In Silico Design of Engineering Optimization via OptHandle for Effective Synthesis of Adipic Acid Precursor, α-Aminoadipate
Abstract
1. Introduction
2. Methods and Materials
2.1. The Optimization Procedure of OptHandle
- (1)
- Screening candidate reactions to be regulated by flux variability analysis (FVA)
- (2)
- Reconstructing reaction equations by Atom–Atom Mapping (AAM)
- (3)
- Identifying key reactions to be regulated
2.2. Media, Plasmids, Strains and Culture Conditions
2.3. HPLC Analysis of Metabolites
3. Results and Discussion
3.1. Assessment of OptHandle via Tow Cases
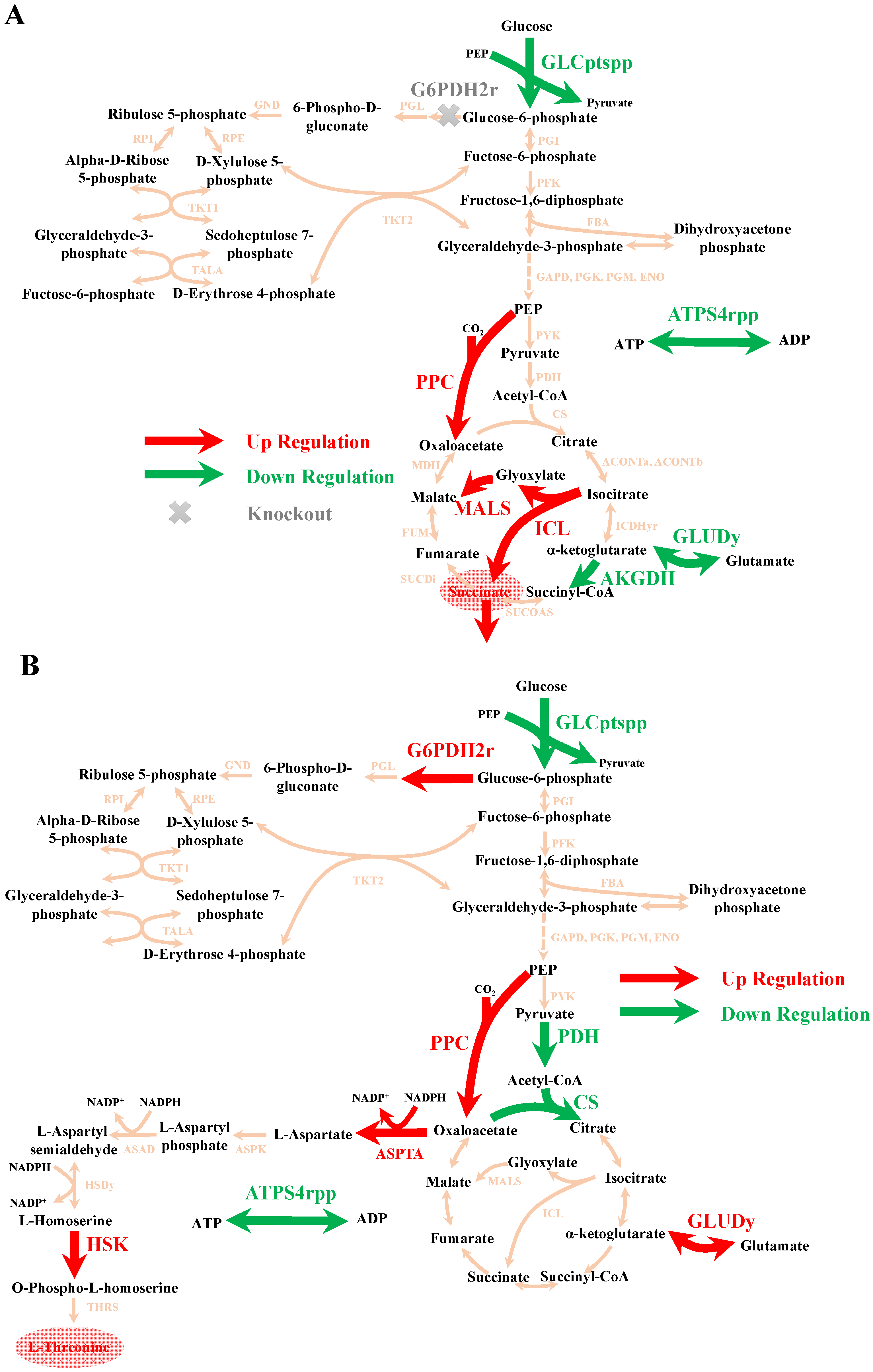
3.1.1. Case 1: Succinate Overproduction in E. coli
3.1.2. Case 2: L-Threonine Overproduction in E. coli
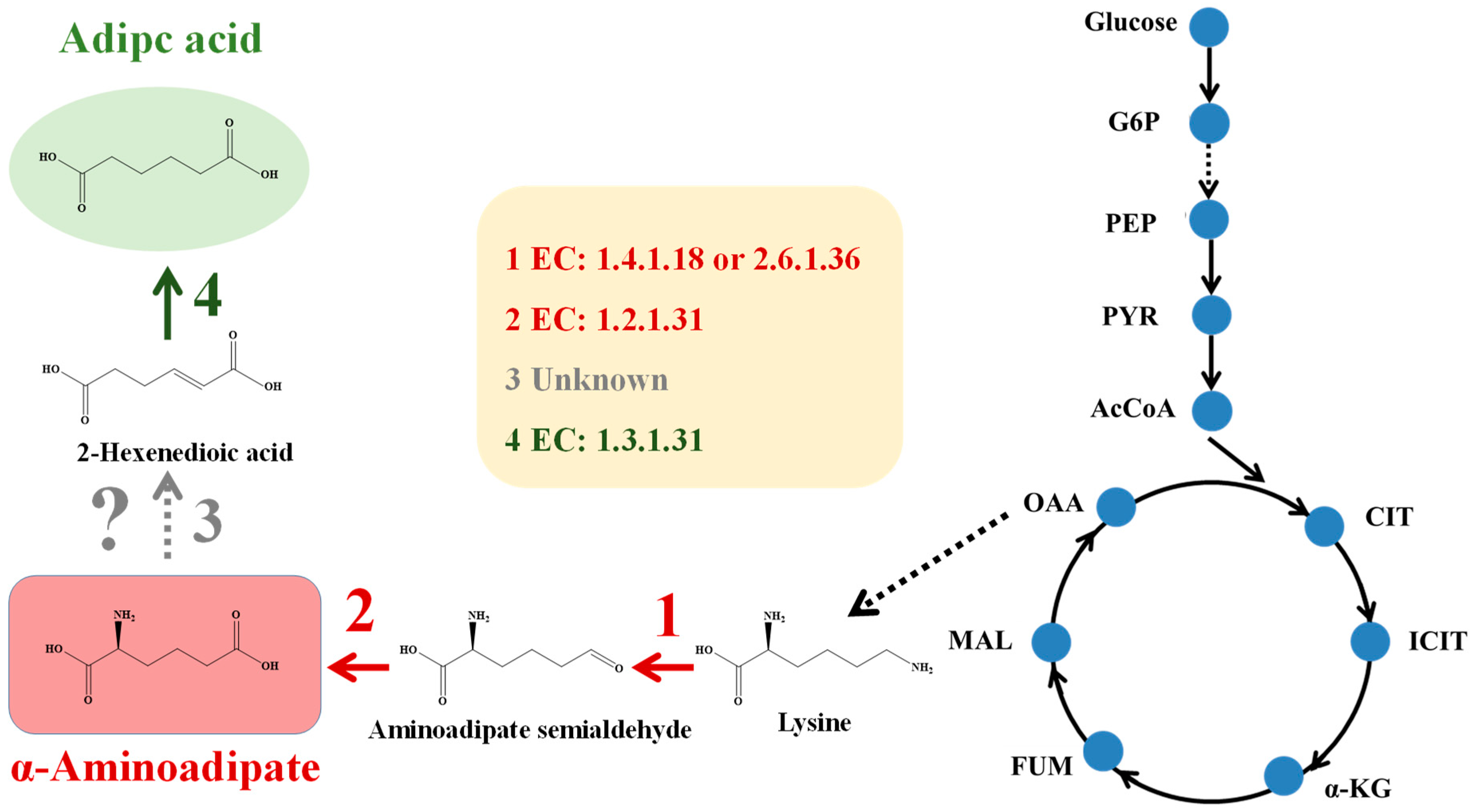
3.2. Overproduction of α-Aminoadipate via OptHandle in E. coli
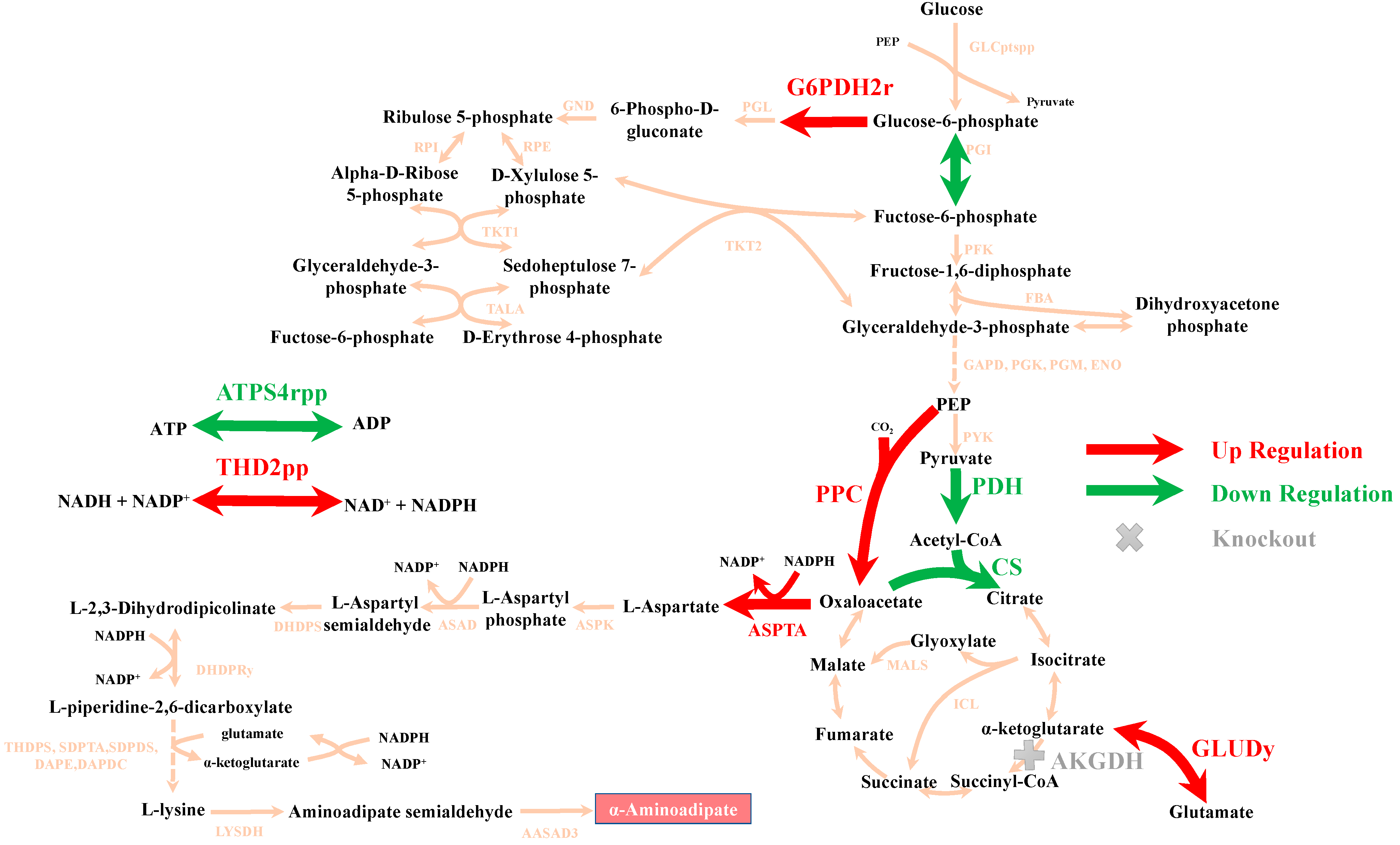
3.2.1. In Silico Design of an Optimal E. coli α-Aminoadipate Producer via OptHandle
3.2.2. Tuning the Architecture of the Biosynthetic Pathway and Selecting Gene Orthologues to Increase α-Aminoadipate Production
3.2.3. Strengthening Synthesis of Precursors to Increase α-Aminoadipate Production
3.2.4. Regulating the Flux of the Key Node to Increase α-Aminoadipate Production
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Luttrell, W.E.; Klaassen, G.R. Adipic acid. J. Chem. Health Saf. 2016, 23, 44–46. [Google Scholar] [CrossRef]
- Skoog, E.; Shin, J.H.; Saez-Jimenez, V.; Mapelli, V.; Olsson, L. Biobased adipic acid-The challenge of developing the production host. Biotechnol. Adv. 2018, 36, 2248–2263. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Ma, L.; Mao, Y. Biological production of adipic acid from renewable substrates: Current and future methods. Biochem. Eng. J. 2016, 105, 16–26. [Google Scholar] [CrossRef]
- Saitoh, M.; Fujisaki, S.; Ishii, Y.; Nishiguchi, T. Convenient selective monoesterification of α, ω-dicarboxylic acids catalyzed by ion-exchange resins. Tetrahedron Lett. 1996, 37, 6733–6736. [Google Scholar] [CrossRef]
