Novel Applications of Lead Acetate and Flow Cytometry Methods for Detection of Sulfur-Containing Molecules
Abstract
1. Introduction
2. Experimental Design
2.1. Detection of Released H2S Using Agar Trap or Lead Acetate Strip Test (LAST)
2.2. Measuring of Sulfane Sulfur Species Using SSP4 and Flow Cytometry in Comparison to Fluorescence Microscopy
2.3. Materials
- Agar (AppliChem, Darmstadt, Germany; Cat. no.: A0949, 1000)
- Aminooxyacetic acid hemihydrochloride, AOAA (Aldrich, St.Louis, MO, USA; Cat. no.: C13408)
- Cetyltrimethylammonium bromide, CTAB (Sigma, St.Louis, MO, USA; Cat. no.: H9151)
- Diallyl trisulfide, DATS (Cayman Chemical, Ann Arbor, MI, USA; Cat. no.: 2050-87-5)
- Dimethyl sulfoxide, DMSO (Sigma-Aldrich, St.Louis, MO, USA; Cat. no.: W387520)
- DL-cysteine, Cys (Aldrich, St.Louis, MO, USA; Cat. no.: 861677)
- DL-propargylglycine, PAG (Sigma, St.Louis, MO, USA; Cat. no.: P7888)
- Dulbecco’s Modified Eagle’s Medium, DMEM (Sigma, St.Louis, MO, USA; Cat. no.: D4947)
- Dulbecco’s Phosphate-Buffered Saline, PBS (Microtech Srl, Pozzuoli, Italy; Cat. no.: L0615-500)
- Fetal Bovine Serum (South America), FBS (Microtech Srl, Pozzuoli, Italy; Cat. no.: S1810-500)
- Human Endothelial Somatic Cell Hybrid, EA.hy926 (American Type Culture Collection, Manassas, VA, USA; Cat. no.: CRL-2922TM)
- Hydrochloric Acid, HCl (Sigma, St.Louis, MO, USA; Cat. no.: H1758)
- Iron (III) chloride, FeCl3 (Sigma-Aldrich, St.Louis, MO, USA; Cat. no.: 157740)
- Lead acetate strips test (Sigma-Aldrich, St.Louis, MO, USA; Cat. no.: 06728)
- L-glutamine (Microtech Srl, Pozzuoli, Italy; Cat. no.: X0550-100)
- L-glutathione reduced, GSH (Sigma, St.Louis, MO, USA; Cat. no.: G6013)
- N,N-dimethyl-p-phenylenediamine sulfate salt, NDPS (Aldrich, St.Louis, MO, USA; Cat. no.: 186384)
- Penicillin–streptomycin (Microtech Srl, Pozzuoli, Italy; Cat. no.: L0022-100)
- Pyridoxine hydrochloride, B6 (Sigma, St.Louis, MO, USA; Cat. no.: P6280)
- S-adenosyl-methionine, SAM (New England Biolabs Inc, Ipswich, MA, USA; Cat. no.: B9003S)
- Sodium hydrosulfide hydrate, NaHS (Sigma-Aldrich, St.Louis, MO, USA; Cat. no.: 161527)
- Sodium hydroxide pellets, NaOH (Carlo Erba, Rodano, Italy, Cat. no.: 480507)
- Sulfane Sulfur Probe 4, SSP4 (Dojindo Laboratories, Kumamoto, Japan; Cat. no.: SB10)
- Trypsin-EDTA (Microtech Srl, Pozzuoli, Italy; Cat. no.: L0940-100)
- Zinc acetate dihydrate, ZnAc (Sigma, St.Louis, MO, USA; Cat. no.: 383058-500G)
2.4. Equipment
- Multimode Microplate Reader “Tecan Infinite” (Tecan Group Ltd, Männedorf, Switzerland; Cat. no.: M200)
- Confocal Laser Scanning Microscope (Zeiss, Milan, Italy; Cat. no.: LSM 710)
- Flow cytometer Accuri™ C6 Plus (BD Biosciences, San Jose, CA, USA)
- UV/VIS Spectrophotometer Lambda 25 (Perkin Elmer, Waltham, MA, USA; Cat. no.: L6020060)
- Multifunctional InkJet printer (Brother Industries, Ltd, Nagoya, Japan, Cat. no.: DCP-197C)
- Water Jacketed Incubator Thermo Forma 3851 (Thermo Fisher Scientific, Waltham, MA, USA)
- Incubator Heraeus B6 (Kendro, Langenselbold, Germany, Cat. no.: 63505)
- Laboratory sterile hood (STERIL spa, Milan, Italy, Cat. no.: VBH 48 MP/99)
- Centrifuge (Eppendorf, Hamburg, Germany, Cat. no.: 5810 R)
3. Procedure
- 5 mM Cys, 5 mM B6, 10 μM SAM;
- 2 mM GSH;
- 150 μM DATS; and
- 2 mM PAG, 2 mM AOAA.
