Improving Tomographic Reconstruction from Limited Data Using Mixed-Scale Dense Convolutional Neural Networks
Abstract
1. Introduction
2. Materials and Methods
2.1. Problem Definition
2.2. Tomographic Reconstruction
2.3. Deep Neural Networks for Improving Reconstructed Images
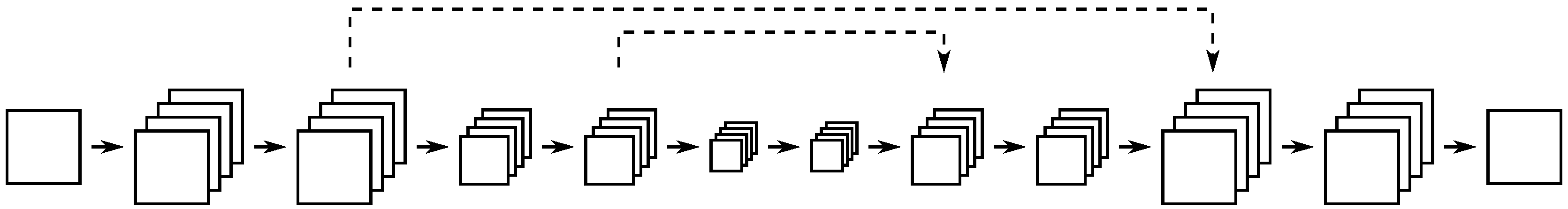
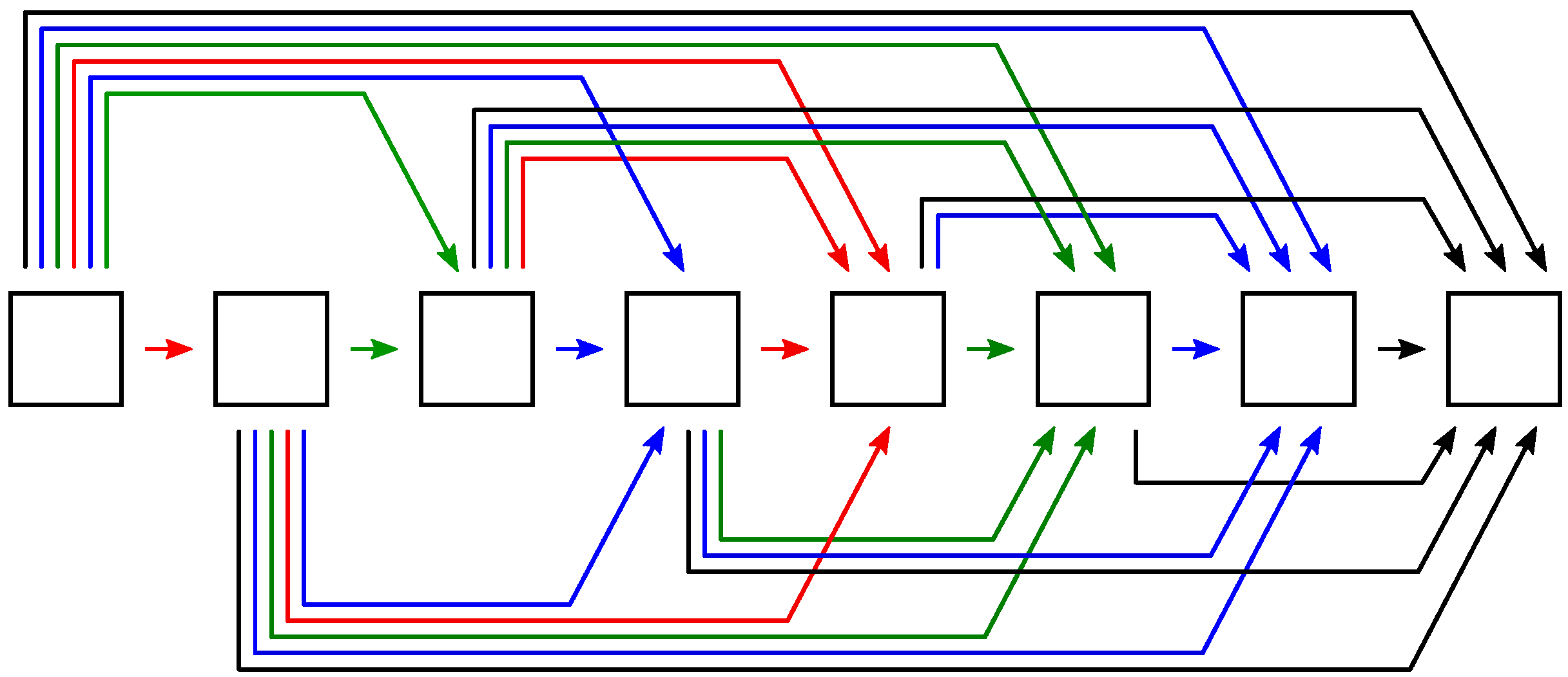
2.4. Mixed-Scale Dense Convolutional Neural Networks
3. Results and Discussion
3.1. Setup
3.2. Simulations
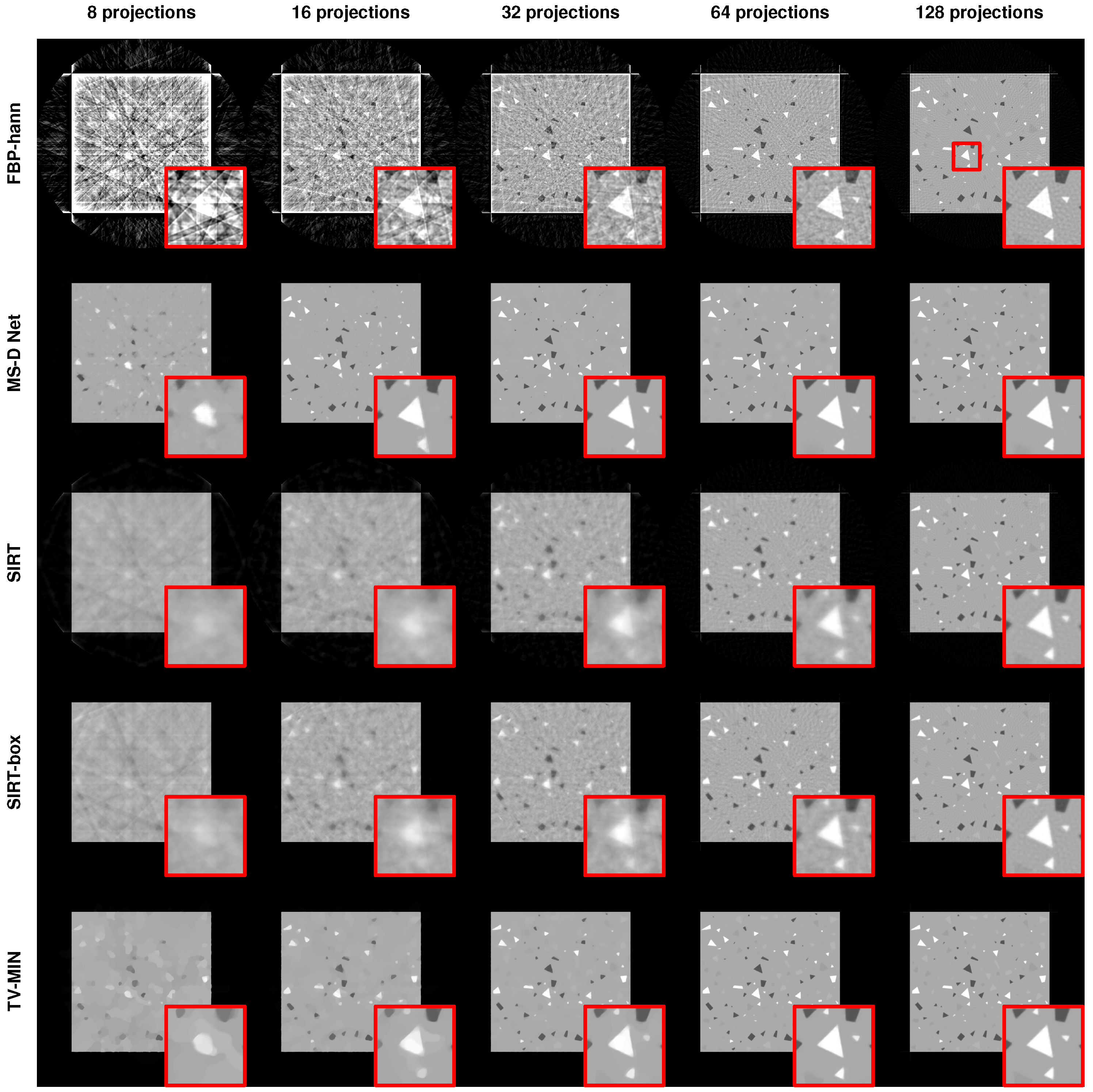
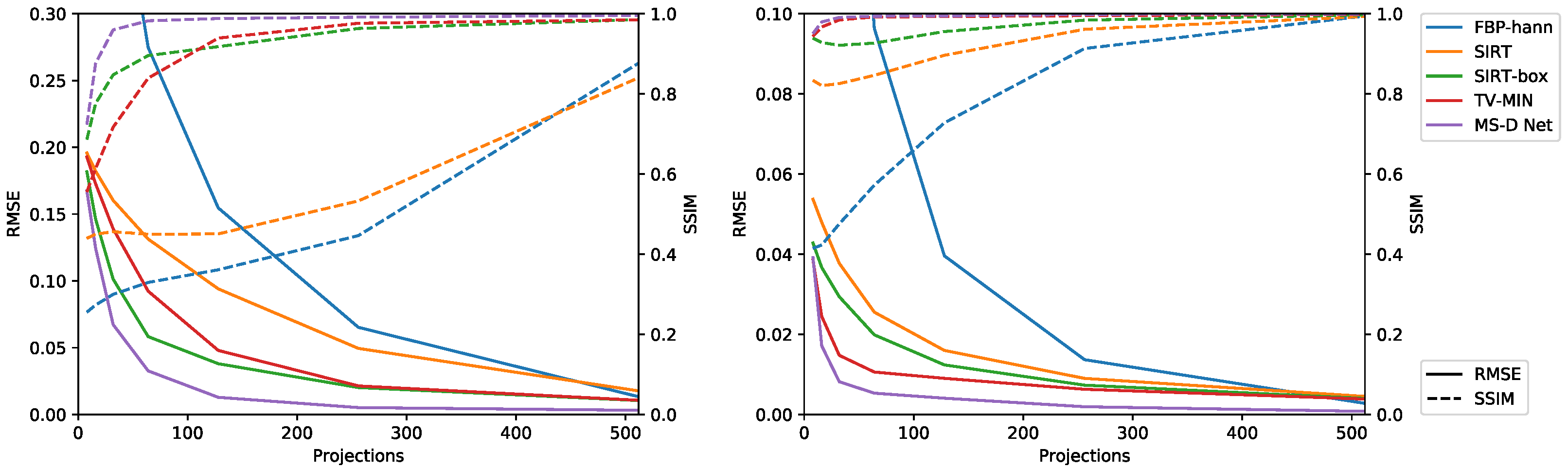
3.2.1. Limited Number of Projections
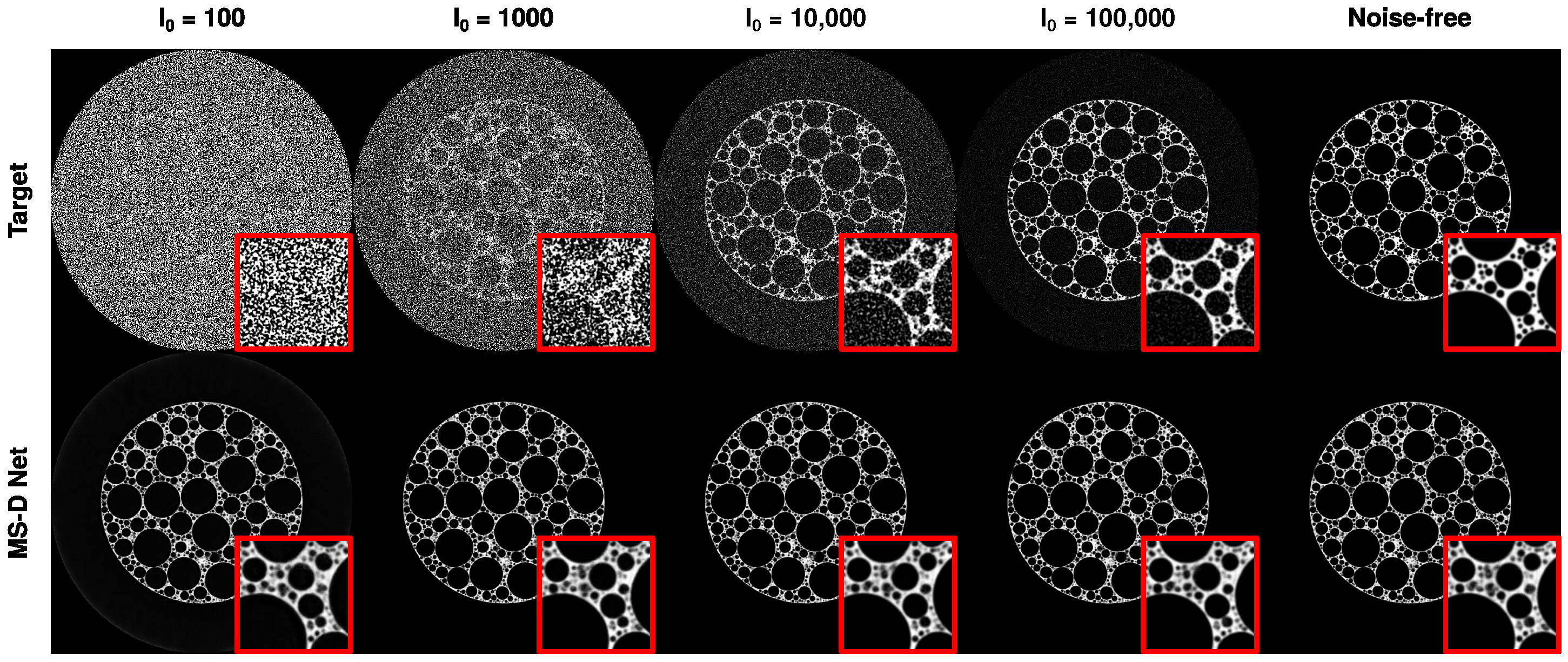
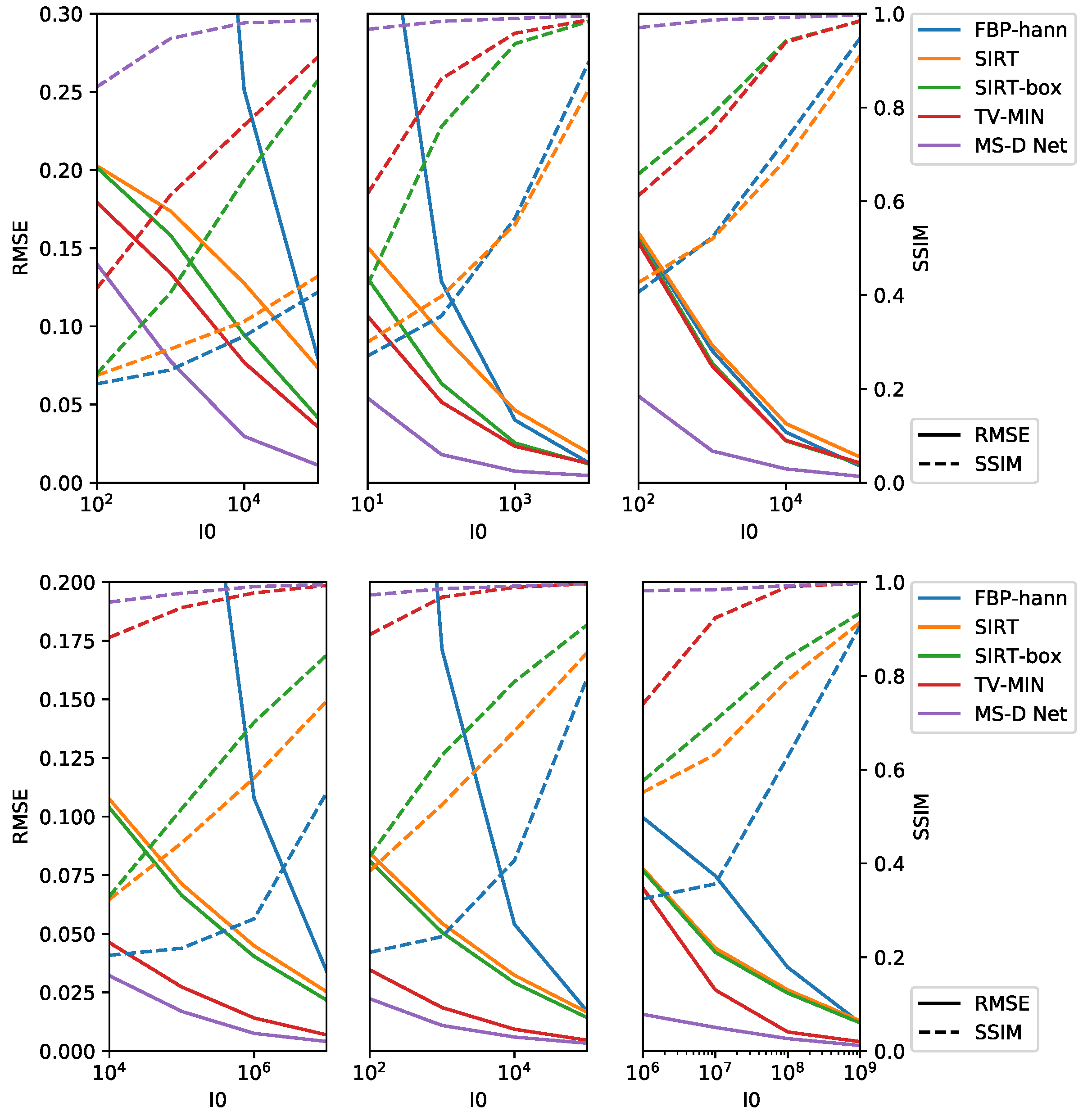
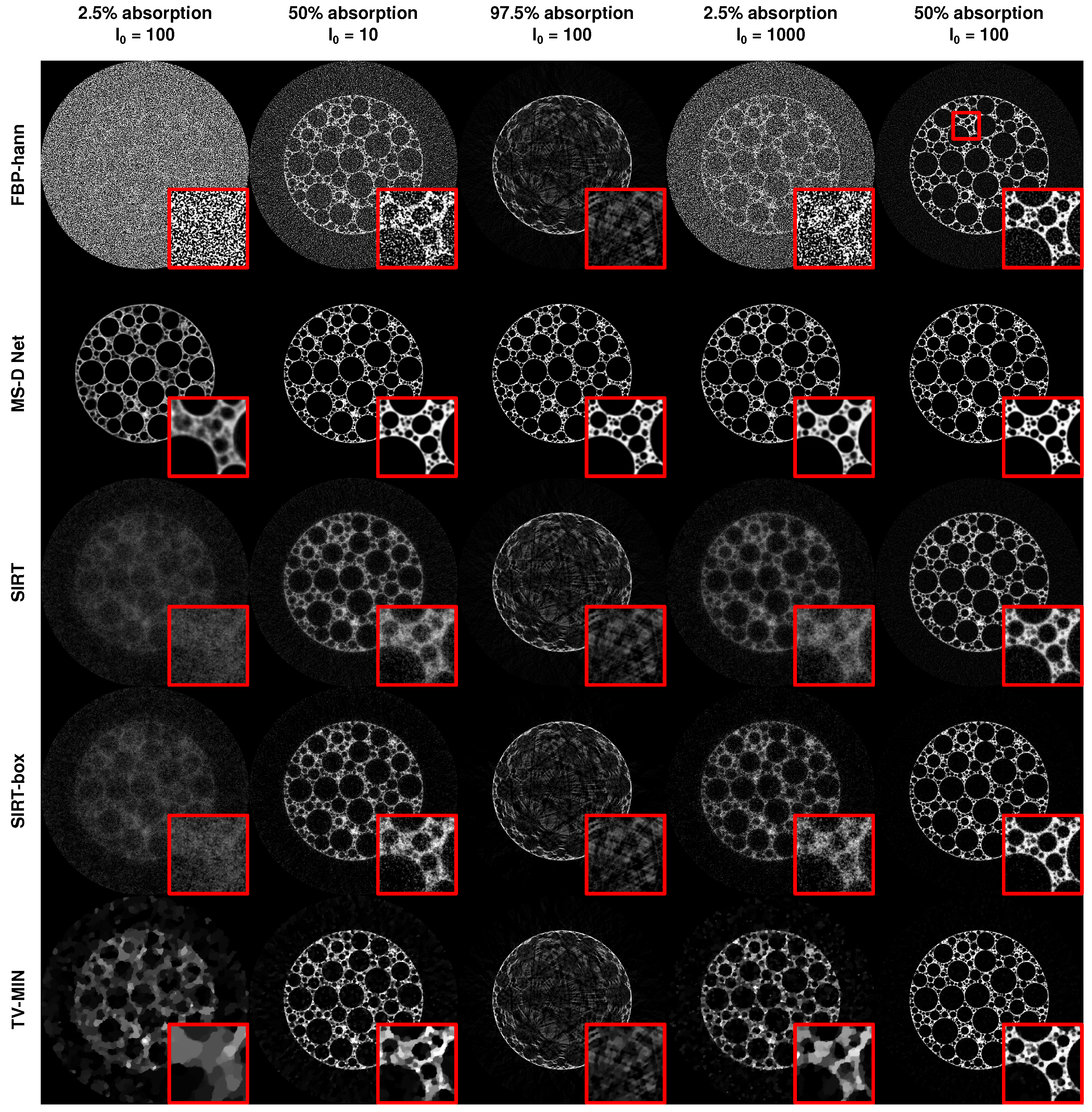
3.2.2. Limited Exposure Time
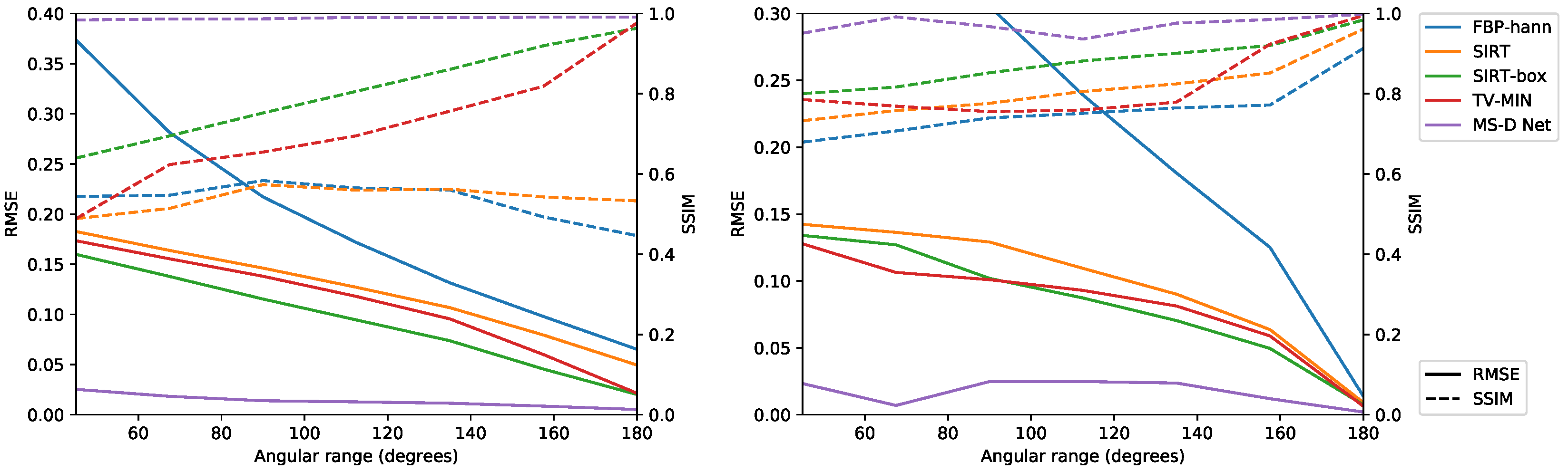
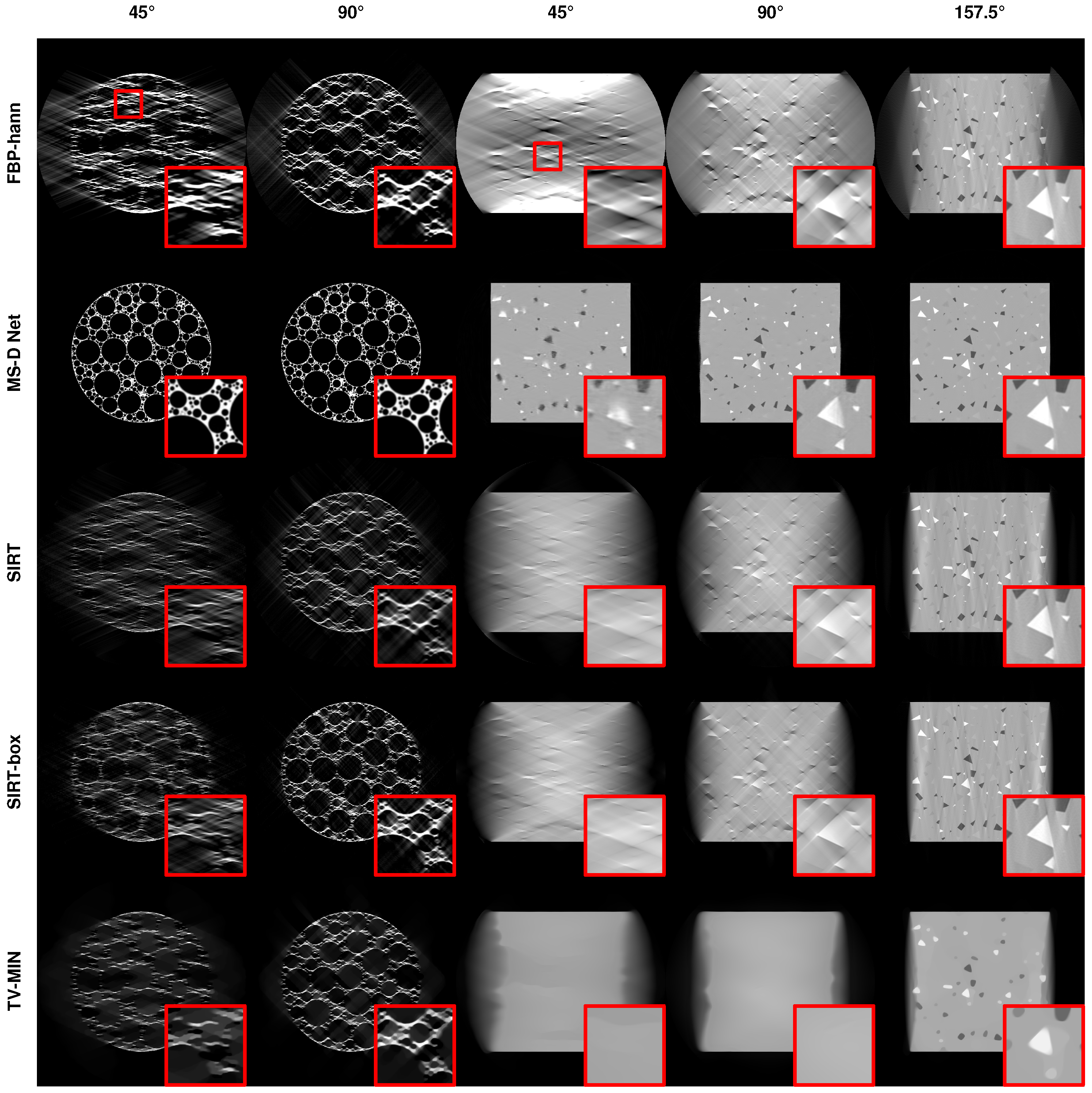
3.2.3. Limited Angular Range
3.2.4. Quality of Training Images
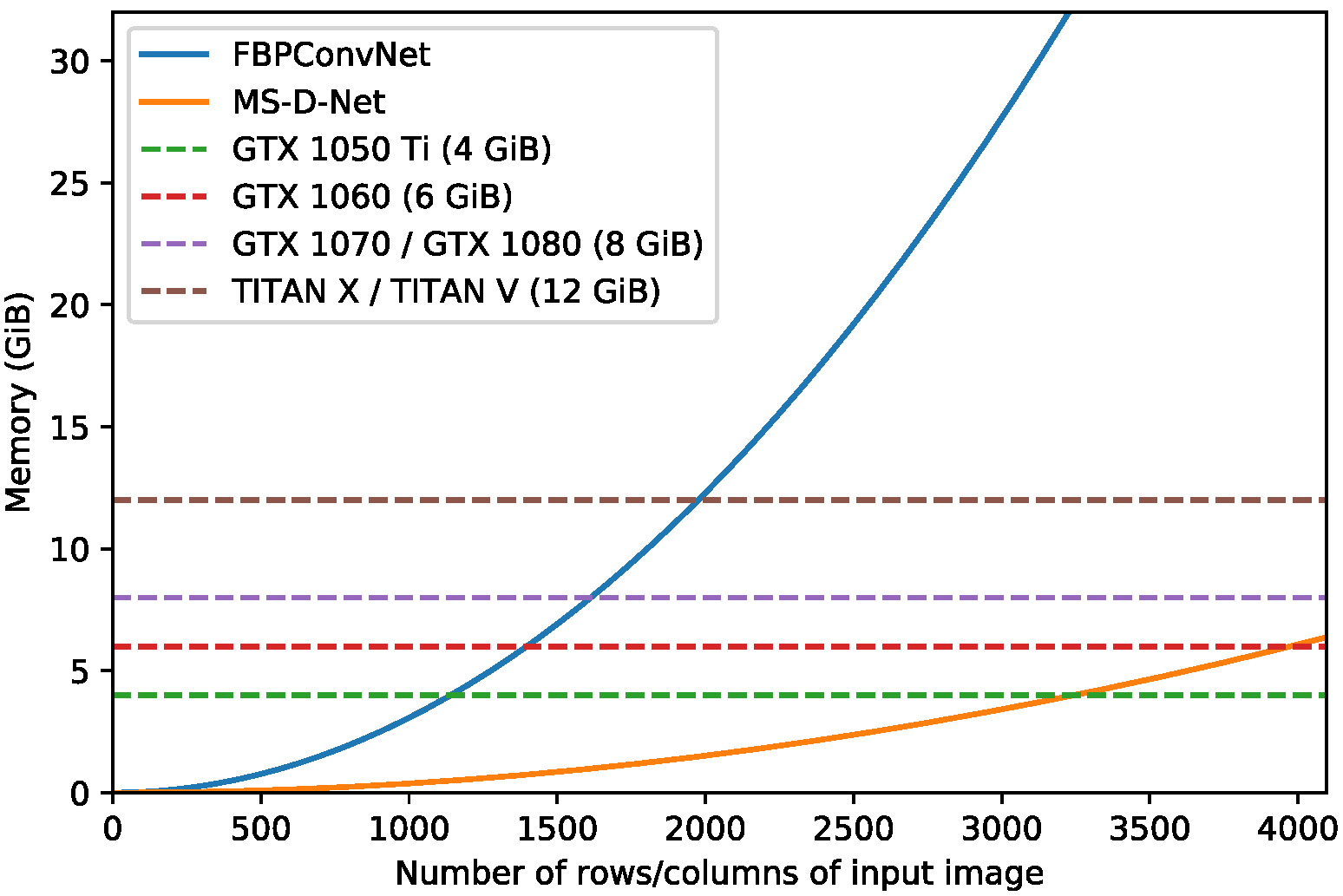
3.2.5. Comparison with Other Networks
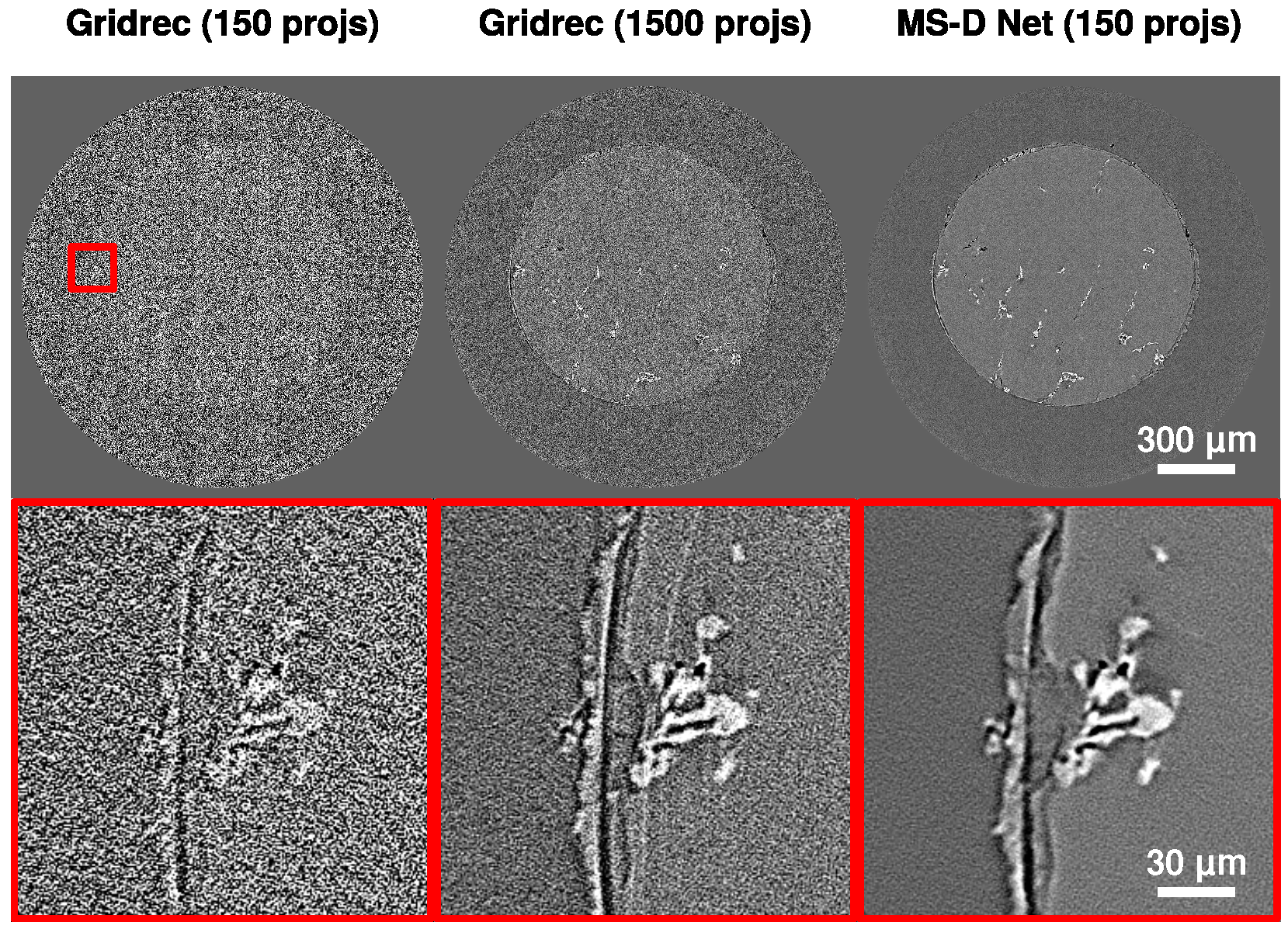
3.3. Experimental Data
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Appendix A


References
- Donoghue, P.C.; Bengtson, S.; Dong, X.P.; Gostling, N.J.; Huldtgren, T.; Cunningham, J.A.; Yin, C.; Yue, Z.; Peng, F.; Stampanoni, M. Synchrotron X-ray tomographic microscopy of fossil embryos. Nature 2006, 442, 680–683. [Google Scholar] [CrossRef] [PubMed]
