A Novel System for Spinal Muscular Atrophy Screening in Newborns: Japanese Pilot Study
Abstract
:1. Introduction
2. Materials and Methods
2.1. Objectives and Ethics
2.2. SMA Patients and Non-SMA Controls
2.3. Newborn Infants
2.4. SMN1-Deletion Detection System
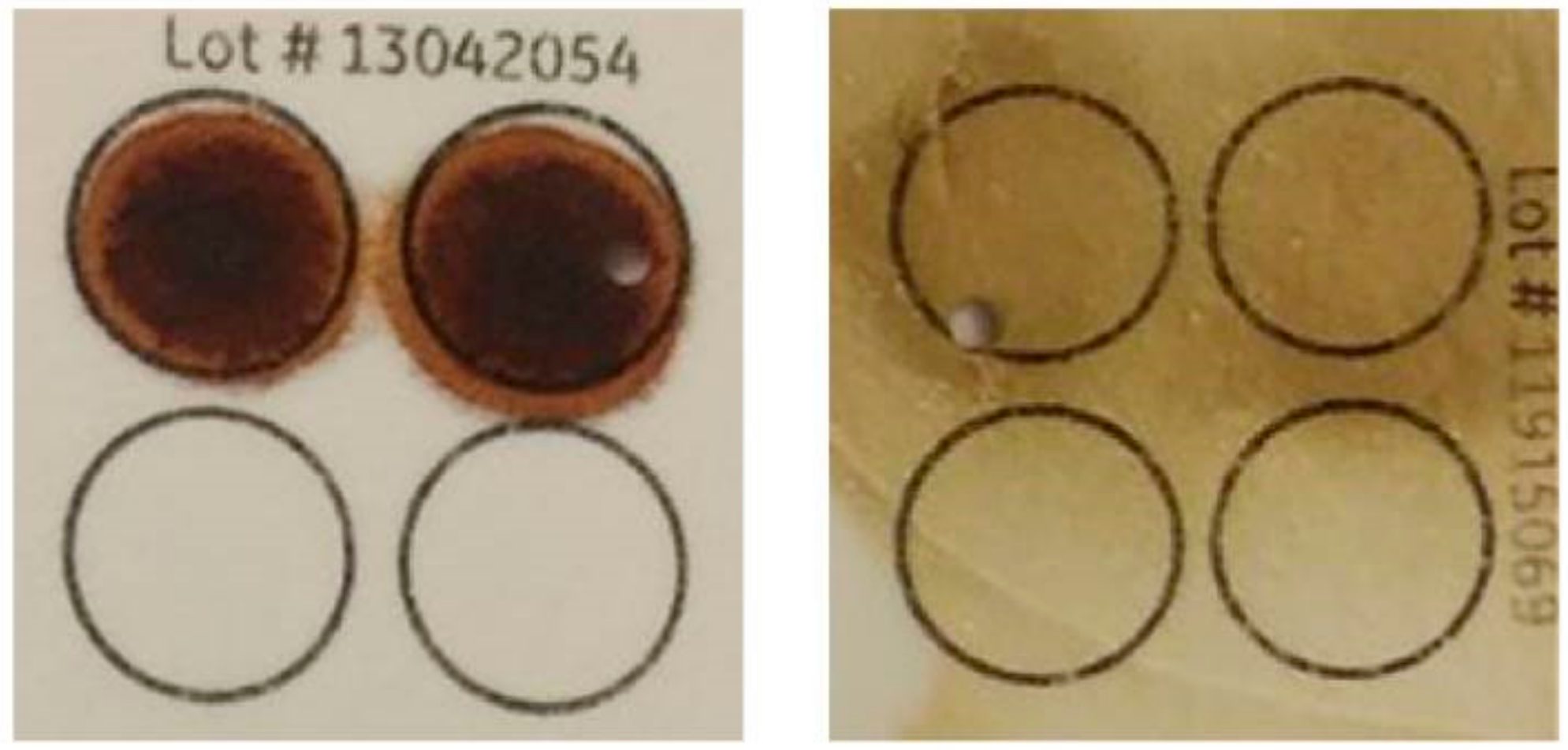
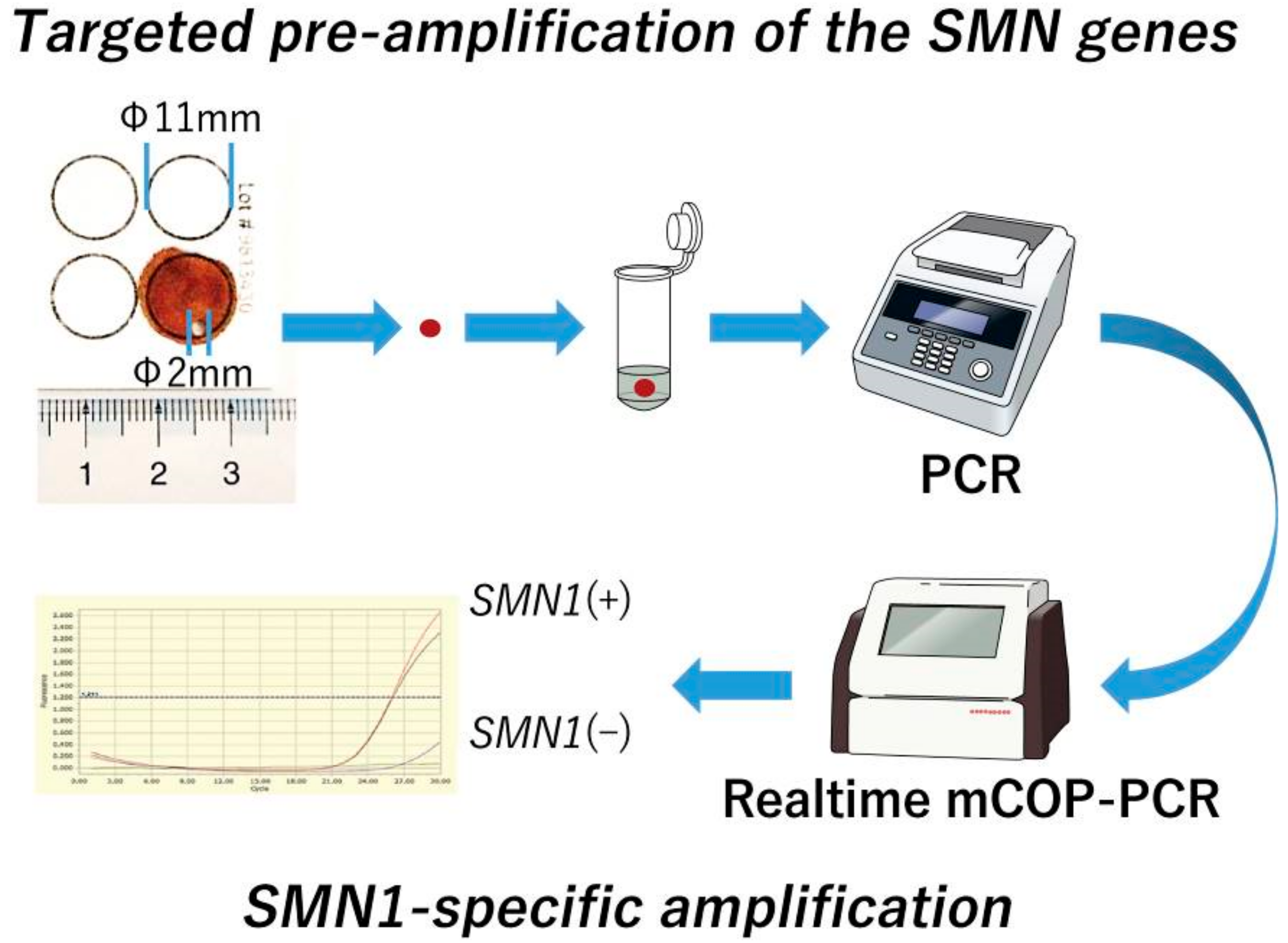
2.4.1. Targeted Pre-Amplification of the SMN Genes
2.4.2. Gene-Specific Amplification of SMN1 Exon 7
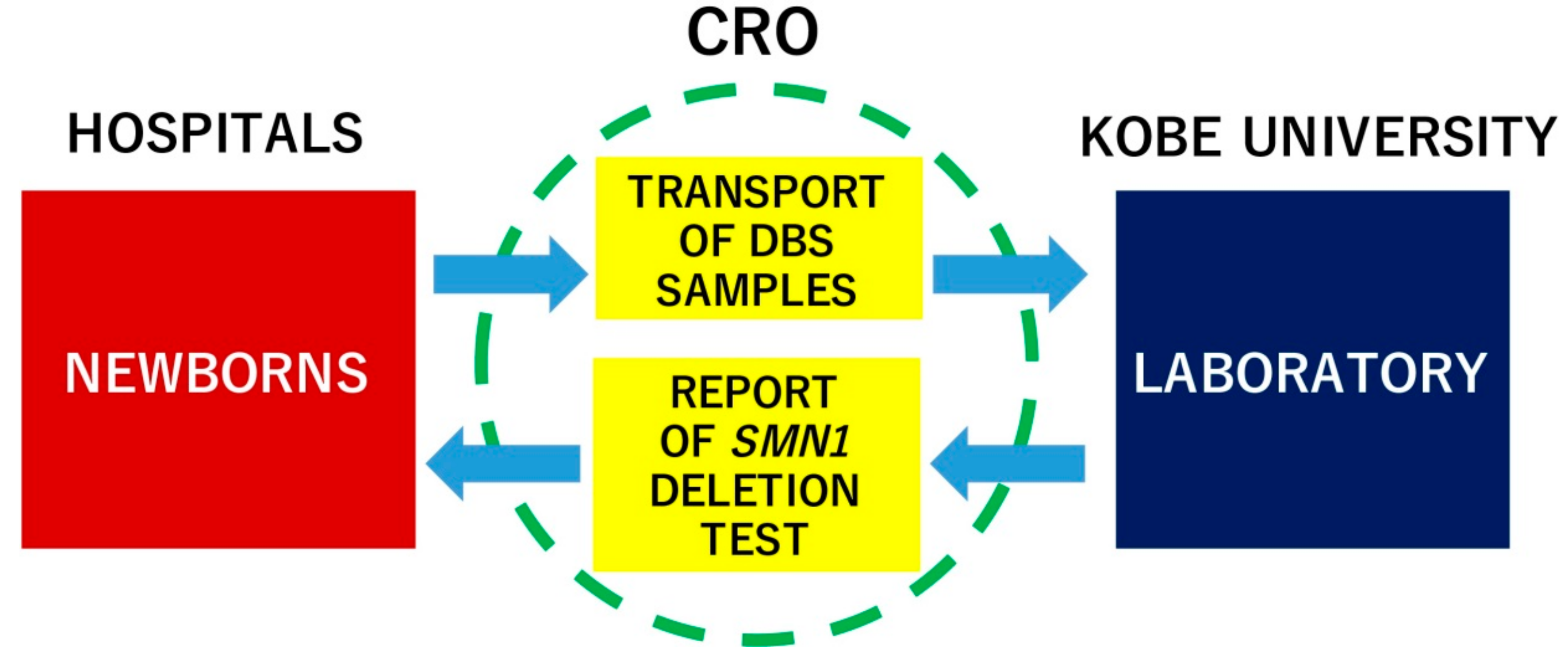
2.5. Newborn Screening Study Design
2.6. Follow-Up Study of the Infants Screened for SMA
2.7. Statistical Analysis
3. Results
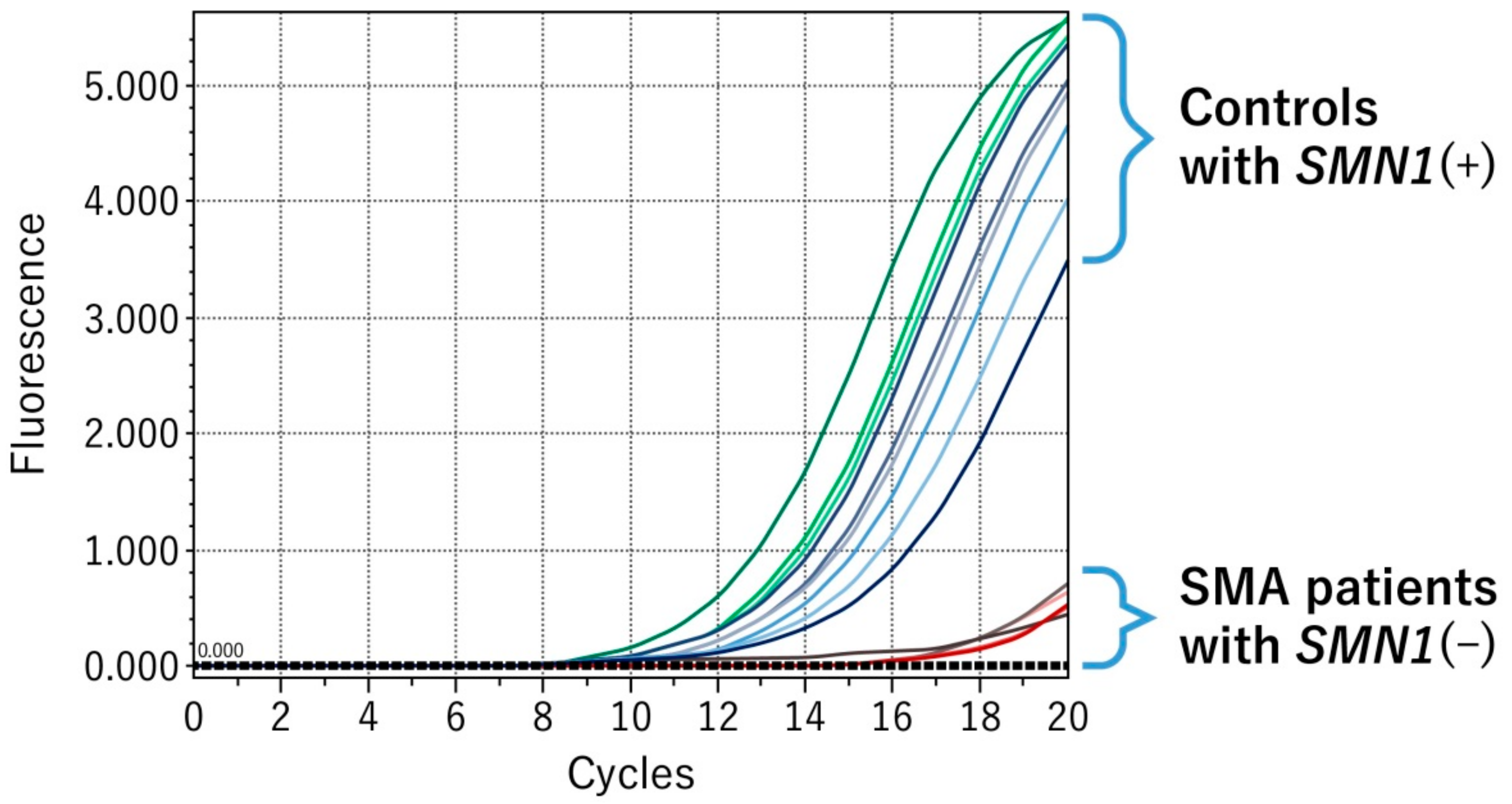
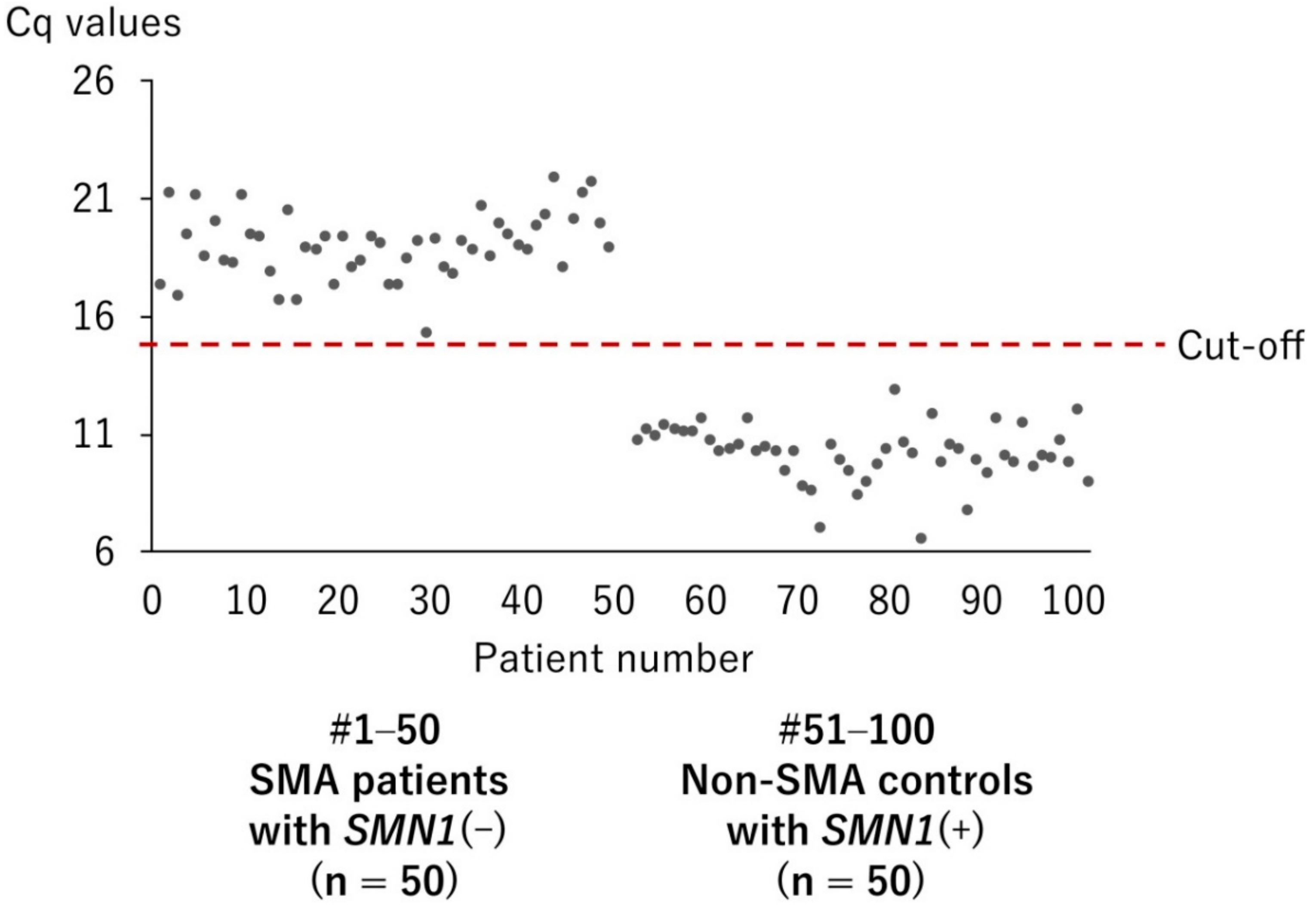
3.1. Validation Study
3.2. Newborn Screening for SMA
3.3. Follow-Up Study of the Infants Screened for SMA
4. Discussion
4.1. Targeted Pre-Amplification of SMN1/SMN2 Sequence
4.2. Modified Competitive Oligonucleotide Priming-PCR (mCOP-PCR)
4.3. Accurate Detection System for SMN1 Deletion
4.4. Robust System for SMA Newborn Screening Using DBS
4.5. Limitations of SMN1-Deletion Detection as an SMA Screening Strategy
5. Conclusions
6. Patents
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
- Japanese Red Cross Katsushika Maternity Hospital: Sakae Kumasaka, Chisako Mitsuishi;
- Matsuyama Red Cross Hospital: Yoichi Kondo;
- Chibune General Hospital: Akihiro Takatera;
- St. Marianna University School of Medicine: Isamu Hokuto;
- Kurashiki Medical Center: Tetsuro Fujino, Nobuyoshi Mimaki, Noriko Yanazaki;
- National Hospital Organization Okayama Medical Center: Makoto Nakamura, Akihito Takeuchi;