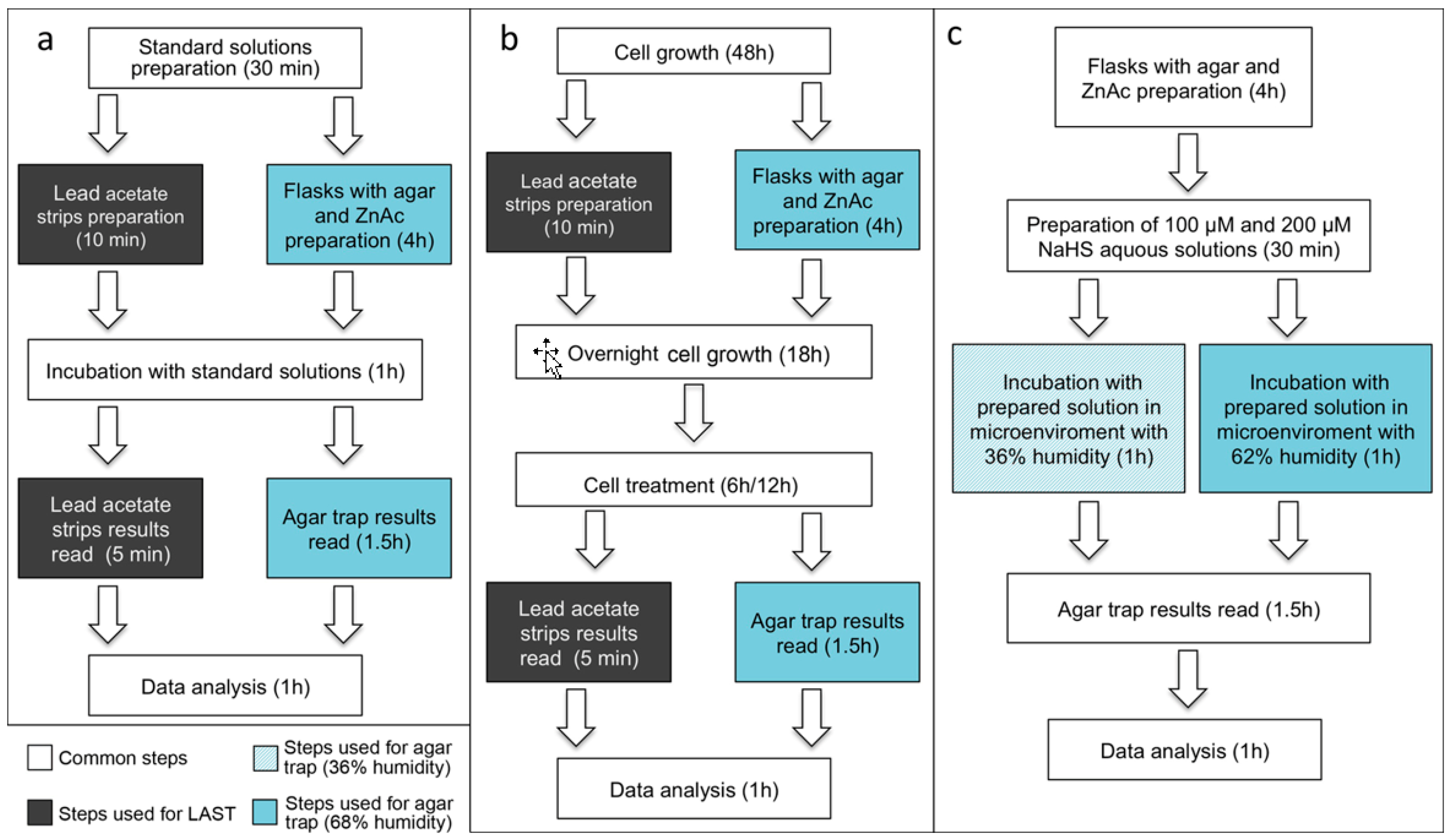
3.1. LAST in Comparison to Agar Trap Method. Time for Completion: Four Days
3.1.1. Agar Trap Method
3.1.2. LAST
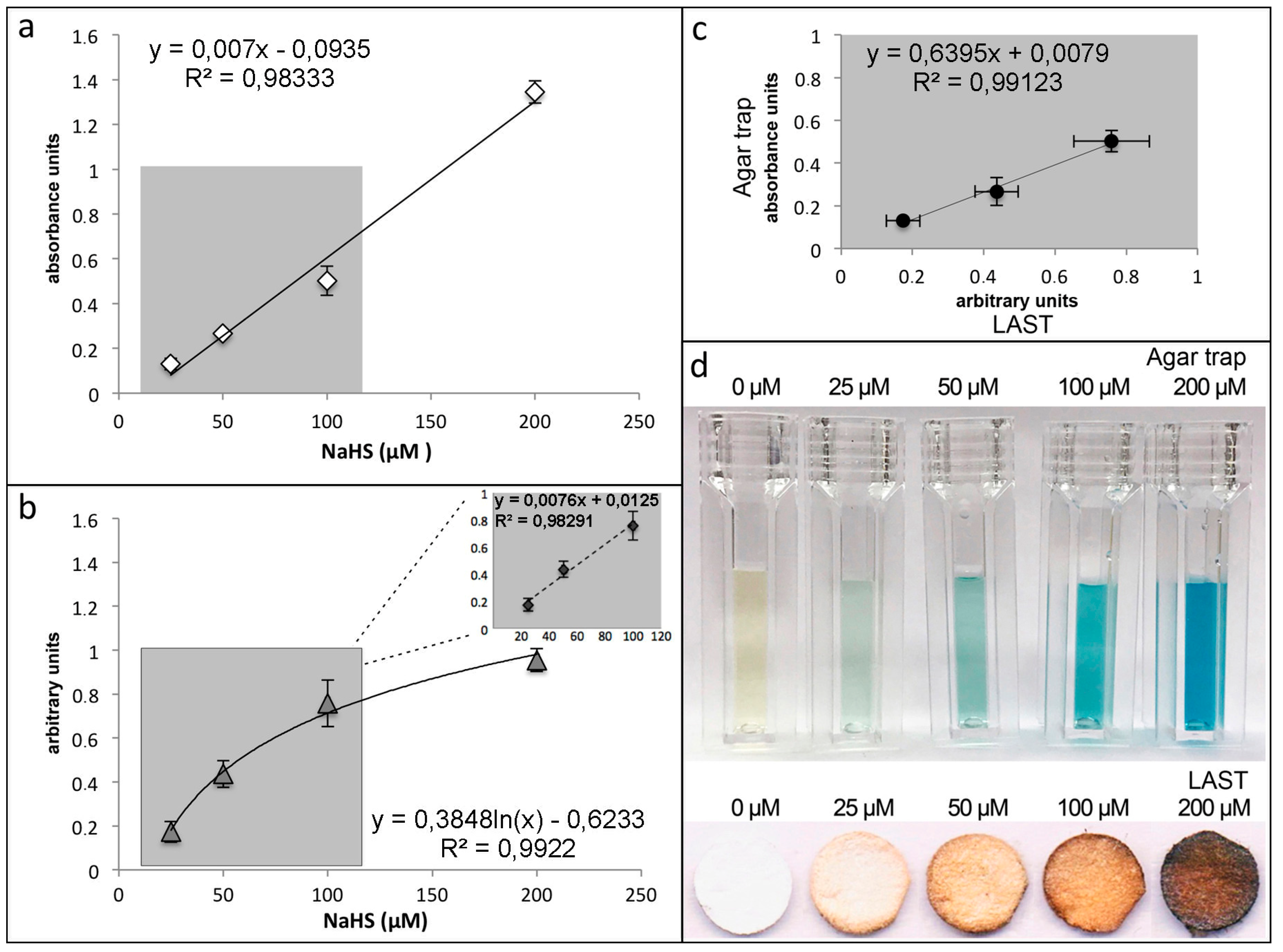
Standard Curve. Time for Completion: 02:45 Hours
- Cut lead acetate strips using a paper perforator;
- Place test strips into the internal side of each flask lid, using a double-sided adhesive tape.