- Metscher, B.D. MicroCT for comparative morphology: Simple staining methods allow high-contrast 3D imaging of diverse non-mineralized animal tissues. BMC Physiol. 2009, 9, 11. [Google Scholar] [CrossRef] [PubMed]
- Midgley, P.A.; Dunin-Borkowski, R.E. Electron tomography and holography in materials science. Nat. Mater. 2009, 8, 271. [Google Scholar] [CrossRef] [PubMed]
- Kak, A.C.; Slaney, M. Principles of Computerized Tomographic Imaging; Society of Industrial and Applied Mathematics: Philadelphia, PA, USA, 2001. [Google Scholar]
- Beck, A.; Teboulle, M. Fast gradient-based algorithms for constrained total variation image denoising and deblurring problems. IEEE Trans. Image Process. 2009, 18, 2419–2434. [Google Scholar] [CrossRef] [PubMed]
- Lovric, G.; Barré, S.F.; Schittny, J.C.; Roth-Kleiner, M.; Stampanoni, M.; Mokso, R. Dose optimization approach to fast X-ray microtomography of the lung alveoli. J. Appl. Crystallogr. 2013, 46, 856–860. [Google Scholar] [CrossRef] [PubMed]
- Sipila, H. Moving-object computer tomography for luggage inspection. In Proceedings of the International Society for Optics and Photonics, Applications of Signal and Image Processing in Explosives Detection Systems, Boston, MA, USA, 1 April 1993; Volume 1824, pp. 39–41. [Google Scholar]
