The Conservation and Study of Macromycetes in the Komarov Botanical Institute Basidiomycetes Culture Collection—Their Taxonomical Diversity and Biotechnological Prospects
Abstract
:1. Introduction
2. Materials and Methods Applied in the LE-BIN Collection
2.1. Origination and Isolation of Fungi
2.2. Preservation Methods
2.3. Strains Verification Methods
2.3.1. Cultivation Media
2.3.2. Morphological Methods
2.3.3. Molecular Methods
2.3.4. Preparing of Substrates for Fruiting in Culture
2.4. Growth and Enzymatic Activity Methods
2.4.1. Lineal Growth in Petri Dishes
2.4.2. Cultivation on Liquid Media
2.4.3. Express Method for Oxidative Enzymes
2.4.4. Enzymatic Activity by Application Tests
2.4.5. Analysis of Lipids
2.4.6. Analysis of Secondary Metabolites
2.5. Taxonomical Analysis
3. Results and Discussion
3.1. Ex Situ Conservation of Macromycetes Diversity
3.2. Verification Results
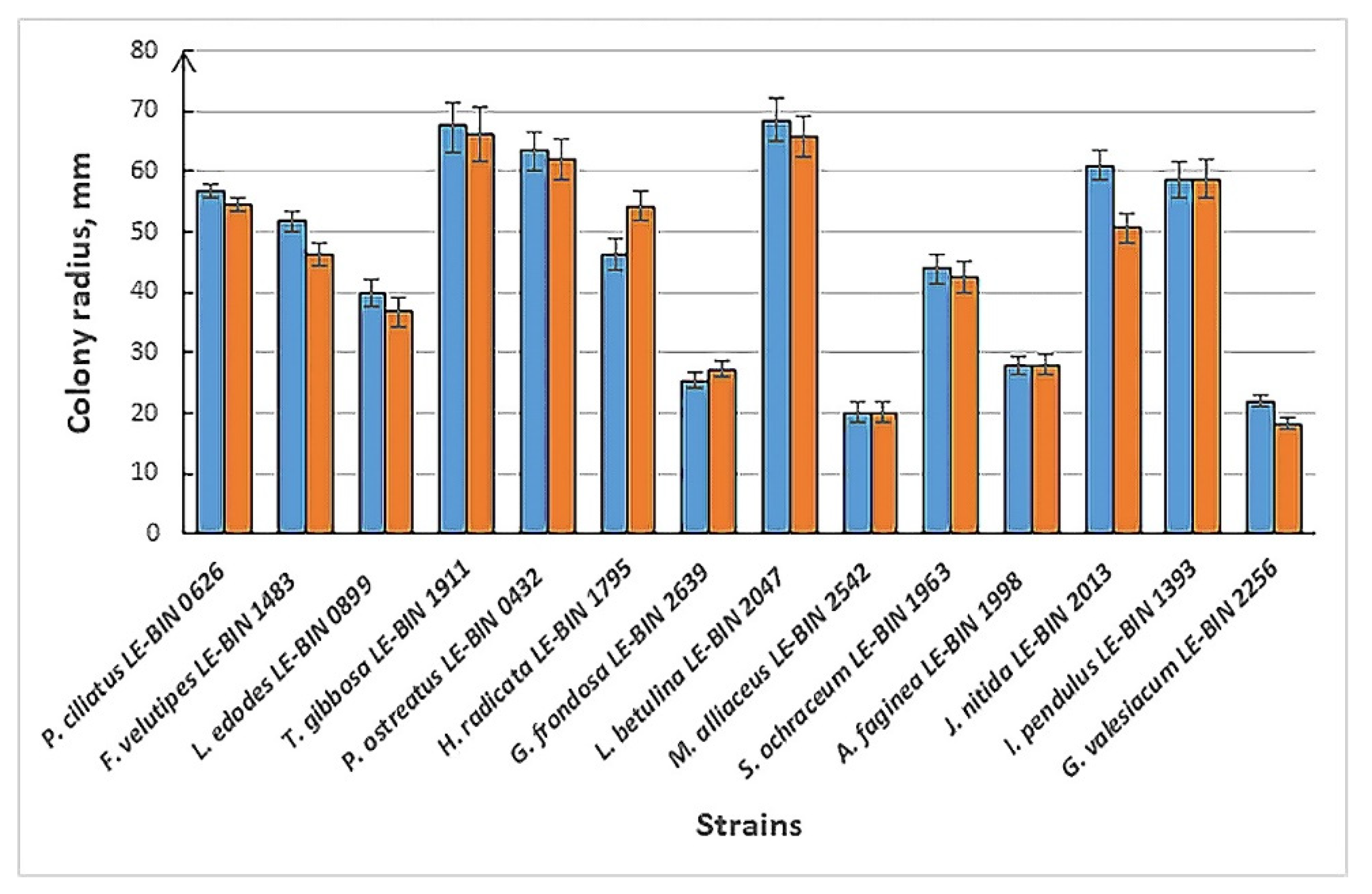
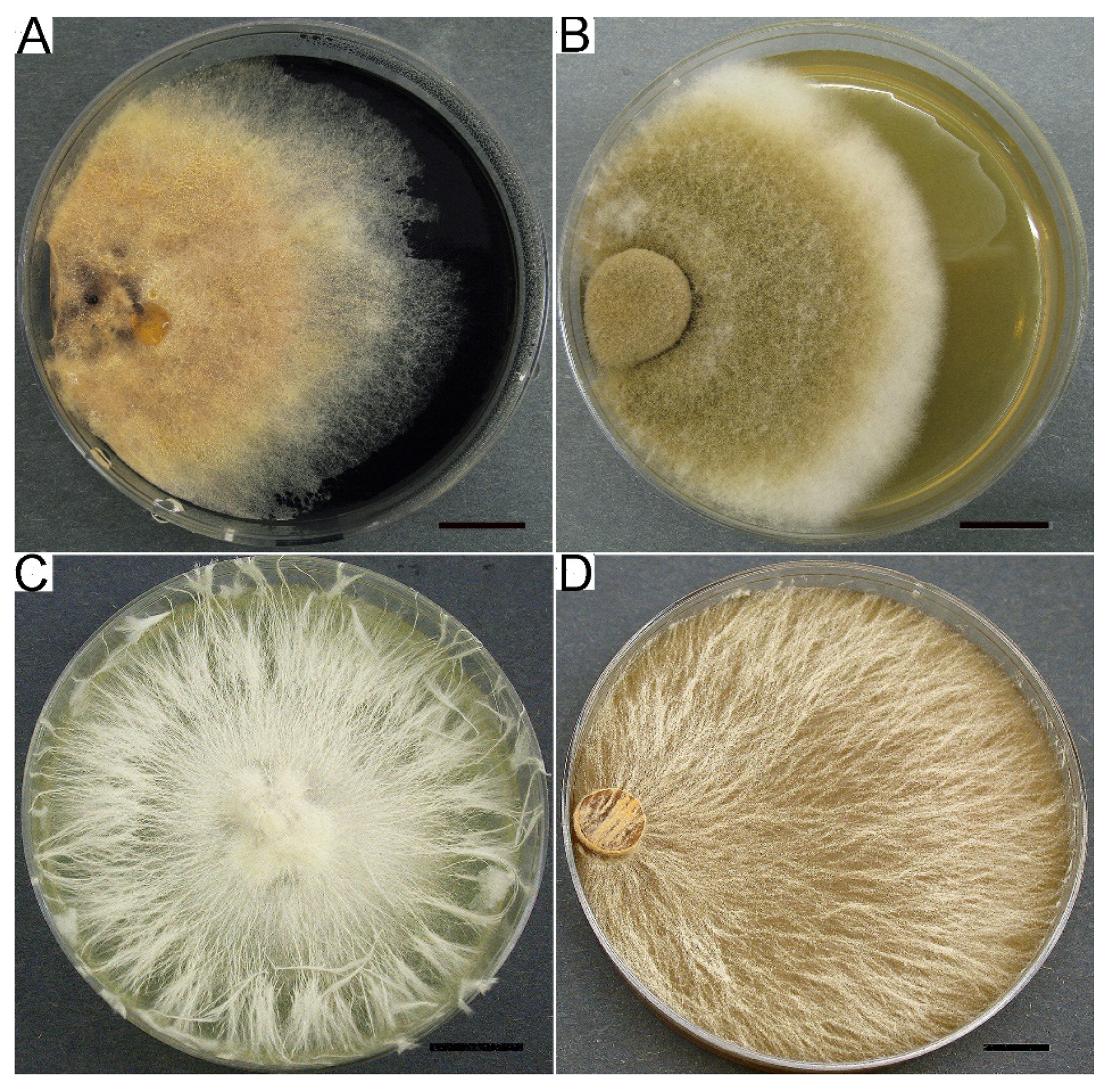
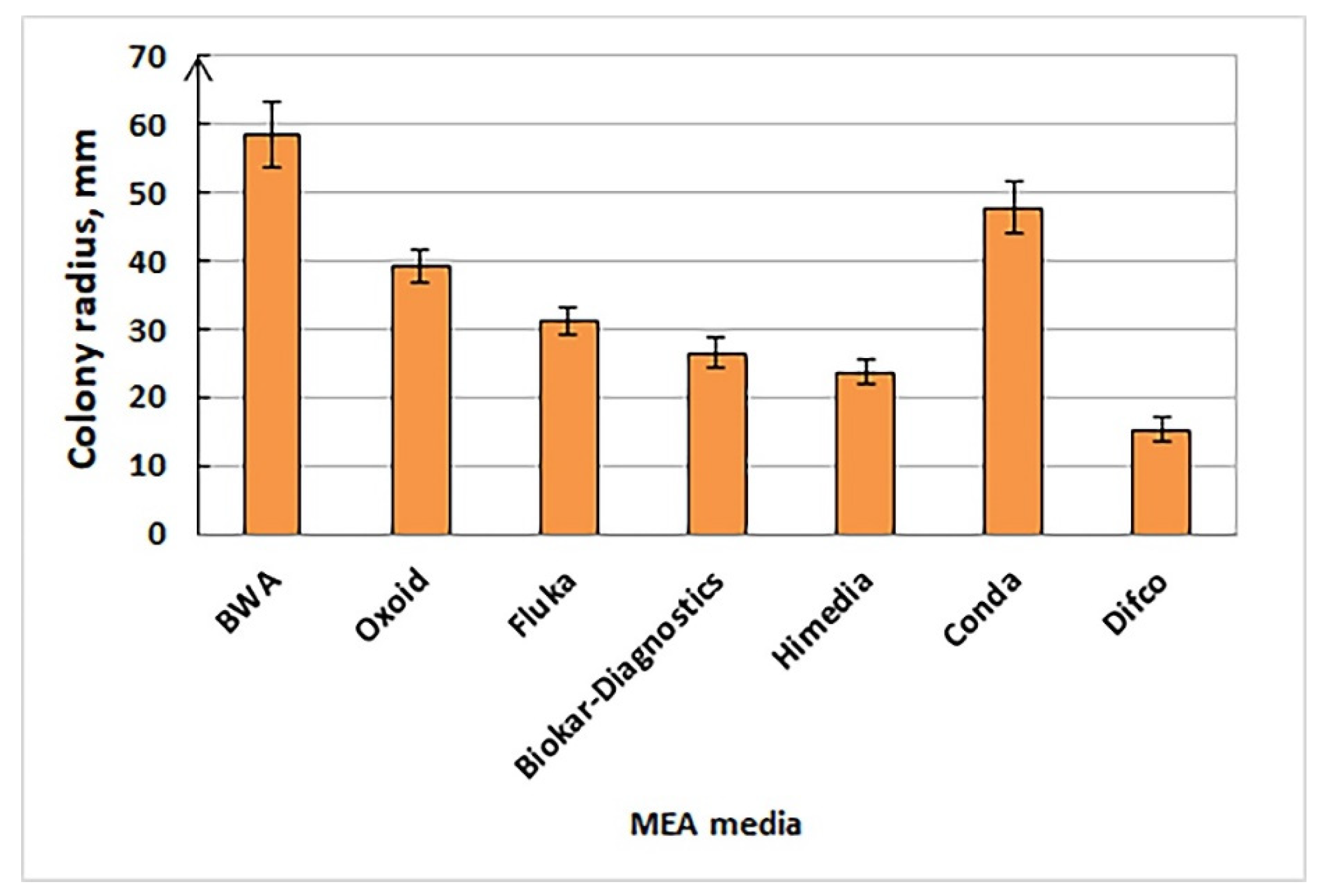
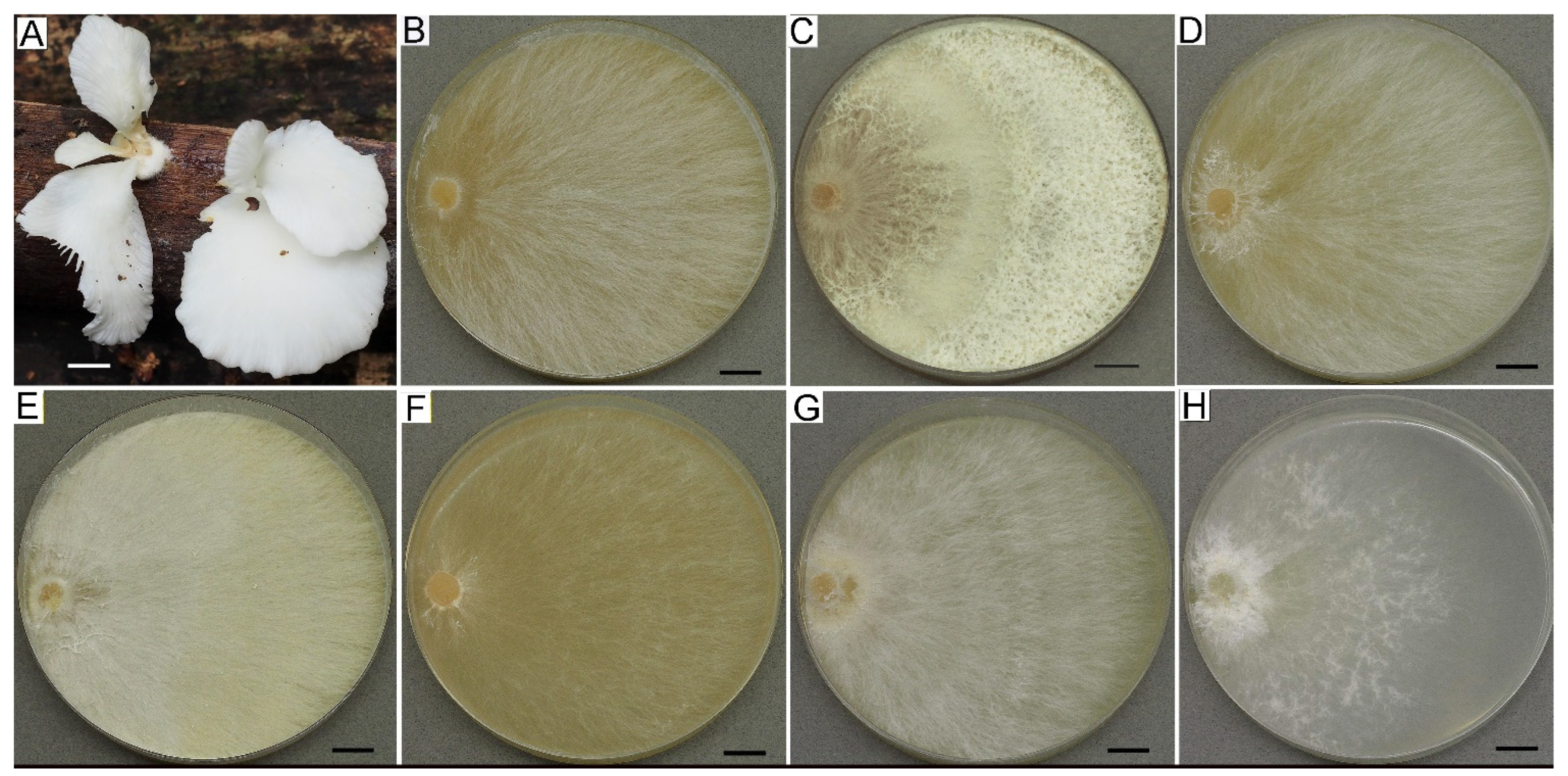
3.2.1. Cultural Characteristics
3.2.2. Fruiting in Culture
3.2.3. Molecular Analysis
