The Application of an On-Line Optical Sensor to Measure Biomass of a Filamentous Bioprocess
Abstract
:1. Introduction
2. Experimental Section
2.1. Organism
2.2. Culture Media
2.3. Culture Viability
2.4. Bioprocess
2.5. CPPs
2.6. BugEye® 100
2.7. Sensor Comparison Analysis
3. Results

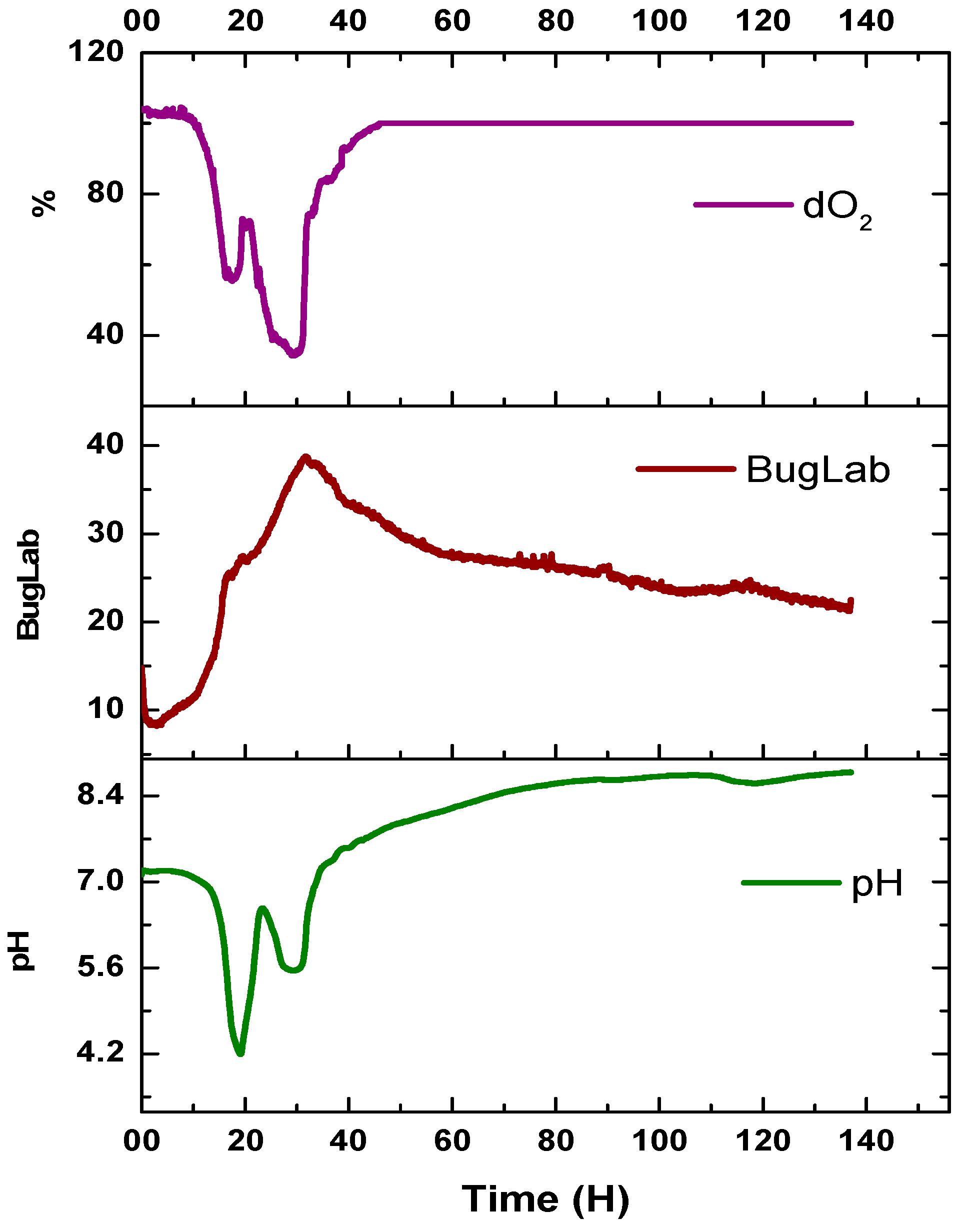
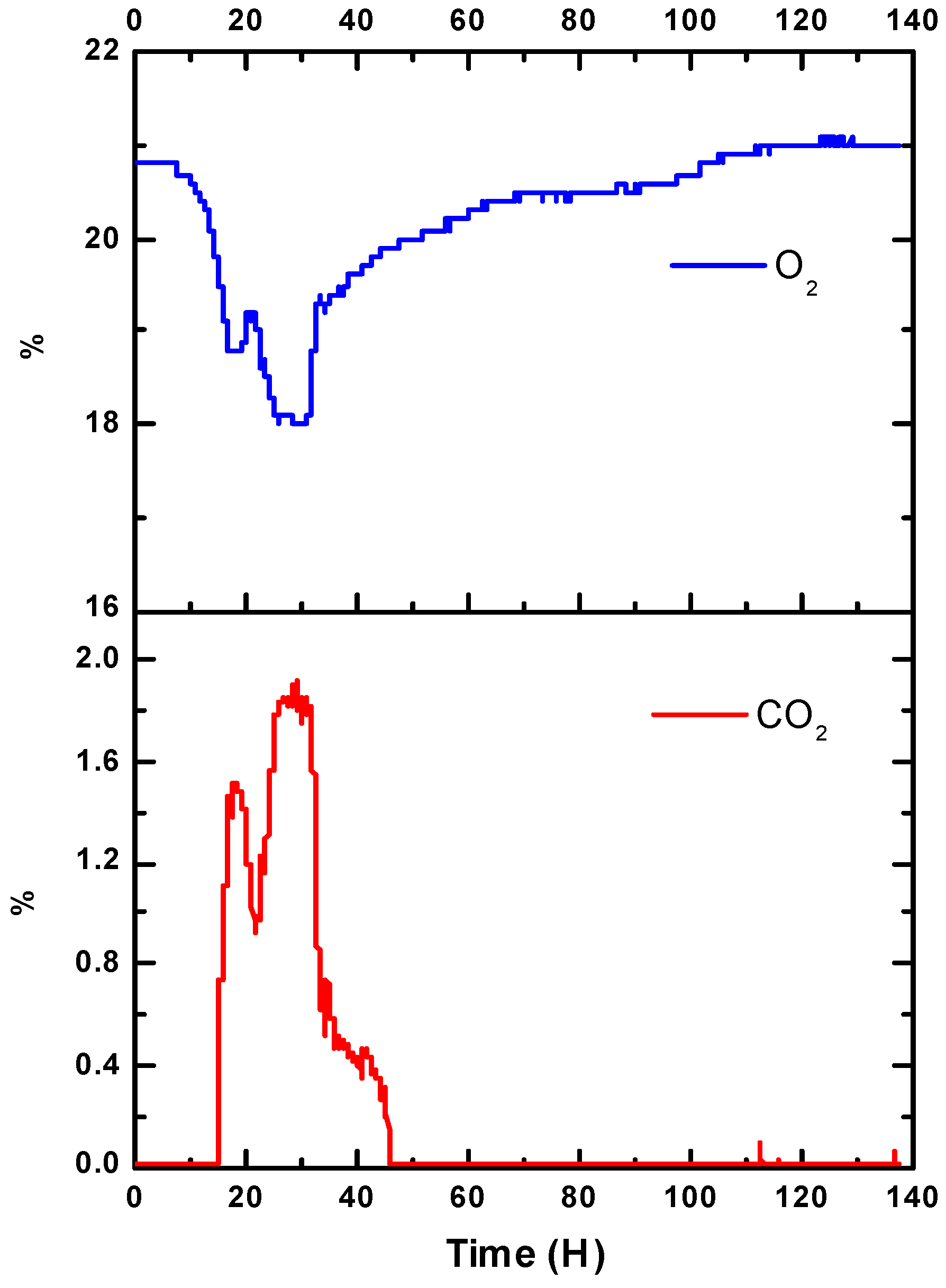
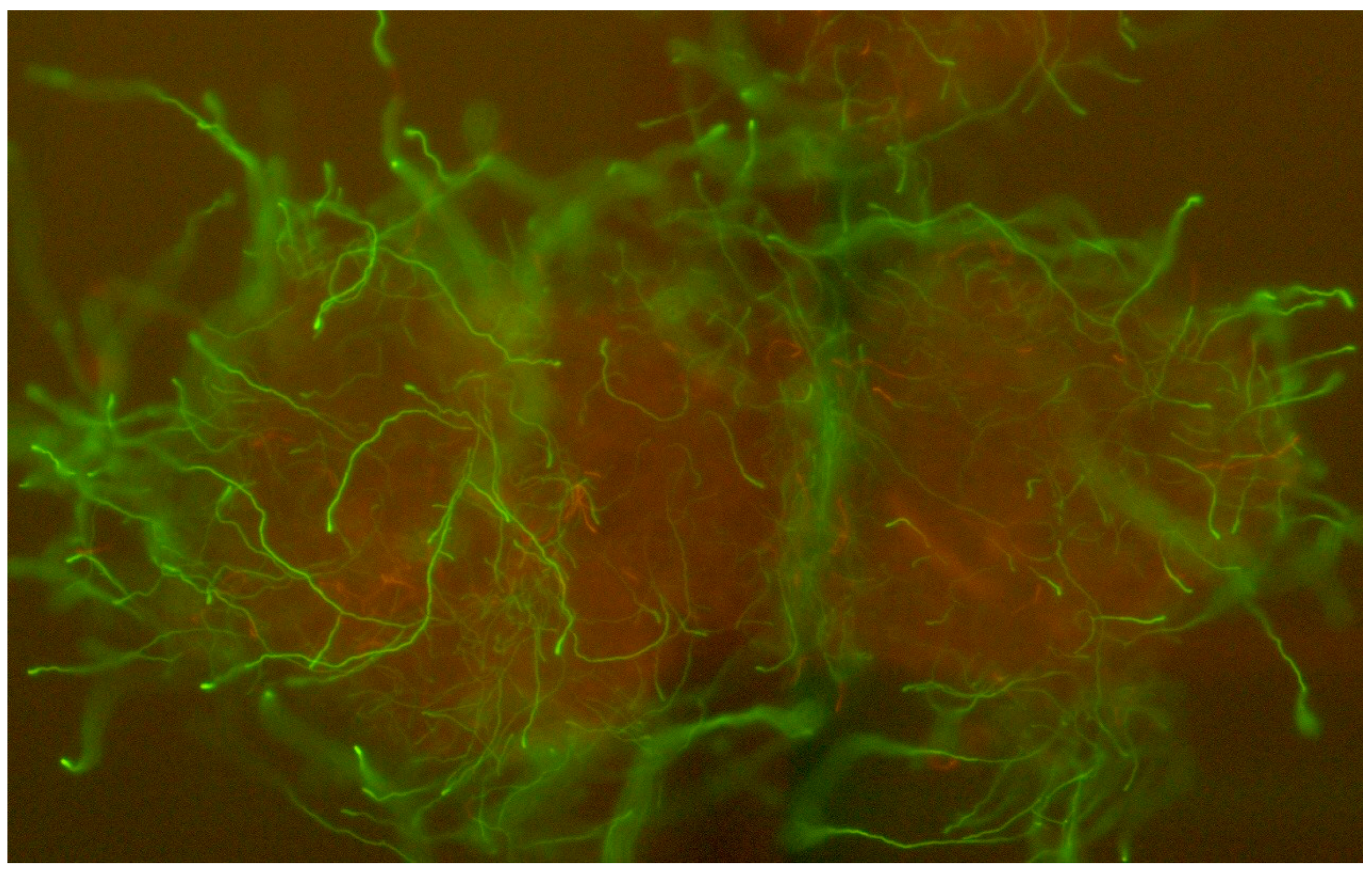
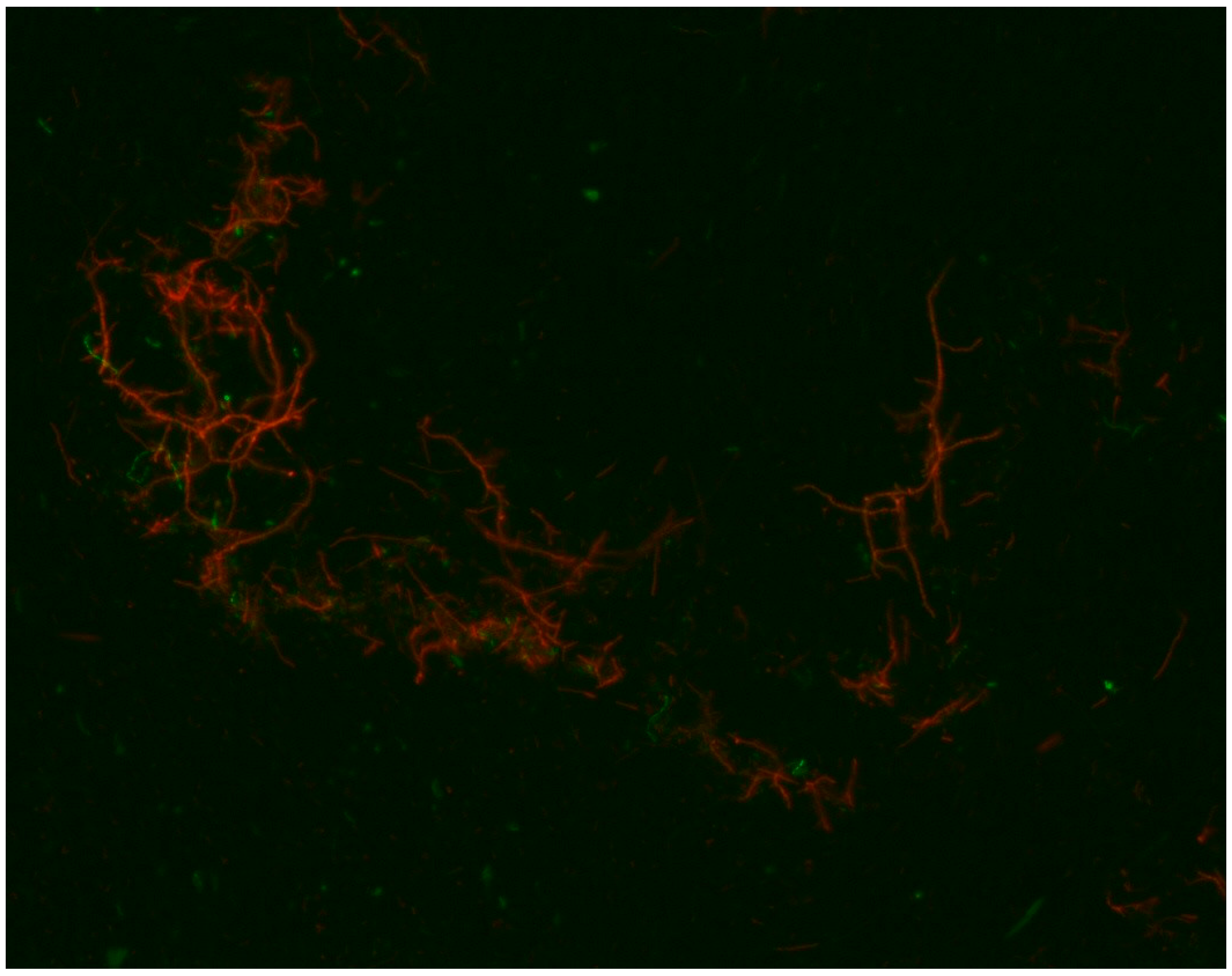
4. Discussion
5. Conclusions
Author Contributions
Conflicts of Interest
References
- Harms, P.; Kostov, Y.; Rao, G. Review: Bioprocess monitoring. Curr. Opin. Biotechnol. 2002, 13, 124–127. [Google Scholar] [CrossRef]
