The Putative Role of the NAC Transcription Factor EjNACL47 in Cell Enlargement of Loquat (Eriobotrya japonica Lindl.)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. EjNACL47 Cloning
2.3. Bioinformatics Analysis
2.4. Real-Time qRT-PCR Analysis
2.5. Subcellular Localization
2.6. Transcriptional Activity Assay
2.7. Genetic Transformation in Arabidopsis
2.8. β-Glucuronidase (GUS) Histochemical Identification and Cell-Area Statistics
3. Results
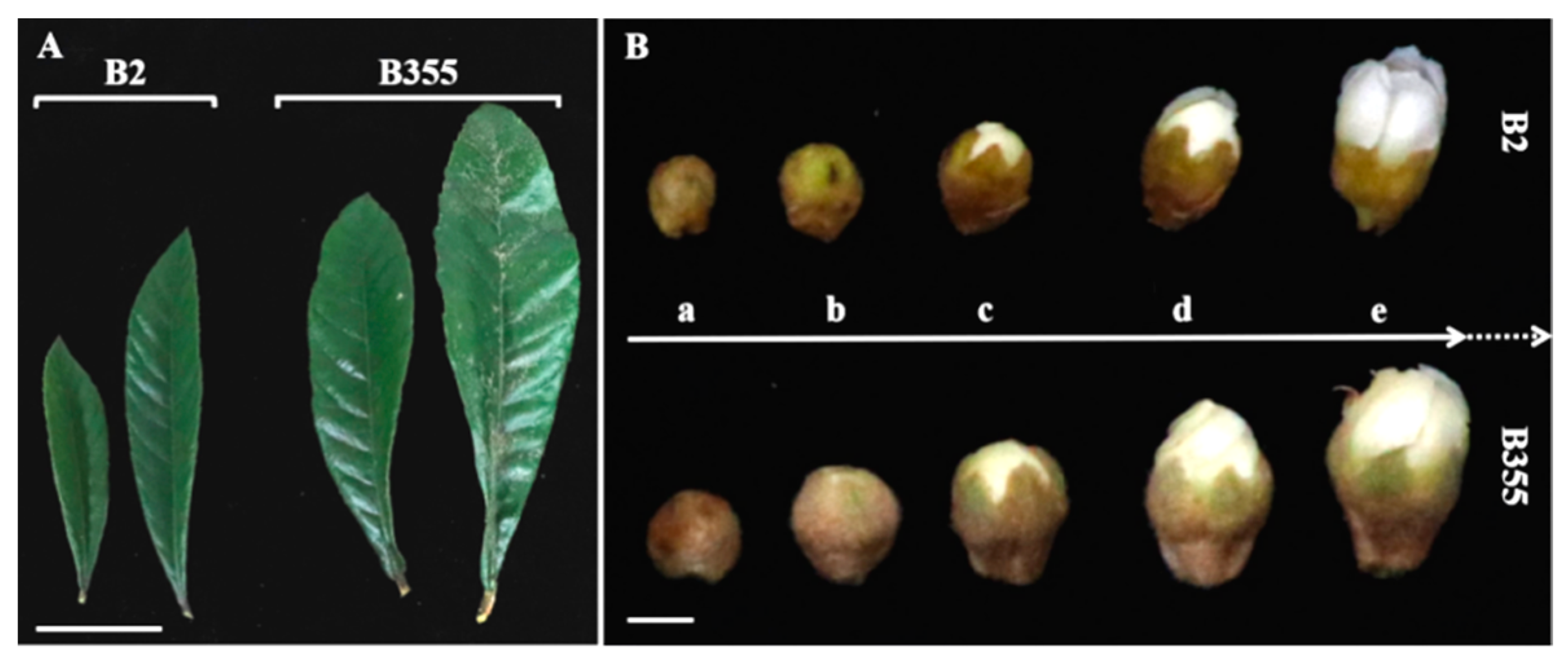
3.1. EjNACL47 Expression Is Positively Correlated with Organ Size
3.2. EjNACL47 Isolation and Sequence Analysis
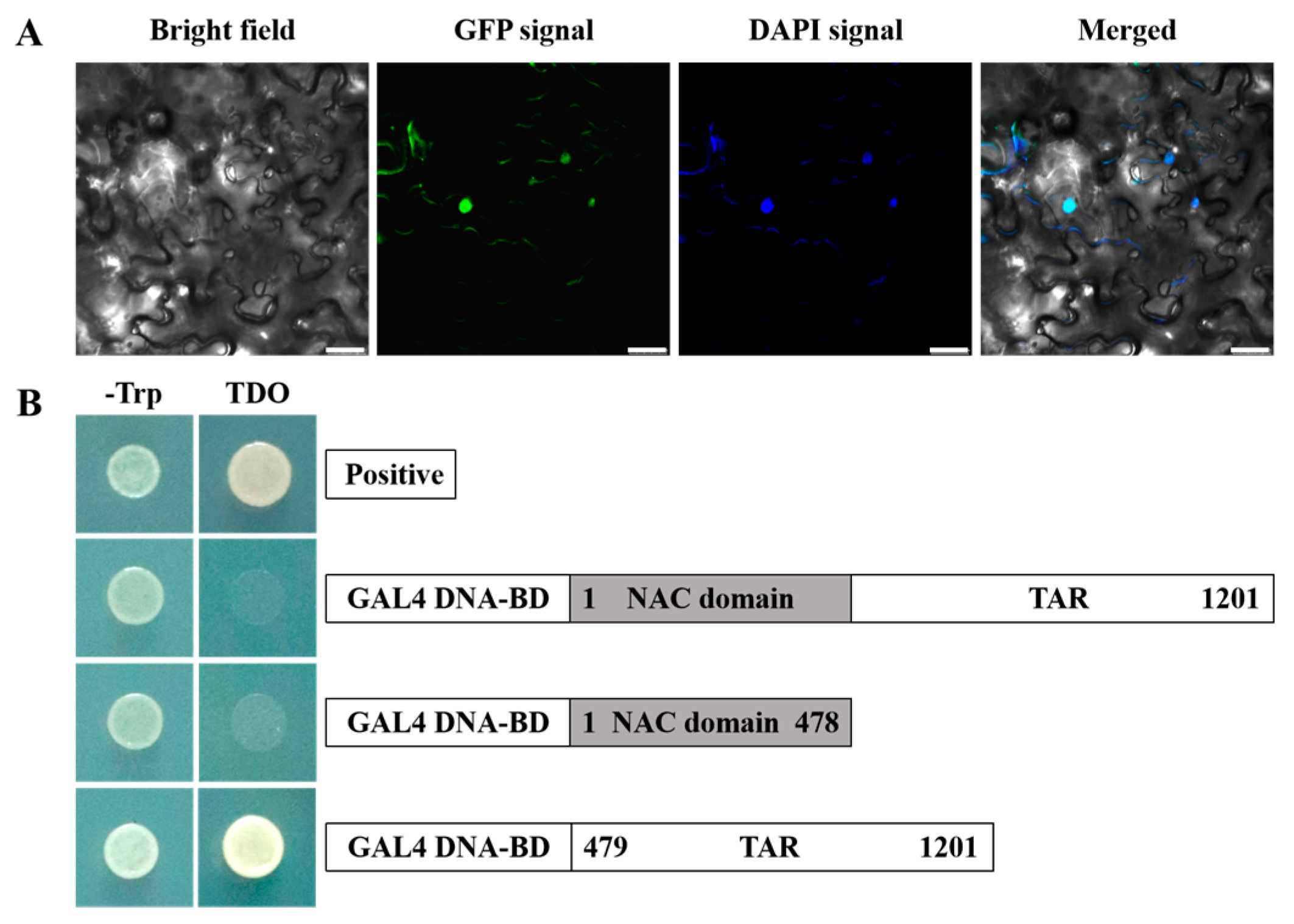
3.3. Subcellular Localization of EjNACL47
3.4. EjNACL47 Transcriptional Activity
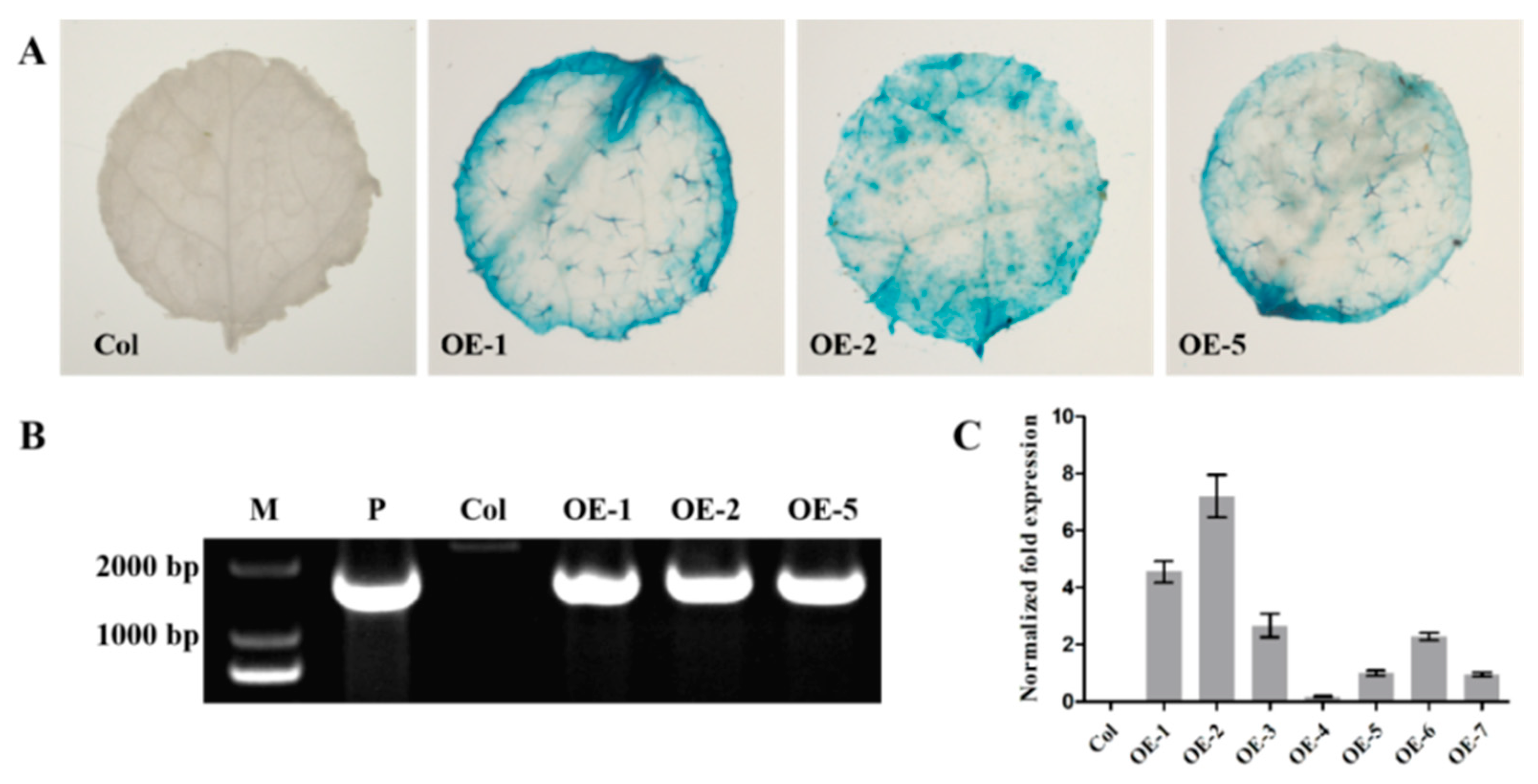
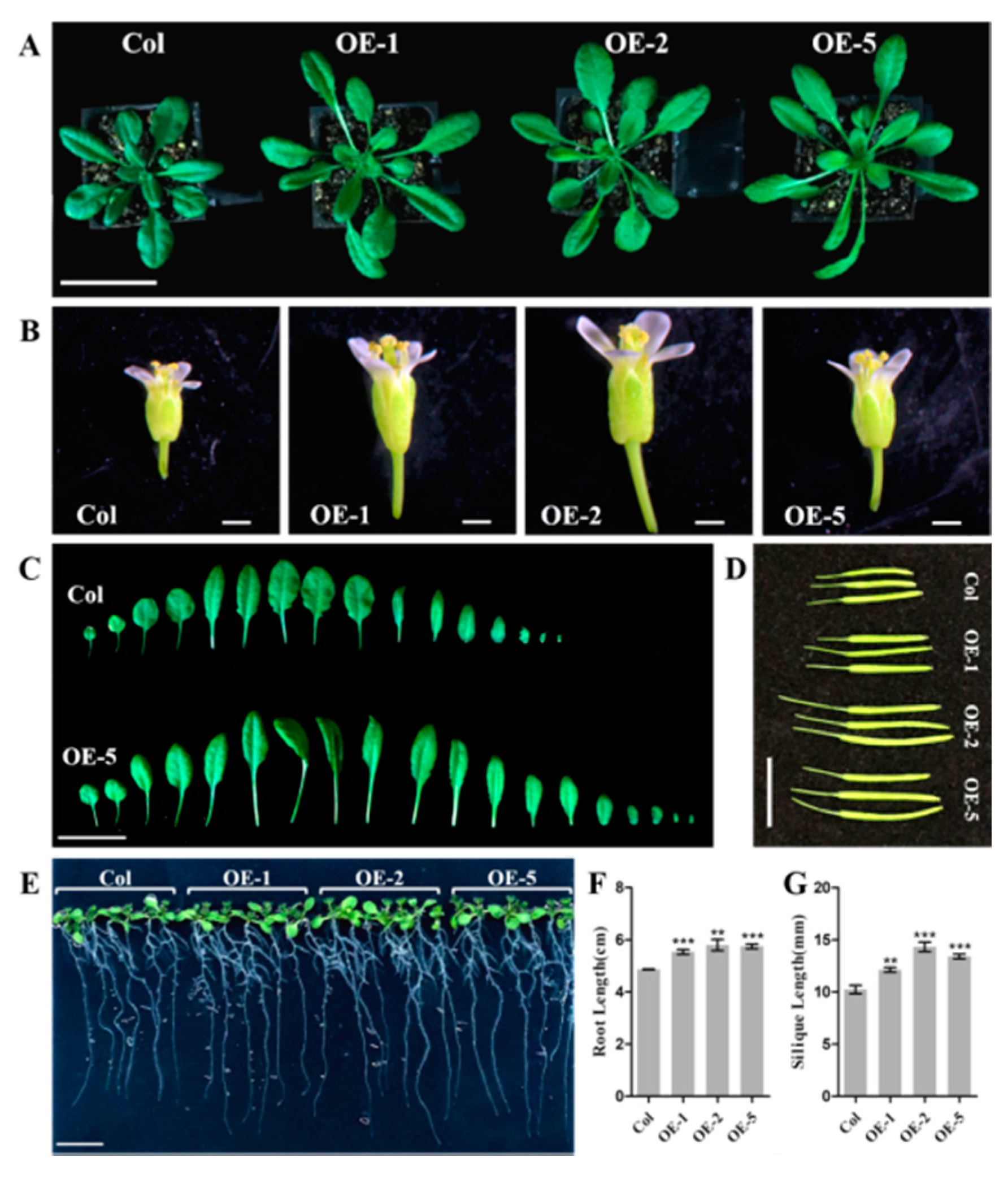
3.5. EjNACL47 Promotes Organ Enlargement in Arabidopsis
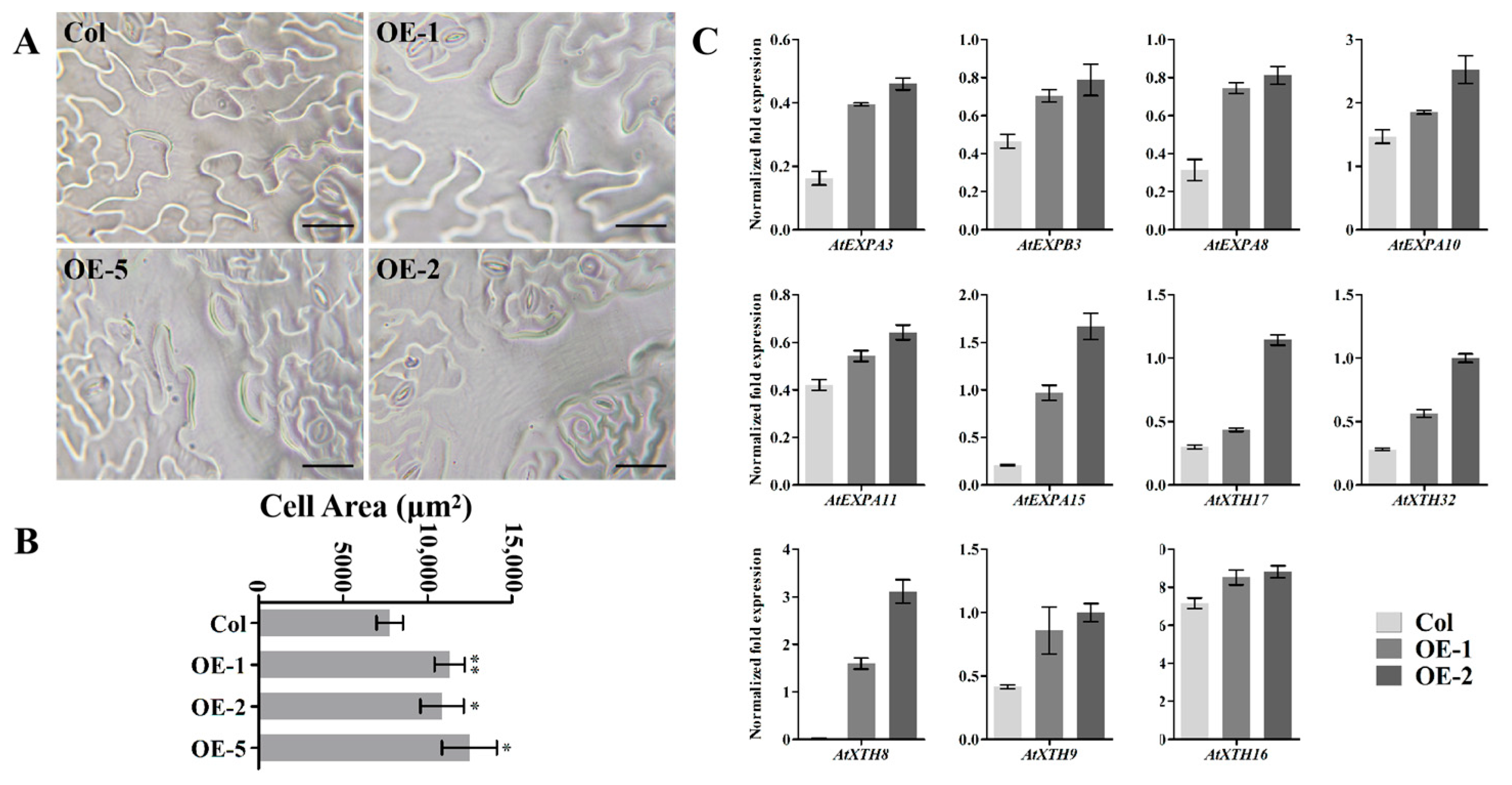
3.6. EjNACL47 Promotes Cell Expansion in Arabidopsis by Enhancing the Expression of Expansin and Xyloglucan Endotransgluco Sylase/Hydrolase (XTH) Genes
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Horiguchi, G.; Ferjani, A.; Fujikura, U.; Tsukaya, H. Coordination of cell proliferation and cell expansion in the control of leaf size in Arabidopsis thaliana. J. Plant Res. 2006, 119, 37–42. [Google Scholar] [CrossRef]
- Huang, X.; Wei, X.; Sang, T.; Zhao, Q.; Feng, Q.; Zhao, Y.; Li, C.; Zhu, C.; Lu, T.; Zhang, Z.; et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 2010, 42, 961–967. [Google Scholar] [CrossRef]
- Zhao, K.; Tung, C.W.; Eizenga, G.C.; Wright, M.H.; Ali, M.L.; Price, A.H.; Norton, G.J.; Islam, M.R.; Reynolds, A.; Mezey, J.; et al. Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat. Commun. 2011, 2, 467. [Google Scholar] [CrossRef] [PubMed]
