Fast PET Scan Tumor Segmentation Using Superpixels, Principal Component Analysis and K-Means Clustering
Abstract
:1. Introduction
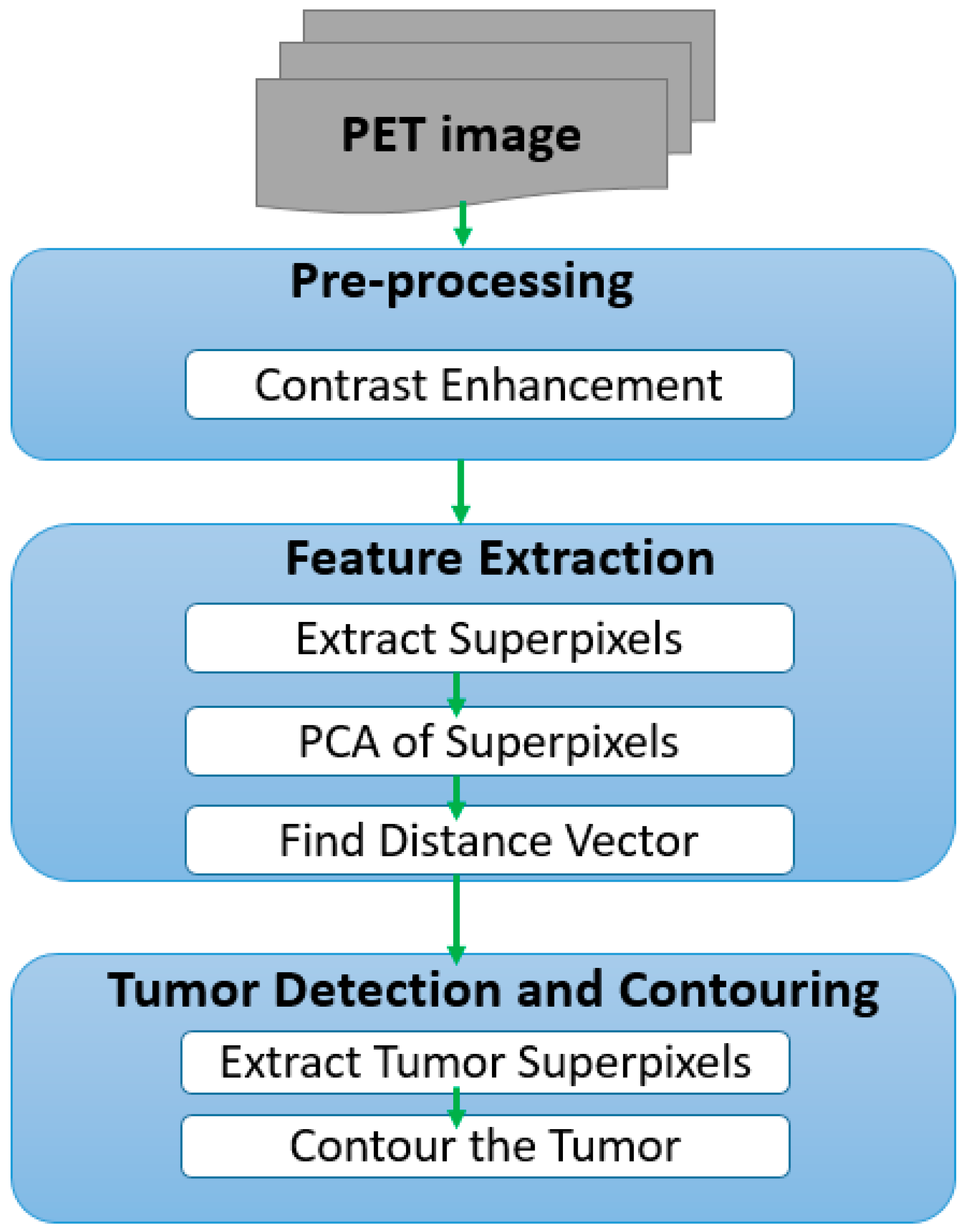
2. Implementation
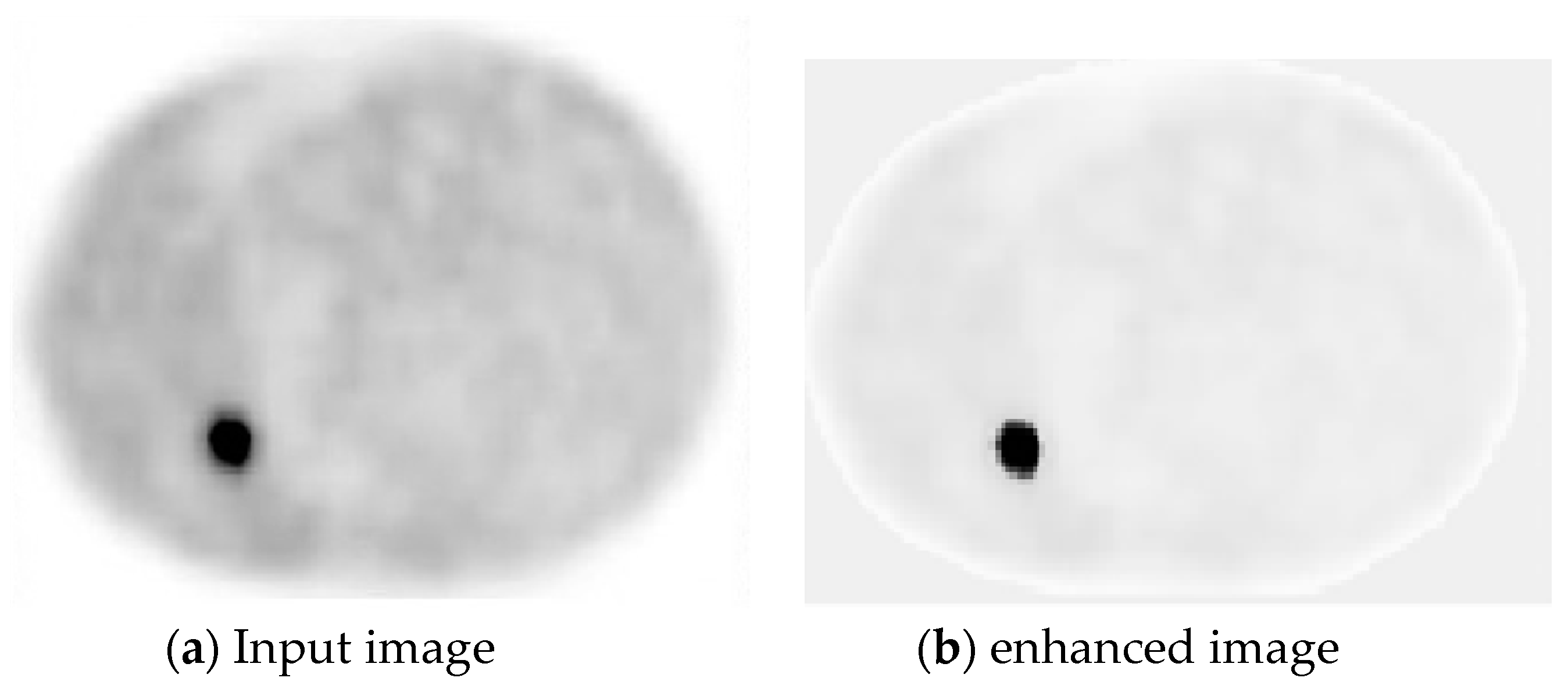
2.1. Pre-Processing
2.2. Feature Extraction
- (1)
- We computed the average size of the superpixel as shown in Equation (2).where N is the number of superpixels, and ni is the number of pixels in ith superpixel. Then, M is an average number of pixels per superpixel.
- (2)
- Then, the size of each superpixel is made same as that of the average one by padding some pixel value to the smaller size superpixel and removing some intensity value from the large size superpixels. Instead of appending random intensity values to smaller sized superpixels, we pad by repeating the last pixels value of the superpixel itself. Finally, the superpixel matrix is generated as shown in Equation (3)where each column represents a superpixel pixel, M is in Equation (2) and N is the number of superpixels.