- Shimane Prefectural Central Hospital: Fumihide Kato;
- Department of Community Medicine and Social Healthcare Science, Kobe University Graduate School of Medicine: Emma Tabe Eko Niba, Hisahide Nishio, Masakazu Shinohara, Izumi Takayama, Yogik Onky Silvana Wijaya;
- Department of Pediatrics, Kobe University Graduate School of Medicine: Kazumichi Fujioka, Kazumoto Iijima;
- Kurashiki Central Hospital: Akihito Takahashi;
- Okayama University Hospital: Daisaku Morimoto, Yosuke Washio;
- Japan Community Health Care Organization Kyushu Hospital: Yasuhiko Takahashi, Junko Yamamoto;
- Fukuoka Children’s Hospital: Yasushi Takahata;
- Nihon University School of Medicine: Ichiro Morioka, Nobuhiko Nagano;
- Hyogo Prefectural Kobe Children’s Hospital Perinatal Center: Hideto Nakao, Tomoyuki Yokota, Seiji Yoshimoto;
- Tosei General Hospital: Kuniko Ieda;
- Hiroshima University Graduate School of Biomedical and Health Sciences: Norioki Ohno;
- Hokkaido University Hospital: Kazutoshi Cho, Hideaki Shiraishi;
- Toyohashi municipal hospital: Norihisa Koyama, Mari Sugimoto;
- Saga University: Manabu Iwanaga, Muneaki Matsuo;
- Jichi Medical School: Hitoshi Osaka, Hironori Shimozawa, Takanori Yamagata;
- Almeida Memorial Hospital: Naoki Fukushima;
- Toyooka Hospital: Masaaki Ueda;
- Omihachiman Community Medical Center: Shinobu Yoshida;
- Hyogo College of Medicine: Hideki Shimomura, Yasuhiro Takeshima;
- Nagasaki University Hospital: Fumiko Kinoshita, Tatsuharu Sato;
- Juntendo University Nerima Hospital: Shinichi Niijima, Noboru Yoshida;