- Chae, T.U.; Ahn, J.H.; Ko, Y.-S.; Kim, J.W.; Lee, J.A.; Lee, E.H.; Lee, S.Y. Metabolic engineering for the production of dicarboxylic acids and diamines. Metab. Eng. 2020, 58, 2–16. [Google Scholar] [CrossRef]
- Li, G.; Huang, D.; Sui, X.; Li, S.; Huang, B.; Zhang, X.; Wu, H.; Deng, Y. Advances in microbial production of medium-chain dicarboxylic acids for nylon materials. React. Chem. Eng. 2020, 5, 221–238. [Google Scholar] [CrossRef]
- Xu, J.; Han, M.; Zhang, J.; Guo, Y.; Zhang, W. Metabolic engineering Corynebacterium glutamicum for the L-lysine production by increasing the flux into L-lysine biosynthetic pathway. Amino Acids 2014, 46, 2165–2175. [Google Scholar] [CrossRef] [PubMed]
- Ying, H.; He, X.; Li, Y.; Chen, K.; Ouyang, P. Optimization of culture conditions for enhanced lysine production using engineered Escherichia coli. Appl. Biochem. Biotechnol. 2014, 172, 3835–3843. [Google Scholar] [CrossRef]
- Rohles, C.M.; Gläser, L.; Kohlstedt, M.; Gießelmann, G.; Pearson, S.; del Campo, A.; Becker, J.; Wittmann, C. A bio-based route to the carbon-5 chemical glutaric acid and to bionylon-6, 5 using metabolically engineered Corynebacterium glutamicum. Green Chem. 2018, 20, 4662–4674. [Google Scholar] [CrossRef]
- Kind, S.; Neubauer, S.; Becker, J.; Yamamoto, M.; Völkert, M.; von Abendroth, G.; Zelder, O.; Wittmann, C. From zero to hero-production of bio-based nylon from renewable resources using engineered Corynebacterium glutamicum. Metab. Eng. 2014, 25, 113–123. [Google Scholar] [CrossRef]
- Nishi, K.; Endo, S.; Mori, Y.; Totsuka, K.; Hirao, Y. Method for Producing Cadaverine Dicarboxylate and Its Use for the Production of Nylon. European Patent EP1482055 (B1), 19 October 2006. [Google Scholar]
- Sigman, M.S.; Vachal, P.; Jacobsen, E.N. A general catalyst for the asymmetric Strecker reaction. Angew. Chem. 2000, 112, 1336–1338. [Google Scholar] [CrossRef]
- Karlsson, E.; Shin, J.H.; Westman, G.; Eriksson, L.A.; Olsson, L.; Mapelli, V. In silico and in vitro studies of the reduction of unsaturated α, β bonds of trans-2-hexenedioic acid and 6-amino-trans-2-hexenoic acid-Important steps towards biobased production of adipic acid. PLoS ONE 2018, 13, e0193503. [Google Scholar] [CrossRef]
