel, F.; et al. Development, cytokine profile and function of human interleukin 17–producing helper T cells. Nat. Immunol. 2007, 8, 950–957. [Google Scholar] [CrossRef]
- Watanabe, T.; Nishimoto, T.; Mlakar, L.; Heywood, J.; Malaab, M.; Hoffman, S.; Feghali-Bostwick, C. Optimization of a murine and human tissue model to recapitulate dermal and pulmonary features of systemic sclerosis. PLoS ONE 2017, 12, 0179917. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, Y.; Takihara, T.; Chambers, R.A.; Veraldi, K.L.; Larregina, A.T.; Feghali-Bostwick, C.A. A peptide derived from endostatin ameliorates organ fibrosis. Sci. Transl. Med. 2012, 4, 136ra71. [Google Scholar] [CrossRef] [PubMed]
- Yasuoka, H.; Larregina, A.T.; Yamaguchi, Y.; Feghali-Bostwick, C.A. Human skin culture as an ex vivo model for assessing the fibrotic effects of insulin-like growth factor binding proteins. Open Rheumatol. J. 2008, 2, 17–22. [Google Scholar] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)). Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Ghoreschi, K.; Laurence, A.; Yang, X.P.; Tato, C.M.; McGeachy, M.J.; Konkel, J.E.; Ramos, H.L.; Wei, L.; Davidson, T.S.; Bouladoux, N.; et al. Generation of pathogenic T(H)17 cells in the absence of TGF-beta signalling. Nature 2010, 467, 967–971. [Google Scholar] [CrossRef] [PubMed]
- McGeachy, M.J.; Chen, Y.; Tato, C.M.; Laurence, A.; Joyce-Shaikh, B.; Blumenschein, W.M.; McClanahan, T.K.; O’Shea, J.J.; Cua, D.J. The interleukin 23 receptor is essential for the terminal differentiation of interleukin 17–producing effector T helper cells in vivo. Nat. Immunol. 2009, 10, 314–324. [Google Scholar] [CrossRef]
- McGeachy, M.J.; Bak-Jensen, K.S.; Chen, Y.I.; Tato, C.M.; Blumenschein, W.; McClanahan, T.; Cua, D.J. TGF-beta and IL-6 drive the production of IL-17 and IL-10 by T cells and restrain T(H)-17 cell-mediated pathology. Nat. Immunol. 2007, 8, 1390–1397. [Google Scholar] [CrossRef]
- Stadhouders, R.; Lubberts, E.; Hendriks, R.W. A cellular and molecular view of T helper 17 cell plasticity in autoimmunity. J. Autoimmun. 2017, 87, 1–15. [Google Scholar] [CrossRef]
- Zielinski, C.E.; Mele, F.; Aschenbrenner, D.; Jarrossay, D.; Ronchi, F.; Gattorno, M.; Monticelli, S.; Lanzavecchia, A.; Sallusto, F. Pathogen-induced human TH17 cells produce IFN-gamma or IL-10 and are regulated by IL-1beta. Nature 2012, 484, 514–518. [Google Scholar] [CrossRef]
- Vogel, C.; Marcotte, E.M. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat. Rev. Microbiol. 2012, 13, 227–232. [Google Scholar] [CrossRef]
- Velez-Bermudez, I.; Schmidt, W. The conundrum of discordant protein and mRNA expression. Are plants special? Front. Plant Sci. 2014, 5, 619. [Google Scholar]
- De Sousa Abreu, R.; Penalva, L.O.; Marcotte, E.M.; Vogel, C. Global signatures of protein and mRNA expression levels. Mol. Biosyst. 2009, 5, 1512–1526. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arnone, J.T.; Robbins-Pianka, A.; Arace, J.R.; Kass-Gergi, S.; McAlear, M.A. The adjacent positioning of co-regulated gene pairs is widely conserved across eukaryotes. BMC Genom. 2012, 13, 546. [Google Scholar] [CrossRef] [PubMed]
- Michalak, P. Coexpression, coregulation, and cofunctionality of neighboring genes in eukaryotic genomes. Genomics 2008, 91, 243–248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Speeckaert, R.; Lambert, J.; Grine, L.; Van Gele, M.; De Schepper, S.; Van Geel, N. The many faces of interleukin-17 in inflammatory skin diseases. Br. J. Dermatol. 2016, 175, 892–901. [Google Scholar] [CrossRef] [PubMed]
- Smith, S.H.; Peredo, C.E.; Takeda, Y.; Bui, T.; Neil, J.; Rickard, D.; Millerman, E.; Therrien, J.P.; Nicodeme, E.; Brusq, J.M.; et al. Development of a topical treatment for psoriasis targeting RORgamma: From bench to skin. PLoS ONE 2016, 11, e0147979. [Google Scholar]