3.3. Taxonomical Structure of the LE-BIN Fund
3.4. Biosynthetic and Biotechnological Potential
3.4.1. Oxidase, Cellulase, and Lignin Degradation
3.4.2. Proteinases
3.4.3. Lipids
3.4.4. Secondary Metabolites
3.5. Medical and Veterinary Aspects of LE-BIN Strains Application
3.6. Information on the LE-BIN Fungi (Databases and Catalogs)
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Singh, S.K. Ex situ Conservation of Fungi: A Review. In Developments in Fungal Biology and Applied Mycology; Satyanarayana, T., Deshmukh, S., Johri, B., Eds.; Springer: Singapore, 2017; pp. 543–562. [Google Scholar] [CrossRef]
- Lofgren, L.A.; Stajich, J.E. Fungal biodiversity and conservation mycology in light of new technology, big data, and changing attitudes. Curr. Biol. 2021, 31, R1312–R1325. [Google Scholar] [CrossRef]
- Smith, D.; Ryan, M.J.; Stackebrandt, E. The ex situ conservation of microorganisms: Aiming at a certified quality management. In Encyclopedia of Life Sciences Support; EoLSS: Oxford, UK, 2008; Available online: https://www.eolss.net/sample-chapters/c17/E6-58-13-04.pdf (accessed on 15 September 2023).