- Skoog, E. Biobased Adipic Acid-Challenges in Establishing a Cell Factory; Chalmers Tekniska Hogskola: Gothenburg, Sweden, 2019. [Google Scholar]
- Feist, A.M.; Palsson, B.Ø. The growing scope of applications of genome-scale metabolic reconstructions using Escherichia coli. Nat. Biotechnol. 2008, 26, 659–667. [Google Scholar] [CrossRef]
- Oberhardt, M.A.; Palsson, B.Ø.; Papin, J.A. Applications of genome-scale metabolic reconstructions. Mol. Syst. Biol. 2009, 5, 320. [Google Scholar] [CrossRef] [PubMed]
- Burgard, A.P.; Pharkya, P.; Maranas, C.D. Optknock: A bilevel programming framework for identifying gene knockout strategies for microbial strain optimization. Biotechnol. Bioeng. 2003, 84, 647–657. [Google Scholar] [CrossRef]
- Pharkya, P.; Burgard, A.P.; Maranas, C.D. OptStrain: A computational framework for redesign of microbial production systems. Genome Res. 2004, 14, 2367–2376. [Google Scholar] [CrossRef] [PubMed]
- Ranganathan, S.; Suthers, P.F.; Maranas, C.D. OptForce: An optimization procedure for identifying all genetic manipulations leading to targeted overproductions. PLoS Comput. Biol. 2010, 6, e1000744. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Otero-Muras, I.; Banga, J.R.; Wang, Y.; Kaiser, M.; Krasnogor, N. OptDesign: Identifying Optimum Design Strategies in Strain Engineering for Biochemical Production. ACS Synth. Biol. 2022, 11, 1531–1541. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Reed, J.L.; Maravelias, C.T. Large-scale bi-level strain design approaches and mixed-integer programming solution techniques. PLoS ONE 2011, 6, e24162. [Google Scholar] [CrossRef] [PubMed]
- Cheng, F.; Yu, H.; Stephanopoulos, G. Engineering Corynebacterium glutamicum for high-titer biosynthesis of hyaluronic acid. Metab. Eng. 2019, 55, 276–289. [Google Scholar] [CrossRef]
- Mahadevan, R.; Schilling, C.H. The effects of alternate optimal solutions in constraint-based genome-scale metabolic models. Metab. Eng. 2003, 5, 264–276. [Google Scholar] [CrossRef]