- Frary, A.; Nesbitt, T.C.; Grandillo, S.; Knaap, E.; Cong, B.; Liu, J.; Meller, J.; Elber, R.; Alpert, K.B.; Tanksley, S.D. fw2.2: A quantitative trait locus key to the evolution of tomato fruit size. Science 2000, 289, 85–88. [Google Scholar] [CrossRef] [Green Version]
- Chakrabarti, M.; Zhang, N.; Sauvage, C.; Munos, S.; Blanca, J.; Canizares, J.; Diez, M.J.; Schneider, R.; Mazourek, M.; McClead, J.; et al. A cytochrome P450 regulates a domestication trait in cultivated tomato. Proc. Natl. Acad. Sci. USA 2013, 110, 17125–17130. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.; He, L.; Li, J.; Zhao, J.; Li, Z.; He, C. Regulatory change at Physalis Organ Size 1 correlates to natural variation in tomatillo reproductive organ size. Nat. Commun. 2014, 5, 4271. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mu, Q.; Huang, Z.; Chakrabarti, M.; Illa-Berenguer, E.; Liu, X.; Wang, Y.; Ramos, A.; van der Knaap, E. Fruit weight is controlled by Cell Size Regulator encoding a novel protein that is expressed in maturing tomato fruits. PLoS Genet. 2017, 13, e1006930. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Jiang, L.; Che, G.; Pan, Y.; Li, Y.; Hou, Y.; Zhao, W.; Zhong, Y.; Ding, L.; Yan, S.; et al. A Functional Allele of CsFUL1 Regulates Fruit Length through Repressing CsSUP and Inhibiting Auxin Transport in Cucumber. Plant Cell 2019, 31, 1289–1307. [Google Scholar] [CrossRef]
- Liu, Y.X.; Song, H.W.; Liu, Z.L.; Hu, G.B.; Lin, S.Q. Molecular characterization of loquat EjAP1 gene in relation to flowering. Plant Growth Regul. 2013, 70, 287–296. [Google Scholar] [CrossRef]
- Jiang, Y.; Peng, J.; Zhu, Y.; Su, W.; Zhang, L.; Jing, Y.; Lin, S.; Gao, Y. The Role of EjSOC1s in Flower Initiation in Eriobotrya japonica. Front. Plant Sci. 2019, 10, 253. [Google Scholar] [CrossRef] [Green Version]