- Xu, H.; Fonseca, M.; Wolner, Z.; Chung, E.; Wu, X.; Geller, S.; Dusza, S.W.; DeRosa, A.P.; Marghoob, A.A.; Busam, K.J.; et al. Reference values for skin microanatomy: A systematic review and meta-analysis of ex-vivo studies. J. Am. Acad. Dermatol. 2017, 77, 1133–1144. [Google Scholar] [CrossRef]
- Sandby-Møller, J.; Poulsen, T.; Wulf, H.C. Epidermal thickness at different body sites: Relationship to age, gender, pigmentation, blood content, skin type and smoking habits. Acta Dermato Venereol. 2003, 83, 410–413. [Google Scholar] [CrossRef]
- Ramezanpour, M.; Moraitis, S.; Smith, J.L.P.; Wormald, P.J.; Vreugde, S. Th17 cytokines disrupt the airway mucosal barrier in chronic rhinosinusitis. Mediat. Inflamm. 2016, 2016, 9798206. [Google Scholar] [CrossRef]
- Cohen, S.; Barer, F.; Itzhak, I.; Silverman, M.H.; Fishman, P. Inhibition of IL-17 and IL-23 in human keratinocytes by the A3 adenosine receptor agonist piclidenoson. J. Immunol. Res. 2018, 2018, 2310970. [Google Scholar] [CrossRef]
- Cua, D.J.; Tato, C.M. Innate IL-17-producing cells: The sentinels of the immune system. Nat. Rev. Immunol. 2010, 10, 479–489. [Google Scholar] [CrossRef] [PubMed]
- Li, T.J.; Jiang, Y.M.; Hu, Y.F.; Huang, L.; Yu, J.; Zhao, L.Y.; Deng, H.J.; Mou, T.Y.; Liu, H.; Yang, Y.; et al. Interleukin-17-producing neutrophils link inflammatory stimuli to disease progression by promoting angiogenesis in gastric cancer. Clin. Cancer Res. 2017, 23, 1575–1585. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Pan, H.F.; Ye, D.Q. Tc17 cells in immunity and systemic autoimmunity. Int. Rev. Immunol. 2015, 34, 318–331. [Google Scholar] [CrossRef] [PubMed]
- Matsuzaki, G.; Umemura, M. Interleukin-17 family cytokines in protective immunity against infections: Role of hematopoietic cell-derived and non-hematopoietic cell-derived interleukin-17s. Microbiol. Immunol. 2018, 62, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Parks, O.B.; Pociask, D.A.; Hodzic, Z.; Kolls, J.K.; Good, M. Interleukin-22 signaling in the regulation of intestinal health and disease. Front. Cell Dev. Biol. 2015, 3, 85. [Google Scholar] [CrossRef] [PubMed]
- Truchetet, M.E.; Brembilla, N.C.; Montanari, E.; Lonati, P.; Raschi, E.; Zeni, S.; Fontao, L.; Meroni, P.L.; Chizzolini, C. Interleukin-17A+ cell counts are increased in systemic sclerosis skin and their number is inversely correlated with the extent of skin involvement. Arthritis Rheum. 2013, 65, 1347–1356. [Google Scholar] [CrossRef] [PubMed]
| Abdominal | Breast | Glutes | Arms | Multiple Sources |
|---|---|---|---|---|
| 12/24 (50%) | 8/24 (33.3%) | 1/24 (4.2%) | 1/24 (4.2%) | 2/24 (8.3%) |
| Component (Final Concentration) | Cocktail 4 | Cocktail 8 | Cocktail 9 |
|---|---|---|---|
| αCD3 (1 μg/mL) | √ | √ | √ |
| αCD28 (1 μg/mL) | √ | √ | √ |
| IL-23 (10 ng/mL) | √ | √ | √ |
| IL-1β (10 ng/mL) | √ | √ | |
| IFNγ (1 μg/mL) | √ | √ | |
| IL-4 (1 μg/mL) | √ | √ | |
| IL-6 (10 ng/mL) | √ | √ | |
| IL-21 (10 ng/mL) | √ | √ | |
| TGFβ (1 ng/mL) | √ | √ |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Garrett, S.M.; Zhao, Q.; Feghali-Bostwick, C. Induction of a Th17 Phenotype in Human Skin—A Mimic of Dermal Inflammatory Diseases. Methods Protoc. 2019, 2, 45. https://doi.org/10.3390/mps2020045
Garrett SM, Zhao Q, Feghali-Bostwick C. Induction of a Th17 Phenotype in Human Skin—A Mimic of Dermal Inflammatory Diseases. Methods and Protocols. 2019; 2(2):45. https://doi.org/10.3390/mps2020045
Chicago/Turabian StyleGarrett, Sara M., Qihong Zhao, and Carol Feghali-Bostwick. 2019. "Induction of a Th17 Phenotype in Human Skin—A Mimic of Dermal Inflammatory Diseases" Methods and Protocols 2, no. 2: 45. https://doi.org/10.3390/mps2020045
APA StyleGarrett, S. M., Zhao, Q., & Feghali-Bostwick, C. (2019). Induction of a Th17 Phenotype in Human Skin—A Mimic of Dermal Inflammatory Diseases. Methods and Protocols, 2(2), 45. https://doi.org/10.3390/mps2020045