- Psurtseva, N.V. Ex situ fungal conservation: The role of culture collections. Mycol. Balc. 2010, 7, 29–35. [Google Scholar]
- Stalpers, J.A. Identification of wood-inhabiting fungi in pure culture. Stud. Mycol. 1978, 16, 1–248. [Google Scholar]
- Clémençon, H. Cytology and plectology of the Hymenomycetes. Bibl. Mycol. 2004, 199, 1–488. [Google Scholar]
- Gardes, M.; Bruns, T.D. ITS primers with enhanced specification of mycorrhizae and rusts. Mol. Ecol. 1993, 2, 113–118. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Chew, A.L.C.; Desjardin, D.E.; Tan, Y.-S.; Musa, M.Y.; Sabaratnam, V. Bioluminescent fungi from Peninsular Malaysia—A taxonomic and phylogenetic overview. Fungal Divers. 2014, 70, 149–187. [Google Scholar] [CrossRef]
- Cano, A.; Maestre, A.B.; Hernández-Ruiz, J.; Arnao, M.B. ABTS/TAC Methodology: Main Milestones and Recent Applications. Processes 2023, 11, 185. [Google Scholar] [CrossRef]
- Astrup, T.; Mullertz, S. The fibrin plate method for estimating fibrinolytic activity. Arch. Biochem. Biophys. 1952, 40, 346–351. [Google Scholar] [CrossRef]