- Varoquaux, F.; Blanvillain, R.; Delseny, M.; Gallois, P. Less is better: New approaches for seedless fruit production. Trends Biotechnol. 2000, 18, 233–242. [Google Scholar] [CrossRef]
- Agusti, M.; Juan, M.; Almela, V.; Gariglio, N. Loquat fruit size is increased through the thinning effect of naphthaleneacetic acid. Plant Growth Regul. 2000, 31, 167–171. [Google Scholar] [CrossRef]
- Reig, C.; Mesejo, C.; Martinez-Fuentes, A.; Agusti, M. Synthetic auxin 3,5,6-TPA increases fruit size of loquat (Eriobotrya japonica Lindl.) by reducing cell turgor pressure. Sci. Hortic. 2016, 210, 213–219. [Google Scholar] [CrossRef]
- Su, W.B.; Zhu, Y.M.; Zhang, L.; Yang, X.H.; Gao, Y.S.; Lin, S.Q. The cellular physiology of loquat (Eriobotrya japonica Lindl.) fruit with a focus on how cell division and cell expansion processes contribute to pome morphogenesis. Sci. Hortic. 2017, 224, 142–149. [Google Scholar] [CrossRef]
- Su, W.B.; Shao, Z.K.; Wang, M.; Gan, X.Q.; Yang, X.H.; Lin, S.Q. EjBZR1 represses fruit enlargement by binding to the EjCYP90 promoter in loquat. Hortic. Res. 2021, 8, 152. [Google Scholar] [CrossRef]
- Souer, E.; van Houwelingen, A.; Kloos, D.; Mol, J.; Koes, R. The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 1996, 85, 159–170. [Google Scholar] [CrossRef] [Green Version]
- Aida, M.; Ishida, T.; Fukaki, H.; Fujisawa, H.; Tasaka, M. Genes involved in organ separation in Arabidopsis: An analysis of the cup-shaped cotyledon mutant. Plant Cell 1997, 9, 841–857. [Google Scholar] [CrossRef] [Green Version]
- Fu, B.L.; Wang, W.Q.; Liu, X.F.; Duan, X.W.; Allan, A.C.; Grierson, D.; Yin, X.R. An ethylene-hypersensitive methionine sulfoxide reductase regulated by NAC transcription factors increases methionine pool size and ethylene production during kiwifruit ripening. New Phytol. 2021. [Google Scholar] [CrossRef]
