Functional Investigation of Mutant Vespa tropica Hyaluronidase Produced in Pichia pastoris: In Silico Studies and Potential Applications
Abstract
1. Introduction
2. Materials and Methods
2.1. Strains, Reagents, and Media
2.2. Plasmid Construction via the Golden Gate-Derived P. pastoris Cloning System (GoldenPiCS)
2.3. Screening of P. pastoris Production and Bioreactor Cultivation
2.4. Protein Purification
2.5. SDS-PAGE and Western Blotting
2.6. LC-MS/MS Analyses
2.7. Hyaluronidase Activity Assay
2.8. Development of mVesT2a Protein for Hyaluronic Acid Detection
2.9. In Silico Molecular Docking and Dynamics Simulations
3. Results and Discussion
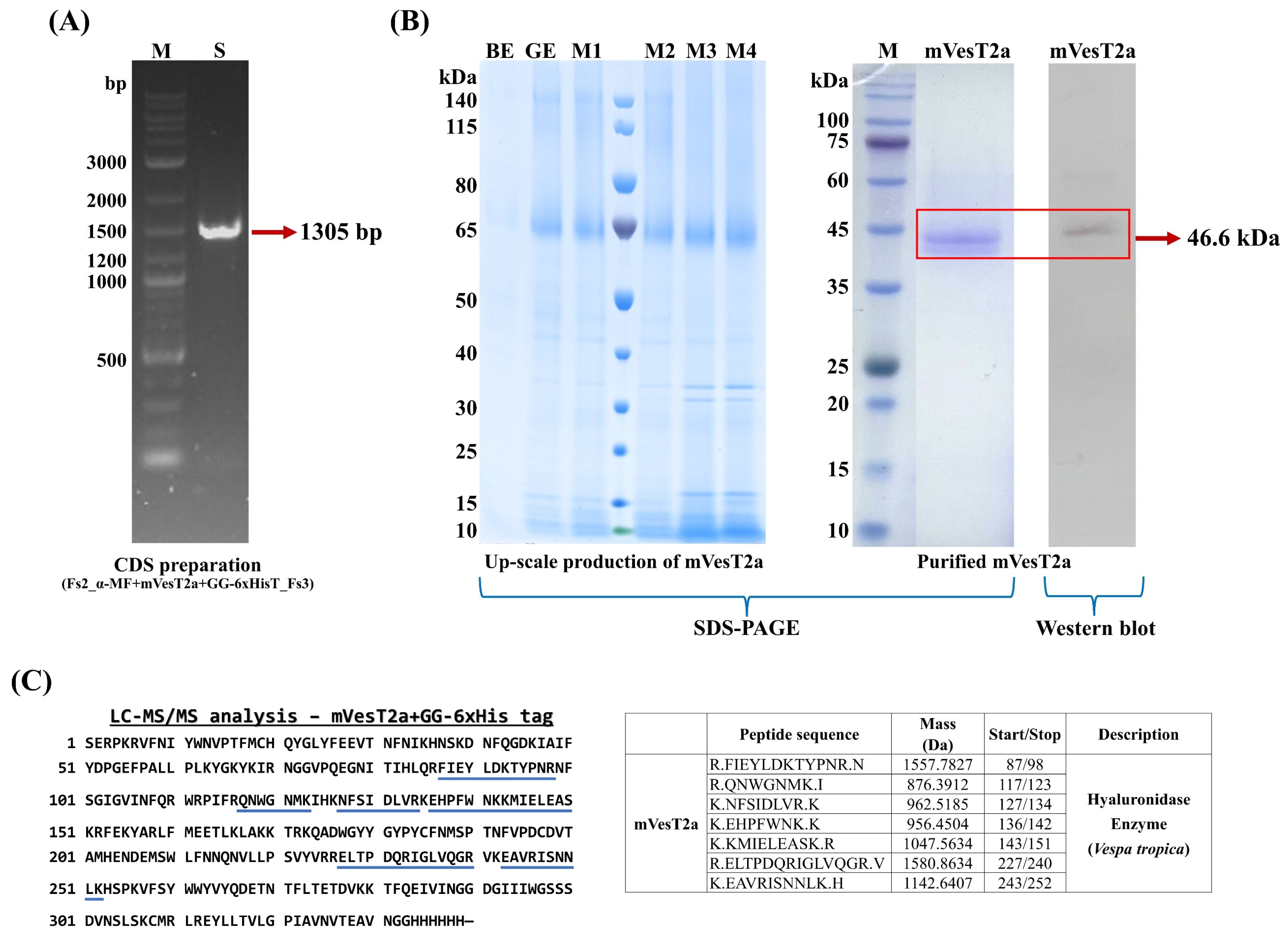
3.1. Recombinant Plasmids Construction and Small-Scale Production of mVesT2a Protein
3.2. Upscaled Production, Detection, and Purification of mVestT2a Produced by P. pastoris
3.3. Hyaluronidase Activity Assay and HA Detection Assay
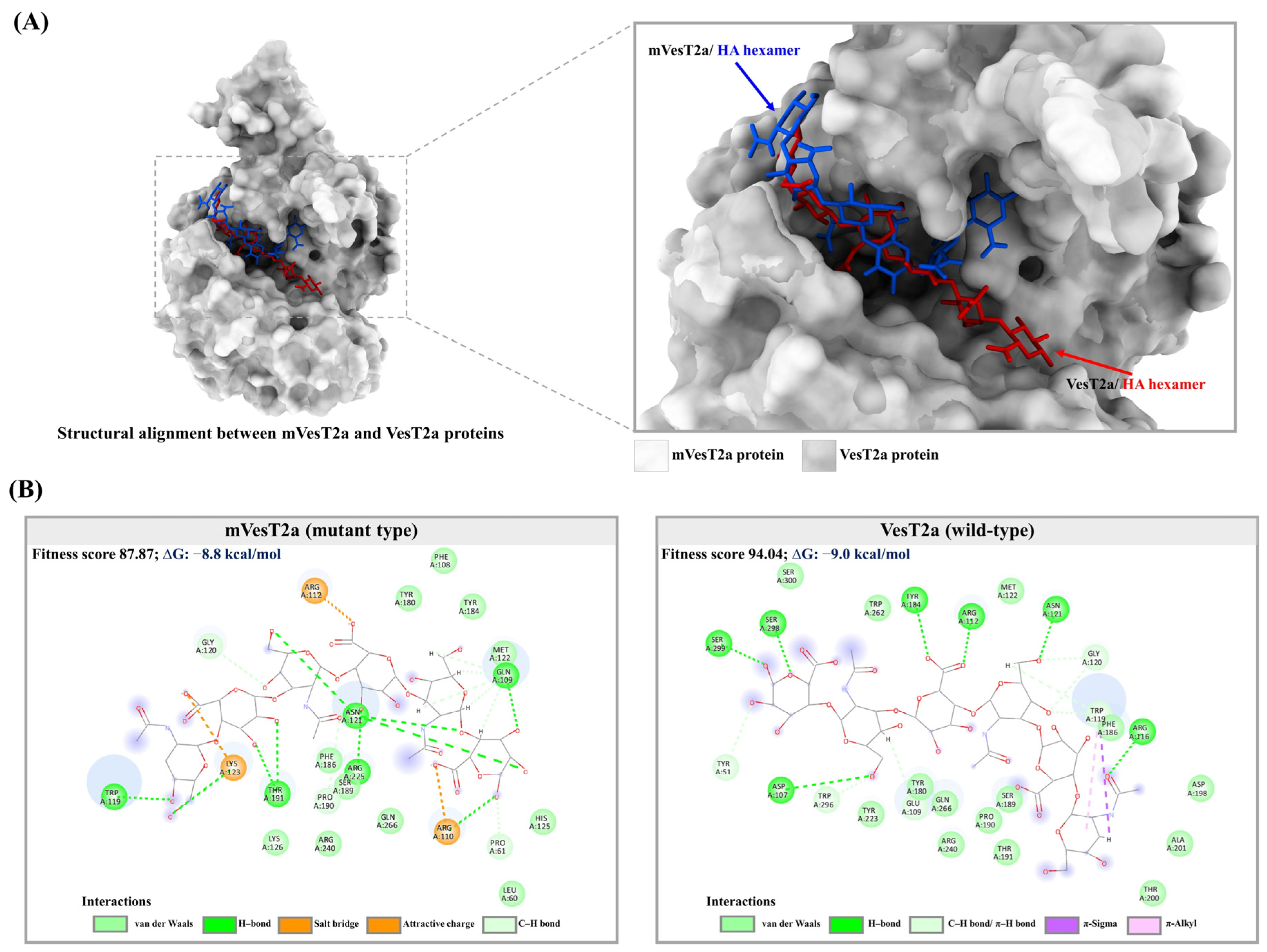
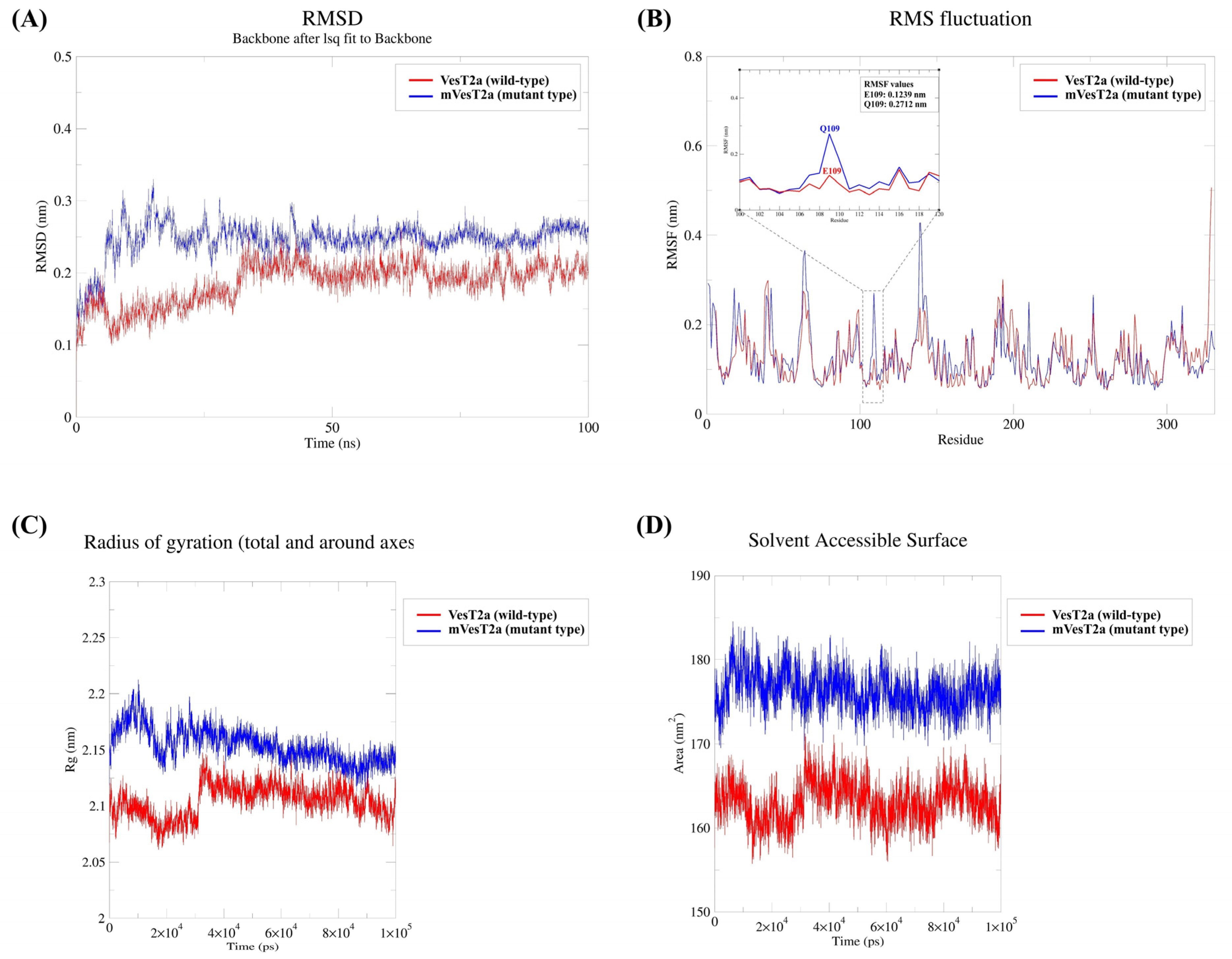
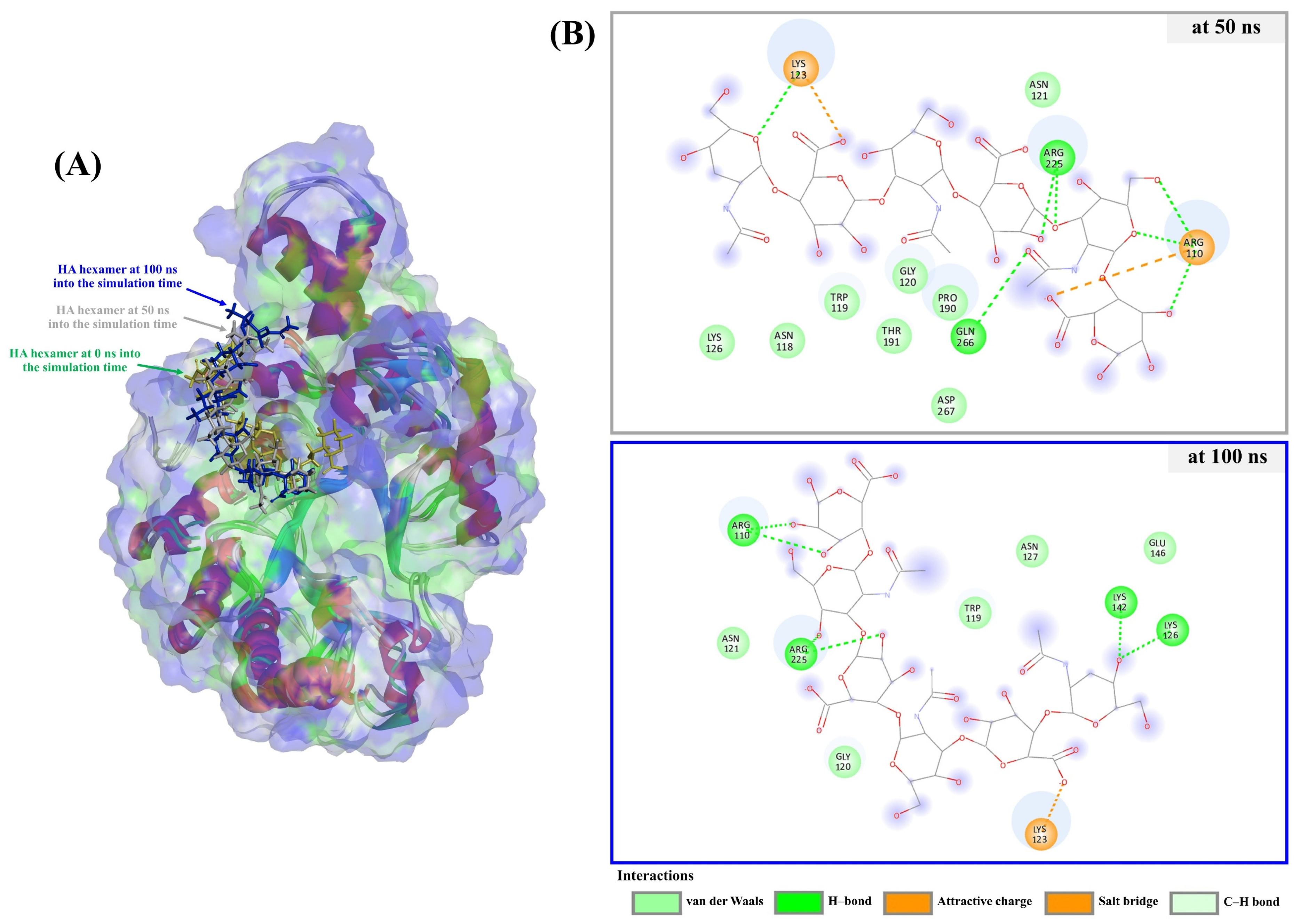
3.4. Molecular Docking and Dynamics Simulation
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- El-safory, N.S.; Fazary, A.E.; Lee, C. Hyaluronidases, a Group of glycosidases: Current and future perspectives. Carbohydr. Polym. 2010, 81, 165–181. [Google Scholar] [CrossRef]
- Girish, K.S.; Kemparaju, K. The magic glue hyaluronan and its eraser hyaluronidase: A biological overview. Life Sci. 2007, 80, 1921–1943. [Google Scholar] [CrossRef]
- Šimek, M.; Lemr, K.; Hermannová, M.; Havlíček, V. Analysis of Hyaluronan and Its Derivatives Using Chromatographic and Mass Spectrometric Techniques. Carbohydr. Polym. 2020, 250, 117014. [Google Scholar] [CrossRef]
