Quantib Prostate Compared to an Expert Radiologist for the Diagnosis of Prostate Cancer on mpMRI: A Single-Center Preliminary Study
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patient Population
2.2. MR Imaging
2.3. Pathological Validation
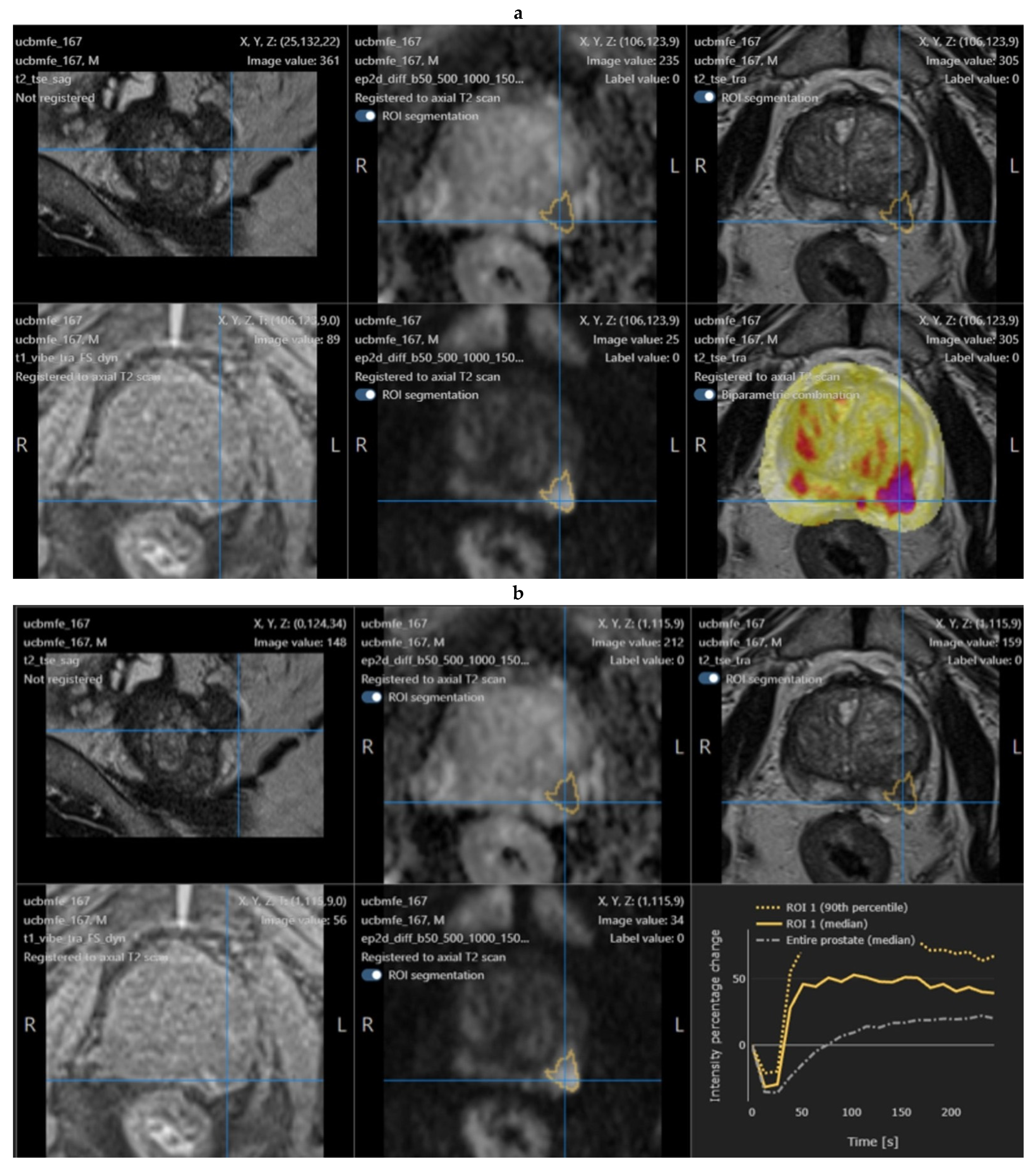
2.4. Quantib Prostate Software
- PSA density analysis, with the automatic segmentation of the prostate, which can be reviewed and modified by the radiologist;
- The multiparametric MRI analysis, where standardized assessment of MRI includes the ability to add, edit, and inspect ROIs and score them according to PI-RADS general scoring, finding, reviewing, and approving the results;
- Results are exported and a standardized report is created.
