Same Fragments, Different Diseases: Analysis of Identical tRNA Fragments Across Diseases Utilizing Functional and Abundance-Based Databases
Abstract
1. Introduction
- (1)
- To catalog and compare instances of repeated tRFs associated with diseases;
- (2)
- To check the consistency of reported targets through literature- and database-supported motif verification to investigate if these overlaps signify biologically significant convergence or merely incidental sequence reuse;
- (3)
- To assess whether enrichment patterns from abundance-based databases align with disease-specific roles inferred from experimental validation in disease contexts.
2. Results
2.1. Candidate tRFs
2.2. Data Extraction and Annotation
2.3. Database Cross-Referencing of tRF Sequences
2.4. Cross-Referencing of Target Genes
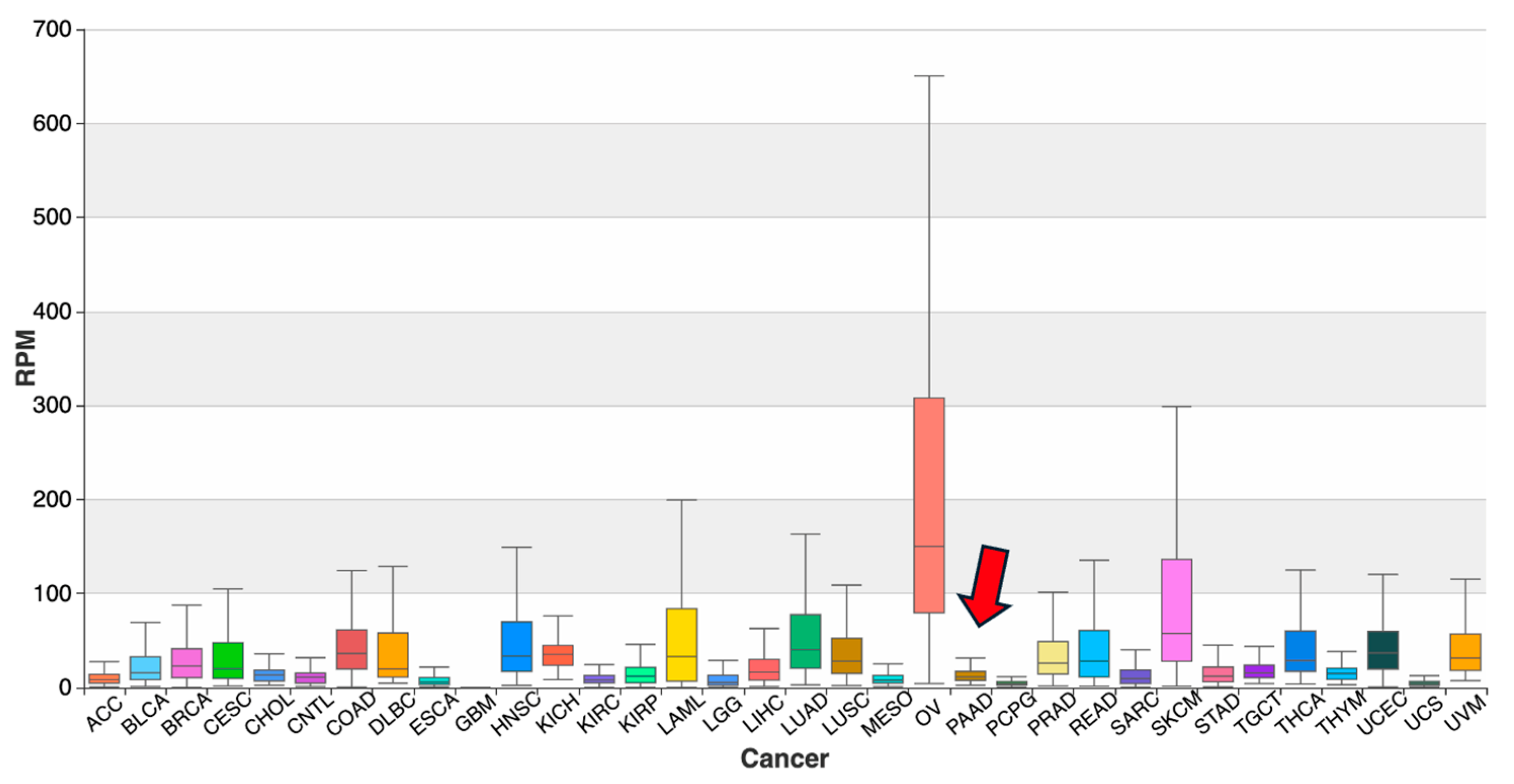
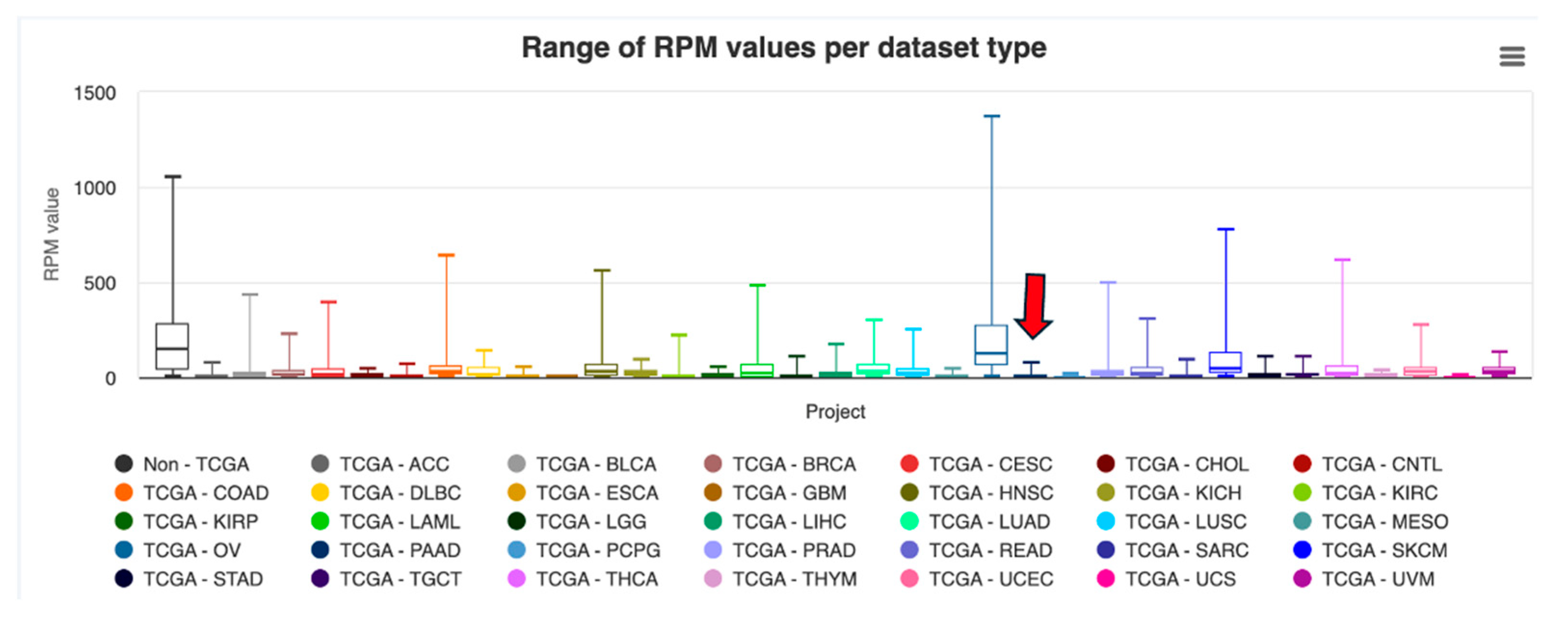
2.5. Abundance Data on tRFs in OncotRF and MINTbase
3. Discussion
4. Materials and Methods
4.1. Selection of Candidate tRFs
- The tRF sequence had to be exactly identical (full sequence match) across at least two independently published studies.
- Each matched tRF sequence had to be associated with distinct disease contexts, indicating its potential multifunctional or pleiotropic role across diseases.
- Studies reporting functional overlaps with divergent (non-identical) tRF sequences were explicitly excluded to ensure a clear and unambiguous focus on exact sequence matches and their implications.