- Sterilize flask lids using UV light.
PAUSE STEP Sterile lids can be stored at 4 °C until use for weeks.
- For standard curve, add 3 mL aqueous solution containing 25, 50, 100, and 200 μM NaHS to flasks and tightly close with the strip containing lids, following incubation for 1 h at 37 °C.
- At the end of the experiment, place test strips on Whatman paper sheets, and scan using a multifunctional printer.
CRITICAL STEP Scanning should be performed within 24 h after experiment, since, within 3–5 days after experiment, the color can fade away.
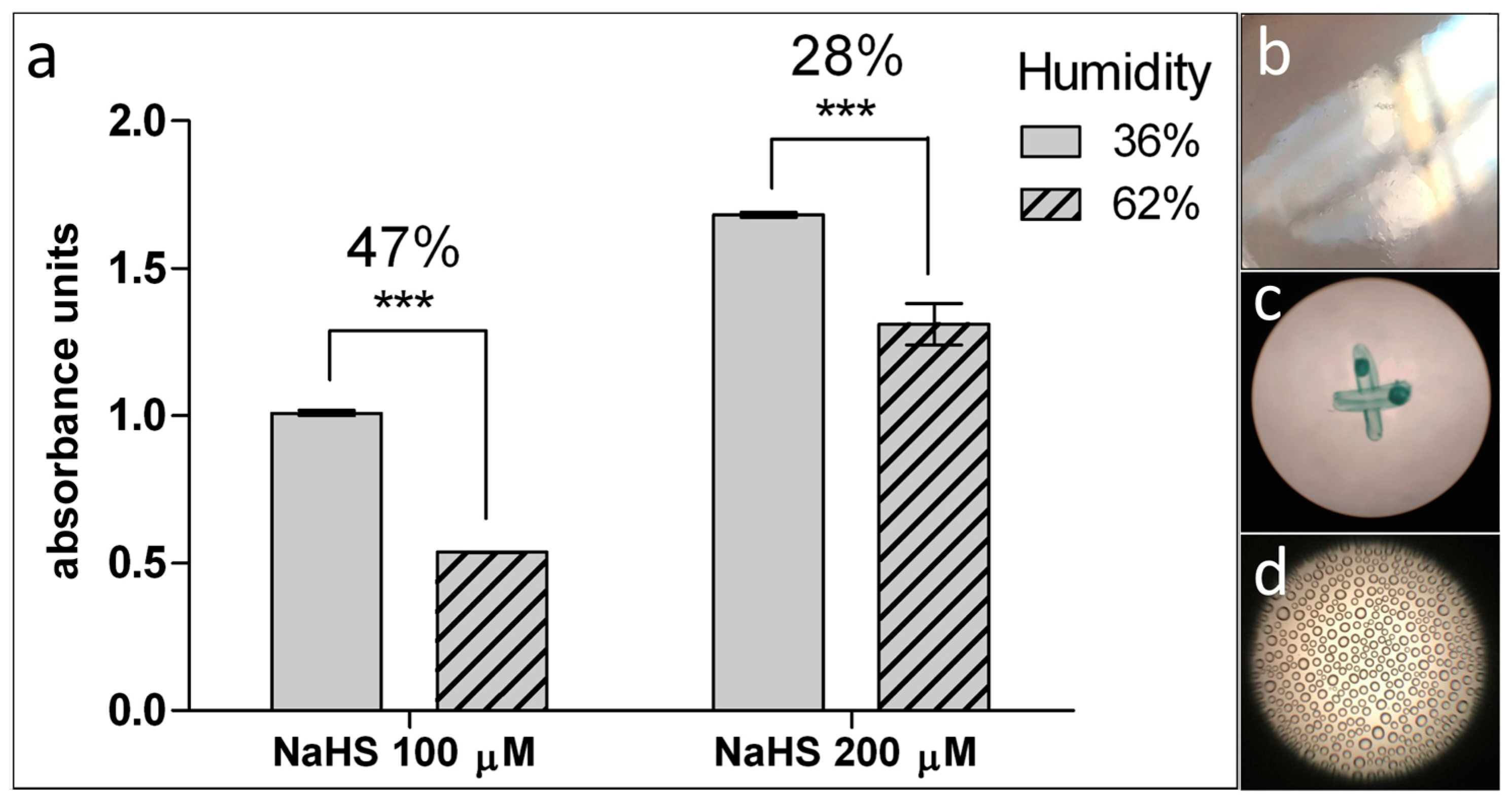
- Image analysis is accomplished using ImageJ software (National Institutes of Health, Bethesda, MD, USA): Analyze > Measure and Analyze > Histogram. Set fixed reading slit for each series of assay measurements (see Figure 3b,d).
LAST in Cell Culture. Time for Completion: Four Days
- Treat cell culture in flasks as described above and tight lids containing test strips. Incubate for 6 h and/or 12 h in 5% CO2 atmosphere.
- At the end of experiment, repeat Steps 5 and 6 as in the previous paragraph.
