Denoising of X-ray Images Using the Adaptive Algorithm Based on the LPA-RICI Algorithm
Abstract
:1. Introduction
2. The ICI Rule and Its Modification
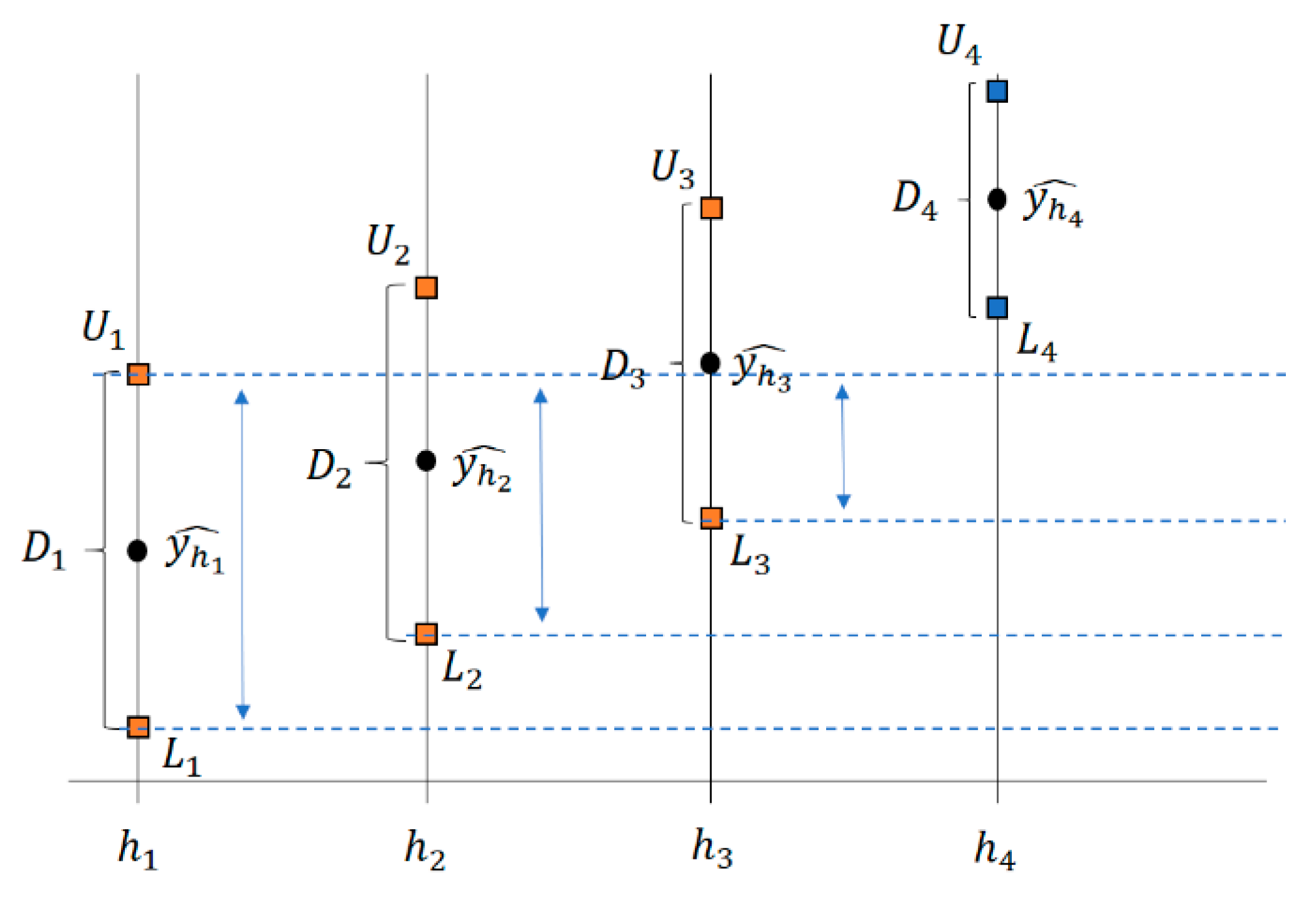
2.1. The ICI Algorithm
2.2. The RICI Algorithm