- D’Inca, E.; Cazzaniga, S.; Foresti, C.; Vitulo, N.; Bertini, E.; Galli, M.; Gallavotti, A.; Pezzotti, M.; Battista Tornielli, G.; Zenoni, S. VviNAC33 promotes organ de-greening and represses vegetative growth during the vegetative-to-mature phase transition in grapevine. New Phytol. 2021, 231, 726–746. [Google Scholar] [CrossRef]
- Martin-Pizarro, C.; Vallarino, J.G.; Osorio, S.; Meco, V.; Urrutia, M.; Pillet, J.; Casanal, A.; Merchante, C.; Amaya, I.; Willmitzer, L.; et al. The NAC transcription factor FaRIF controls fruit ripening in strawberry. Plant Cell 2021, 33, 1574–1593. [Google Scholar] [CrossRef]
- Yu, M.; Liu, J.; Du, B.; Zhang, M.; Wang, A.; Zhang, L. NAC Transcription Factor PwNAC11 Activates ERD1 by Interaction with ABF3 and DREB2A to Enhance Drought Tolerance in Transgenic Arabidopsis. Int. J. Mol. Sci. 2021, 22, 6952. [Google Scholar] [CrossRef] [PubMed]
- Pei, H.; Ma, N.; Tian, J.; Luo, J.; Chen, J.; Li, J.; Zheng, Y.; Chen, X.; Fei, Z.; Gao, J. An NAC transcription factor controls ethylene-regulated cell expansion in flower petals. Plant Physiol. 2013, 163, 775–791. [Google Scholar] [CrossRef] [Green Version]
- Ma, X.; Zhang, Y.; Tureckova, V.; Xue, G.P.; Fernie, A.R.; Mueller-Roeber, B.; Balazadeh, S. The NAC Transcription Factor SlNAP2 Regulates Leaf Senescence and Fruit Yield in Tomato. Plant Physiol. 2018, 177, 1286–1302. [Google Scholar] [CrossRef] [Green Version]
- Xu, Q.; Wang, W.Q.; Zeng, J.K.; Zhang, J.; Grierson, D.; Li, X.; Yin, X.R.; Chen, K.S. A NAC transcription factor, EjNAC1, affects lignification of loquat fruit by regulating lignin. Postharvest Biol. Technol. 2015, 102, 25–31. [Google Scholar] [CrossRef]