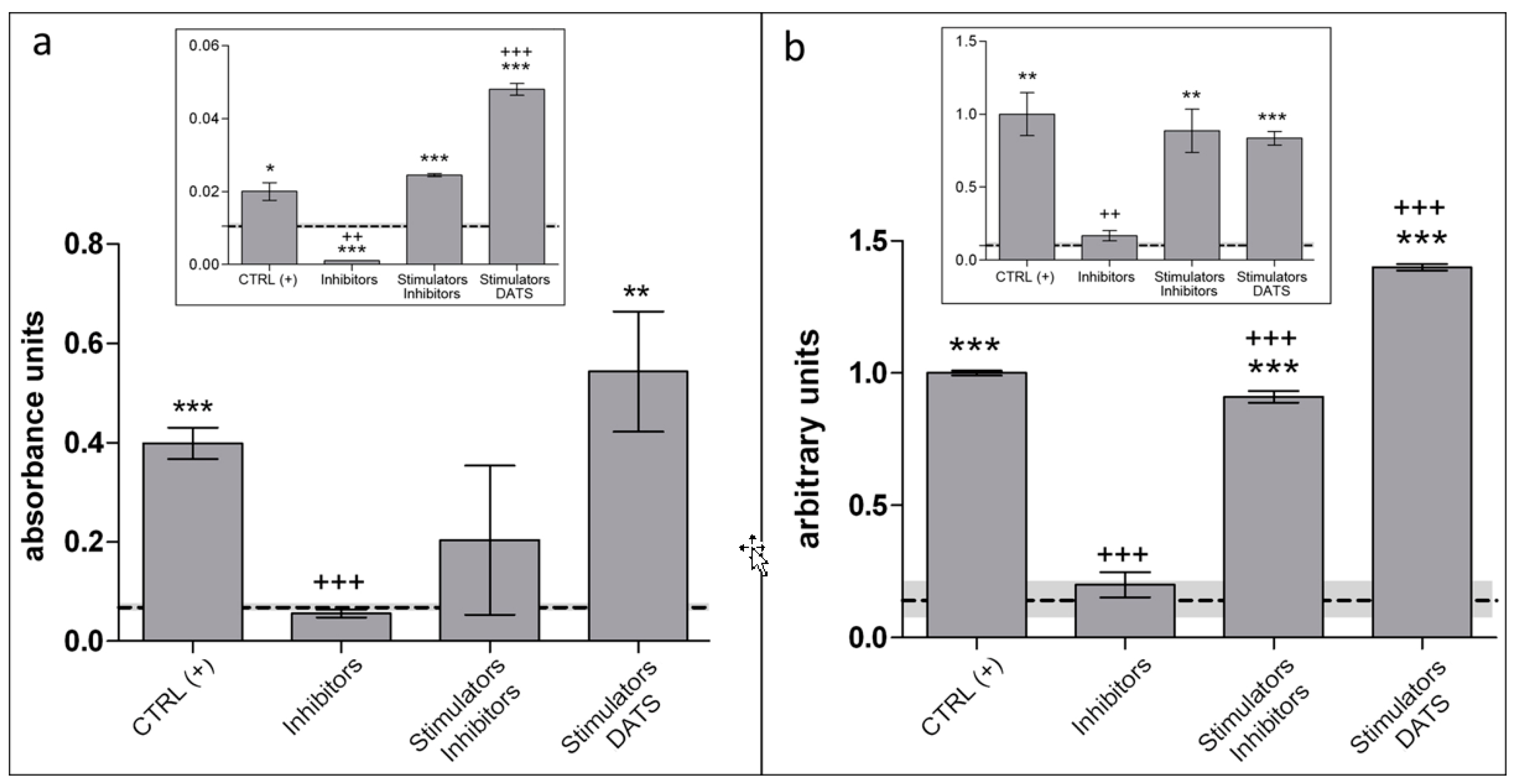
- Data are analyzed with ImageJ software, using GraphPad Prism 5 for statistics (GraphPad Software, La Jolla, CA, USA) and applying Student’s two-tailed t-test (Figure 4b).
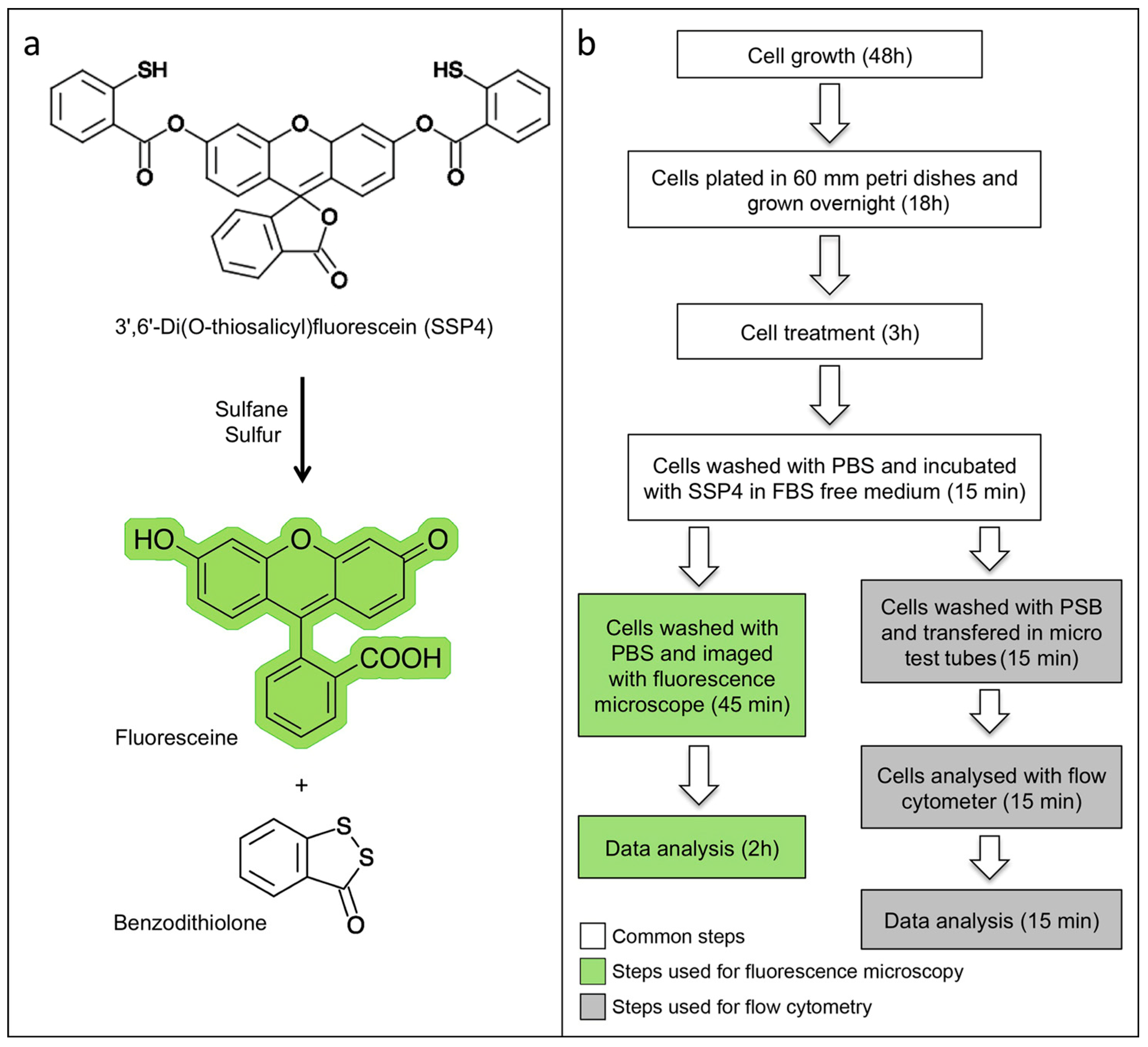
3.2. Sulfane Sulfur Quantitation with SSP4 Using Flow Cytometry in Comparison to Fluorescence Microscopy. Time for Completion: 00:45 Hours
3.2.1. Sulfane Sulfur Quantitation with SSP4 Using Fluorescence Microscopy
3.2.2. Sulfane Sulfur Quantitation Using SSP4 Labelig and Detection by Flow Cytometry
- Treatment duration is 3 h, then wash the cells twice with PBS and leave for 15 min in 2 mL of FBS-free DMEM with 1.5 μM SSP4 and 150 μM CTAB.