- Kotlova, E.R.; Senik, S.V.; Kücher, T.; Shavarda, A.L.; Kiyashko, A.A.; Psurtseva, N.V.; Sinyutina, N.F.; Zubarev, R.A. Alterations in the composition of membrane glycero- and sphingolipids in the course of Flammulina velutipes surface culture development. Microbiology 2009, 78, 193–201. [Google Scholar] [CrossRef]
- Kotlova, E.R.; Senik, S.V.; Manzhieva, B.S.; Kiyashko, A.A.; Shakhova, N.V.; Puzansky, R.K.; Volobuev, S.V.; Misharev, A.D.; Serebryakov, E.B.; Psurtseva, N.V. Diversity of ESI-MS based phosphatidylcholine profiles in basidiomycetes. J. Fungi 2022, 8, 177. [Google Scholar] [CrossRef] [PubMed]
- Sazanova, K.V.; Psurtseva, N.V.; Shavarda, A.L. Cultural and metabolomic studies of a new phthalids producer Lignomyces vetlinianus. Int. J. Med. Mushrooms 2018, 20, 1031–1045. [Google Scholar] [CrossRef] [PubMed]
- Sazanova, K.; Psurtseva, N.; Shavarda, A. Metabolomic Changes in Wood Inhabiting Filamentous Fungi during Ontogenesis. In Metabolomics. Methodology and Applications in Medical Sciences and Life Sciences; IntechOpen: London, UK, 2021; pp. 137–156. Available online: https://www.intechopen.com/online-first/metabolomic-changes-in-wood-inhabiting-filamentous-fungi-during-ontogenesis (accessed on 15 September 2023).