- Justo Jacomini, D.L.; Campos Pereira, F.D.; dos Santos Pinto, J.R.A.; dos Santos, L.D.; da Silva Neto, A.J.; Giratto, D.T.; Palma, M.S.; de Lima Zollner, R.; Brochetto Braga, M.R. Hyaluronidase from the venom of the social wasp Polybia paulista (Hymenoptera, Vespidae): Cloning, structural modeling, purification, and immunological analysis. Toxicon 2013, 64, 70–80. [Google Scholar] [CrossRef]
- Bordon, K.C.F.; Wiezel, G.A.; Amorim, F.G.; Arantes, E.C. Arthropod venom hyaluronidases: Biochemical properties and potential applications in medicine and biotechnology. J. Venom. Anim. Toxins Incl. Trop. Dis. 2015, 21, 4. [Google Scholar] [CrossRef]
- Zhang, L.; Bharadwaj, A.G.; Casper, A.; Barkley, J.; Barycki, J.J.; Simpson, M.A. Hyaluronidase activity of human Hyal1 requires active site acidic and tyrosine residues. J. Biol. Chem. 2009, 284, 9433–9442. [Google Scholar] [CrossRef]
- Arming, S.; Strobl, B.; Wechselberger, C.; Kreil, G. In vitro mutagenesis of PH-20 hyaluronidase from human sperm. Eur. J. Biochem. 1997, 247, 810–814. [Google Scholar] [CrossRef]
- Rungsa, P.; Janpan, P.; Saengkun, Y.; Jangpromma, N.; Klaynongsruang, S.; Patramanon, R.; Uawonggul, N.; Daduang, J.; Daduang, S. Heterologous expression and mutagenesis of recombinant Vespa affinis hyaluronidase protein (rVesA2a). J. Venom. Anim. Toxins Incl. Trop. Dis. 2019, 25, e20190030. [Google Scholar] [CrossRef] [PubMed]
- Rungsa, P.; Incamnoi, P.; Sukprasert, S.; Uawonggul, N.; Klaynongsruang, S.; Daduang, J.; Patramanon, R.; Roytrakul, S.; Daduang, S. Comparative proteomic analysis of two wasps venom, Vespa tropica and Vespa affinis. Toxicon 2016, 119, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Marković-Housley, Z.; Miglierini, G.; Soldatova, L.; Rizkallah, P.J.; Müller, U.; Schirmer, T. Crystal structure of hyaluronidase, a major allergen of bee venom. Structure 2000, 8, 1025–1035. [Google Scholar] [CrossRef]