- Ge, H.; Zhang, J.; Zhang, Y.J.; Li, X.; Yin, X.R.; Grierson, D.; Chen, K.S. EjNAC3 transcriptionally regulates chilling-induced lignification of loquat fruit via physical interaction with an atypical CAD-like gene. J. Exp. Bot. 2017, 68, 5129–5136. [Google Scholar] [CrossRef] [Green Version]
- Rauf, M.; Arif, M.; Fisahn, J.; Xue, G.P.; Balazadeh, S.; Mueller-Roeber, B. NAC transcription factor speedy hyponastic growth regulates flooding-induced leaf movement in Arabidopsis. Plant Cell 2013, 25, 4941–4955. [Google Scholar] [CrossRef] [Green Version]
- Jiang, S.; An, H.; Xu, F.; Zhang, X. Chromosome-level genome assembly and annotation of the loquat (Eriobotrya japonica) genome. Gigascience 2020, 9, giaa015. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Confidence-Limits on Phylogenies—An Approach Using the Bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Nei, M.; Rogozin, I.B.; Piontkivska, H. Purifying selection and birth-and-death evolution in the ubiquitin gene family. Proc. Natl. Acad. Sci. USA 2000, 97, 10866–10871. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(-Delta Delta C) method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [Green Version]
- Jefferson, R.A.; Kavanagh, T.A.; Bevan, M.W. GUS fusions: Beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 1987, 6, 3901–3907. [Google Scholar] [CrossRef] [PubMed]
- Olsen, A.N.; Ernst, H.A.; Leggio, L.L.; Skriver, K. NAC transcription factors: Structurally distinct, functionally diverse. Trends Plant Sci. 2005, 10, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Ernst, H.A.; Olsen, A.N.; Larsen, S.; Lo Leggio, L. Structure of the conserved domain of ANAC, a member of the NAC family of transcription factors. EMBO Rep. 2004, 5, 297–303. [Google Scholar] [CrossRef] [PubMed]