2.5. Study Design and Statistical Analysis
- -
- Prostatic zones: transition zone (TZ) or peripherical zone (PZ);
- -
- Side and localization: right or left; apex, equatorial, or base;
- -
- Largest axial dimension;
- -
- Lesion volume (calculated by Quantib);
- -
- PI-RADS.
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kumar, V.; Bora, G.S.; Kumar, R.; Jagannathan, N.R. Multiparametric (mp) MRI of prostate cancer. Prog. Nucl. Magn. Reson. Spectrosc. 2018, 105, 23–40. [Google Scholar] [CrossRef] [PubMed]
- Mottet, N.; van den Bergh, R.C.N.; Briers, E.; van den Broeck, T.; Cumberbatch, M.G.; De Santis, M.; Fanti, S.; Fossati, N.; Gandaglia, G.; Gillessen, S.; et al. EAU-EANM-ESTRO-ESUR-SIOG Guidelines on Prostate Cancer—2020 Update. Part 1: Screening, Diagnosis, and Local Treatment with Curative Intent. Eur. Urol. 2020, 79, 243–262. [Google Scholar] [CrossRef] [PubMed]
- Wagaskar, V.G.; Mittal, A.; Sobotka, S.; Ratnani, P.; Lantz, A.; Falagario, U.G.; Martini, A.; Dovey, Z.; Treacy, P.-J.; Pathak, P.; et al. Hood Technique for Robotic Radical Prostatectomy—Preserving Periurethral Anatomical Structures in the Space of Retzius and Sparing the Pouch of Douglas, Enabling Early Return of Continence Without Compromising Surgical Margin Rates. Eur. Urol. 2020, 80, 213–221. [Google Scholar] [CrossRef] [PubMed]
- Drost, F.-J.H.; Osses, D.; Nieboer, D.; Bangma, C.H.; Steyerberg, E.W.; Roobol, M.J.; Schoots, I.G. Prostate Magnetic Resonance Imaging, with or Without Magnetic Resonance Imaging-targeted Biopsy, and Systematic Biopsy for Detecting Prostate Cancer: A Cochrane Systematic Review and Meta-analysis. Eur. Urol. 2020, 77, 78–94. [Google Scholar] [CrossRef] [PubMed]
- Fütterer, J.J.; Briganti, A.; De Visschere, P.; Emberton, M.; Giannarini, G.; Kirkham, A.; Taneja, S.S.; Thoeny, H.; Villeirs, G.; Villers, A. Can Clinically Significant Prostate Cancer Be Detected with Multiparametric Magnetic Resonance Imaging? A Systematic Review of the Literature. Eur. Urol. 2015, 68, 1045–1053. [Google Scholar] [CrossRef] [PubMed]
- Purysko, A.S.; Baroni, R.H.; Giganti, F.; Costa, D.; Renard-Penna, R.; Kim, C.K.; Raman, S.S. PI-RADS Version 2.1: A Critical Review, From the AJR Special Series on Radiology Reporting and Data Systems. Am. J. Roentgenol. 2021, 216, 20–32. [Google Scholar] [CrossRef] [PubMed]