- Rungsa, P.; Incamnoi, P.; Sukprasert, S.; Uawonggul, N.; Klaynongsruang, S.; Daduang, J.; Patramanon, R.; Roytrakul, S.; Daduang, S. Cloning, structural modelling and characterization of VesT2s, a wasp venom hyaluronidase (HAase) from Vespa tropica. J. Venom. Anim. Toxins Incl. Trop. Dis. 2016, 22, 28. [Google Scholar] [CrossRef] [PubMed]
- Martins, J.R.M.; Passerotti, C.C.; Maciel, R.M.B.; Sampaio, L.O.; Dietrich, C.P.; Nader, H.B. Practical determination of hyaluronan by a new noncompetitive fluorescence-based assay on serum of normal and cirrhotic patients. Anal. Biochem. 2003, 319, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Rössler, A. An Ultrasensitive, nonisotopic immunoassay for hyaluronan using the streptavidin-biotin system. Clin. Chim. Acta 1998, 270, 101–114. [Google Scholar] [CrossRef]
- Barone, G.D.; Emmerstorfer-Augustin, A.; Biundo, A.; Pisano, I.; Coccetti, P.; Mapelli, V.; Camattari, A. Industrial production of proteins with Pichia pastoris—Komagataella phaffii. Biomolecules 2023, 13, 441. [Google Scholar] [CrossRef]
- Spohner, S.C.; Müller, H.; Quitmann, H.; Czermak, P. Expression of enzymes for the usage in food and feed industry with Pichia pastoris. J. Biotechnol. 2015, 202, 118–134. [Google Scholar] [CrossRef]
- Prielhofer, R.; Barrero, J.J.; Steuer, S.; Gassler, T.; Zahrl, R.; Baumann, K.; Sauer, M.; Mattanovich, D.; Gasser, B.; Marx, H. GoldenPiCS: A golden gate-derived modular cloning system for applied synthetic biology in the yeast Pichia pastoris. BMC Syst. Biol. 2017, 11, 123. [Google Scholar] [CrossRef]
- Janpan, P.; Schmelzer, B.; Klamrak, A.; Tastub, P.; Upathanpreecha, T.; Rahman, S.S.; Nabnueangsap, J.; Saengkun, Y.; Rungsa, P.; Mattanovich, D.; et al. Production of Vespa tropica hyaluronidase by Pichia pastoris. J. Fungi 2024, 10, 854. [Google Scholar] [CrossRef]
- Zavec, D.; Gasser, B.; Mattanovich, D. Characterization of methanol utilization negative Pichia pastoris for secreted protein production: New cultivation strategies for current and future applications. Biotechnol. Bioeng. 2020, 117, 1394–1405. [Google Scholar] [CrossRef]