- Wash twice again with PBS and leave for 3 min in the presence of 0.5 mL Trypsin-EDTA at 37 °C.
- Transfer detached cell samples in 1.5 mL micro test tubes and wash once again with PBS.
CRITICAL STEP The subsequent analysis should be performed within 5–15 min.
- Detect SSP4-emitted fluorescence using a flow cytometer (Accuri™ C6 Plus).
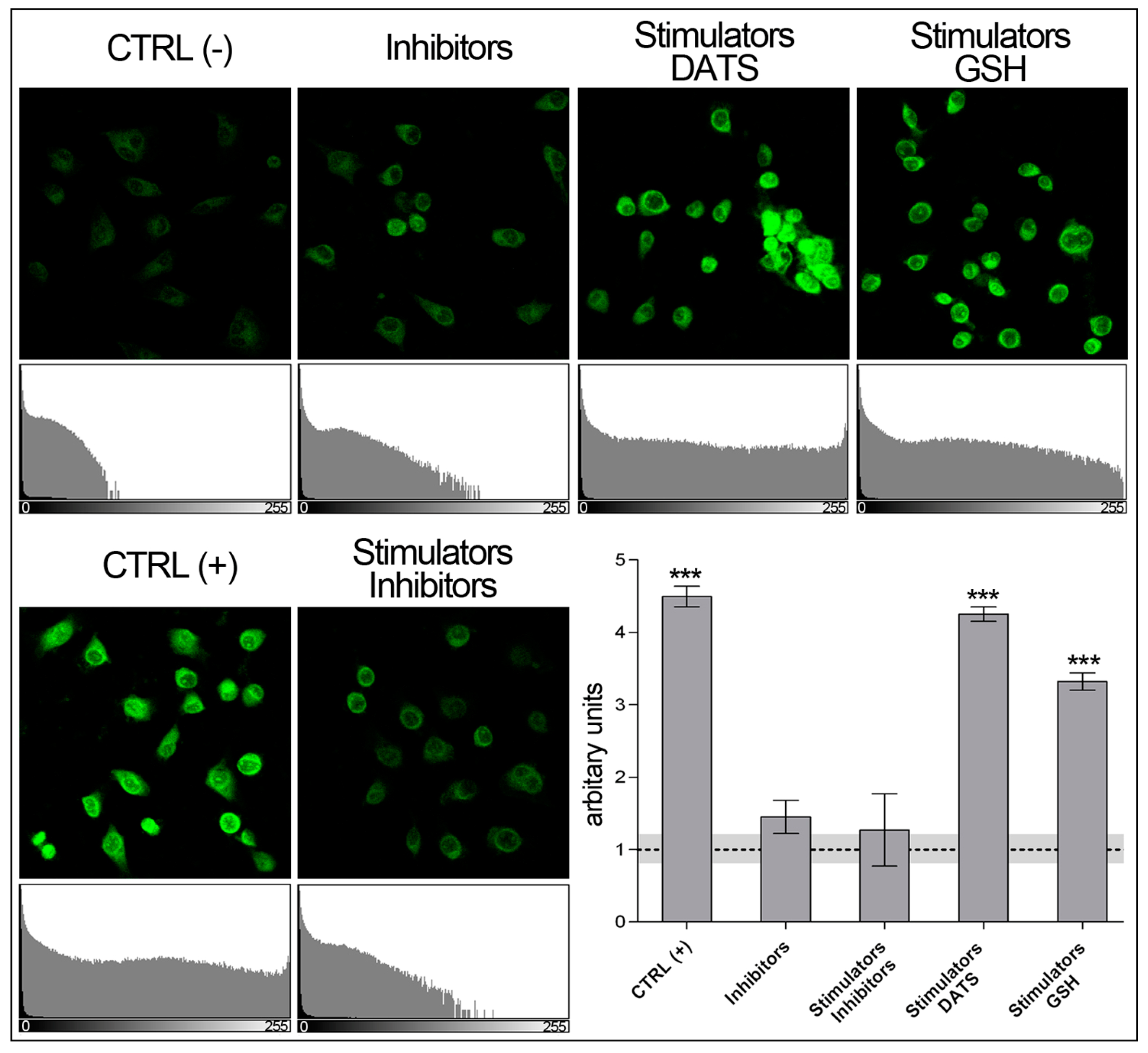
- Analyze data using Accuri™ C6 Plus and GraphPad Prism 5 software (Figure 7).
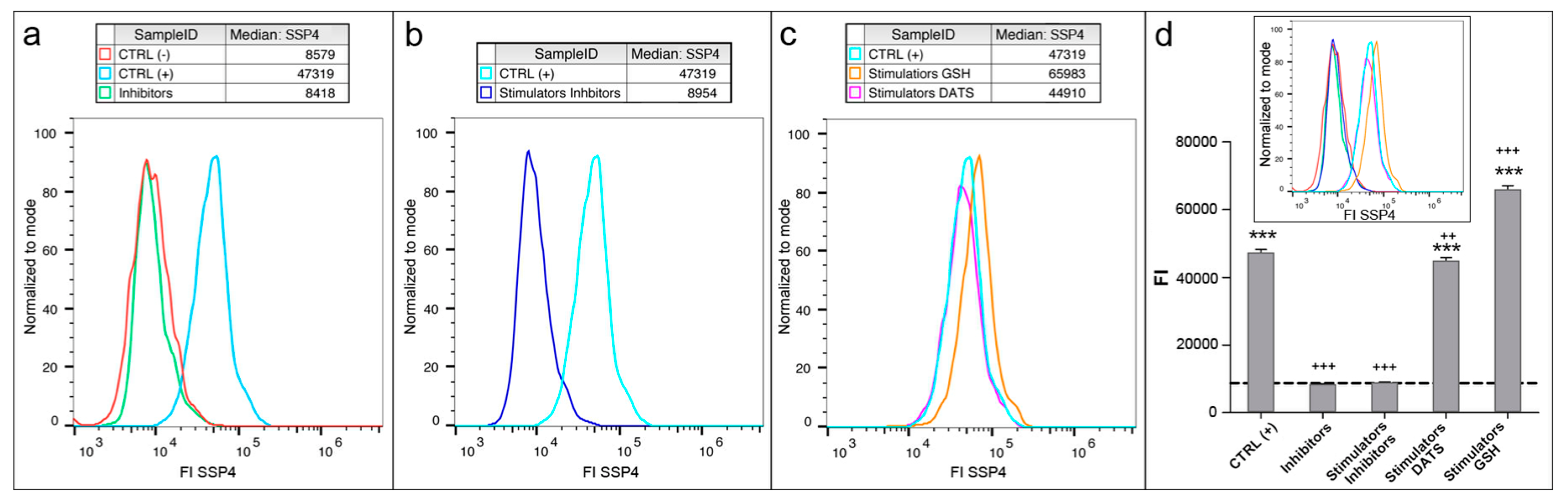
4. Expected Results
- (a)
- The data obtained with flow cytometer are much more precise: the quantity of reads is identical for each sample (3000 reads), resulting in a negligible SD, indeed.