- Psurtseva, N.V.; Sazanova, K.V.; Kiyashko, A.A.; Shavarda, A.L. Contribution to studies on rare medicinal mushroom Sparassis crispa (Agaricomycetes) in culture and its production of phenol compounds under various cultivation conditions. Int. J. Med. Mushrooms 2022, 24, 71–79. [Google Scholar] [CrossRef]
- Senik, S.V.; Shakhova, N.V.; Kiyashko, A.A.; Shavarda, A.L. Triterpenoid acids of Agaricometes (Basidiomycota). Chem. Nat. Compd 2024, 59. in print. [Google Scholar]
- He, M.Q.; Zhao, R.L.; Hyde, K.D.; Begerow, D.; Kemler, M.; Yurkov, A.; McKenzie, E.H.C.; Raspé, O.; Kakishima, M.; Sánchez-Ramírez, S.; et al. Notes, outline and divergence times of Basidiomycota. Fungal Divers. 2019, 99, 105–367. [Google Scholar] [CrossRef]
- Wijayawardene, N.N.; Hyde, K.D.; Dai, D.Q.; Sanchez-Garcia, M.; Goto, B.T.; Magurno, F. Outline of Fungi and Fungus-Like Taxa—2021. Mycosphere 2022, 13, 53–453. Available online: https://www.mycosphere.org/pdf/MYCOSPHERE_13_1_2.pdf (accessed on 15 September 2023). [CrossRef]
- Kalichman, J.; Kirk, P.M.; Matheny, P.B. A compendium of generic names of agarics and Agaricales. Taxon 2020, 69, 425–447. [Google Scholar] [CrossRef]
- Audet, S. Mushrooms Nomenclatural Novelties. 2018. Available online: http://www.indexfungorum.org/Publications/PDF/Mushrooms%20nomenclatural%20novelties%20no.%2012.pdf (accessed on 10 November 2023).
- Lücking, R.; Aime, M.C.; Robbertse, B.; Miller, A.N.; Ariyawansa, H.A.; Aoki, T.; Cardinali, G.; Crous, P.W.; Druzhinina, I.S.; Geiser, D.M.; et al. Unambiguous identification of fungi: Where do we stand and how accurate and precise is fungal DNA barcoding? IMA Fungus 2020, 11, 14. [Google Scholar] [CrossRef]
- Olariaga, I.; Huhtinen, S.; Læssøe, T.; Petersen, J.; Hansen, K. Phylogenetic origins and family classification of typhuloid fungi, with emphasis on Ceratellopsis, Macrotyphula and Typhula (Basidiomycota). Stud. Mycol. 2020, 96, 155–184. [Google Scholar] [CrossRef]
- Sun, Y.-F.; Costa-Rezende, D.H.; Xing, J.-H.; Zhou, J.-L.; Zhang, B.; Gibertoni, T.B.; Gates, G.; Glen, M.; Dai, Y.-C.; Cui, B.-K. Multi-gene phylogeny and taxonomy of Amauroderma s. lat. (Ganodermataceae). Pers. Mol. Phylogeny Evol. Fungi 2020, 44, 206–239. [Google Scholar] [CrossRef]
- Tian, E.-J.; Matheny, P.B. A phylogenetic assessment of Pholiota and the new genus Pyrrhulomyces. Mycologia 2021, 113, 146–167. [Google Scholar] [CrossRef] [PubMed]
- Vizzini, A.; Consiglio, G.; Marchetti, M.; Borovička, J.; Campo, E.; Cooper, J.; Lebeuf, R.; Ševčíková, H. New data in Porotheleaceae and Cyphellaceae: Epitypification of Prunulus scabripes Murrill, the status of Mycopan Redhead, Moncalvo & Vilgalys and a new combination in Pleurella Horak emend. Mycol. Prog. 2022, 21, 44. [Google Scholar] [CrossRef]
- Liu, S.; Chen, Y.Y.; Sun, Y.F.; He, X.L.; Song, C.G.; Si, J.; Liu, D.M.; Gates, G.; Cui, B.K. Systematic classification and phylogenetic relationships of the brown rot fungi within the Polyporales. Fungal Divers. 2023, 118, 1–94. [Google Scholar] [CrossRef]
- Liu, S.; Zhou, J.L.; Song, J.; Sun, Y.F.; Dai, Y.C.; Cui, B.K. Climacocystaceae fam. nov. and Gloeoporellaceae fam. nov., two new families of Polyporales (Basidiomycota). Front. Microbiol. 2023, 14, 1115761. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.-L.; Wei, H.-W.; Zhou, L.-W. Xenasmatellales ord. nov. and Xenasmatellaceae fam. nov. for Xenasmatella (Agaricomycetes, Basidiomycota). Mycology 2023, 14, 175–189. [Google Scholar] [CrossRef]