- Ooka, H.; Satoh, K.; Doi, K.; Nagata, T.; Otomo, Y.; Murakami, K.; Matsubara, K.; Osato, N.; Kawai, J.; Carninci, P.; et al. Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res. 2003, 10, 239–247. [Google Scholar] [CrossRef]
- Hao, Y.J.; Song, Q.X.; Chen, H.W.; Zou, H.F.; Wei, W.; Kang, X.S.; Ma, B.; Zhang, W.K.; Zhang, J.S.; Chen, S.Y. Plant NAC-type transcription factor proteins contain a NARD domain for repression of transcriptional activation. Planta 2010, 232, 1033–1043. [Google Scholar] [CrossRef] [PubMed]
- Puranik, S.; Bahadur, R.P.; Srivastava, P.S.; Prasad, M. Molecular cloning and characterization of a membrane associated NAC family gene, SiNAC from foxtail millet [Setaria italica (L.) P. Beauv]. Mol. Biotechnol. 2011, 49, 138–150. [Google Scholar] [CrossRef]
- Lu, P.L.; Chen, N.Z.; An, R.; Su, Z.; Qi, B.S.; Ren, F.; Chen, J.; Wang, X.C. A novel drought-inducible gene, ATAF1, encodes a NAC family protein that negatively regulates the expression of stress-responsive genes in Arabidopsis. Plant Mol. Biol. 2007, 63, 289–305. [Google Scholar] [CrossRef]
- Yoshii, M.; Yamazaki, M.; Rakwal, R.; Kishi-Kaboshi, M.; Miyao, A.; Hirochika, H. The NAC transcription factor RIM1 of rice is a new regulator of jasmonate signaling. Plant J. 2010, 61, 804–815. [Google Scholar] [CrossRef]
- Kim, H.S.; Park, B.O.; Yoo, J.H.; Jung, M.S.; Lee, S.M.; Han, H.J.; Kim, K.E.; Kim, S.H.; Lim, C.O.; Yun, D.J.; et al. Identification of a calmodulin-binding NAC protein as a transcriptional repressor in Arabidopsis. J. Biol. Chem. 2007, 282, 36292–36302. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Delessert, C.; Kazan, K.; Wilson, I.W.; Van Der Straeten, D.; Manners, J.; Dennis, E.S.; Dolferus, R. The transcription factor ATAF2 represses the expression of pathogenesis-related genes in Arabidopsis. Plant J. 2005, 43, 745–757. [Google Scholar] [CrossRef] [PubMed]