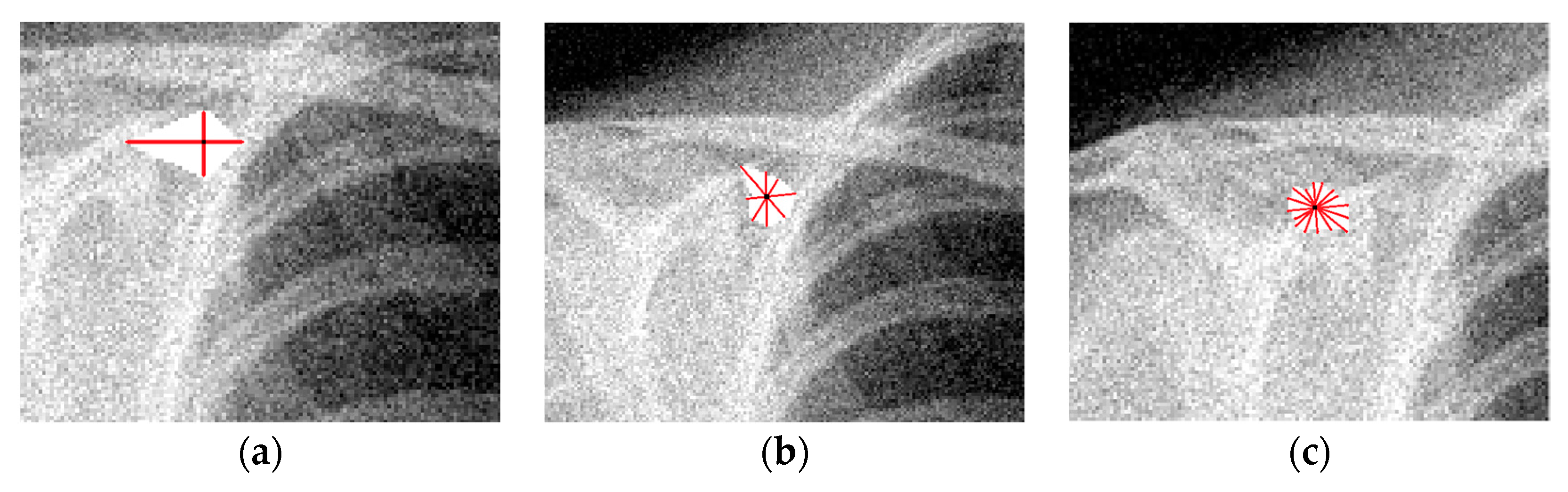
2.3. Extending the ICI and RICI Algorithm to Image Processing
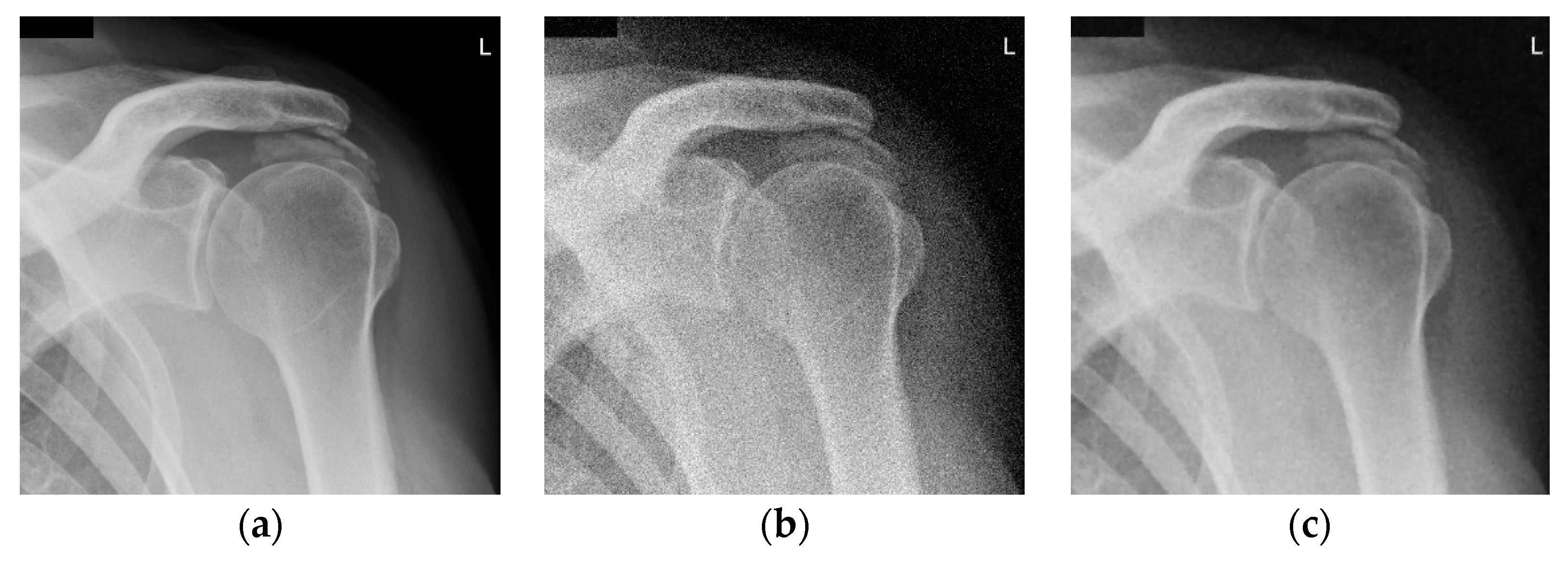
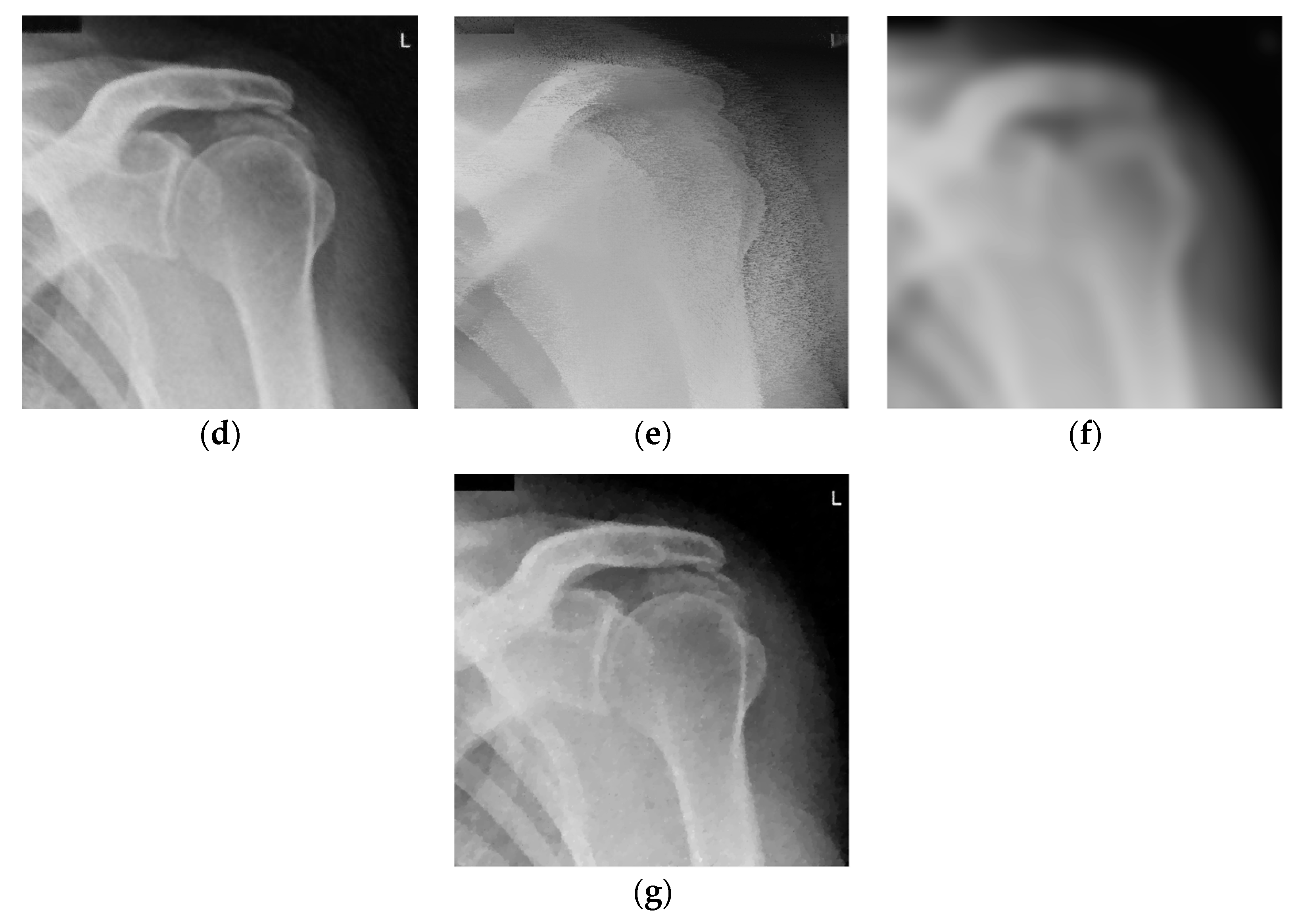
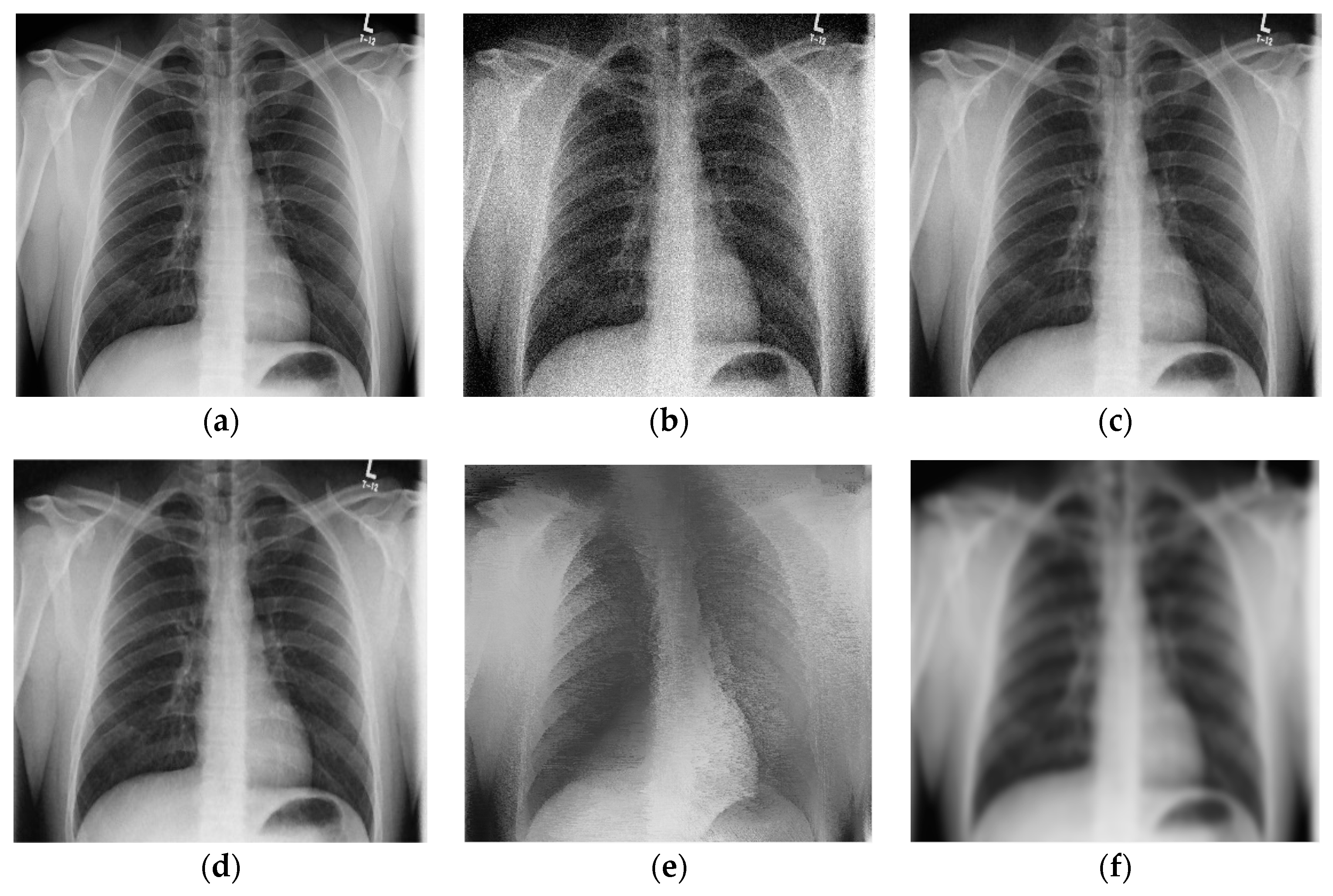
3. Results and Discussion
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Buades, A.; Coll, B.; Morel, J.M. On Image Denoising Methods. CMLA Preprint. 2004, Volume 5. Available online: http://www3.cs.stonybrook.edu/~cse577/image.smoothing.morel.pdf (accessed on 4 February 2018).