- (b)
- The quantity of sulfane sulfurs in cells treated with GSH plus stimulators is over 20% higher than that detected in stimulated controls, CTRL (+), while no significant difference can be detected by microscopy.
Author Contributions
Funding
Conflicts of Interest
References
- Perna, A.F.; Glorieux, G.; Zacchia, M.; Trepiccione, F.; Capolongo, G.; Vigorito, C.; Anishchenko, E.; Ingrosso, D. The role of the intestinal microbiota in uremic solute accumulation: A focus on sulfur compounds. JNEP 2019, in press. [Google Scholar] [CrossRef] [PubMed]
- Perna, A.; Anishchenko, E.; Vigorito, C.; Zacchia, M.; Trepiccione, F.; D’Aniello, S.; Ingrosso, D. Zebrafish, a Novel Model System to Study Uremic Toxins: The Case for the Sulfur Amino Acid Lanthionine. Int. J. Mol. Sci. 2018, 19, 1323. [Google Scholar] [CrossRef] [PubMed]
- Perna, A.F.; Di Nunzio, A.; Amoresano, A.; Pane, F.; Fontanarosa, C.; Pucci, P.; Vigorito, C.; Cirillo, G.; Zacchia, M.; Trepiccione, F.; et al. Divergent behavior of hydrogen sulfide pools and of the sulfur metabolite lanthionine, a novel uremic toxin, in dialysis patients. Biochimie 2016, 126, 97–107. [Google Scholar] [CrossRef] [PubMed]
- Goodwin, L.R.; Francom, D.; Dieken, F.P.; Taylor, J.D.; Warenycia, M.W.; Reiffenstein, R.J.; Dowling, G. Determination of sulfide in brain tissue by gas dialysis/ion chromatography: Postmortem studies and two case reports. J. Anal. Toxicol. 1989, 13, 105–109. [Google Scholar] [CrossRef] [PubMed]
- Warenycia, M.W.; Goodwin, L.R.; Benishin, C.G.; Reiffenstein, R.J.; Francom, D.M.; Taylor, J.D.; Dieken, F.P. Acute hydrogen sulfide poisoning. Demonstration of selective uptake of sulfide by the brainstem by measurement of brain sulfide levels. Biochem. Pharmacol. 1989, 38, 973–981. [Google Scholar] [CrossRef]
- Qu, K.; Lee, S.W.; Bian, J.S.; Low, C.-M.; Wong, P.T.-H. Hydrogen sulfide: Neurochemistry and neurobiology. Neurochem. Int. 2008, 52, 155–165. [Google Scholar] [CrossRef]
- Szabó, C. Hydrogen sulphide and its therapeutic potential. Nat. Rev. Drug Discov. 2007, 6, 917–935. [Google Scholar] [CrossRef]
- Yang, G.; Wu, L.; Jiang, B.; Yang, W.; Qi, J.; Cao, K.; Meng, Q.; Mustafa, A.K.; Mu, W.; Zhang, S.; et al. H2S as a Physiologic Vasorelaxant: Hypertension in Mice with Deletion of Cystathionine-Lyase. Science 2008, 322, 587–590. [Google Scholar] [CrossRef]
- Vandiver, M.S.; Snyder, S.H. Hydrogen sulfide: A gasotransmitter of clinical relevance. J. Mol. Med. 2012, 90, 255–263. [Google Scholar] [CrossRef]
- Wang, X.-H.; Wang, F.; You, S.-J.; Cao, Y.-J.; Cao, L.-D.; Han, Q.; Liu, C.-F.; Hu, L.-F. Dysregulation of cystathionine γ-lyase (CSE)/hydrogen sulfide pathway contributes to ox-LDL-induced inflammation in macrophage. Cell. Signal. 2013, 25, 2255–2262. [Google Scholar] [CrossRef]
- Zhang, H.; Guo, C.; Wu, D.; Zhang, A.; Gu, T.; Wang, L.; Wang, C. Hydrogen sulfide inhibits the development of atherosclerosis with suppressing CX3CR1 and CX3CL1 expression. PLoS ONE 2012, 7, e41147. [Google Scholar] [CrossRef] [PubMed]
- Qian, L.-L.; Liu, X.-Y.; Chai, Q.; Wang, R.-X. Hydrogen Sulfide in Diabetic Complications: Focus on Molecular Mechanisms. Endocr. Metab. Immune Disord. Drug Targets 2018, 18, 470–476. [Google Scholar] [CrossRef] [PubMed]
- Szijártó, I.A.; Markó, L.; Filipovic, M.R.; Miljkovic, J.L.; Tabeling, C.; Tsvetkov, D.; Wang, N.; Rabelo, L.A.; Witzenrath, M.; Diedrich, A.; et al. Cystathionine γ-Lyase-Produced Hydrogen Sulfide Controls Endothelial NO Bioavailability and Blood Pressure. Hypertension 2018, 71, 1210–1217. [Google Scholar] [CrossRef]
- Bibli, S.-I.; Luck, B.; Zukunft, S.; Wittig, J.; Chen, W.; Xian, M.; Papapetropoulos, A.; Hu, J.; Fleming, I. A selective and sensitive method for quantification of endogenous polysulfide production in biological samples. Redox Biol. 2018, 18, 295–304. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Zhang, F.; Munske, G.; Zhang, H.; Xian, M. Isotope dilution mass spectrometry for the quantification of sulfane sulfurs. Free Radic. Biol. Med. 2014, 76, 200–207. [Google Scholar] [CrossRef]
- Kabil, O.; Vitvitsky, V.; Xie, P.; Banerjee, R. The quantitative significance of the transsulfuration enzymes for H2S production in murine tissues. Antioxid. Redox Signal. 2011, 15, 363–372. [Google Scholar] [CrossRef]
- Kartha, R.V.; Zhou, J.; Hovde, L.B.; Cheung, B.W.Y.; Schröder, H. Enhanced detection of hydrogen sulfide generated in cell culture using an agar trap method. Anal. Biochem. 2012, 423, 102–108. [Google Scholar] [CrossRef]
- Jarosz, A.P.; Yep, T.; Mutus, B. Microplate-based colorimetric detection of free hydrogen sulfide. Anal. Chem. 2013, 85, 3638–3643. [Google Scholar] [CrossRef]