- Moldovan, P.C.; Broeck, T.V.D.; Sylvester, R.; Marconi, L.; Bellmunt, J.; Bergh, R.C.V.D.; Bolla, M.; Briers, E.; Cumberbatch, M.G.; Fossati, N.; et al. What Is the Negative Predictive Value of Multiparametric Magnetic Resonance Imaging in Excluding Prostate Cancer at Biopsy? A Systematic Review and Meta-analysis from the European Association of Urology Prostate Cancer Guidelines Panel. Eur. Urol. 2017, 72, 250–266. [Google Scholar] [CrossRef] [PubMed]
- Hietikko, R.; Kilpeläinen, T.P.; Kenttämies, A.; Ronkainen, J.; Ijäs, K.; Lind, K.; Marjasuo, S.; Oksala, J.; Oksanen, O.; Saarinen, T.; et al. Expected impact of MRI-related interreader variability on ProScreen prostate cancer screening trial: A pre-trial validation study. Cancer Imaging 2020, 20, 72. [Google Scholar] [CrossRef] [PubMed]
- Rudolph, M.M.; Baur, A.D.J.; Cash, H.; Haas, M.; Mahjoub, S.; Hartenstein, A.; Hamm, C.A.; Beetz, N.L.; Konietschke, F.; Hamm, B.; et al. Diagnostic performance of PI-RADS version 2.1 compared to version 2.0 for detection of peripheral and transition zone prostate cancer. Sci. Rep. 2020, 10, 15982. [Google Scholar] [CrossRef] [PubMed]
- Rosenkrantz, A.B.; Ayoola, A.; Hoffman, D.; Khasgiwala, A.; Prabhu, V.; Smereka, P.; Somberg, M.; Taneja, S.S. The Learning Curve in Prostate MRI Interpretation: Self-Directed Learning Versus Continual Reader Feedback. Am. J. Roentgenol. 2017, 208, W92–W100. [Google Scholar] [CrossRef] [PubMed]
- Twilt, J.; van Leeuwen, K.; Huisman, H.; Fütterer, J.; de Rooij, M. Artificial Intelligence Based Algorithms for Prostate Cancer Classification and Detection on Magnetic Resonance Imaging: A Narrative Review. Diagnostics 2021, 11, 959. [Google Scholar] [CrossRef] [PubMed]
- Hambrock, T.; Vos, P.C.; De Kaa, C.A.H.; Barentsz, J.O.; Huisman, H. Prostate Cancer: Computer-aided Diagnosis with Multiparametric 3-T MR Imaging—Effect on Observer Performance. Radiology 2013, 266, 521–530. [Google Scholar] [CrossRef] [PubMed]
- Greer, M.D.; Lay, N.; Shih, J.H.; Barrett, T.; Bittencourt, L.K.; Borofsky, S.; Kabakus, I.; Law, Y.M.; Marko, J.; Shebel, H.; et al. Computer-aided diagnosis prior to conventional interpretation of prostate mpMRI: An international multi-reader study. Eur. Radiol. 2018, 28, 4407–4417. [Google Scholar] [CrossRef] [PubMed]