- Xiao, X.; Liu, H.; Wang, L.; De Carlo, F. Density measurement of samples under high pressure using synchrotron microtomography and diamond anvil cell techniques. J. Synchrotron Radiat. 2010, 17, 360–366. [Google Scholar] [CrossRef] [PubMed]
- Ritschl, L.; Bergner, F.; Fleischmann, C.; Kachelrieß, M. Improved total variation-based CT image reconstruction applied to clinical data. Phys. Med. Biol. 2011, 56, 1545. [Google Scholar] [CrossRef] [PubMed]
- Bicer, T.; Gürsoy, D.; De Andrade, V.; Kettimuthu, R.; Scullin, W.; De Carlo, F.; Foster, I.T. Trace: A high-throughput tomographic reconstruction engine for large-scale datasets. Adv. Struct. Chem. Imaging 2017, 3, 6. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Delving deep into rectifiers: Surpassing human-level performance on imagenet classification. In Proceedings of the IEEE International Conference on Computer Vision, Santiago, Chile, 7–13 December 2015; pp. 1026–1034. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015; Springer: Berlin, Germany, 2015; pp. 234–241. [Google Scholar]
- Dong, C.; Loy, C.C.; He, K.; Tang, X. Image super-resolution using deep convolutional networks. IEEE Trans. Pattern Anal. Mach. Intell. 2016, 38, 295–307. [Google Scholar] [CrossRef] [PubMed]