- Rahman, S.A.; Torrance, G.; Baldacci, L.; Martínez Cuesta, S.; Fenninger, F.; Gopal, N.; Choudhary, S.; May, J.W.; Holliday, G.L.; Steinbeck Thornton, J.M. Reaction Decoder Tool (RDT): Extracting features from chemical reactions. Bioinformatics 2016, 32, 2065–2066. [Google Scholar] [CrossRef]
- Li, W.; Ma, L.; Shen, X.; Wang, J.; Feng, Q.; Liu, L.; Zheng, G.; Yan, Y.; Sun, X.; Yuan, Q. Targeting metabolic driving and intermediate influx in lysine catabolism for high-level glutarate production. Nat. Commun. 2019, 10, 3337. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, A.; Zomorrodi, A.R.; Maranas, C.D. k-OptForce: Integrating kinetics with flux balance analysis for strain design. PLoS Comput. Biol. 2014, 10, e1003487. [Google Scholar] [CrossRef]
- Khodayari, A.; Chowdhury, A.; Maranas, C.D. Succinate overproduction: A case study of computational strain design using a comprehensive Escherichia coli kinetic model. Front. Bioeng. Biotechnol. 2015, 2, 76. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lin, H.; Bennett, G.N.; San, K.Y. Fed-batch culture of a metabolically engineered Escherichia coli strain designed for high-level succinate production and yield under aerobic conditions. Biotechnol. Bioeng. 2005, 90, 775–779. [Google Scholar] [CrossRef]
- Lin, H.; Bennett, G.N.; San, K.Y. Chemostat culture characterization of Escherichia coli mutant strains metabolically engineered for aerobic succinate production: A study of the modified metabolic network based on metabolite profile, enzyme activity, and gene expression profile. Metab. Eng. 2005, 7, 337–352. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Baba, T.; Mori, H.; Shimizu, K. Effect of zwf gene knockout on the metabolism of Escherichia coli grown on glucose or acetate. Metab. Eng. 2004, 6, 164–174. [Google Scholar] [CrossRef] [PubMed]
- Hager, P.W.; Calfee, M.W.; Phibbs, P.V. The Pseudomonas aeruginosa devB/SOL homolog, pgl, is a member of the hex regulon and encodes 6-phosphogluconolactonase. J. Bacteriol. 2000, 182, 3934–3941. [Google Scholar] [CrossRef]