- Bansal, M.; Devi, M.; Jain, N.; Kukreja, C. A proposed approach for biomedical image denoising using PCA-NLM. Int. J. Bio-Sci. Bio-Technol. 2014, 6, 13–20. [Google Scholar]
- Samundeeswari, E.S.; Saranya, P.K.; Manavalan, R. M2 filter for speckle noise suppression in breast ultrasound images. ICTACT J. Image Video Process. 2015, 6, 1137–1144. [Google Scholar]
- Vijay, M.; Suhha, S.V.; Karthik, K. Adaptive spatial and wavelet multiscale products thresholding method for medical image denoising. In Proceedings of the International Conference on Computing, Electronics and Electrical Technologies, Kumaracoil, India, 21–22 March 2002; pp. 1109–1114. [Google Scholar]
- Ali, H.M. MRI medical image denoising by fundamental filters. SCIREA J. Comput. 2017, 12, 2–26. [Google Scholar]
- Katkovnik, V.; Egiazarian, K.; Astola, J. A spatially adaptive nonparametric regression image deblurring. IEEE Trans. Image Process. 2005, 14, 1469–1478. [Google Scholar] [CrossRef] [PubMed]
- Katkovnik, V.; Egiazarian, K.; Astola, J. Adaptive window size image de-noising based on intersection of confidence intervals (ICI) rule. J. Math. Imaging Vis. 2002, 16, 223–235. [Google Scholar] [CrossRef]
- Segon, G.; Lerga, J.; Sucic, V. Improved LPA-ICI-based estimators embedded in a signal denoising virtual instrument. Signal Image Video Process. 2017, 11, 211–218. [Google Scholar] [CrossRef]
- Katkovnik, V.; Egiazarian, K.; Astola, J. Local Approximation Techniques in Signal and Image Processing; SPIE Publications—The International Society for Optical Engineering: Washington, WA, USA, 2006. [Google Scholar]
- Lerga, J.; Vrankic, M.; Sucic, V. A signal denoising method based on the improved ICI rule. IEEE Signal Process. Lett. 2008, 15, 601–604. [Google Scholar] [CrossRef]
- Lerga, J.; Sucic, V.; Grbac, E. An adaptive method for video denoising based on the ICI rule. Eng. Rev. 2012, 32, 33–40. [Google Scholar]
- Rudin, L.I.; Osher, S.; Fatemi, E. Nonlinear total variation based noise removal algorithms. Phys. D Nonlinear Phenom. 1992, 60, 259–268. [Google Scholar] [CrossRef]

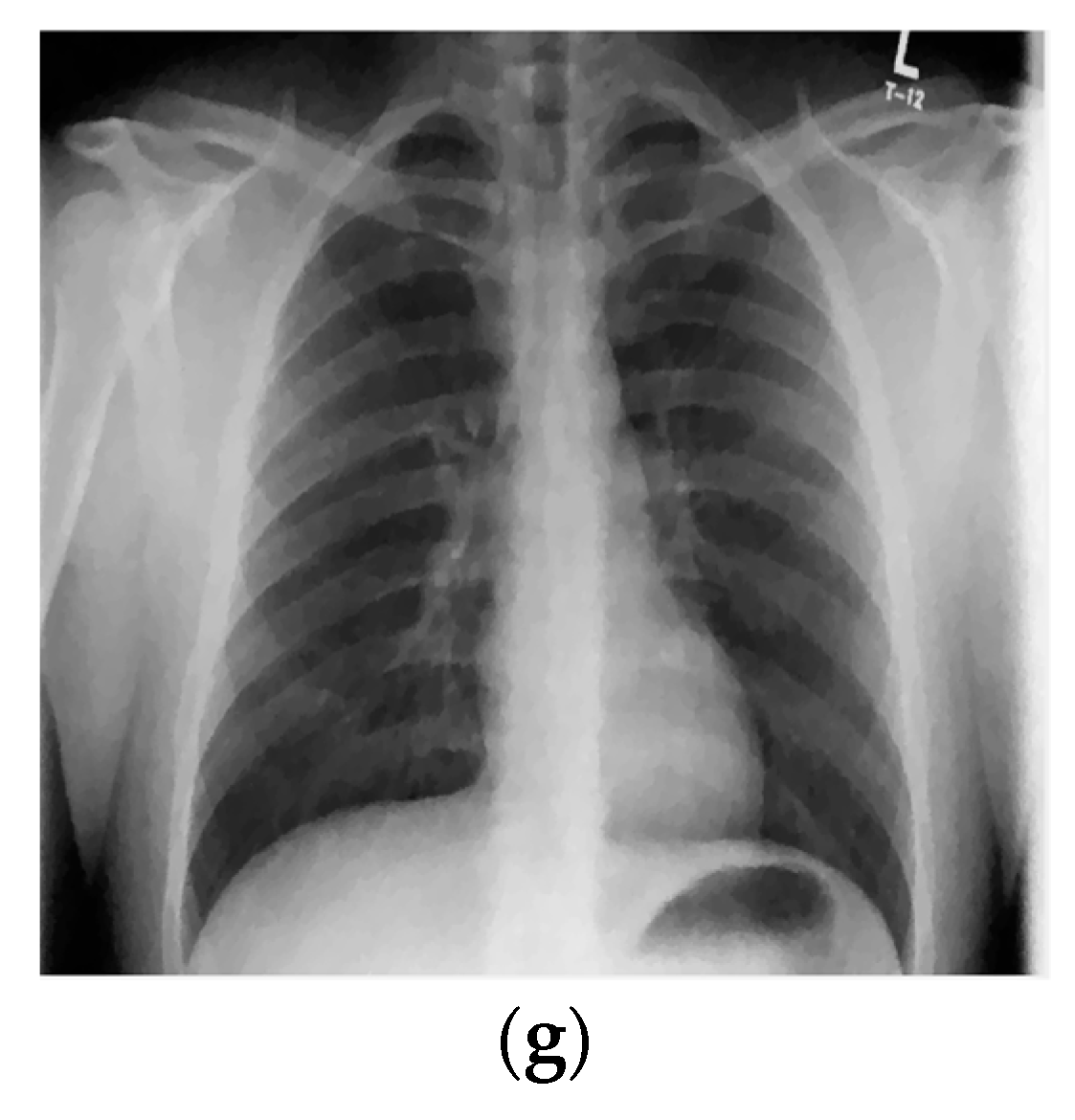
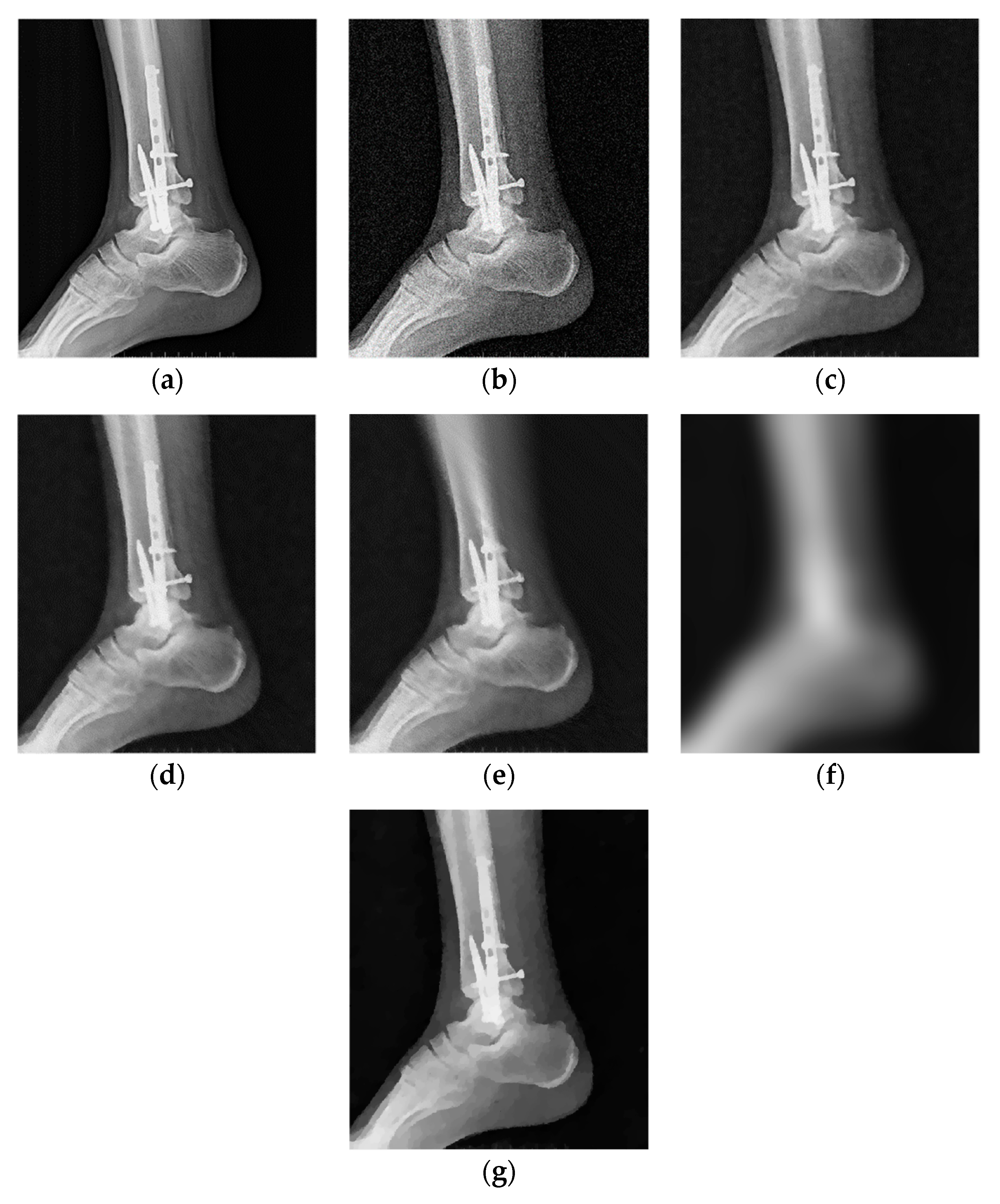
| Parameters | PSNR (dB) | |||||||
|---|---|---|---|---|---|---|---|---|
| Region | Г | Rc | σ | Noisy Image | RICI | Fixed Size Filters | Gaussian Smoothing | Total Variation |
| Quadrilateral | 1.8 | 0.8 | 20 | 29.82 | 38.73 | 33.4 | 34.69 | 35.02 |
| Octagonal | 1.8 | 0.8 | 20 | 29.82 | 39.37 | 33.4 | 34.69 | 35.02 |
| Hexadecagonal | 1.8 | 0.8 | 20 | 29.82 | 33.61 | 33.4 | 34.69 | 35.02 |
| Quadrilateral | 1.7 | 0.9 | 20 | 29.82 | 38.63 | 33.4 | 34.69 | 35.02 |
| Octagonal | 1.7 | 0.9 | 20 | 29.82 | 39.1 | 33.4 | 34.69 | 35.02 |
| Hexadecagonal | 1.7 | 0.9 | 20 | 29.82 | 33.42 | 33.4 | 34.69 | 35.02 |
| Quadrilateral | 1.6 | 0.6 | 20 | 29.82 | 38.72 | 33.4 | 34.69 | 35.02 |
| Octagonal | 1.6 | 0.6 | 20 | 29.82 | 39.19 | 33.4 | 34.69 | 35.02 |
| Hexadecagonal | 1.6 | 0.6 | 20 | 29.82 | 33.5 | 33.4 | 34.69 | 35.02 |