- Nordon, A.; littlejohn, D.; Dann, A.S.; Jeffkins, P.A.; Richardson, M.D.; Stimpson, S.L. In situ monitoring of the seed stage of a fermentation process using non-invasive NIR spectrometry. Analyst 2007, 133, 660–666. [Google Scholar] [CrossRef]
- Tiwari, S.; Suraishkumar, G.K.; Chandavarkar, A. Robust near-ifra-red spectroscopic probe for dynamic monitoring of critical nutrient ration in microbial fermentation processes. Biochem. Eng. J. 2013, 71, 47–56. [Google Scholar] [CrossRef]
- Landgrebe, D.; Haake, C.; Höpfner, T.; Beutel, S.; Hitzmann, B.; Scheper, T.; Rhiel, M.; Reardon, K. On-line infrared spectroscopy for bioprocess monitoring. Appl. Microbiol. Biotechnol. 2010, 88, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, S.; Suraishkumar, G.K.; Chandavarkar, A. Robust near-infra-red spectroscopic probe for dynamic monitoring of critical nutrient ratio in microbial fermentation processes. Biochem. Eng. J. 2013, 71, 47–56. [Google Scholar] [CrossRef]
- Cervera, A.E.; Petersen, N.; Lantz, A.E.; Larsen, A.; Gernaey, K.V. Application of near-infrared spectroscopy for monitoring and control of cell culture and fermentation. Biotechnol. Prog. 2009, 25, 1561–1581. [Google Scholar] [CrossRef] [PubMed]
- Bentley, S.D.; Chater, K.F.; Cerdeno-Tarraga, A.M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3 (2). Nature 2002, 417, 141–145. [Google Scholar] [CrossRef] [PubMed]