- Tokyo Metropolitan Ohtsuka Hospital: Ken Masunaga;
- National Hospital Organization Nagasaki Medical Center: Mikihiro Aoki;
- Kakogawa Central City Hospital: Takeshi Morisawa;
- Niigata City General Hospital: Yoshihisa Nagayama;
- Chiba Children’s Hospital: Kei Murayama, Tomoko Tsuruoka;
- Yokosuka General Hospital Uwamachi: Tomoyuki Miyamoto;
- Kouseiren Takaoka Hospital: Hiroaki Imamura, Jiro Ogawa;
- Hokkaido Medical Center for Child Health and Rehabilitation: Hideomi Asanuma, Shuku Ishikawa;
- Sapporo Medical University Hospital: Masaki Kobayashi;
- Japanese Red Cross Nagoya Daini Hospital: Taihei Tanaka, Takaharu Yamada;
- Akita University Graduate School of Medicine: Hiroyuki Adachi, Atsuko Noguchi, Tsutomu Takahashi;
- Japanese Red Cross Saitama Hospital: Yuko Sato;
- Graduate School of Medicine, University of the Ryukyus: Koichi Nakanishi, Tomohide Yoshida;
- Numazu City Hospital: Masao Murabayashi;
- Sapporo City General Hospital: Masato Mizushima, Tatsuo Satomi;
- Maternal & Child Health Center Aiiku Hospital: Shinya Hayashida;
- Hiroshima City Hiroshima Citizens Hospital: Yutaka Nishimura;
- Juntendo University Shizuoka Hospital: Masato Kantake;
- Yamaguchi University Hospital: Sasagu Kimura, Kazumasa Takahashi;
- National Center for Child Health and Development: Go Tajima;
- Kobe Pharmaceutical University: Atsuko Takeuchi
References