- Benedict, R.G.; Tyler, V.E., Jr.; Brady, L.R. Studies on spore germination and growth of some mycorrhizal-associated Basidiomycetes. Mycopathologia 1967, 31, 319–326. [Google Scholar] [CrossRef]
- Hutchison, L.J. Description and identification of cultures of ectomycorrhizal fungi found in North America. Mycotaxon 1991, 42, 387–504. [Google Scholar]
- Siri-in, J.; Kumla, J.; Suwannarach, N.; Lumyong, S. Culture Conditions and Some Properties of Pure Culture of Ectomycorrhizal Fungus, Scleroderma sinnamariense. Chiang Mai J. Sci. 2014, 41, 275–285. Available online: http://epg.science.cmu.ac.th/ejournal/ (accessed on 15 September 2023).
- Johnson, D.; Voller, F. Ectomycorrhizal Fungal Growth Rates and Biomass in Pure Culture in Response to Organic and Inorganic Nutrient Additions. NERC EDS Environmental Information Data Centre. (Dataset). 2022. Available online: https://catalogue.ceh.ac.uk/documents/8eb1b693-8f56-4d97-b42d-f2c206dd33c0 (accessed on 15 September 2023).
- Sangtiean, T.; Schmidt, S. Growth of subtropical ECM fungi with different nitrogen sources using a new floating culture technique. Mycol. Res. 2002, 106, 74–85. [Google Scholar] [CrossRef]
- Fransson, P.M.A.; Anderson, I.C.; Alexander, I.J. Ectomycorrhizal fungi in culture respond differently to increased carbon availability. FEMS Microbiol. Ecol. 2007, 61, 246–257. [Google Scholar] [CrossRef]
- Shishikura, M.; Sugawara, R.; Takemura, Y.; Sotome, K.; Maekawa, N.; Nakagiri, A.; Endo, N. First successful isolation of Entoloma clypeatum species complex from basidiospores. Mycoscience 2019, 60, 221–227. [Google Scholar] [CrossRef]
- Fries, N. Germination of spores of Cantharellus cibarius. Mycologia 1979, 71, 216–219. [Google Scholar] [CrossRef]
- Danell, E. Chanterelle Mycelium. U.S. Patent N 6173525 B1, 16 January 2001. [Google Scholar]
- Ogawa, W.; Endo, N.; Takeda, Y.; Kodaira, M.; Fukuda, M.; Yamada, A. Efficient establishment of pure cultures of yellow chanterelle Cantharellus anzutake from ectomycorrhizal root tips, and morphological characteristics of ectomycorrhizae and cultured mycelium. Mycoscience 2019, 60, 45–53. [Google Scholar] [CrossRef]
- Yamada, A. Cultivation studies of edible ectomycorrhizal mushrooms: Successful establishment of ectomycorrhizal associations in vitro and efficient production of fruiting bodies. Mycoscience 2022, 63, 235–246. [Google Scholar] [CrossRef] [PubMed]
- Guerin-Laguette, A. Successes and challenges in the sustainable cultivation of edible mycorrhizal fungi—Furthering the dream. Mycoscience 2021, 62, 10–28. [Google Scholar] [CrossRef]
- Kamelin, R.V.; Novikov, V.S. Red Book of Russian Federation (plants and fungi). KMK 2008, 885, 779–780. [Google Scholar]
- Yadav, D.; Negi, P.D. Bioactive components of mushrooms: Processing effects and health benefits. Food Res. Int. 2021, 148, 110599. [Google Scholar] [CrossRef]
- Venturella, G.; Ferraro, V.; Cirlincione, F.; Gargano, M.L. Medicinal mushrooms: Bioactive compounds, use, and clinical trials. Int. J. Mol. Sci. 2021, 22, 634. [Google Scholar] [CrossRef]
- Llanaj, X.; Törős, G.; Hajdú, P.; Abdalla, N.; El-Ramady, H.; Kiss, A.; Solberg, S.Ø.; Prokisch, J. Biotechnological applications of mushrooms under the water-energy-food nexus: Crucial aspects and prospects from farm to pharmacy. Foods 2023, 12, 2671. [Google Scholar] [CrossRef]
- Vasilenko, A.; Ivanushkina, N.; Kochkina, G.; Ozerskaya, S. Fungi in microbial Culture Collections and their metabolites. Diversity 2022, 14, 507. [Google Scholar] [CrossRef]