- Liu, Y.; Yang, X.; Gan, J.; Chen, S.; Xiao, Z.X.; Cao, Y. CB-Dock2: Improved protein-ligand blind docking by integrating cavity detection, docking and homologous template fitting. Nucleic Acids Res. 2022, 50, 159–164. [Google Scholar] [CrossRef]
- Amin, M.; Barzegari, E.; Pourshohod, A.; Zeinali, M.; Jamalan, M. 3D structure prediction, dynamic investigation and rational construction of an epitope-masked thermostable bovine hyaluronidase. Int. J. Biol. Macromol. 2021, 187, 544–553. [Google Scholar] [CrossRef] [PubMed]
- Fu, H.; Liang, Y.; Zhong, X.; Pan, Z.L.; Huang, L.; Zhang, H.L.; Xu, Y.; Zhou, W.; Liu, Z. Codon optimization with deep learning to enhance protein expression. Sci. Rep. 2020, 10, 17617. [Google Scholar] [CrossRef]
- Xu, Y.; Liu, K.; Han, Y.; Xing, Y.; Zhang, Y.; Yang, Q.; Zhou, M. Codon usage bias regulates gene expression and protein conformation in yeast expression system P. pastoris. Microb. Cell Fact. 2021, 20, 91. [Google Scholar] [CrossRef]
- Yuankai, H.; Chen, Y.; Mo, D.; Cong, P.; He, Z. Attenuated secretion of the thermostable xylanase XynB from Pichia pastoris using synthesized sequences optimized from the preferred codon usage in yeast. J. Microbiol. Biotechnol. 2011, 22, 316–325. [Google Scholar] [CrossRef]
- Naseem, M.U.; Tajti, G.; Gaspar, A.; Szanto, T.G.; Borrego, J.; Panyi, G. Optimization of Pichia pastoris expression system for high-level production of margatoxin. Front. Pharmacol. 2021, 12, 733610. [Google Scholar] [CrossRef] [PubMed]
- Nordén, K.; Agemark, M.; Danielson, J.J.T.; Alexandersson, E.; Kjellbom, P.; Johanson, U. Increasing gene dosage greatly enhances recombinant expression of aquaporins in Pichia pastoris. BMC Biotechnol. 2011, 11, 47. [Google Scholar] [CrossRef] [PubMed]