- Arnold, W.D.; Kassar, D.; Kissel, J.T. Spinal muscular atrophy: Diagnosis and management in a new therapeutic era. Muscle Nerve 2015, 51, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Verhaart, I.E.C.; Robertson, A.; Wilson, I.J.; Aartsma-Rus, A.; Cameron, S.; Jones, C.C.; Cook, S.F.; Lochmüller, H. Prevalence, incidence and carrier frequency of 5q-linked spinal muscular atrophy—A literature review. Orphanet J. Rare Dis. 2017, 12, 124. [Google Scholar] [CrossRef] [PubMed]
- Muralidharan, K.; Wilson, R.B.; Ogino, S.; Nagan, N.; Curtis, C.; Schrijver, I. Population carrier screening for spinal muscular atrophy a position statement of the association for molecular pathology. J. Mol. Diagn. 2011, 13, 3–6. [Google Scholar] [CrossRef] [PubMed]
- Finkel, R.S.; McDermott, M.P.; Kaufmann, P.; Darras, B.T.; Chung, W.K.; Sproule, D.M.; Kang, P.B.; Foley, A.R.; Yang, M.L.; Martens, W.B.; et al. Observational study of spinal muscular atrophy type I and implications for clinical trials. Neurology 2014, 83, 810–817. [Google Scholar] [CrossRef] [PubMed]
- Zerres, K.; Rudnik-Schöneborn, S. Natural history in proximal spinal muscular atrophy. Clinical analysis of 445 patients and suggestions for a modification of existing classifications. Arch. Neurol. 1995, 52, 518–523. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, S.; Bürglen, L.; Reboullet, S.; Clermont, O.; Burlet, P.; Viollet, L.; Benichou, B.; Cruaud, C.; Millasseau, P.; Zeviani, M.; et al. Identification and characterization of a spinal muscular atrophy-determining gene. Cell 1995, 80, 155–165. [Google Scholar] [CrossRef]