- Yoshida, A.; Yoshimura, M.; Ohara, N.; Yoshimura, S.; Nagashima, S.; Takehara, T.; Nakayama, K. Hydrogen sulfide production from cysteine and homocysteine by periodontal and oral bacteria. J. Periodontol. 2009, 80, 1845–1851. [Google Scholar] [CrossRef]
- Velázquez-Moyado, J.A.; Navarrete, A. The detection and quantification, in vivo and in real time, of hydrogen sulfide in ethanol-induced lesions in rat stomachs using an ion sensitive electrode. J. Pharmacol. Toxicol. Methods 2018, 89, 54–58. [Google Scholar] [CrossRef]
- Nagy, P.; Pálinkás, Z.; Nagy, A.; Budai, B.; Tóth, I.; Vasas, A. Chemical aspects of hydrogen sulfide measurements in physiological samples. Biochim. Biophys. Acta 2014, 1840, 876–891. [Google Scholar] [CrossRef] [PubMed]
- Whiteman, M.; Le Trionnaire, S.; Chopra, M.; Fox, B.; Whatmore, J. Emerging role of hydrogen sulfide in health and disease: Critical appraisal of biomarkers and pharmacological tools. Clin. Sci. 2011, 121, 459–488. [Google Scholar] [CrossRef]
- Hydrogen Sulfide Test Strips 06728. Available online: https://www.sigmaaldrich.com/catalog/product/sial/06728 (accessed on 1 November 2018).
- Lead Acetate Test Paper—Precision Laboratories. Available online: https://preclaboratories.com/product/lead-acetate-test-paper/ (accessed on 1 November 2018).
- Sulfite Test Analytical Strips Vial 10-500 PPM. Available online: https://www.bartovation.com/water-quality/sulfite-test-analytical-strips-vial-10-500-ppm (accessed on 1 November 2018).
- Han, L.; Shi, R.; Xin, C.; Ci, Q.; Ge, J.; Liu, J.; Wu, Q.; Zhang, C.; Li, L.; Huang, W. Mitochondrial Specific H2Sn Fluorogenic Probe for Live Cell Imaging by Rational Utilization of a Dual-Functional-Photocage Group. ACS Sens. 2018, 3, 1622–1626. [Google Scholar] [CrossRef] [PubMed]
- Gao, M.; Wang, R.; Yu, F.; Chen, L. Evaluation of sulfane sulfur bioeffects via a mitochondria-targeting selenium-containing near-infrared fluorescent probe. Biomaterials 2018, 160, 1–14. [Google Scholar] [CrossRef]
- Han, X.; Song, X.; Li, B.; Yu, F.; Chen, L. A near-infrared fluorescent probe for sensitive detection and imaging of sulfane sulfur in living cells and in vivo. Biomater. Sci. 2018, 6, 672–682. [Google Scholar] [CrossRef] [PubMed]
- Takano, M.; Echizen, M.; Hanaoka, K. Fluorescent probes and selective inhibitors for biological studies of H2S- and polysulfide-mediated signaling. Antioxid. Redox Signal. 2017, 27, 669–683. [Google Scholar] [CrossRef]
- Chen, W.; Rosser, E.; Matsunaga, T.; Pacheco, A.; Akaike, T.; Xian, M. The Development of Fluorescent Probes for Visualizing Intracellular Hydrogen Polysulfides. Angew. Chem. Int. Ed. 2015, 127, 14167–14171. [Google Scholar] [CrossRef]
- Yu, F.; Gao, M.; Li, M.; Chen, L. A dual response near-infrared fluorescent probe for hydrogen polysulfides and superoxide anion detection in cells and in vivo. Biomaterials 2015, 63, 93–101. [Google Scholar] [CrossRef]
- Huang, Y.; Yu, F.; Wang, J.; Chen, L. Near-Infrared Fluorescence Probe for in Situ Detection of Superoxide Anion and Hydrogen Polysulfides in Mitochondrial Oxidative Stress. Anal. Chem. 2016, 88, 4122–4129. [Google Scholar] [CrossRef]
- Umezawa, K.; Kamiya, M.; Urano, Y. A Reversible Fluorescent Probe for Real-Time Live-Cell Imaging and Quantification of Endogenous Hydropolysulfides. Angew. Chem. Int. Ed. 2018, 57, 9346–9350. [Google Scholar] [CrossRef]
- Du, Z.; Song, B.; Zhang, W.; Duan, C.; Wang, Y.-L.; Liu, C.; Zhang, R.; Yuan, J. Quantitative Monitoring and Visualization of Hydrogen Sulfide in Vivo Using a Luminescent Probe Based on a Ruthenium(II) Complex. Angew. Chem. Int. Ed. Engl. 2018, 57, 3999–4004. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Liu, C.; Peng, B.; Zhao, Y.; Pacheco, A.; Xian, M. New fluorescent probes for sulfane sulfurs and the application in bioimaging. Chem. Sci. 2013, 4, 2892–2896. [Google Scholar] [CrossRef] [PubMed]
- Takano, Y.; Hanaoka, K.; Shimamoto, K.; Miyamoto, R.; Komatsu, T.; Ueno, T.; Terai, T.; Kimura, H.; Nagano, T.; Urano, Y. Development of a reversible fluorescent probe for reactive sulfur species, sulfane sulfur, and its biological application. Chem. Commun. 2017, 53, 1064–1067. [Google Scholar]
- Chiku, T.; Padovani, D.; Zhu, W.; Singh, S.; Vitvitsky, V.; Banerjee, R. H2S biogenesis by human cystathionine gamma-lyase leads to the novel sulfur metabolites lanthionine and homolanthionine and is responsive to the grade of hyperhomocysteinemia. J. Biol. Chem. 2009, 284, 11601–11612. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Padovani, D.; Leslie, R.A.; Chiku, T.; Banerjee, R. Relative contributions of cystathionine beta-synthase and gamma-cystathionase to H2S biogenesis via alternative trans-sulfuration reactions. J. Biol. Chem. 2009, 284, 22457–22466. [Google Scholar] [CrossRef]