- Cameron, A.; Khalvati, F.; Haider, M.A.; Wong, A. MAPS: A Quantitative Radiomics Approach for Prostate Cancer Detection. IEEE Trans. Biomed. Eng. 2015, 63, 1145–1156. [Google Scholar] [CrossRef] [PubMed]
- Khalvati, F.; Zhang, J.; Chung, A.G.; Shafiee, M.J.; Wong, A.; Haider, M.A. MPCaD: A multi-scale radiomics-driven framework for automated prostate cancer localization and detection. BMC Med. Imaging 2018, 18, 16. [Google Scholar] [CrossRef] [PubMed]
- Penzias, G.; Singanamalli, A.; Elliott, R.; Gollamudi, J.; Shih, N.; Feldman, M.; Stricker, P.; Delprado, W.; Tiwari, S.; Böhm, M.; et al. Identifying the morphologic basis for radiomic features in distinguishing different Gleason grades of prostate cancer on MRI: Preliminary findings. PLoS ONE 2018, 13, e0200730. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wu, C.-J.; Bao, M.-L.; Zhang, J.; Wang, X.-N.; Zhang, Y.-D. Machine learning-based analysis of MR radiomics can help to improve the diagnostic performance of PI-RADS v2 in clinically relevant prostate cancer. Eur. Radiol. 2017, 27, 4082–4090. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.; Bao, M.-L.; Wu, C.-J.; Zhang, J.; Zhang, Y.-D.; Shi, H.-B. A radiomics machine learning-based redefining score robustly identifies clinically significant prostate cancer in equivocal PI-RADS score 3 lesions. Abdom. Radiol. 2020, 45, 4223–4234. [Google Scholar] [CrossRef] [PubMed]
- Sellers, J.; Wagstaff, R.G.; Helo, N.; de Riese, W.T.W. Quantitative measurements of prostatic zones by MRI and their dependence on prostate size: Possible clinical implications in prostate cancer. Ther. Adv. Urol. 2021, 13, 17562872211000852. [Google Scholar] [CrossRef] [PubMed]

| Group | A | B | C |
|---|---|---|---|
| Patients | 73 (67.6%) | 14 (13%) | 21 (19.4%) |
| Notes | positive mpMRI positive biopsy (positive target ± positive random biopsies) | positive mpMRI negative biopsy | negative mpMRI no biopsy confirmation |
| Group A (n = 73) | Group B (n = 14) | p-Value | Group C (n = 21) | |
|---|---|---|---|---|
| Age (years) | 67.7 (52.4–84) | 66.5 (54.9–75.2) | 0.8 | 64.3 (56.2–73) |
| PSA (ng/mL) | 8.2 (2.7–25) | 7.6 (3–13.2) | 0.74 | 6.3 (1.8–9.2) |
| Prostate volume (mL) | 56.6 (21–137.9) | 93.3 (44.4–182.7) | <0.01 * | 81.1 (28.7–157) |
| Group A | Group B | p Value | Group C | |
|---|---|---|---|---|
| Lesion Volume (mL) Mean, median (range) | 0.71; 0.56 (0.06–3.69) | 0.65; 0.5 (0.02–2.06) | 0.72 | 0.24; 0.2 (0.04–0.62) |