- Singh, R.N.; Howell, M.D.; Ottesen, E.W.; Singh, N.N. Diverse role of survival motor neuron protein. Biochim. Biophys. Acta Gene Regul. Mech. 2017, 1860, 299–315. [Google Scholar] [CrossRef] [PubMed]
- Feldkötter, M.; Schwarzer, V.; Wirth, R.; Wienker, T.F.; Wirth, B. Quantitative analyses of SMN1 and SMN2 based on real-time lightCycler PCR: Fast and highly reliable carrier testing and prediction of severity of spinal muscular atrophy. Am. J. Hum. Genet. 2002, 70, 358–368. [Google Scholar] [CrossRef] [PubMed]
- Schrank, B.; Götz, R.; Gunnersen, J.M.; Ure, J.M.; Toyka, K.V.; Smith, A.G.; Sendtner, M. Inactivation of the survival motor neuron gene, a candidate gene for human spinal muscular atrophy, leads to massive cell death in early mouse embryos. Proc. Natl. Acad. Sci. USA 1997, 94, 9920–9925. [Google Scholar] [CrossRef] [PubMed]
- Hsieh-Li, H.M.; Chang, J.G.; Jong, Y.J.; Wu, M.H.; Wang, N.M.; Tsai, C.H.; Li, H. A mouse model for spinal muscular atrophy. Nat. Genet. 2000, 24, 66–70. [Google Scholar] [CrossRef] [PubMed]
- Finkel, R.S.; Mercuri, E.; Darras, B.T.; Connolly, A.M.; Kuntz, N.L.; Kirschner, J.; Chiriboga, C.A.; Saito, K.; Servais, L.; Tizzano, E.; et al. Nusinersen versus sham control in infantile-onset spinal muscular atrophy. N. Engl. J. Med. 2017, 377, 1723–1732. [Google Scholar] [CrossRef] [PubMed]
- Mercuri, E.; Darras, B.T.; Chiriboga, C.A.; Day, J.W.; Campbell, C.; Connolly, A.M.; Iannaccone, S.T.; Kirschner, J.; Kuntz, N.L.; Saito, K.; et al. Nusinersen versus sham control in later-onset spinal muscular atrophy. N. Engl. J. Med. 2018, 378, 625–635. [Google Scholar] [CrossRef] [PubMed]