- Potvin, G.; Ahmad, A.; Zhang, Z. Bioprocess engineering aspects of heterologous protein production in Pichia pastoris: A review. Biochem. Eng. J. 2012, 64, 91–105. [Google Scholar] [CrossRef]
- Waterham, H.R.; Digan, M.E.; Koutz, P.J.; Lair, S.V.; Cregg, J.M.; Bernardi, A. Isolation of the Pichia pastoris glyceraldehyde-3-phosphate dehydrogenase gene and regulation and use of its promoter. Gene 1997, 186, 37–44. [Google Scholar] [CrossRef]
- Anane, E.; van Rensburg, E.; Görgens, J.F. Comparison of constitutive and inducible β-fructofuranosidase production by recombinant Pichia pastoris in fed-batch culture using defined and semi-defined media. S. Afr. J. Chem. Eng. 2016, 22, 17–22. [Google Scholar] [CrossRef]
- Kim, S.; Warburton, S.; Boldogh, I.; Svensson, C.; Pon, L.; d’Anjou, M.; Stadheim, T.A.; Choi, B.K. Regulation of alcohol oxidase 1 (AOX1) promoter and peroxisome biogenesis in different fermentation processes in Pichia pastoris. J. Biotechnol. 2013, 166, 174–181. [Google Scholar] [CrossRef]
- Jia, L.; Tu, T.; Huai, Q.; Sun, J.; Chen, S.; Li, X.; Shi, Z.; Ding, J. Enhancing monellin production by Pichia pastoris at low cell induction concentration via effectively regulating methanol metabolism patterns and energy utilization efficiency. PLoS ONE 2017, 12, e0184602. [Google Scholar] [CrossRef]
- Hartner, F.S.; Glieder, A. Regulation of methanol utilisation pathway genes in yeasts. Microb. Cell Fact. 2006, 5, 39. [Google Scholar] [CrossRef] [PubMed]
- Gidijala, L.; Uthoff, S.; Kampen, S.J.; Steinbüchel, A.; Verhaert, R.M.D. Presence of protein production enhancers results in significantly higher methanol-induced protein production in Pichia pastoris. Microb. Cell Fact. 2018, 17, 112. [Google Scholar] [CrossRef] [PubMed]
- Karbalaei, M.; Rezaee, S.A.; Farsiani, H. Pichia pastoris: A highly successful expression system for optimal synthesis of heterologous proteins. J. Cell. Physiol. 2020, 235, 5867–5881. [Google Scholar] [CrossRef]