4.2. Data Extraction and Annotation
4.3. Database Cross-Referencing of tRF Sequences
4.4. Cross-Referencing of Target Genes and Motif-Level Confirmation
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| Abbreviation | Cancer Type (Full Name) |
|---|---|
| ACC | Adrenocortical Carcinoma |
| BLCA | Bladder Urothelial Carcinoma |
| BRCA | Breast Invasive Carcinoma |
| CESC | Cervical Squamous Cell Carcinoma and Endocervical Adenocarcinoma |
| CHOL | Cholangiocarcinoma |
| COAD | Colon Adenocarcinoma |
| DLBC | Diffuse Large B-cell Lymphoma |
| ESCA | Esophageal Carcinoma |
| GBM | Glioblastoma Multiforme |
| HNSC | Head and Neck Squamous Cell Carcinoma |
| KICH | Kidney Chromophobe |
| KIRC | Kidney Renal Clear Cell Carcinoma |
| KIRP | Kidney Renal Papillary Cell Carcinoma |
| LAML | Acute Myeloid Leukemia |
| LGG | Lower Grade Glioma |
| LIHC | Liver Hepatocellular Carcinoma |
| LUAD | Lung Adenocarcinoma |
| LUSC | Lung Squamous Cell Carcinoma |
| MESO | Mesothelioma |
| OV | Ovarian Serous Cystadenocarcinoma |
| PAAD | Pancreatic Adenocarcinoma |
| PCPG | Pheochromocytoma and Paraganglioma |
| PRAD | Prostate Adenocarcinoma |
| READ | Rectum Adenocarcinoma |
| SARC | Sarcoma |
| SKCM | Skin Cutaneous Melanoma |
| STAD | Stomach Adenocarcinoma |
| TGCT | Testicular Germ Cell Tumors |
| THCA | Thyroid Carcinoma |
| THYM | Thymoma |
| UCEC | Uterine Corpus Endometrial Carcinoma |
| UCS | Uterine Carcinosarcoma |
| UVM | Uveal Melanoma |
| CNTL | Control (non-cancer samples) |
| Non-TCGA | Non-TCGA dataset |
References
- Guan, L.; Lam, V.; Grigoriev, A. Large-Scale Computational Discovery of Binding Motifs in tRNA Fragments. Front. Mol. Biosci. 2021, 8, 647449. [Google Scholar] [CrossRef]
- Kuscu, C.; Kumar, P.; Kiran, M.; Su, Z.; Malik, A.; Dutta, A. tRNA fragments (tRFs) guide Ago to regulate gene expression post-transcriptionally in a Dicer-independent manner. RNA 2018, 24, 1093–1105. [Google Scholar] [CrossRef] [PubMed]
- Vaidhyanathan, S.; Durbin, M.; Adetowubo, A.A.; Do, L.H.; Kavehmoghaddam, S.; Jonnalagadda, S.A.; Aguilar, B.R.; Ortiz-Gomez, T.; Lin, Y.X.; Dave, A.; et al. Mapping Current Studies of tRNA Fragments onto Disease Landscape. Biomolecules 2025, 15, 512. [Google Scholar] [CrossRef] [PubMed]
- Pliatsika, V.; Loher, P.; Magee, R.; Telonis, A.G.; Londin, E.; Shigematsu, M.; Kirino, Y.; Rigoutsos, I. MINTbase v2.0: A comprehensive database for tRNA-derived fragments that includes nuclear and mitochondrial fragments from all The Cancer Genome Atlas projects. Nucleic Acids Res. 2018, 46, D152–D159. [Google Scholar] [CrossRef]
- Helwak, A.; Kudla, G.; Dudnakova, T.; Tollervey, D. Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell 2013, 153, 654–665. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Anaya, J.; Mudunuri, S.B.; Dutta, A. Meta-analysis of tRNA derived RNA fragments reveals that they are evolutionarily conserved and associate with AGO proteins to recognize specific RNA targets. BMC Biol. 2014, 12, 78. [Google Scholar] [CrossRef]
- Guan, L.; Karaiskos, S.; Grigoriev, A. Inferring targeting modes of Argonaute-loaded tRNA fragments. RNA Biol. 2020, 17, 1070–1080. [Google Scholar] [CrossRef]
- Moore, M.J.; Scheel, T.K.; Luna, J.M.; Park, C.Y.; Fak, J.J.; Nishiuchi, E.; Rice, C.M.; Darnell, R.B. miRNA-target chimeras reveal miRNA 3′-end pairing as a major determinant of Argonaute target specificity. Nat. Commun. 2015, 6, 8864. [Google Scholar] [CrossRef]
- Guan, L.; Grigoriev, A. tatDB: A database of Ago1-mediated targets of transfer RNA fragments. Nucleic Acids Res. 2023, 51, D297–D305. [Google Scholar] [CrossRef]