- Finkel, R.S.; Chiriboga, C.A.; Vajsar, J.; Day, J.W.; Montes, J.; De Vivo, D.C.; Yamashita, M.; Rigo, F.; Hung, G.; Schneider, E.; et al. Treatment of infantile-onset spinal muscular atrophy with nusinersen: A phase 2, open-label, dose-escalation study. Lancet 2016, 388, 3017–3026. [Google Scholar] [CrossRef]
- Swoboda, K.J.; Prior, T.W.; Scott, C.B.; McNaught, T.P.; Wride, M.C.; Reyna, S.P.; Bromberg, M.B. Natural history of denervation in SMA: Relation to age, SMN2 copy number, and function. Ann. Neurol. 2005, 57, 704–712. [Google Scholar] [CrossRef] [PubMed]
- Glascock, J.; Sampson, J.; Haidet-Phillips, A.; Connolly, A.; Darras, B.; Day, J.; Finkel, R.; Howell, R.R.; Klinger, K.; Kuntz, N.; et al. Treatment algorithm for infants diagnosed with spinal muscular atrophy through newborn screening. J. Neuromuscul Dis. 2018, 5, 145–158. [Google Scholar] [CrossRef] [PubMed]
- Kraszewski, J.N.; Kay, D.M.; Stevens, C.F.; Koval, C.; Haser, B.; Ortiz, V.; Albertorio, A.; Cohen, L.L.; Jain, R.; Andrew, S.P.; et al. Pilot study of population-based newborn screening for spinal muscular atrophy in New York state. Genet. Med. 2018, 20, 608–613. [Google Scholar] [CrossRef] [PubMed]
- Kemper, A.R.; Lam, K.K.; Comeau, A.M.; Kwon, J.; Green, N.S.; Ojodu, J.; Grosse, S.; Prosser, L.A.; Jones, E.; Tanksley, S.; et al. Evidence-Based Review of Newborn Screening for Spinal Muscular Atrophy (SMA): Final Report (v5.2). Available online: https://www.hrsa.gov/sites/default/files/hrsa/advisory-committees/heritable-disorders/reports-recommendations/sma-final-report.pdf (accessed on 9 August 2019).