- (1)
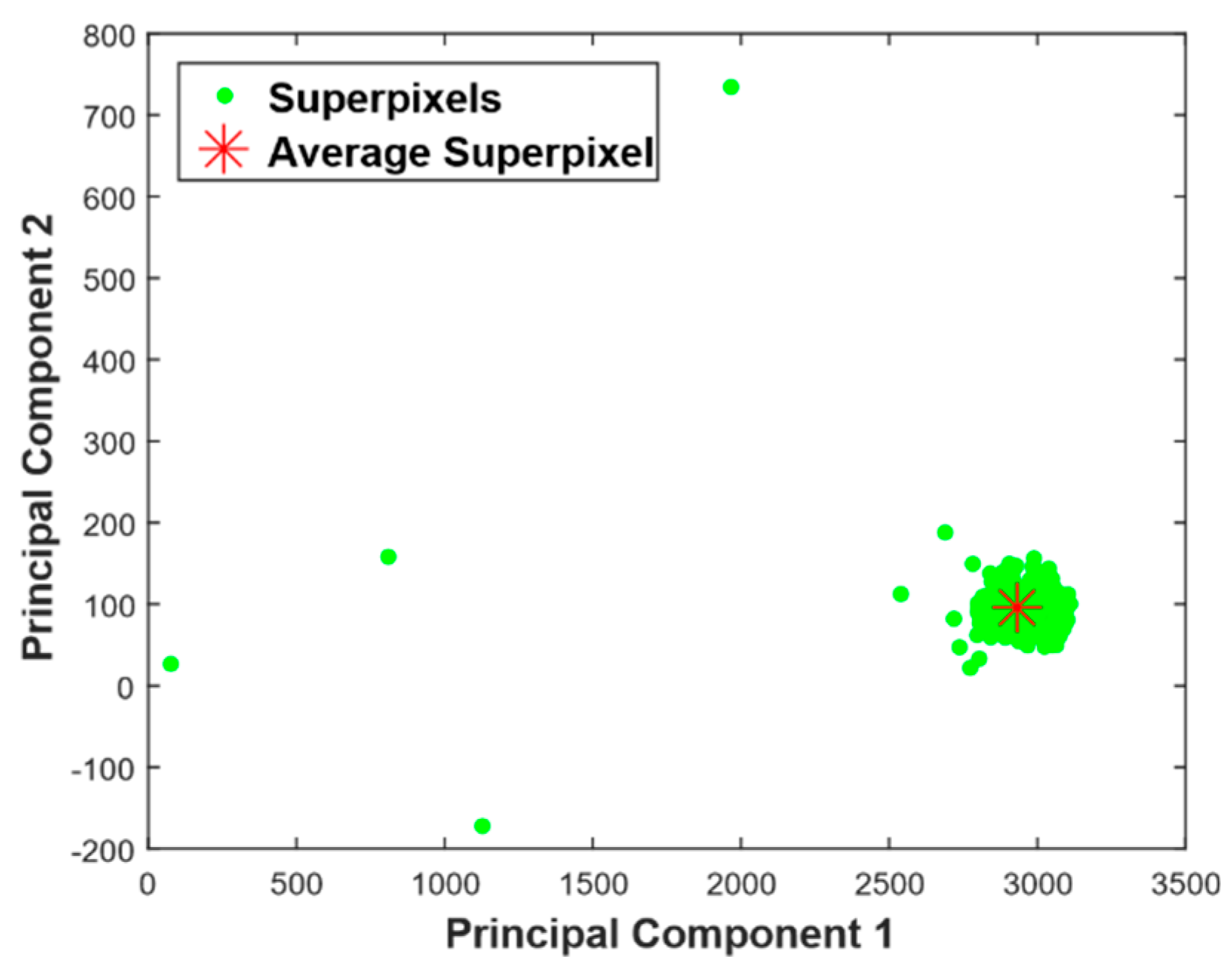
- Compute average superpixel.where Si is the ith superpixel and Sa average superpixel.
- (2)
- Determine the covariance of superpixels (Cs)where Y is the superpixel matrix after average superpixel padding and Yat is the mean of transpose of Y.
- (3)
- Calculate the eigensuperpixels (eigenvectors) and eigenvalues of the covariance matrixwhere P is matrix with eigensuperpixels as column and Σ is diagonal matrix of eigenvalues, λ1, λ2, …, λN, where, λ1 ≥ λ2 ≥ λ3…≥ λN.The magnitude of eigenvalue shows the variance of the data in the direction of its corresponding eigensuperpixel. For N superpixels in Equation (3) above, total variance of intensities of the M- dimensional superpixels can be computed in terms of eigenvalues from Equation (7).where V is the total variance.