- Pelt, D.M.; Batenburg, K.J. Fast Tomographic Reconstruction From Limited Data Using Artificial Neural Networks. IEEE Trans. Image Process. 2013, 22, 5238–5251. [Google Scholar] [CrossRef] [PubMed]
- Bladt, E.; Pelt, D.M.; Bals, S.; Batenburg, K.J. Electron tomography based on highly limited data using a neural network reconstruction technique. Ultramicroscopy 2015, 158, 81–88. [Google Scholar] [CrossRef] [PubMed]
- Boublil, D.; Elad, M.; Shtok, J.; Zibulevsky, M. Spatially-adaptive reconstruction in computed tomography using neural networks. IEEE Trans. Med. Imaging 2015, 34, 1474–1485. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; De Carlo, F.; Phatak, C.; Gürsoy, D. A convolutional neural network approach to calibrating the rotation axis for X-ray computed tomography. J. Synchrotron Radiat. 2017, 24, 469–475. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Zhang, Y.; Zhang, W.; Liao, P.; Li, K.; Zhou, J.; Wang, G. Low-dose CT via convolutional neural network. Biomed. Opt. Express 2017, 8, 679–694. [Google Scholar] [CrossRef] [PubMed]
- Jin, K.H.; McCann, M.T.; Froustey, E.; Unser, M. Deep Convolutional Neural Network for Inverse Problems in Imaging. IEEE Trans. Image Process. 2017, 26, 4509–4522. [Google Scholar] [CrossRef] [PubMed]