- DeLeon, E.R.; Gao, Y.; Huang, E.; Olson, K.R. Garlic oil polysulfides: H2S- and O2-independent prooxidants in buffer and antioxidants in cells. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2016, 310, R1212–R1225. [Google Scholar] [CrossRef] [PubMed]
- Martelli, A.; Testai, L.; Citi, V.; Marino, A.; Pugliesi, I.; Barresi, E.; Nesi, G.; Rapposelli, S.; Taliani, S.; Settimo, F.; et al. Arylthioamides as H2S Donors:l-Cysteine-Activated Releasing Properties and Vascular Effects in Vitro and in Vivo. ACS Med. Chem. Lett. 2013, 4, 904–908. [Google Scholar] [CrossRef] [PubMed]
- Barresi, E.; Nesi, G.; Citi, V.; Piragine, E.; Piano, I.; Taliani, S.; Settimo, F.; Rapposelli, S.; Testai, L.; Breschi, M.; et al. Iminothioethers as Hydrogen Sulfide Donors: From the Gasotransmitter Release to the Vascular Effects. J. Med. Chem. 2017, 60, 7512–7523. [Google Scholar] [CrossRef] [PubMed]
- Ida, T.; Sawa, T.; Ihara, H.; Tsuchiya, Y.; Watanabe, Y.; Kumagai, Y.; Suematsu, M.; Motohashi, H.; Fujii, S.; Matsunaga, T.; et al. Reactive cysteine persulfides and S-polythiolation regulate oxidative stress and redox signaling. Proc. Natl. Acad. Sci. USA 2014, 111, 7606–7611. [Google Scholar] [CrossRef] [PubMed]
- Basic, A.; Blomqvist, S.; Carlén, A.; Dahlén, G. Estimation of bacterial hydrogen sulfide production in vitro. J. Oral Microbiol. 2015, 7, 28166. [Google Scholar] [CrossRef]
- SulfoBiotics-SSP4|Dojindo. Available online: https://www.dojindo.eu.com/store/p/864-SulfoBiotics-SSP4.aspx (accessed on 7 November 2018).
- Sitdikova, G.F.; Fuchs, R.; Kainz, V.; Weiger, T.M.; Hermann, A. Phosphorylation of BK channels modulates the sensitivity to hydrogen sulfide (H2S). Front. Physiol. 2014, 5, 431. [Google Scholar] [CrossRef]
- Sakaguchi, M.; Marutani, E.; Shin, H.-S.; Chen, W.; Hanaoka, K.; Xian, M.; Ichinose, F. Sodium thiosulfate attenuates acute lung injury in mice. Anesthesiology 2014, 121, 1248–1257. [Google Scholar] [CrossRef]
- Olson, K.R.; Gao, Y.; DeLeon, E.R.; Arif, M.; Arif, F.; Arora, N.; Straub, K.D. Catalase as a sulfide-sulfur oxido-reductase: An ancient (and modern?) regulator of reactive sulfur species (RSS). Redox Biol. 2017, 12, 325–339. [Google Scholar] [CrossRef]
- Olson, K.R.; Gao, Y.; Arif, F.; Arora, K.; Patel, S.; DeLeon, E.; Straub, K.D. Fluorescence quenching by metal centered porphyrins and poryphyrin enzymes. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2017, 313, R340–R346. [Google Scholar] [CrossRef] [PubMed]
- Olson, K.R.; Gao, Y.; Arif, F.; Arora, K.; Patel, S.; DeLeon, E.R.; Sutton, T.R.; Feelisch, M.; Cortese-Krott, M.M.; Straub, K.D. Metabolism of hydrogen sulfide (HS) and Production of Reactive Sulfur Species (RSS) by superoxide dismutase. Redox Biol. 2018, 15, 74–85. [Google Scholar] [CrossRef] [PubMed]
- Marutani, E.; Sakaguchi, M.; Chen, W.; Sasakura, K.; Liu, J.; Xian, M.; Hanaoka, K.; Nagano, T.; Ichinose, F. Cytoprotective effects of hydrogen sulfide-releasing-methyl-d-aspartate receptor antagonists are mediated by intracellular sulfane sulfur. Medchemcomm 2014, 5, 1577–1583. [Google Scholar] [CrossRef]
- Pitchaimani, A.; Nguyen, T.D.T.; Koirala, M.; Zhang, Y.; Aryal, S. Impact of cell adhesion and migration on nanoparticle uptake and cellular toxicity. Toxicol. In Vitro 2017, 43, 29–39. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Anishchenko, E.; Vigorito, C.; Mele, L.; Lombari, P.; Perna, A.F.; Ingrosso, D. Novel Applications of Lead Acetate and Flow Cytometry Methods for Detection of Sulfur-Containing Molecules. Methods Protoc. 2019, 2, 13. https://doi.org/10.3390/mps2010013
Anishchenko E, Vigorito C, Mele L, Lombari P, Perna AF, Ingrosso D. Novel Applications of Lead Acetate and Flow Cytometry Methods for Detection of Sulfur-Containing Molecules. Methods and Protocols. 2019; 2(1):13. https://doi.org/10.3390/mps2010013
Chicago/Turabian StyleAnishchenko, Evgeniya, Carmela Vigorito, Luigi Mele, Patrizia Lombari, Alessandra F. Perna, and Diego Ingrosso. 2019. "Novel Applications of Lead Acetate and Flow Cytometry Methods for Detection of Sulfur-Containing Molecules" Methods and Protocols 2, no. 1: 13. https://doi.org/10.3390/mps2010013
APA StyleAnishchenko, E., Vigorito, C., Mele, L., Lombari, P., Perna, A. F., & Ingrosso, D. (2019). Novel Applications of Lead Acetate and Flow Cytometry Methods for Detection of Sulfur-Containing Molecules. Methods and Protocols, 2(1), 13. https://doi.org/10.3390/mps2010013