- Zhou, Y.; Peng, H.; Cui, Q.; Zhou, Y. tRFTar: Prediction of tRF-target gene interactions via systemic re-analysis of Argonaute CLIP-seq datasets. Methods 2021, 187, 57–67. [Google Scholar] [CrossRef]
- Yao, D.; Sun, X.; Zhou, L.; Amanullah, M.; Pan, X.; Liu, Y.; Liang, M.; Liu, P.; Lu, Y. OncotRF: An online resource for exploration of tRNA-derived fragments in human cancers. RNA Biol. 2020, 17, 1081–1091. [Google Scholar] [CrossRef] [PubMed]
- Tu, M.; Zuo, Z.; Chen, C.; Zhang, X.; Wang, S.; Chen, C.; Sun, Y. Transfer RNA-derived small RNAs (tsRNAs) sequencing revealed a differential expression landscape of tsRNAs between glioblastoma and low-grade glioma. Gene 2023, 855, 147114. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Gu, Y.; Ge, J.; Xie, Y.; Yu, X.; Wu, X.; Sun, D.; Zhang, X.; Guo, J.; Guo, J. tRF-33-P4R8YP9LON4VDP inhibits gastric cancer progression via modulating STAT3 signaling pathway in an AGO2-dependent manner. Oncogene 2024, 43, 2160–2171. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Hu, B.; Hu, G.W.; Chen, C.Y.; Niu, X.; Liu, J.; Zhou, S.M.; Zhang, C.Q.; Wang, Y.; Deng, Z.F. tRNA-Derived Small Non-Coding RNAs in Response to Ischemia Inhibit Angiogenesis. Sci. Rep. 2016, 6, 20850. [Google Scholar] [CrossRef]
- Wu, W.; Lee, I.; Spratt, H.; Fang, X.; Bao, X. tRNA-Derived Fragments in Alzheimer’s Disease: Implications for New Disease Biomarkers and Neuropathological Mechanisms. J. Alzheimer’s Dis. 2021, 79, 793–806. [Google Scholar] [CrossRef]
- Chang, X.; Du, M.; Wei, J.; Zhang, Y.; Feng, X.; Deng, B.; Liu, P.; Wang, Y. Serum tsncRNAs reveals novel potential therapeutic targets of Salvianolic Acid B on atherosclerosis. Phytomedicine 2024, 134, 155994. [Google Scholar] [CrossRef]
- Herrero-Lorenzo, M.; Pérez-Pérez, J.; Escaramís, G.; Martínez-Horta, S.; Pérez-González, R.; Rivas-Asensio, E.; Kulisevsky, J.; Gámez-Valero, A.; Martí, E. Small RNAs in plasma extracellular vesicles define biomarkers of premanifest changes in Huntington’s disease. J. Extracell. Vesicles 2024, 13, e12522. [Google Scholar] [CrossRef]
- Lan, S.; Liu, S.; Wang, K.; Chen, W.; Zheng, D.; Zhuang, Y.; Zhang, S. tRNA-derived RNA fragment, tRF-18-8R6546D2, promotes pancreatic adenocarcinoma progression by directly targeting ASCL2. Gene 2024, 927, 148739. [Google Scholar] [CrossRef]
- Sui, S.; Wang, Z.; Cui, X.; Jin, L.; Zhu, C. The biological behavior of tRNA-derived fragment tRF-Leu-AAG in pancreatic cancer cells. Bioengineered 2022, 13, 10617–10628. [Google Scholar] [CrossRef]
- Winek, K.; Lobentanzer, S.; Nadorp, B.; Dubnov, S.; Dames, C.; Jagdmann, S.; Moshitzky, G.; Hotter, B.; Meisel, C.; Greenberg, D.S.; et al. Transfer RNA fragments replace microRNA regulators of the cholinergic poststroke immune blockade. Proc. Natl. Acad. Sci. USA 2020, 117, 32606–32616. [Google Scholar] [CrossRef]
- Liang, Y.; Ji, D.; Ying, X.; Ma, R.; Ji, W. tsRNA modifications: An emerging layer of biological regulation in disease. J. Adv. Res. 2025, 74, 403–414. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.-H.; Chen, W.-X.; Mei, S.-Q.; Yang, Y.-D.; Yang, J.-H.; Qu, L.-H.; Zheng, L.-L. tsRFun: A comprehensive platform for decoding human tsRNA expression, functions and prognostic value by high-throughput small RNA-Seq and CLIP-Seq data. Nucleic Acids Res. 2021, 50, D421–D431. [Google Scholar] [CrossRef] [PubMed]



| tRF ID | Sequence (5′–3′) | Disease Contexts | Validated Target(s) | Reported Functional Role |
|---|---|---|---|---|