- Matos, F.A.; Ferreira, D.R.; Carvalho, P.J.; Contributors, J. Deep learning for plasma tomography using the bolometer system at JET. Fusion Eng. Des. 2017, 114, 18–25. [Google Scholar] [CrossRef]
- Yang, X.; De Andrade, V.; Scullin, W.; Dyer, E.L.; Kasthuri, N.; De Carlo, F.; Gürsoy, D. Low-dose X-ray tomography through a deep convolutional neural network. Sci. Rep. 2018, 8, 2575. [Google Scholar] [CrossRef] [PubMed]
- Pelt, D.M.; Sethian, J.A. A mixed-scale dense convolutional neural network for image analysis. Proc. Natl. Acad. Sci. USA 2018, 115, 254–259. [Google Scholar] [CrossRef] [PubMed]
- Adler, J.; Öktem, O. Learned primal-dual reconstruction. IEEE Trans. Med. Imaging 2018, 37, 1322–1332. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 2014, 15, 1929–1958. [Google Scholar]
- Farquhar, T.; Chatziioannou, A.; Chinn, G.; Dahlbom, M.; Hoffman, E. An investigation of filter choice for filtered back-projection reconstruction in PET. IEEE Trans. Nucl. Sci. 1998, 45, 1133–1137. [Google Scholar] [CrossRef]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning; MIT Press: Cambridge, MA, USA, 2016. [Google Scholar]
- Pan, X.; Sidky, E.Y.; Vannier, M. Why do commercial CT scanners still employ traditional, filtered back-projection for image reconstruction? Inverse Probl. 2009, 25, 123009. [Google Scholar] [CrossRef] [PubMed]