- Ar Rochmah, M.; Harahap, N.I.F.; Niba, E.T.E.; Nakanishi, K.; Awano, H.; Morioka, I.; Iijima, K.; Saito, T.; Saito, K.; Lai, P.S.; et al. Genetic screening of spinal muscular atrophy using a real-time modified COP-PCR technique with dried blood-spot DNA. Brain Dev. 2017, 39, 774–782. [Google Scholar] [CrossRef] [PubMed]
- Van der Steege, G.; Grootscholten, P.M.; van der Vlies, P.; Draaijers, T.G.; Osinga, J.; Cobben, J.M.; Scheffer, H.; Buys, C.H. PCR-based DNA test to confirm clinical diagnosis of autosomal recessive spinal muscular atrophy. Lancet 1995, 345, 985–986. [Google Scholar] [CrossRef]
- Takeuchi, A.; Tode, C.; Nishino, M.; Wijaya, Y.O.S.; Niba, E.T.E.; Awano, H.; Takeshima, Y.; Saito, T.; Saito, K.; Lai, P.S.; et al. Newborn screening for spinal muscular atrophy: DNA preparation from dried blood spot and DNA polymerase selection in PCR. Kobe J. Med. Sci. 2019, 65, E95–E99. [Google Scholar]
- Kato, N.; Sa’Adah, N.; Ar Rochmah, M.; Harahap, N.I.; Nurputra, D.K.; Sato, H.; Sadewa, A.H.; Astuti, I.; Haryana, S.M.; Saito, T.; et al. SMA screening system using dried blood spots on filter paper: Application of COP-PCR to the SMN1 deletion test. Kobe J. Med. Sci. 2015, 60, E78–E85. [Google Scholar] [PubMed]
- Pezzullo, J.C. 2-way Contingency Table Analysis (Japanese). Available online: http://www.grade-jpn.com/2x2.html (accessed on 27 September 2019).
- Pezzullo, J.C. 2-way Contingency Table Analysis (English). Available online: http://statpages.info/ctab2x2.html (accessed on 27 September 2019).