- Kyselova, L.; Kreitmayer, D.; Kremling, A.; Bettenbrock, K. Type and capacity of glucose transport influences succinate yield in two-stage cultivations. Microb. Cell Fact. 2018, 17, 132. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, D.; Cai, N.; Han, C.; Mao, Q.; Wang, T.; Zhou, Q.; Chen, N.; Xie, X. Betaine supplementation improved L-threonine fermentation of Escherichia coli THRD by upregulating zwf (glucose-6-phosphate dehydrogenase) expression. Electron. J. Biotechnol. 2019, 39, 67–73. [Google Scholar] [CrossRef]
- Dong, X.; Quinn, P.J.; Wang, X. Metabolic engineering of Escherichia coli and Corynebacterium glutamicum for the production of L-threonine. Biotechnol. Adv. 2011, 29, 11–23. [Google Scholar] [CrossRef]
- Lee, K.H.; Park, J.H.; Kim, T.Y.; Kim, H.U.; Lee, S.Y. Systems metabolic engineering of Escherichia coli for L-threonine production. Mol. Syst. Biol. 2007, 3, 149. [Google Scholar] [CrossRef]
- Liu, J.; Li, H.; Xiong, H.; Xie, X.; Chen, N.; Zhao, G.; Caiyin, Q.; Zhu, H.; Qiao, J. Two-stage carbon distribution and cofactor generation for improving L-threonine production of Escherichia coli. Biotechnol. Bioeng. 2019, 116, 110–120. [Google Scholar] [CrossRef]
- Lee, J.H.; Sung, B.H.; Kim, M.S.; Blattner, F.R.; Yoon, B.H.; Kim, J.H.; Kim, S.C. Metabolic engineering of a reduced-genome strain of Escherichia coli for L-threonine production. Microb. Cell Fact. 2009, 8, 2. [Google Scholar] [CrossRef]
- Kim, Y.-H.; Park, J.-S.; Cho, J.-Y.; Cho, K.M.; Park, Y.-H.; Lee, J. Proteomic response analysis of a threonine-overproducing mutant of Escherichia coli. Biochem. J. 2004, 381, 823–829. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Lu, Y.; Yang, J.; Fang, Y.; Zhu, L.; Ding, Z.; Wang, C.; Ma, W.; Hu, X.; Wang, X. Expression regulation of multiple key genes to improve L-threonine in Escherichia coli. Microb. Cell Fact. 2020, 19, 46. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, M.; Cai, B.; He, K.; Wang, M.; Chen, B.; Tan, T. De novo biosynthesis of α-aminoadipate via multi-strategy metabolic engineering in Escherichia coli. MicrobiologyOpen 2022, 11, e1301. [Google Scholar] [CrossRef]
- Zhang, Y.; An, N.; Zhao, Y.; Li, X.; Shen, X.; Wang, J.; Sun, X.; Yuan, Q. Efficient biosynthesis of α-aminoadipic acid via lysine catabolism in Escherichia coli. Biotechnol. Bioeng. 2023, 120, 312–317. [Google Scholar] [CrossRef] [PubMed]