- Nair, V.; Hinton, G.E. Rectified linear units improve restricted boltzmann machines. In Proceedings of the 27th International Conference on Machine Learning (ICML-10), Haifa, Israel, 21–24 June 2010; pp. 807–814. [Google Scholar]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar]
- Yu, F.; Koltun, V. Multi-scale context aggregation by dilated convolutions. arXiv, 2015; arXiv:1511.07122. [Google Scholar]
- Klöckner, A.; Pinto, N.; Lee, Y.; Catanzaro, B.; Ivanov, P.; Fasih, A. PyCUDA and PyOpenCL: A Scripting-Based Approach to GPU Run-Time Code Generation. Parallel Comput. 2012, 38, 157–174. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, L. ADAM: A method for stochastic optimization. In Proceedings of the International Conference on Learning Representations, San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- van Aarle, W.; Palenstijn, W.J.; De Beenhouwer, J.; Altantzis, T.; Bals, S.; Batenburg, K.J.; Sijbers, J. The ASTRA Toolbox: A platform for advanced algorithm development in electron tomography. Ultramicroscopy 2015, 157, 35–47. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Bovik, A.C.; Sheikh, H.R.; Simoncelli, E.P. Image quality assessment: From error visibility to structural similarity. IEEE Trans. Image Process. 2004, 13, 600–612. [Google Scholar] [CrossRef] [PubMed]