- Xie, Q.; Frugis, G.; Colgan, D.; Chua, N.H. Arabidopsis NAC1 transduces auxin signal downstream of TIR1 to promote lateral root development. Genes Dev. 2000, 14, 3024–3036. [Google Scholar] [CrossRef] [Green Version]
- Peng, H.; Yu, X.; Cheng, H.; Shi, Q.; Zhang, H.; Li, J.; Ma, H. Cloning and characterization of a novel NAC family gene CarNAC1 from chickpea (Cicer arietinum L.). Mol. Biotechnol. 2010, 44, 30–40. [Google Scholar] [CrossRef]
- Powell, A.E.; Lenhard, M. Control of organ size in plants. Curr. Biol. 2012, 22, R360–R367. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsukaya, H. Interpretation of mutants in leaf morphology: Genetic evidence for a compensatory system in leaf morphogenesis that provides a new link between cell and organismal theories. Int. Rev. Cytol. 2002, 217, 1–39. [Google Scholar] [CrossRef]
- Czesnick, H.; Lenhard, M. Size control in plants--lessons from leaves and flowers. Cold Spring Harb. Perspect. Biol. 2015, 7, a019190. [Google Scholar] [CrossRef]
- Kurepa, J.; Wang, S.; Li, Y.; Zaitlin, D.; Pierce, A.J.; Smalle, J.A. Loss of 26S proteasome function leads to increased cell size and decreased cell number in Arabidopsis shoot organs. Plant Physiol. 2009, 150, 178–189. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wolf, S.; Hematy, K.; Hofte, H. Growth control and cell wall signaling in plants. Annu. Rev. Plant Biol. 2012, 63, 381–407. [Google Scholar] [CrossRef] [Green Version]
- Goh, H.H.; Sloan, J.; Malinowski, R.; Fleming, A. Variable expansin expression in Arabidopsis leads to different growth responses. J. Plant Physiol. 2014, 171, 329–339. [Google Scholar] [CrossRef] [PubMed]