- (4)
- Project the superpixels onto eigensuperpixels that contain most of variance of the data. In Equation (6), the number of principal components is same as the number of superpixels. As stated in [17], the eigenvectors or principal components that contain at least 95% of the variance of superpixels can represent the whole image with confidence and this is computed as shown in Equation (8). It reduces the dimensional space, as most of the information is contained in the first two or three largest eigenvalues.In our test images, 95% of the variance of superpixels was in the top two (K = 2) principal components, with most of them reducing M-dimensional superpixels to a 2D point. Once the dominant vectors are found, for feature extraction, the superpixel matrix is projected onto these vectors using Equation (9).where Pk is eigenvectors matrix that contains at least 95% of the variation in the image and Pproj is projection of superpixel matrix to Pk.
- (5)
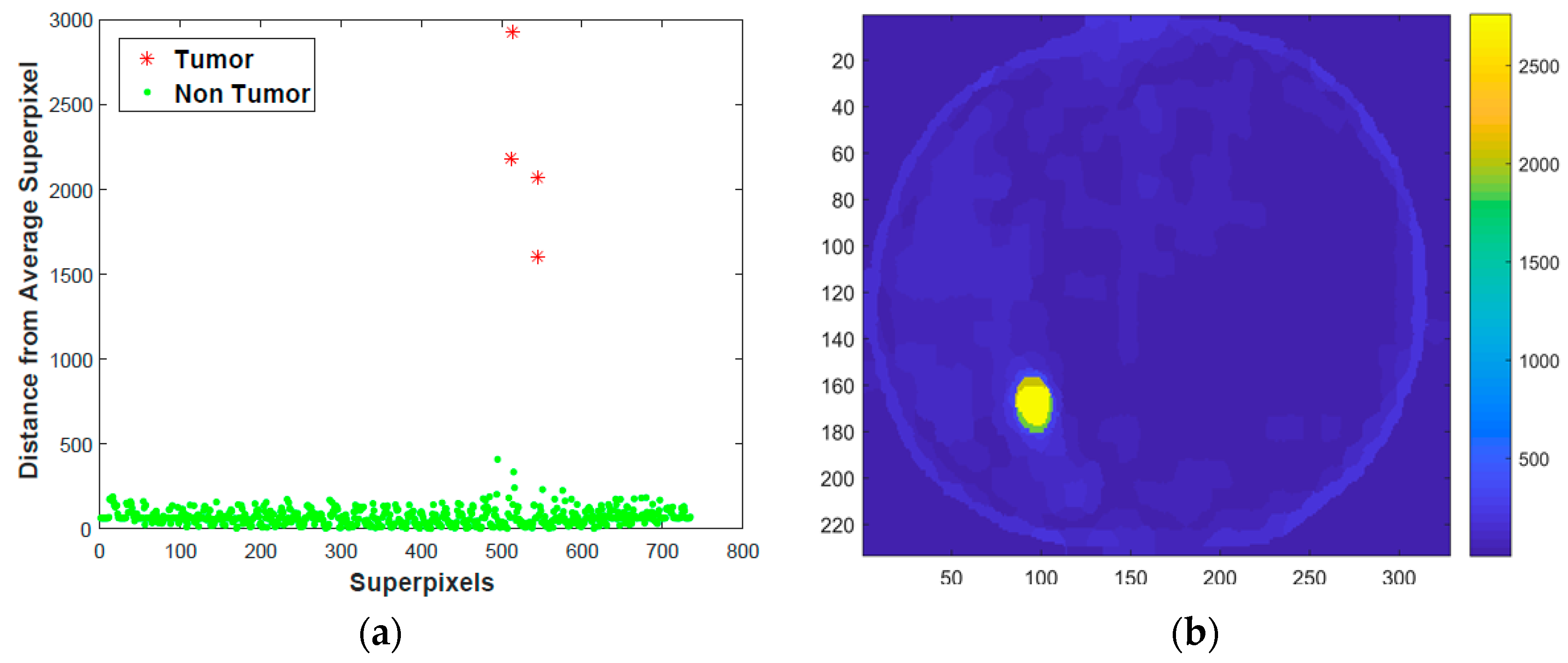
- Calculate the distance of each superpixel to average superpixel. Computing distance should consider the distribution of superpixels in the principal component coordinate system [12]. To incorporate this concept, we computed the distance along the principal components. Mathematically, this will be computing L1 norm distance in the principal components coordinate system as shown in Equation (10) below.where S′i is coordinate of Si relative to Sa in the principal component coordinate system, and D is L1 norm distance.