- Abadi, M.; Barham, P.; Chen, J.; Chen, Z.; Davis, A.; Dean, J.; Devin, M.; Ghemawat, S.; Irving, G.; Isard, M.; et al. Tensorflow: A system for large-scale machine learning. In Proceedings of the 12th USENIX Symposium on Operating Systems Design and Implementation (OSDI), Savannah, GA, USA, 2–4 November 2016; Volume 16, pp. 265–283. [Google Scholar]
- De Carlo, F.; Gürsoy, D.; Ching, D.J.; Batenburg, K.J.; Ludwig, W.; Mancini, L.; Marone, F.; Mokso, R.; Pelt, D.M.; Sijbers, J.; et al. TomoBank: A tomographic data repository for computational X-ray science. Meas. Sci. Technol. 2018, 29, 034004. [Google Scholar] [CrossRef]
- Gürsoy, D.; De Carlo, F.; Xiao, X.; Jacobsen, C. TomoPy: A framework for the analysis of synchrotron tomographic data. J. Synchrotron Radiat. 2014, 21, 1188–1193. [Google Scholar] [CrossRef] [PubMed]
- Marone, F.; Stampanoni, M. Regridding reconstruction algorithm for real-time tomographic imaging. J. Synchrotron Radiat. 2012, 19, 1029–1037. [Google Scholar] [CrossRef] [PubMed]
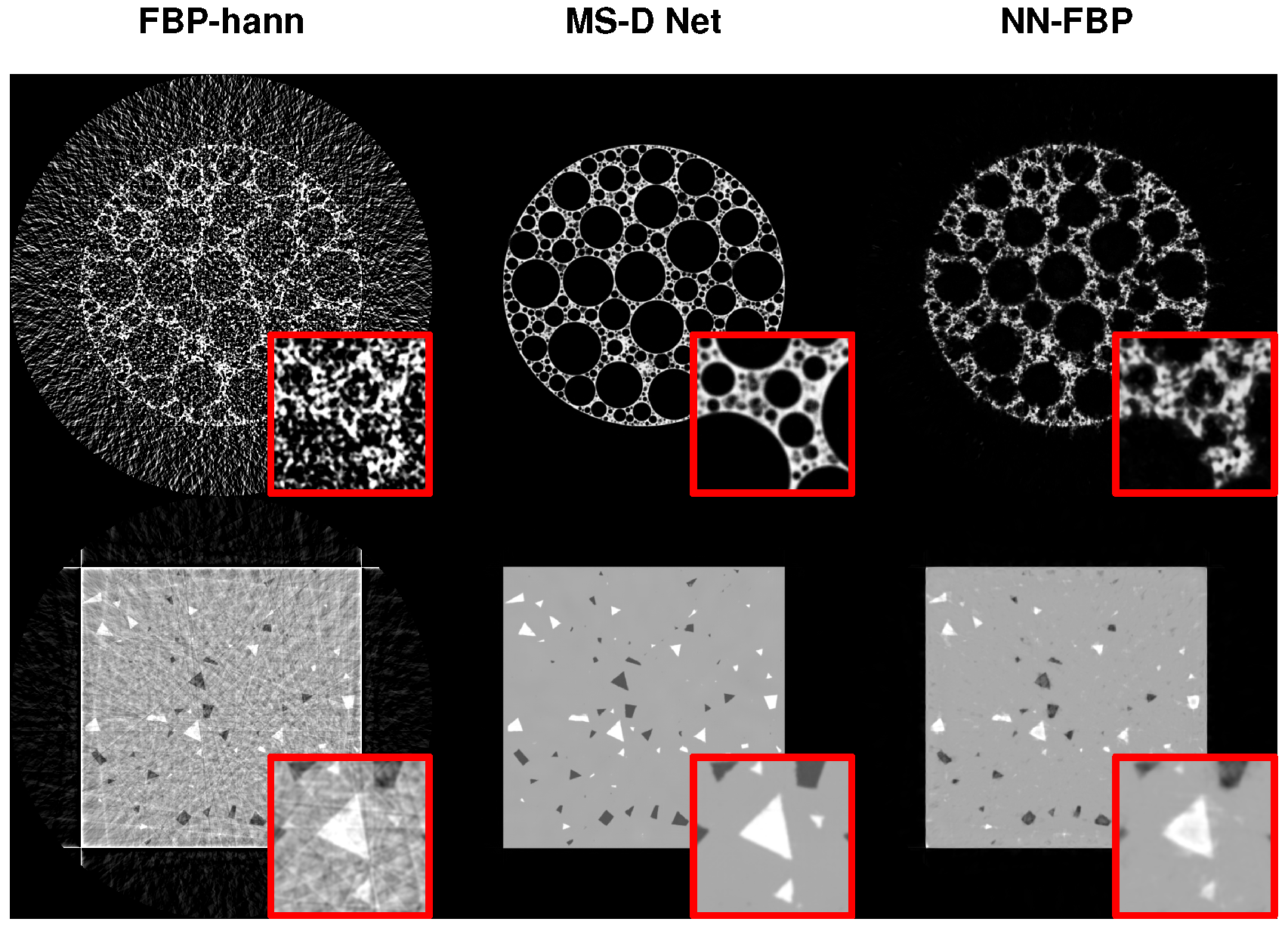




| Method | Metric | 16 Projections | 45° Range | Noisy Target | Noisy Input | |||
|---|---|---|---|---|---|---|---|---|
| Foam | Rock | Foam | Rock | Foam | Rock | Foam | ||