- Fry, S.C.; Smith, R.C.; Renwick, K.F.; Martin, D.J.; Hodge, S.K.; Matthews, K.J. Xyloglucan endotransglycosylase, a new wall-loosening enzyme activity from plants. Biochem. J. 1992, 282 Pt 3, 821–828. [Google Scholar] [CrossRef] [Green Version]
- Nishitani, K.; Tominaga, R. Endoxyloglucan Transferase, a Novel Class of Glycosyltransferase That Catalyzes Transfer of a Segment of Xyloglucan Molecule to Another Xyloglucan Molecule. J. Biol. Chem. 1992, 267, 21058–21064. [Google Scholar] [CrossRef]
- An, J.P.; Yao, J.F.; Xu, R.R.; You, C.X.; Wang, X.F.; Hao, Y.J. An apple NAC transcription factor enhances salt stress tolerance by modulating the ethylene response. Physiol. Plant. 2018, 164, 279–289. [Google Scholar] [CrossRef] [Green Version]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, Q.; Jing, D.; Wang, S.; Xu, F.; Bao, C.; Luo, M.; Guo, Q. The Putative Role of the NAC Transcription Factor EjNACL47 in Cell Enlargement of Loquat (Eriobotrya japonica Lindl.). Horticulturae 2021, 7, 323. https://doi.org/10.3390/horticulturae7090323
Chen Q, Jing D, Wang S, Xu F, Bao C, Luo M, Guo Q. The Putative Role of the NAC Transcription Factor EjNACL47 in Cell Enlargement of Loquat (Eriobotrya japonica Lindl.). Horticulturae. 2021; 7(9):323. https://doi.org/10.3390/horticulturae7090323
Chicago/Turabian StyleChen, Qian, Danlong Jing, Shuming Wang, Fan Xu, Chaoya Bao, Ming Luo, and Qigao Guo. 2021. "The Putative Role of the NAC Transcription Factor EjNACL47 in Cell Enlargement of Loquat (Eriobotrya japonica Lindl.)" Horticulturae 7, no. 9: 323. https://doi.org/10.3390/horticulturae7090323
APA StyleChen, Q., Jing, D., Wang, S., Xu, F., Bao, C., Luo, M., & Guo, Q. (2021). The Putative Role of the NAC Transcription Factor EjNACL47 in Cell Enlargement of Loquat (Eriobotrya japonica Lindl.). Horticulturae, 7(9), 323. https://doi.org/10.3390/horticulturae7090323