- Pan, Y.; Yang, J.; Wu, J.; Yang, L.; Fang, H. Current advances of Pichia pastoris as cell factories for production of recombinant proteins. Front. Microbiol. 2022, 13, 1059777. [Google Scholar] [CrossRef]
- Zha, J.; Liu, D.; Ren, J.; Liu, Z.; Wu, X. Advances in metabolic engineering of Pichia pastoris strains as powerful cell factories. J. Fungi 2023, 9, 1027. [Google Scholar] [CrossRef]
- Zou, C.; Lu, L.; Wang, S.; Zhang, C.; Chen, X.; Lin, Y.; Huang, Y. The α-mating factor secretion signals and endogenous signal peptides for recombinant protein secretion in Komagataella phaffii. Biotechnol. Biofuels Bioprod. 2022, 15, 140. [Google Scholar] [CrossRef]
- Ahmad, M.; Hirz, M.; Pichler, H.; Schwab, H. Protein expression in Pichia pastoris: Recent achievements and perspectives for heterologous protein production. Appl. Microbiol. Biotechnol. 2014, 98, 5301–5317. [Google Scholar] [CrossRef]
- Schwaighofer, A.; Ablasser, S.; Lux, L.; Kopp, J.; Herwig, C.; Spadiut, O.; Lendl, B.; Slouka, C. Production of active recombinant hyaluronidase inclusion bodies from Apis mellifera in E. coli BL21(DE3) and characterization by FT-IR spectroscopy. Int. J. Mol. Sci. 2020, 21, 3881. [Google Scholar] [CrossRef]
- Justo Jacomini, D.L.; Gomes Moreira, S.M.; Campos Pereira, F.D.; Zollner, R.D.L.; Brochetto Braga, M.R. Reactivity of IgE to the allergen hyaluronidase from Polybia paulista (Hymenoptera, Vespidae) venom. Toxicon 2014, 82, 104–111. [Google Scholar] [CrossRef]
- Amorim, F.G.; Boldrini-França, J.; de Castro Figueiredo Bordon, K.; Cardoso, I.A.; De Pauw, E.; Quinton, L.; Kashima, S.; Arantes, E.C. Heterologous expression of rTsHyal-1: The first recombinant hyaluronidase of scorpion venom produced in Pichia pastoris system. Appl. Microbiol. Biotechnol. 2018, 102, 3145–3158. [Google Scholar] [CrossRef]