| Quadrilateral | 1.8 | 0.8 | 25 | 29.32 | 38.45 | 32.24 | 33.99 | 36.69 |
| Octagonal | 1.8 | 0.8 | 25 | 29.32 | 38.96 | 32.34 | 33.99 | 36.69 |
| Hexadecagonal | 1.8 | 0.8 | 25 | 29.32 | 33.51 | 32.34 | 33.99 | 36.69 |
| Quadrilateral | 1.7 | 0.9 | 25 | 29.32 | 38.17 | 32.24 | 33.99 | 36.69 |
| Octagonal | 1.7 | 0.9 | 25 | 29.32 | 38.82 | 32.24 | 33.99 | 36.69 |
| Hexadecagonal | 1.7 | 0.9 | 25 | 29.32 | 33.5 | 32.24 | 33.99 | 36.69 |
| Quadrilateral | 1.6 | 0.6 | 25 | 29.32 | 38.27 | 32.24 | 33.99 | 36.69 |
| Octagonal | 1.6 | 0.6 | 25 | 29.32 | 38.84 | 32.24 | 33.99 | 36.69 |
| Hexadecagonal | 1.6 | 0.6 | 25 | 29.32 | 33.57 | 32.24 | 33.99 | 36.69 |
| Quadrilateral | 1.8 | 0.8 | 30 | 28.99 | 38.09 | 31.5 | 33.41 | 35.62 |
| Octagonal | 1.8 | 0.8 | 30 | 28.99 | 38.76 | 31.5 | 33.41 | 35.62 |
| Hexadecagonal | 1.8 | 0.8 | 30 | 28.99 | 33.68 | 31.5 | 33.41 | 35.62 |
| Quadrilateral | 1.7 | 0.9 | 30 | 28.99 | 37.78 | 31.5 | 33.41 | 35.62 |
| Octagonal | 1.7 | 0.9 | 30 | 28.99 | 38.58 | 31.5 | 33.41 | 35.62 |
| Hexadecagonal | 1.7 | 0.9 | 30 | 28.99 | 33.4 | 31.5 | 33.41 | 35.62 |
| Quadrilateral | 1.6 | 0.6 | 30 | 28.99 | 37.82 | 31.5 | 33.41 | 35.62 |
| Octagonal | 1.6 | 0.6 | 30 | 28.99 | 39.49 | 31.5 | 33.41 | 35.62 |
| Hexadecagonal | 1.6 | 0.6 | 30 | 28.99 | 33.66 | 31.5 | 33.41 | 35.62 |
| Parameters | PSNR (dB) | ||||||
|---|---|---|---|---|---|---|---|
| Region | Г | Rc | σ | RICI vs. Noisy | RICI vs. Fixed | RICI vs. GAUSSIAN | RICI vs. Total Variation |
| Quadrilateral | 1.8 | 0.8 | 20 | 8.91 | 5.33 | 4.04 | 3.71 |
| Octagonal | 1.8 | 0.8 | 20 | 9.55 | 5.97 | 4.68 | 4.35 |
| Hexadecagonal | 1.8 | 0.8 | 20 | 3.79 | 0.21 | −1.08 | −1.41 |
| Quadrilateral | 1.7 | 0.9 | 20 | 8.81 | 5.23 | 3.94 | 3.61 |
| Octagonal | 1.7 | 0.9 | 20 | 9.28 | 5.7 | 4.41 | 4.08 |
| Hexadecagonal | 1.7 | 0.9 | 20 | 3.6 | 0.02 | −1.27 | −1.6 |
| Quadrilateral | 1.6 | 0.6 | 20 | 8.9 | 5.32 | 4.03 | 3.7 |
| Octagonal | 1.6 | 0.6 | 20 | 9.37 | 5.79 | 4.5 | 4.17 |
| Hexadecagonal | 1.6 | 0.6 | 20 | 3.68 | 0.1 | −1.19 | −1.52 |
| Quadrilateral | 1.8 | 0.8 | 25 | 9.13 | 6.21 | 4.46 | 1.76 |
| Octagonal | 1.8 | 0.8 | 25 | 9.64 | 6.62 | 4.97 | 2.27 |
| Hexadecagonal | 1.8 | 0.8 | 25 | 4.19 | 1.17 | −0.48 | −3.18 |
| Quadrilateral | 1.7 | 0.9 | 25 | 8.85 | 5.93 | 4.18 | 1.48 |
| Octagonal | 1.7 | 0.9 | 25 | 9.5 | 6.58 | 4.83 | 2.13 |
| Hexadecagonal | 1.7 | 0.9 | 25 | 4.18 | 1.26 | −0.49 | −3.19 |
| Quadrilateral | 1.6 | 0.6 | 25 | 8.95 | 6.03 | 4.28 | 1.58 |
| Octagonal | 1.6 | 0.6 | 25 | 9.52 | 6.6 | 4.85 | 2.15 |
| Hexadecagonal | 1.6 | 0.6 | 25 | 4.25 | 1.33 | −0.42 | −3.12 |
| Quadrilateral | 1.8 | 0.8 | 30 | 9.1 | 6.59 | 4.68 | 2.47 |
| Octagonal | 1.8 | 0.8 | 30 | 9.77 | 7.26 | 5.35 | 3.14 |
| Hexadecagonal | 1.8 | 0.8 | 30 | 4.69 | 2.18 | 0.27 | −1.94 |
| Quadrilateral | 1.7 | 0.9 | 30 | 8.79 | 6.28 | 4.37 | 2.16 |
| Octagonal | 1.7 | 0.9 | 30 | 9.59 | 7.08 | 5.17 | 2.96 |
| Hexadecagonal | 1.7 | 0.9 | 30 | 4.41 | 1.9 | −0.01 | −2.22 |
| Quadrilateral | 1.6 | 0.6 | 30 | 8.83 | 6.32 | 4.41 | 2.2 |