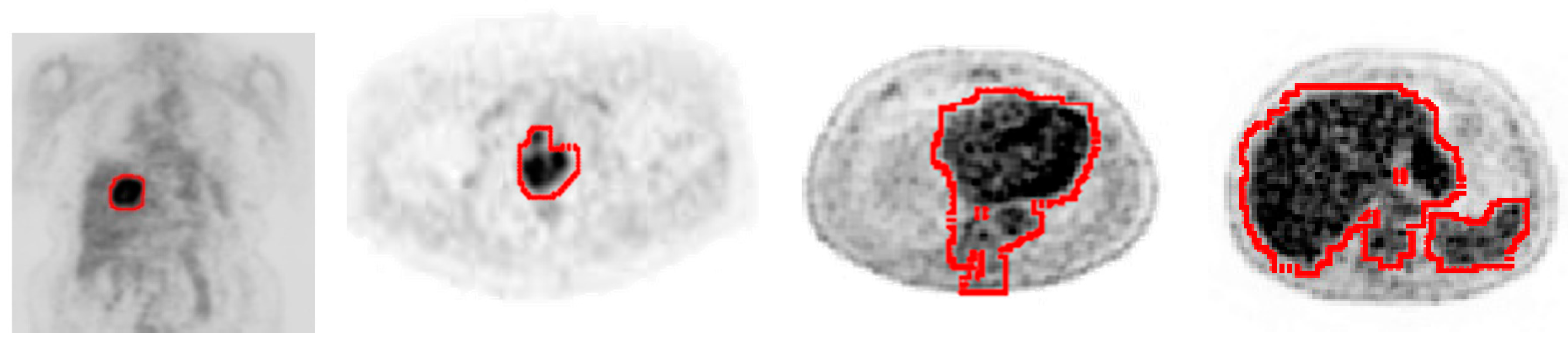
2.3. Tumor Detection and Contouring
3. Result and Discussion
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Sun, C.-C.; Ruan, S.-J.; Shie, M.-C.; Pai, T.-W. Dnamic Contrast enhancement based on histogram specification. IEEE Trans. Consum. Electron. 2005, 51, 1300–1305. [Google Scholar]
- Hanzouli, H.; Lapuyade-Lahorgue, J.; Monfrini, E.; Delso, G.; Pieczynski, W.; Visvikis, D.; Hatt, M. PET/CT image denoising and segmentation based on a multi observation and a multi scale Markov tree model. In Proceedings of the Nuclear Science Symposium and Medical Imaging Conference (NSS/MIC), Seoul, Korea, 27 October–2 November 2013; pp. 1–4. [Google Scholar]
- Xu, Z.; Bagci, U.; Gao, M.; Mollura, D. Improved PET image quantification via iterative denoising and partial volume correction. J. Nucl. Med. 2015, 56, 1744. [Google Scholar]
- Slavine, N.; McColl, R.; Kulkarni, P. Iterative deconvolution method for preclinical PET/CT image enhancement: Application in imaging Alzheimer’s plaque deposition in ad transgenic mice. J. Nucl. Med. 2016, 57, 1985. [Google Scholar]
- Saha, P.K.; Udupa, J.K. Scalebased diffusive image filtering preserving boundary sharpness and fine structures. IEEE Trans. Med. Imaging 2001, 20, 1140–1155. [Google Scholar] [CrossRef] [PubMed]
- Foster, B.; Bagci, U.; Mansoor, A.; Xu, Z.; Mollura, D.J. A review on segmentation of positron emission tomography images. Comput. Boil. Med. 2014, 50, 76–96. [Google Scholar] [CrossRef] [PubMed]
- Zaidi, H.; El Naqa, I. Pet-guided delineation of radiation therapy treatment volumes: A survey of image segmentation techniques. Eur. J. Nucl. Med. Mol. Imaging 2010, 37, 2165–2187. [Google Scholar] [CrossRef] [PubMed]
- Bagci, U.; Udupa, J.K.; Yao, J.; Mollura, D.J. Co-segmentation of functional and anatomical images. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Berlin/Heidelberg, Germany, 2012; pp. 459–467. [Google Scholar]
- Nishio, M.; Kono, A.K.; Kubo, K.; Koyama, H.; Nishii, T.; Sugimura, K. Tumor segmentation on 18F FDG-PET images using graph cut and local spatial information. Open J. Med. Imaging 2015, 5, 174. [Google Scholar] [CrossRef]
- Zhang, K.; Zhang, L.; Song, H.; Zhou, W. Active contours with selective local or global segmentation: A new formulation and level set method. Image Vis. Comput. 2010, 28, 668–676. [Google Scholar] [CrossRef]
- Meena, A.; Raja, K. Segmentation of Alzheimer’s Disease in PET scan datasets using MATLAB. Int. J. Inf. Sci. Comput. arXiv 2012, arXiv:1302.6426. [Google Scholar]