| GlyGCC-001-N-5i-1-33 | GCATGGGTGGTTCAGTGGTAGAATTCTCGCCTG | Gastric cancer [13]; Glioblastoma [12] | STAT3 None | Suppression of tumor progression via STAT3 inhibition |
| GlyCCC-001-N-5i-1-32 | GCATTGGTGGTTCAGTGGTAGAATTCTCGCCT | Alzheimer’s disease [15]; Ischemia [14] | None | Inhibition of angiogenesis via modulation of endothelial cell function; biogenesis linked to angiogenin activity |
| GluCTC-001-N-5p-1-31 | TCCCTGGTGGTCTAGTGGTTAGGATTCGGCG | Huntington’s disease [17]; Atherosclerosis [16] | SRF; ARRB | Regulation of neuronal and vascular inflammation pathways |
| LeuAAG-001-N-3p-68-85 | ATCCCACCGCTGCCACCA | Pancreatic cancer [19]; Stroke [20] | UPF1; None | Promotion of tumor proliferation via UPF1 suppression |
| AlaAGC-002-N-3p-58-75 | TCCCCGGCACCTCCACCA | Pancreatic adenocarcinoma [18]; Stroke [20] | ASCL2 None | Promotion of tumor proliferation via ASCL2 inhibition |
| Guide Sequence (5′–3′) | Target Gene |
|---|---|
| GCATGGGTGGTTCAGTGGTAGAATTCTCGCCTG | EIF2AK1 |
| GCATGGGTGGTTCAGTGGTAGAATTCTCGCCTGC | EEF1A1 |
| GCATGGGTGGTTCAGTGGTAGAATTCTCGCCTGC | FASN |
| GCATGGGTGGTTCAGTGGTAGAATTCTCGCCTGC | GAPDH |
| GCATTGGTGGTTCAGTGGTAGAATTCTCGCCTCCCAC | NEO1 |
| Data Type | Description |
|---|---|
| tRF Nucleotide Sequence | Full tRF sequence (5′–3′) extracted from original publications. |
| Disease Context | Disease(s) in which each tRF was reported to play a role. |
| Experimentally Validated Targets | Genes confirmed as targets using qRT-PCR, luciferase reporter assays, or Western blot. |
| Seed Region or Motif | Reported functional motifs or seed regions relevant to target interaction. |
| Validation Metadata | Position of interaction and evidence tier (e.g., high-confidence vs. putative). |
| Database | Purpose |
|---|---|
| tatDB [9] | Provided CLASH-based evidence for tRF–mRNA interactions, motif matches, and hybrid structures in AGO1-loaded complexes. |
| tRFTar [10] | Offered computational target predictions using machine learning trained on CLASH and CLEAR-CLIP datasets. |
| tsRFun [22] | Enabled functional enrichment analysis of tRFs and associated targets in human tissues. |
| OncotRF [11] | Displayed tumor–normal abundance data (RPM) across TCGA datasets; no target predictions included. |
| MINTbase v2.0 [4] | Visualized tRF distribution across TCGA and non-TCGA projects using sequence queries and abundance filters (e.g., RPM ≥ 1). |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Adetowubo, A.; Vaidhyanathan, S.; Grigoriev, A. Same Fragments, Different Diseases: Analysis of Identical tRNA Fragments Across Diseases Utilizing Functional and Abundance-Based Databases. Non-Coding RNA 2025, 11, 63. https://doi.org/10.3390/ncrna11050063
Adetowubo A, Vaidhyanathan S, Grigoriev A. Same Fragments, Different Diseases: Analysis of Identical tRNA Fragments Across Diseases Utilizing Functional and Abundance-Based Databases. Non-Coding RNA. 2025; 11(5):63. https://doi.org/10.3390/ncrna11050063
Chicago/Turabian StyleAdetowubo, Adesupo, Sathyanarayanan Vaidhyanathan, and Andrey Grigoriev. 2025. "Same Fragments, Different Diseases: Analysis of Identical tRNA Fragments Across Diseases Utilizing Functional and Abundance-Based Databases" Non-Coding RNA 11, no. 5: 63. https://doi.org/10.3390/ncrna11050063
APA StyleAdetowubo, A., Vaidhyanathan, S., & Grigoriev, A. (2025). Same Fragments, Different Diseases: Analysis of Identical tRNA Fragments Across Diseases Utilizing Functional and Abundance-Based Databases. Non-Coding RNA, 11(5), 63. https://doi.org/10.3390/ncrna11050063