| Largest axial diameter (mm) Mean, median (range) | 14.8; 14.4 (4.6–40.9) | 13.2; 12.8 (3.9–26.7) | 0.81 | 10.1; 9.9 (5.2–16.6) |
| Sensitivity | PPV | |||
|---|---|---|---|---|
| Radiologist | Quantib | Radiologist | Quantib | |
| LOCATION | ||||
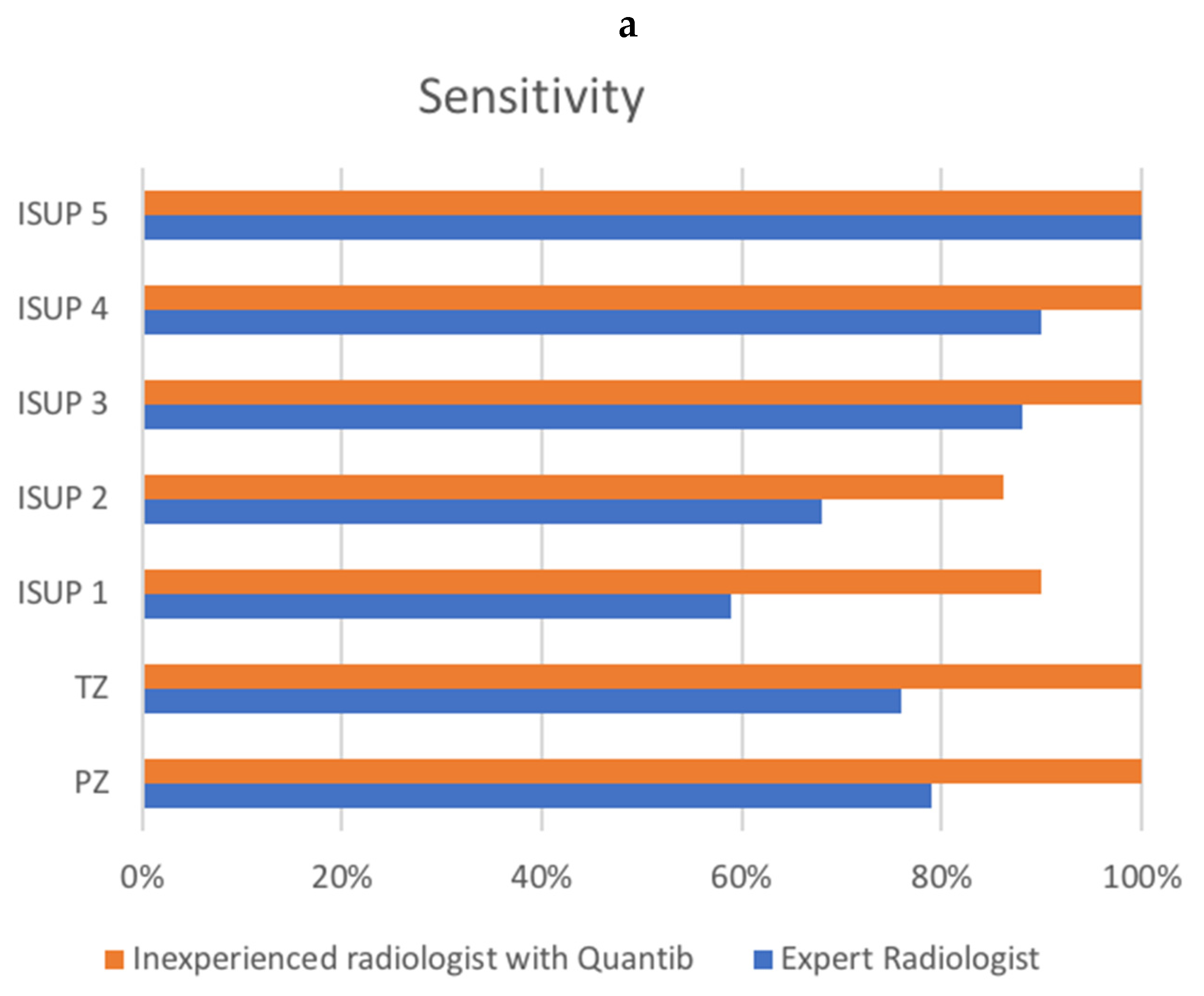
| PZ | 51/65 (78.5%) | 67/67 (100%) | 51/55 (92.7%) | 67/72 (93.1%) |
| TZ | 30/39 (76.9%) | 42/42 (100%) | 30/41 (73.2%) | 42/49 (85.7%) |
| Gleason score | ||||
| ISUP 1 | 23/39 (59%) | 36/40 (90%) | ||
| ISUP 2 | 25/37 (67.6%) | 32/37 (86.5%) | ||
| ISUP 3 | 21/24 (87.5%) | 26/26 (100%) | ||
| ISUP 4 | 9/10 (90%) | 12/12 (100%) | ||
| ISUP 5 | 3/3 (100%) | 3/3 (100%) | ||
| PIRADS | ||||
| PIRADS 3 | 10/17 (58.8%) | 1/1 (100%) | ||
| PIRADS 4 | 48/54 (88.9%) | 56/65 (86.2%) | ||
| PIRADS 5 | 23/25 (92%) | 52/55 (94.5%) | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Faiella, E.; Vertulli, D.; Esperto, F.; Cordelli, E.; Soda, P.; Muraca, R.M.; Moramarco, L.P.; Grasso, R.F.; Beomonte Zobel, B.; Santucci, D. Quantib Prostate Compared to an Expert Radiologist for the Diagnosis of Prostate Cancer on mpMRI: A Single-Center Preliminary Study. Tomography 2022, 8, 2010-2019. https://doi.org/10.3390/tomography8040168
Faiella E, Vertulli D, Esperto F, Cordelli E, Soda P, Muraca RM, Moramarco LP, Grasso RF, Beomonte Zobel B, Santucci D. Quantib Prostate Compared to an Expert Radiologist for the Diagnosis of Prostate Cancer on mpMRI: A Single-Center Preliminary Study. Tomography. 2022; 8(4):2010-2019. https://doi.org/10.3390/tomography8040168
Chicago/Turabian StyleFaiella, Eliodoro, Daniele Vertulli, Francesco Esperto, Ermanno Cordelli, Paolo Soda, Rosa Maria Muraca, Lorenzo Paolo Moramarco, Rosario Francesco Grasso, Bruno Beomonte Zobel, and Domiziana Santucci. 2022. "Quantib Prostate Compared to an Expert Radiologist for the Diagnosis of Prostate Cancer on mpMRI: A Single-Center Preliminary Study" Tomography 8, no. 4: 2010-2019. https://doi.org/10.3390/tomography8040168
APA StyleFaiella, E., Vertulli, D., Esperto, F., Cordelli, E., Soda, P., Muraca, R. M., Moramarco, L. P., Grasso, R. F., Beomonte Zobel, B., & Santucci, D. (2022). Quantib Prostate Compared to an Expert Radiologist for the Diagnosis of Prostate Cancer on mpMRI: A Single-Center Preliminary Study. Tomography, 8(4), 2010-2019. https://doi.org/10.3390/tomography8040168