- Iliás, A.; Liliom, K.; Greiderer-Kleinlercher, B.; Reitinger, S.; Lepperdinger, G. Unbinding of hyaluronan accelerates the enzymatic activity of bee hyaluronidase. J. Biol. Chem. 2011, 286, 35699–35707. [Google Scholar] [CrossRef]
- Kohl, T.O.; Ascoli, C.A. Immunometric double-antibody sandwich enzyme-linked immunosorbent assay. Cold Spring Harb. Protoc. 2017, 2017, 458–462. [Google Scholar] [CrossRef] [PubMed]
- Anagu, L.O.; Andoh, N.E. Vaccine development: From the laboratory to the field. In Vaccinology and Methods in Vaccine Research; Ashfield, R., Ed.; Elsevier: London, UK, 2022; pp. 95–131. [Google Scholar] [CrossRef]
- Vuong, T.V.; Wilson, D.B. Glycoside hydrolases: Catalytic base/nucleophile diversity. Biotechnol. Bioeng. 2010, 107, 195–205. [Google Scholar] [CrossRef] [PubMed]
- Jitonnom, J. Carbohydrate hydrolysis by glycosidase: Mechanistic aspects and applications. KKU Sci. J. 2013, 41, 582–594. [Google Scholar]

| Name | Sequence (5′ to 3′) | Purpose |
|---|---|---|
| F_mVesT2a | GATCGGTCTCGTCAGAAAGGCCCAAAAGAGTGTTTAACATTTACTG | mVesT2a gene |
| R_mVesT2a | GATCGGTCTCCAAGCCTATTAGTGATGGTGGTGGTGATGTCCAC | mVesT2a gene |
| F_BB1 | CAGGAAACAGCTATGAC | Sequencing |
| R_BB1 | GTAAAACGACGGCCAGTT | Sequencing |
| F_GAP | ACCAGAATCGAATATAAA | Sequencing |
| F_AOX1 | CTTTCATAATTGCGACTGGTTC | Sequencing |
| R_BB3 | CGAGCGTCCCAAAACC | Sequencing |
| Promoter | Protein conc. (mg/L) | |
|---|---|---|
| Small-scale production | PGAP | 0.42 |
| PAOX1 | 1.50 | |
| Upscaled production | PAOX1 | 34.03 ± 2.31 |
| mVesT2a (Mutant Type) | VesT2a (Wild Type) | |
|---|---|---|
| Specific activity (U/mg) | 119.28 ± 10.75 (This study) | 4238.37 ± 135.65 [17] |
| Parameters | Developed Method (Indirect ELISA) | Conventional Method (CTAB Assay) |
|---|---|---|
| Linearity and range | 0.050–0.400 µg/mL | 25–500 µg/mL |
| Linear equation | Y = 0.0606X + 0.0107 | Y = 0.0015X + 0.0524 |
| R-squared | R2 = 0.9994 | R2 = 0.9911 |