- Xiong, B.; Zhu, Y.; Tian, D.; Jiang, S.; Fan, X.; Ma, Q.; Wu, H.; Xie, X. Flux redistribution of central carbon metabolism for efficient production of L-tryptophan in Escherichia coli. Biotechnol. Bioeng. 2021, 118, 1393–1404. [Google Scholar] [CrossRef]



| Serial Number | Reaction Sets of OptHandle | Total Flux of Reaction Sets | Priority Regulated Reaction | Flux Ratio | Regulation Strategy | Validation of Experiments or Other Algorithms |
|---|---|---|---|---|---|---|
| 1 | ATPS4rpp a + ADSS + SADT2 + BPNT + PRATPP | 49.32 | ATPS4rpp | 98.69% | Down-regulation | [20] |
| 2 | CYTBO3_4pp | 29.05 | CYTBO3_4pp | 100.00% | Down-regulation | - |
| 3 | PPC | 8.31 | PPC | 100.00% | Up-regulation | [19,26,27,29] |
| 4 | GLUDy + DDPA | 6.50 | GLUDy | 95.83% | Down-regulation | [19] |
| 5 | ACCOAC + AKGDH | 5.38 | AKGDH | 67.29% | Down-regulation | - |
| 6 | G6PDH2r | 3.51 | G6PDH2r | 100.00% | Knockout | [30] |
| 7 | ICL | 2.66 | ICL | 100.00% | Up-regulation | [26,27,28] |
| 8 | MALS | 2.66 | MALS | 100.00% | Up-regulation | [26,27] |
| 9 | CBMKr + HCO3E | 2.54 | HCO3E | 82.03% | Down-regulation | - |
| 10 | GLCptspp | 2.53 | GLCptspp | 100.00% | Down-regulation | [32] |
| Serial Number | Reaction Sets of OptHandle | Total Flux of Reaction Sets | Priority Regulated Reaction | Flux Ratio | Regulation Strategy | Validation of Experiments or Other Algorithms |
|---|---|---|---|---|---|---|
| 1 | ATPS4rpp a | 42.55 | ATPS4rpp | 100.00% | Down-regulation | - |
| 2 | CYTBO3_4pp | 22.86 | CYTBO3_4pp | 100.00% | Down-regulation | - |
| 3 | G6PDH2r | 15.22 | G6PDH2r | 100.00% | Up-regulation | [33] |
| 4 | PDH + AKGDH | 12.03 | PDH | 65.46% | Down-regulation | [39] |
| 5 | HSK | 11.85 | HSK | 100.00% | Up-regulation | [37] |
| 6 | ASPTA | 9.76 | ASPTA | 100.00% | Up-regulation | [35] |
| 7 | PPC | 9.73 | PPC | 100.00% | Up-regulation | [35] |
| 8 | GLCptspp | 8.18 | GLCptspp | 100.00% | Down-regulation | [37] |
| 9 | CS | 5.10 | CS | 100.00% | Down-regulation | [38] |
| 10 | GLUDy | 4.83 | GLUDy | 100.00% | Up-regulation | [36] |