| Octagonal | 1.6 | 0.6 | 30 | 10.5 | 7.99 | 6.08 | 3.87 |
| Hexadecagonal | 1.6 | 0.6 | 30 | 4.67 | 2.16 | 0.25 | −1.96 |
| Parameters | PSNR (dB) | |||||||
|---|---|---|---|---|---|---|---|---|
| Region | Г | Rc | σ | Noisy Image | RICI | Fixed Size Filters | Gaussian Smoothing | Total Variation |
| Quadrilateral | 1.8 | 0.8 | 20 | 29.36 | 36.8 | 33.23 | 34.18 | 35.34 |
| Octagonal | 1.8 | 0.8 | 20 | 29.36 | 38.09 | 33.23 | 34.18 | 35.34 |
| Hexadecagonal | 1.8 | 0.8 | 20 | 29.36 | 33.76 | 33.23 | 34.18 | 35.34 |
| Quadrilateral | 1.7 | 0.9 | 20 | 29.36 | 36.74 | 33.23 | 34.18 | 35.34 |
| Octagonal | 1.7 | 0.9 | 20 | 29.36 | 38.23 | 33.23 | 34.18 | 35.34 |
| Hexadecagonal | 1.7 | 0.9 | 20 | 29.36 | 33.81 | 33.23 | 34.18 | 35.34 |
| Quadrilateral | 1.6 | 0.6 | 20 | 29.36 | 36.6 | 33.23 | 34.18 | 35.34 |
| Octagonal | 1.6 | 0.6 | 20 | 29.36 | 38.06 | 33.23 | 34.18 | 35.34 |
| Hexadecagonal | 1.6 | 0.6 | 20 | 29.36 | 33.3 | 33.23 | 34.18 | 35.34 |
| Quadrilateral | 1.8 | 0.8 | 25 | 28.26 | 37.28 | 31.86 | 33.76 | 36.43 |
| Octagonal | 1.8 | 0.8 | 25 | 28.26 | 38.49 | 31.86 | 33.76 | 36.43 |
| Hexadecagonal | 1.8 | 0.8 | 25 | 28.26 | 33.51 | 31.86 | 33.76 | 36.43 |
| Quadrilateral | 1.7 | 0.9 | 25 | 28.26 | 37.66 | 31.86 | 33.76 | 36.43 |
| Octagonal | 1.7 | 0.9 | 25 | 28.26 | 38.47 | 31.86 | 33.76 | 36.43 |
| Hexadecagonal | 1.7 | 0.9 | 25 | 28.26 | 32.58 | 31.86 | 33.76 | 36.43 |
| Quadrilateral | 1.6 | 0.6 | 25 | 28.26 | 37.26 | 31.86 | 33.76 | 36.43 |
| Octagonal | 1.6 | 0.6 | 25 | 28.26 | 38.48 | 31.86 | 33.76 | 36.43 |
| Hexadecagonal | 1.6 | 0.6 | 25 | 28.26 | 33.09 | 31.86 | 33.76 | 36.43 |
| Quadrilateral | 1.8 | 0.8 | 30 | 28.87 | 37.74 | 31.01 | 33.17 | 35.22 |
| Octagonal | 1.8 | 0.8 | 30 | 28.87 | 38.47 | 31.01 | 33.17 | 35.22 |
| Hexadecagonal | 1.8 | 0.8 | 30 | 28.87 | 33.01 | 31.01 | 33.17 | 35.22 |
| Quadrilateral | 1.7 | 0.9 | 30 | 28.87 | 36.96 | 31.01 | 33.17 | 35.22 |
| Octagonal | 1.7 | 0.9 | 30 | 28.87 | 38.52 | 31.01 | 33.17 | 35.22 |
| Hexadecagonal | 1.7 | 0.9 | 30 | 28.87 | 33.14 | 31.01 | 33.17 | 35.22 |
| Quadrilateral | 1.6 | 0.6 | 30 | 28.87 | 37.62 | 31.01 | 33.17 | 35.22 |
| Octagonal | 1.6 | 0.6 | 30 | 28.87 | 37.94 | 31.01 | 33.17 | 35.22 |
| Hexadecagonal | 1.6 | 0.6 | 30 | 28.87 | 32.92 | 31.01 | 33.17 | 35.22 |
| Parameters | PSNR (dB) | ||||||
|---|---|---|---|---|---|---|---|
| Region | Г | Rc | σ | RICI vs. Noisy | RICI vs. Fixed | RICI vs. Gaussian | RICI vs. Total Variation |
| Quadrilateral | 1.8 | 0.8 | 20 | 7.44 | 3.57 | 2.62 | 1.46 |
| Octagonal | 1.8 | 0.8 | 20 | 8.73 | 4.86 | 3.91 | 2.75 |
| Hexadecagonal | 1.8 | 0.8 | 20 | 4.4 | 0.53 | −0.42 | −1.58 |
| Quadrilateral | 1.7 | 0.9 | 20 | 7.38 | 3.51 | 2.56 | 1.4 |
| Octagonal | 1.7 | 0.9 | 20 | 8.87 | 5 | 4.05 | 2.89 |
| Hexadecagonal | 1.7 | 0.9 | 20 | 4.45 | 0.58 | −0.37 | −1.53 |
| Quadrilateral | 1.6 | 0.6 | 20 | 7.24 | 3.37 | 2.42 | 1.26 |
| Octagonal | 1.6 | 0.6 | 20 | 8.7 | 4.83 | 3.88 | 2.72 |
| Hexadecagonal | 1.6 | 0.6 | 20 | 3.94 | 0.07 | −0.88 | −2.04 |