| FBPConvNet | RMSE | 0.110 | 0.017 | 0.046 | 0.012 | 0.072 | 0.011 | 0.103 |
| SSIM | 0.734 | 0.963 | 0.982 | 0.981 | 0.630 | 0.920 | 0.782 | |
| MS-D-Net | RMSE | 0.100 | 0.019 | 0.039 | 0.013 | 0.065 | 0.011 | 0.099 |
| SSIM | 0.910 | 0.957 | 0.985 | 0.980 | 0.726 | 0.930 | 0.907 | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pelt, D.M.; Batenburg, K.J.; Sethian, J.A. Improving Tomographic Reconstruction from Limited Data Using Mixed-Scale Dense Convolutional Neural Networks. J. Imaging 2018, 4, 128. https://doi.org/10.3390/jimaging4110128
Pelt DM, Batenburg KJ, Sethian JA. Improving Tomographic Reconstruction from Limited Data Using Mixed-Scale Dense Convolutional Neural Networks. Journal of Imaging. 2018; 4(11):128. https://doi.org/10.3390/jimaging4110128
Chicago/Turabian StylePelt, Daniël M., Kees Joost Batenburg, and James A. Sethian. 2018. "Improving Tomographic Reconstruction from Limited Data Using Mixed-Scale Dense Convolutional Neural Networks" Journal of Imaging 4, no. 11: 128. https://doi.org/10.3390/jimaging4110128
APA StylePelt, D. M., Batenburg, K. J., & Sethian, J. A. (2018). Improving Tomographic Reconstruction from Limited Data Using Mixed-Scale Dense Convolutional Neural Networks. Journal of Imaging, 4(11), 128. https://doi.org/10.3390/jimaging4110128