| Serial Number | Reaction Sets of OptHandle | Total Flux of Reaction Sets | Priority Regulated Reaction | Flux Ratio | Regulation Strategy |
|---|---|---|---|---|---|
| 1 | ATPS4rpp a + ADSS + SADT2 + BPNT + PRATPP | 52.34 | ATPS4rpp | 98.58% | Down-regulation |
| 2 | CYTBO3_4pp | 26.40 | CYTBO3_4pp | 100.00% | Down-regulation |
| 3 | THD2pp | 15.64 | THD2pp | 100.00% | Up-regulation |
| 4 | PTAr + PDH | 9.99 | PDH | 95.15% | Down-regulation |
| 5 | GLUDy | 9.27 | GLUDy | 100.00% | Up-regulation |
| 6 | DDPA + CS | 7.18 | CS | 95.58% | Down-regulation |
| 7 | PGI | 6.49 | PGI | 100.00% | Down-regulation |
| 8 | G6PDH2r | 6.49 | G6PDH2r | 100.00% | Up-regulation |
| 9 | AKGDH | 5.97 | AKGDH | 100.00% | Knockout |
| 10 | ASPTA | 5.76 | ASPTA | 100.00% | Up-regulation |
| 11 | PPC | 5.73 | PPC | 100.00% | Down-regulation |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Cai, B.; Liu, M.; He, K.; Gong, Z.; Bi, H.; Wang, K.; Chen, B.; Wang, M.; Su, H.; et al. In Silico Design of Engineering Optimization via OptHandle for Effective Synthesis of Adipic Acid Precursor, α-Aminoadipate. Fermentation 2023, 9, 859. https://doi.org/10.3390/fermentation9090859
Zhang Y, Cai B, Liu M, He K, Gong Z, Bi H, Wang K, Chen B, Wang M, Su H, et al. In Silico Design of Engineering Optimization via OptHandle for Effective Synthesis of Adipic Acid Precursor, α-Aminoadipate. Fermentation. 2023; 9(9):859. https://doi.org/10.3390/fermentation9090859
Chicago/Turabian StyleZhang, Yang, Bingqi Cai, Meng Liu, Keqin He, Zhijin Gong, Haoran Bi, Kai Wang, Biqiang Chen, Meng Wang, Haijia Su, and et al. 2023. "In Silico Design of Engineering Optimization via OptHandle for Effective Synthesis of Adipic Acid Precursor, α-Aminoadipate" Fermentation 9, no. 9: 859. https://doi.org/10.3390/fermentation9090859
APA StyleZhang, Y., Cai, B., Liu, M., He, K., Gong, Z., Bi, H., Wang, K., Chen, B., Wang, M., Su, H., & Tan, T. (2023). In Silico Design of Engineering Optimization via OptHandle for Effective Synthesis of Adipic Acid Precursor, α-Aminoadipate. Fermentation, 9(9), 859. https://doi.org/10.3390/fermentation9090859