| Quadrilateral | 1.8 | 0.8 | 25 | 9.02 | 5.42 | 3.52 | 0.85 |
| Octagonal | 1.8 | 0.8 | 25 | 10.23 | 6.63 | 4.73 | 2.06 |
| Hexadecagonal | 1.8 | 0.8 | 25 | 5.25 | 1.65 | −0.25 | −2.92 |
| Quadrilateral | 1.7 | 0.9 | 25 | 9.4 | 5.8 | 3.9 | 1.23 |
| Octagonal | 1.7 | 0.9 | 25 | 10.21 | 6.61 | 4.71 | 2.04 |
| Hexadecagonal | 1.7 | 0.9 | 25 | 4.32 | 0.72 | −1.18 | −3.85 |
| Quadrilateral | 1.6 | 0.6 | 25 | 9 | 5.4 | 3.5 | 0.83 |
| Octagonal | 1.6 | 0.6 | 25 | 10.22 | 6.62 | 4.72 | 2.05 |
| Hexadecagonal | 1.6 | 0.6 | 25 | 4.83 | 1.23 | −0.67 | −3.34 |
| Quadrilateral | 1.8 | 0.8 | 30 | 8.87 | 6.73 | 4.57 | 2.52 |
| Octagonal | 1.8 | 0.8 | 30 | 9.6 | 7.46 | 5.3 | 3.25 |
| Hexadecagonal | 1.8 | 0.8 | 30 | 4.14 | 2 | −0.16 | −2.21 |
| Quadrilateral | 1.7 | 0.9 | 30 | 8.09 | 5.95 | 3.79 | 1.74 |
| Octagonal | 1.7 | 0.9 | 30 | 9.65 | 7.51 | 5.35 | 3.3 |
| Hexadecagonal | 1.7 | 0.9 | 30 | 4.27 | 2.13 | −0.03 | −2.08 |
| Quadrilateral | 1.6 | 0.6 | 30 | 8.75 | 6.61 | 4.45 | 2.4 |
| Octagonal | 1.6 | 0.6 | 30 | 9.07 | 6.93 | 4.77 | 2.72 |
| Hexadecagonal | 1.6 | 0.6 | 30 | 4.05 | 1.91 | −0.25 | −2.3 |
| Parameters | PSNR (dB) | |||||||
|---|---|---|---|---|---|---|---|---|
| Region | Г | Rc | σ | Noisy Image | RICI | Fixed Size Filters | Gaussian Smoothing | Total Variation |
| Quadrilateral | 1.8 | 0.8 | 20 | 29.85 | 36.68 | 32.79 | 32.22 | 34.45 |
| Octagonal | 1.8 | 0.8 | 20 | 29.85 | 36.16 | 32.79 | 32.22 | 34.45 |
| Hexadecagonal | 1.8 | 0.8 | 20 | 29.85 | 32.92 | 32.79 | 32.22 | 34.45 |
| Quadrilateral | 1.7 | 0.9 | 20 | 29.85 | 36.97 | 32.79 | 32.22 | 34.45 |
| Octagonal | 1.7 | 0.9 | 20 | 29.85 | 36.28 | 32.79 | 32.22 | 34.45 |
| Hexadecagonal | 1.7 | 0.9 | 20 | 29.85 | 32.83 | 32.79 | 32.22 | 34.45 |
| Quadrilateral | 1.6 | 0.6 | 20 | 29.85 | 36.93 | 32.79 | 32.22 | 34.45 |
| Octagonal | 1.6 | 0.6 | 20 | 29.85 | 36.41 | 32.79 | 32.22 | 34.45 |
| Hexadecagonal | 1.6 | 0.6 | 20 | 29.85 | 32.95 | 32.79 | 32.22 | 34.45 |
| Quadrilateral | 1.8 | 0.8 | 25 | 29.37 | 37.23 | 31.96 | 31.93 | 35.61 |
| Octagonal | 1.8 | 0.8 | 25 | 29.37 | 36.57 | 31.96 | 31.93 | 35.61 |
| Hexadecagonal | 1.8 | 0.8 | 25 | 29.37 | 32.94 | 31.96 | 31.93 | 35.61 |
| Quadrilateral | 1.7 | 0.9 | 25 | 29.37 | 37.07 | 31.96 | 31.93 | 35.61 |
| Octagonal | 1.7 | 0.9 | 25 | 29.37 | 36.67 | 31.96 | 31.93 | 35.61 |
| Hexadecagonal | 1.7 | 0.9 | 25 | 29.37 | 32.98 | 31.96 | 31.93 | 35.61 |
| Quadrilateral | 1.6 | 0.6 | 25 | 29.37 | 37.03 | 31.96 | 31.93 | 35.61 |
| Octagonal | 1.6 | 0.6 | 25 | 29.37 | 36.83 | 31.96 | 31.93 | 35.61 |
| Hexadecagonal | 1.6 | 0.6 | 25 | 29.37 | 32.98 | 31.96 | 31.93 | 35.61 |
| Quadrilateral | 1.8 | 0.8 | 30 | 29.06 | 37.15 | 31.15 | 31.69 | 34.35 |
| Octagonal | 1.8 | 0.8 | 30 | 29.06 | 36.35 | 31.15 | 31.69 | 34.35 |
| Hexadecagonal | 1.8 | 0.8 | 30 | 29.06 | 32.63 | 31.15 | 31.69 | 34.35 |
| Quadrilateral | 1.7 | 0.9 | 30 | 29.06 | 36.97 | 31.15 | 31.69 | 34.35 |
| Octagonal | 1.7 | 0.9 | 30 | 29.06 | 36.28 | 31.15 | 31.69 | 34.35 |
| Hexadecagonal | 1.7 | 0.9 | 30 | 29.06 | 32.83 | 31.15 | 31.69 | 34.35 |