- Sakurai, K.; Yuasa, M.; Ohji, S.; Hosoyama, A.; Sato, M.; Fujita, N.; Kawasaki, H. Gene Mutations in Ganoderma lucidum during long-term preservation by repeated subculturing. Biopreserv. Biobank. 2019, 17, 395–400. [Google Scholar] [CrossRef]
- Danner, C.; Mach, R.L.; Mach-Aigner, A.R. The phenomenon of strain degeneration in biotechnologically relevant fungi. Appl. Microbiol. Biotechnol. 2023, 107, 4745–4758. [Google Scholar] [CrossRef]
- Burdsall, H.H.; Dorworth, E.B. Preserving cultures of wood-decaying Basidiomycotina using sterile distilled water in cryovials. Mycologia 1994, 86, 275–280. [Google Scholar] [CrossRef]
- Richter, D.L. Revival of saprotrophic and mycorrhizal basidiomycete cultures after 20 years in cold storage in sterile water. Can. J. Microbiol. 2008, 54, 595–599. [Google Scholar] [CrossRef]
- Richter, D.L.; Dixon, T.G.; Smith, J.K. Revival of saprotrophic and mycorrhizal basidiomycete cultures after 30 years in cold storage in sterile water. Can. J. Microbiol. 2016, 62, 932–937. [Google Scholar] [CrossRef]
- Johnson, G.C.; Martin, A.K. Survival of wood-inhabiting fungi stored for 10 years in water and under oil. Can. J. Microbiol. 1992, 38, 861–864. [Google Scholar] [CrossRef]
- Danell, E.; Flygh, G. Cryopreservation of the ectomycorrhizal mushroom Cantharellus cibarius. Mycol. Res. 2002, 106, 1340–1342. [Google Scholar] [CrossRef]
- Komissarov, N.S.; Dyakov, M.Y.; Garibova, L.V. Methods for long-term storage of pure cultures of macrofungi. Mikol. Fitopatol. 2023, 57, 155–171. [Google Scholar]
- Senik, S.V.; Kolker, T.L.; Kotlova, E.R.; Vlasov, D.Y.; Shavarda, A.L.; Puzansky, R.K.; Psurtseva, N.V. Lipid and metabolite profiling of Serpula lacrymans under freezing stress. Curr. Microbiol. 2021, 78, 961–966. [Google Scholar] [CrossRef] [PubMed]
- Petersen, R.H.; Hughes, K.W.; Lickey, E.B.; Kovalenko, A.E.; Morozova, O.V.; Psurtseva, N.V. A new genus, Cruentomycena, with Mycena viscidocruenta as type species. Mycotaxon 2008, 105, 119–136. [Google Scholar]
- Vydryakova, G.A.; Van, D.T.; Shoukouhi, P.; Psurtseva, N.V.; Bissett, J. Intergenomic and intragenomic ITS sequence heterogeneity in Neonothopanus nambi (Agaricales) from Vietnam. Mycology 2012, 2, 89–99. [Google Scholar]
- Petersen, R.H.; Psurtseva, N.; Zmitrovich, I.; Chachula, P.; Arslanov, S.; Hughes, K.W. Lignomyces, a new genus of pleurotoid Agaricomycetes. Mycologia 2015, 107, 1045–1054. [Google Scholar] [CrossRef] [PubMed]
- Psurtseva, N.V.; Zmitrovich, I.V.; Malysheva, V.F. Taxonomy and developmental morphology of Rogersiomyces malaysianus comb. nov. (Cantharellales, Agaricomycetes). Botany 2016, 94, 579–592. [Google Scholar] [CrossRef]
- Haelewaters, D.; Dima, B.; Abdel-Hafiz, A., II; Abdel-Wahab, M.A.; Abul-Ezz, S.R.; Acar, I.; Aguirre-Acosta, E.; Aime, M.C.; Aldemir, S.; Ali, M.; et al. Fungal systematics and evolution: FUSE 6. Sydowia 2020, 72, 222–227. [Google Scholar] [CrossRef]
- Psurtseva, N.V.; Zmitrovich, I.V.; Seelan, J.S.S.; Bulakh, E.M.; Hughes, K.W.; Petersen, R.H. New data on morphology, physiology, and geographical distribution of Lignomyces vetlinianus, its identity with Lentinus pilososquamulosus, and sufficient phylogenetic distance from Le. martianoffianus. Mycol. Prog. 2021, 20, 809–821. [Google Scholar] [CrossRef]
- Shakhova, N.V.; Volobuev, S.V. Culture characteristics and enzymatic activity of Sarcodontia crocea (Basidiomycota) strains collected from the Central Russian Upland. Mikol. Fitopatol. 2020, 54, 446–451. [Google Scholar] [CrossRef]
- Volobuev, S.V.; Shakhova, N.V. A comparative study on growth and lignocellulolytic activity of nine Sarcodontia crocea strains in four different media. Divers. Plant World 2020, 3, 45–54. [Google Scholar] [CrossRef]