| Accuracy | 99.34% | 118.34% |
| Precision (%RSD) | 16.61% | 2.44% |
| Limit of Detection (LOD) | 0.095 µg/mL | 1.52 µg/mL |
| Limit of quantitation (LOQ) | 0.197 µg/mL | 4.60 µg/mL |
| Specificity | HA | HA |
| Interferences (% recovery) | ||
| Glucose (0.400 µg/mL) | N.D. | N.D. |
| Chondroitin sulfate (0.400 µg/mL) | Below LOQ | 98.97% |
| Chitosan (0.400 µg/mL) | N.D. | N.D. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Janpan, P.; Schmelzer, B.; Klamrak, A.; Nabnueangsap, J.; Saengkun, Y.; Rahman, S.S.; Rungsa, P.; Mattanovich, D.; Daduang, S. Functional Investigation of Mutant Vespa tropica Hyaluronidase Produced in Pichia pastoris: In Silico Studies and Potential Applications. Fermentation 2025, 11, 497. https://doi.org/10.3390/fermentation11090497
Janpan P, Schmelzer B, Klamrak A, Nabnueangsap J, Saengkun Y, Rahman SS, Rungsa P, Mattanovich D, Daduang S. Functional Investigation of Mutant Vespa tropica Hyaluronidase Produced in Pichia pastoris: In Silico Studies and Potential Applications. Fermentation. 2025; 11(9):497. https://doi.org/10.3390/fermentation11090497
Chicago/Turabian StyleJanpan, Piyapon, Bernhard Schmelzer, Anuwatchakij Klamrak, Jaran Nabnueangsap, Yutthakan Saengkun, Shaikh Shahinur Rahman, Prapenpuksiri Rungsa, Diethard Mattanovich, and Sakda Daduang. 2025. "Functional Investigation of Mutant Vespa tropica Hyaluronidase Produced in Pichia pastoris: In Silico Studies and Potential Applications" Fermentation 11, no. 9: 497. https://doi.org/10.3390/fermentation11090497
APA StyleJanpan, P., Schmelzer, B., Klamrak, A., Nabnueangsap, J., Saengkun, Y., Rahman, S. S., Rungsa, P., Mattanovich, D., & Daduang, S. (2025). Functional Investigation of Mutant Vespa tropica Hyaluronidase Produced in Pichia pastoris: In Silico Studies and Potential Applications. Fermentation, 11(9), 497. https://doi.org/10.3390/fermentation11090497