- Gibbs, R.A.; Nguyen, P.N.; Caskey, C.T. Detection of single DNA base differences by competitive oligonucleotide priming. Nucleic Acids Res. 1989, 17, 2437–2448. [Google Scholar] [CrossRef] [PubMed]
- Yen, T.; Nightingale, B.N.; Burns, J.C.; Sullivan, D.R.; Stewart, P.M. Butyrylcholinesterase (BCHE) genotyping for post-succinylcholine apnea in an Australian population. Clin. Chem. 2003, 49, 1297–1308. [Google Scholar] [CrossRef] [PubMed]
- Chien, Y.H.; Chiang, S.C.; Weng, W.C.; Lee, N.C.; Lin, C.J.; Hsieh, W.S.; Lee, W.T.; Jong, Y.J.; Ko, T.M.; Hwu, W.L. Presymptomatic diagnosis of spinal muscular atrophy through newborn screening. J. Pediatr. 2017, 190, 124–129. [Google Scholar] [CrossRef] [PubMed]
- Czibere, L.; Burggraf, S.; Fleige, T.; Glück, B.; Keitel, L.M.; Landt, O.; Durner, J.; Röschinger, W.; Hohenfellner, K.; Wirth, B.; et al. High-throughput genetic newborn screening for spinal muscular atrophy by rapid nucleic acid extraction from dried blood spots and 384-well qPCR. Eur. J. Hum. Genet. 2019. [Google Scholar] [CrossRef] [PubMed]
- Boemer, F.; Caberg, J.H.; Dideberg, V.; Dardenne, D.; Bours, V.; Hiligsmann, M.; Dangouloff, T.; Servais, L. Newborn screening for SMA in Southern Belgium. Neuromuscul. Disord. 2019, 29, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.L.; Lee, F.K.; Yazdanpanah, G.K.; Staropoli, J.F.; Liu, M.; Carulli, J.P.; Sun, C.; Dobrowolski, S.F.; Hannon, W.H.; Vogt, R.F. Newborn blood spot screening test using multiplexed real-time PCR to simultaneously screen for spinal muscular atrophy and severe combined immunodeficiency. Clin. Chem. 2015, 61, 412–419. [Google Scholar] [CrossRef] [PubMed]
- Strunk, A.; Abbes, A.; Stuitje, A.R.; Hettinga, C.; Sepers, E.M.; Snetselaar, R.; Schouten, J.; Asselman, F.-L.; Cuppen, I.; Lemmink, H.; et al. Validation of a fast, robust, inexpensive, two-tiered neonatal screening test algorithm on dried blood spots for spinal muscular atrophy. Int. J. Neonatal Screen 2019, 5, 21. [Google Scholar] [CrossRef]
- Harada, Y.; Sutomo, R.; Sadewa, A.H.; Akutsu, T.; Takeshima, Y.; Wada, H.; Matsuo, M.; Nishio, H. Correlation between SMN2 copy number and clinical phenotype of spinal muscular atrophy: Three SMN2 copies fail to rescue some patients from the disease severity. J. Neurol. 2002, 249, 1211–1219. [Google Scholar] [CrossRef] [PubMed]

| PCR–RFLP (Fresh Blood) | Total | ||
|---|---|---|---|
| SMN1(–) | SMN1(+) | ||
| Real-time mCOP-PCR (DBS) | |||
| SMN1(–) | 50 | 0 | 50 |
| SMN1(+) | 0 | 50 | 50 |
| Total | 50 | 50 | 100 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shinohara, M.; Niba, E.T.E.; Wijaya, Y.O.S.; Takayama, I.; Mitsuishi, C.; Kumasaka, S.; Kondo, Y.; Takatera, A.; Hokuto, I.; Morioka, I.; et al. A Novel System for Spinal Muscular Atrophy Screening in Newborns: Japanese Pilot Study. Int. J. Neonatal Screen. 2019, 5, 41. https://doi.org/10.3390/ijns5040041
Shinohara M, Niba ETE, Wijaya YOS, Takayama I, Mitsuishi C, Kumasaka S, Kondo Y, Takatera A, Hokuto I, Morioka I, et al. A Novel System for Spinal Muscular Atrophy Screening in Newborns: Japanese Pilot Study. International Journal of Neonatal Screening. 2019; 5(4):41. https://doi.org/10.3390/ijns5040041
Chicago/Turabian StyleShinohara, Masakazu, Emma Tabe Eko Niba, Yogik Onky Silvana Wijaya, Izumi Takayama, Chisako Mitsuishi, Sakae Kumasaka, Yoichi Kondo, Akihiro Takatera, Isamu Hokuto, Ichiro Morioka, and et al. 2019. "A Novel System for Spinal Muscular Atrophy Screening in Newborns: Japanese Pilot Study" International Journal of Neonatal Screening 5, no. 4: 41. https://doi.org/10.3390/ijns5040041
APA StyleShinohara, M., Niba, E. T. E., Wijaya, Y. O. S., Takayama, I., Mitsuishi, C., Kumasaka, S., Kondo, Y., Takatera, A., Hokuto, I., Morioka, I., Ogiwara, K., Tobita, K., Takeuchi, A., Nishio, H., & for the SMA-NBS PILOT STUDY GROUP. (2019). A Novel System for Spinal Muscular Atrophy Screening in Newborns: Japanese Pilot Study. International Journal of Neonatal Screening, 5(4), 41. https://doi.org/10.3390/ijns5040041