| Quadrilateral | 1.6 | 0.6 | 30 | 29.06 | 37.11 | 31.15 | 31.69 | 34.35 |
| Octagonal | 1.6 | 0.6 | 30 | 29.06 | 36.51 | 31.15 | 31.69 | 34.35 |
| Hexadecagonal | 1.6 | 0.6 | 30 | 29.06 | 32.99 | 31.15 | 31.69 | 34.35 |
| Parameters | PSNR (dB) | ||||||
|---|---|---|---|---|---|---|---|
| Region | Г | Rc | σ | RICI vs. Noisy | RICI vs. Fixed | RICI vs. Gaussian | RICI vs. Total Variation |
| Quadrilateral | 1.8 | 0.8 | 20 | 6.83 | 3.89 | 4.46 | 2.23 |
| Octagonal | 1.8 | 0.8 | 20 | 6.31 | 3.37 | 3.94 | 1.71 |
| Hexadecagonal | 1.8 | 0.8 | 20 | 3.07 | 0.13 | 0.70 | −1.53 |
| Quadrilateral | 1.7 | 0.9 | 20 | 7.12 | 4.18 | 4.75 | 2.52 |
| Octagonal | 1.7 | 0.9 | 20 | 6.43 | 3.49 | 4.06 | 1.83 |
| Hexadecagonal | 1.7 | 0.9 | 20 | 2.98 | 0.04 | 0.61 | −1.62 |
| Quadrilateral | 1.6 | 0.6 | 20 | 7.08 | 4.14 | 4.71 | 2.48 |
| Octagonal | 1.6 | 0.6 | 20 | 6.56 | 3.62 | 4.19 | 1.96 |
| Hexadecagonal | 1.6 | 0.6 | 20 | 3.10 | 0.16 | 0.73 | −1.50 |
| Quadrilateral | 1.8 | 0.8 | 25 | 7.86 | 5.27 | 5.30 | 1.62 |
| Octagonal | 1.8 | 0.8 | 25 | 7.20 | 4.61 | 4.64 | 0.96 |
| Hexadecagonal | 1.8 | 0.8 | 25 | 3.57 | 0.98 | 1.01 | −2.67 |
| Quadrilateral | 1.7 | 0.9 | 25 | 7.70 | 5.11 | 5.14 | 1.46 |
| Octagonal | 1.7 | 0.9 | 25 | 7.30 | 4.71 | 4.74 | 1.06 |
| Hexadecagonal | 1.7 | 0.9 | 25 | 3.61 | 1.02 | 1.05 | −2.63 |
| Quadrilateral | 1.6 | 0.6 | 25 | 7.66 | 5.07 | 5.10 | 1.42 |
| Octagonal | 1.6 | 0.6 | 25 | 7.46 | 4.87 | 4.90 | 1.22 |
| Hexadecagonal | 1.6 | 0.6 | 25 | 3.61 | 1.02 | 1.05 | −2.63 |
| Quadrilateral | 1.8 | 0.8 | 30 | 8.09 | 6.00 | 5.46 | 2.80 |
| Octagonal | 1.8 | 0.8 | 30 | 7.29 | 5.20 | 4.66 | 2.00 |
| Hexadecagonal | 1.8 | 0.8 | 30 | 3.57 | 1.48 | 0.94 | −1.72 |
| Quadrilateral | 1.7 | 0.9 | 30 | 7.91 | 5.82 | 5.28 | 2.62 |
| Octagonal | 1.7 | 0.9 | 30 | 7.22 | 5.13 | 4.59 | 1.93 |
| Hexadecagonal | 1.7 | 0.9 | 30 | 3.77 | 1.68 | 1.14 | −1.52 |
| Quadrilateral | 1.6 | 0.6 | 30 | 8.05 | 5.96 | 5.42 | 2.76 |
| Octagonal | 1.6 | 0.6 | 30 | 7.45 | 5.36 | 4.82 | 2.16 |
| Hexadecagonal | 1.6 | 0.6 | 30 | 3.93 | 1.84 | 1.30 | −1.36 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mandić, I.; Peić, H.; Lerga, J.; Štajduhar, I. Denoising of X-ray Images Using the Adaptive Algorithm Based on the LPA-RICI Algorithm. J. Imaging 2018, 4, 34. https://doi.org/10.3390/jimaging4020034
Mandić I, Peić H, Lerga J, Štajduhar I. Denoising of X-ray Images Using the Adaptive Algorithm Based on the LPA-RICI Algorithm. Journal of Imaging. 2018; 4(2):34. https://doi.org/10.3390/jimaging4020034
Chicago/Turabian StyleMandić, Ivica, Hajdi Peić, Jonatan Lerga, and Ivan Štajduhar. 2018. "Denoising of X-ray Images Using the Adaptive Algorithm Based on the LPA-RICI Algorithm" Journal of Imaging 4, no. 2: 34. https://doi.org/10.3390/jimaging4020034
APA StyleMandić, I., Peić, H., Lerga, J., & Štajduhar, I. (2018). Denoising of X-ray Images Using the Adaptive Algorithm Based on the LPA-RICI Algorithm. Journal of Imaging, 4(2), 34. https://doi.org/10.3390/jimaging4020034