- Margolin, R.; Tal, A.; ZelnikManor, L. What makes a patch distinct? In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Portland, OR, USA, 23–28 June 2013; pp. 1139–1146. [Google Scholar]
- Achanta, R.; Shaji, A.; Smith, K.; Lucchi, A.; Fua, P.; Susstrunk, S. SLIC superpixels compared to state-of-the-art superpixel methods. IEEE Trans. Pattern Anal. Mach. Inell. 2010, 34, 2274–2282. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Hu, Z.; Fan, Q. Superpixel-based unsupervised change detection using multi-dimensional change vector analysis and Svm-based classification. ISPRS Ann. Photogramm. Remote Sens. Spat. Inf. Sci. 2012, 7, 257–262. [Google Scholar] [CrossRef]
- Tian, Z.; Liu, L.; Zhang, Z.; Xue, J.; Fei, B. A supervoxel-based segmentation method for prostate MR images. Med. Phys. 2016, 44, 558–569. [Google Scholar] [CrossRef] [PubMed]
- Chung, H.; Lu, G.; Tian, Z.; Wang, D.; Chen, Z.G.; Fei, B. Superpixel-based spectral classification for the detection of head and neck cancer with hyperspectral imaging. Proc. SPIE Int. Soc. Opt. Eng. 2016, 813–978. [Google Scholar] [CrossRef]
- Jackson, D.A. Stopping rules in principal components analysis: A comparison of heuristical and statistical approaches. Ecology 1993, 74, 2204–2214. [Google Scholar] [CrossRef]




| Size | No. Superpixels | Distance Vector Dimension | Execution Time (s) | |
|---|---|---|---|---|
| Image 1 | 233 × 328 | 692 | 692 | 2.2 |
| Image 2 | 233 × 328 | 500 | 500 | 2.4 |
| Image 3 | 681 × 572 | 660 | 660 | 2.55 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hagos, Y.B.; Minh, V.H.; Khawaldeh, S.; Pervaiz, U.; Aleef, T.A. Fast PET Scan Tumor Segmentation Using Superpixels, Principal Component Analysis and K-Means Clustering. Methods Protoc. 2018, 1, 7. https://doi.org/10.3390/mps1010007
Hagos YB, Minh VH, Khawaldeh S, Pervaiz U, Aleef TA. Fast PET Scan Tumor Segmentation Using Superpixels, Principal Component Analysis and K-Means Clustering. Methods and Protocols. 2018; 1(1):7. https://doi.org/10.3390/mps1010007
Chicago/Turabian StyleHagos, Yeman Brhane, Vu Hoang Minh, Saed Khawaldeh, Usama Pervaiz, and Tajwar Abrar Aleef. 2018. "Fast PET Scan Tumor Segmentation Using Superpixels, Principal Component Analysis and K-Means Clustering" Methods and Protocols 1, no. 1: 7. https://doi.org/10.3390/mps1010007
APA StyleHagos, Y. B., Minh, V. H., Khawaldeh, S., Pervaiz, U., & Aleef, T. A. (2018). Fast PET Scan Tumor Segmentation Using Superpixels, Principal Component Analysis and K-Means Clustering. Methods and Protocols, 1(1), 7. https://doi.org/10.3390/mps1010007