- Debreczeny, M.P.; Davies, E.T. Non-invasive Biomass Monitor with Wide Linear Range. 2012. Available online: http://www.buglab.com/docs/OnLineODMeasurement.pdf (accessed on 21 September 2015).
- Dyson, P.J. Streptomyces: Molecular Biology and Biotechnology; Dyson, P., Ed.; Caister Academic: Wymondham, UK, 2011. [Google Scholar]
- Hoskisson, P.A.; Makin, V.; Taylor, H.; Sharples, G.P.; Hobbs, G. Intracellular protease activity during the growth of Streptomyces coelicolor. Biotechnol. Lett. 2001, 23, 185–187. [Google Scholar] [CrossRef]
- Hayes, A.; Hobbs, G.; Smith, C.P.; Oliver, S.G.; Butler, P.R. Environmental signals triggering methylenomycin production by Streptomyces coelicolor A3 (2). J. Bacteriol. 1997, 179, 5511–5515. [Google Scholar] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nakouti, I.; Hobbs, G. The Application of an On-Line Optical Sensor to Measure Biomass of a Filamentous Bioprocess. Fermentation 2015, 1, 79-85. https://doi.org/10.3390/fermentation1010079
Nakouti I, Hobbs G. The Application of an On-Line Optical Sensor to Measure Biomass of a Filamentous Bioprocess. Fermentation. 2015; 1(1):79-85. https://doi.org/10.3390/fermentation1010079
Chicago/Turabian StyleNakouti, Ismini, and Glyn Hobbs. 2015. "The Application of an On-Line Optical Sensor to Measure Biomass of a Filamentous Bioprocess" Fermentation 1, no. 1: 79-85. https://doi.org/10.3390/fermentation1010079