- Shakhova, N.; Volobuev, S. Cultural and enzymatic activity studies of a pathogenic wood-decaying fungus Fomitiporia hippophaeicola (Hymenochaetales, Basidiomycota), recollected in the Eastern Caucasus. Arch. Microbiol. 2023, 205, 249. [Google Scholar] [CrossRef]
- Whaley, A.O.; Whaley, A.K.; Orlova, A.A.; Volobuev, S.V.; Shakhova, N.V.; Smirnov, S.N.; Pustovit, N.V.; Kraeva, L.A. Polyacetylene derivatives from the Polypore fungus (Fistulina hepatica, Agaricomycetes) and their antimicrobial activity. Int. J. Med. Mush. 2023, 25, 43–53. [Google Scholar] [CrossRef]
- Nobles, M.K. Identification of cultures of wood-inhabiting Hymenomycetes. Can. J. Bot. 1965, 43, 1097–1139. [Google Scholar] [CrossRef]
- Moiseenko, K.; Glazunova, O.; Shakhova, N.; Savinova, O.; Vasina, D.; Tyazhelova, T.; Psurtseva, N.; Fedorova, T. Data on the genome analysis of the wood-rotting fungus Steccherinum ochraceum LE-BIN 3174. Data Br. 2020, 29, 105169. [Google Scholar] [CrossRef]
- Zhang, J.; Shen, N.; Li, C.; Xiang, X.; Liu, G.; Gui, Y.; Patev, S.; Hibbett, D.S.; Barry, K.; Andreopoulos, W.; et al. Population genomics provides insights into the genetic basis of adaptive evolution in the mushroom-forming fungus Lentinula edodes. J. Adv. Res. 2022, 38, 91–106. [Google Scholar] [CrossRef]
- Pavlov, A.R.; Tyazhelova, T.V.; Moiseenko, K.V.; Vasina, D.V.; Mosunova, O.V.; Fedorova, T.V.; Maloshenok, L.G.; Landesman, E.O.; Bruskin, S.A.; Psurtseva, N.V.; et al. Draft genome sequence of the fungus Trametes hirsuta 072. Genome Announc. 2015, 3, e01287-15. [Google Scholar] [CrossRef]
- Arregui, L.; Ayala, M.; Gómez-Gil, X.; Gutiérrez-Soto, G.; Hernández-Luna, C.E.; de los Santos, M.H.; Levin, L.; Rojo-Domínguez, A.; Romero-Martínez, D.; Saparrat, M.C.; et al. Laccases: Structure, function, and potential application in water bioremediation. Microb. Cell Fact. 2019, 18, 200. [Google Scholar] [CrossRef]
- Mäkelä, M.R.; Tuomela, M.; Hatakka, A.; Hildén, K. Fungal laccases and their potential in bioremediation applications. In Laccases in Bioremediation and Waste Valorisation. Microbiology Monographs; Schlosser, D., Ed.; Springer: Berlin/Heidelberg, Germany, 2020; p. 33. [Google Scholar] [CrossRef]
- Loi, M.; Glazunova, O.; Fedorova, T.; Logrieco, A.F.; Mulè, G. Fungal laccases: The forefront of enzymes for sustainability. J. Fungi 2021, 7, 1048. [Google Scholar] [CrossRef] [PubMed]
- Viswanath, B.; Rajesh, B.; Janardhan, A.; Kumar, A.P.; Narasimha, G. Fungal laccases and their applications in bioremediation. Enzyme Res. 2014, 2014, 163242. [Google Scholar] [CrossRef] [PubMed]
- Martínez, A.T.; Ruiz-Dueñas, F.J.; Camarero, S.; Serrano, A.; Linde, D.; Lund, H.; Vind, J.; Tovborg, M.; Herold-Majumdar, O.M.; Hofrichter, M.; et al. Oxidoreductases on their way to industrial biotransformations. Biotechnol. Adv. 2017, 35, 815–831. [Google Scholar] [CrossRef]
- Bissaro, B.; Várnai, A.; Røhr, Å.K.; Eijsink, V.G.H. Oxidoreductases and reactive oxygen species in conversion of lignocellulosic biomass. Microbiol. Mol. Biol. Rev. 2018, 82, e00029-18. [Google Scholar] [CrossRef]
- Agustin, M.; Morais de Carvalho, D.; Lahtinen, M.; Hilden, K.; Lundell, T.; Mikkonen, K.S. Laccase as a tool in building advanced lignin-based materials. ChemSusChem 2021, 14, 4615–4635. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K.; Bilal, M.; Jesionowski, T.; Iqbal, H.M.N. Deployment of oxidoreductases for sustainable biocatalytic degradation of selected endocrine-disrupting chemicals. Sustain. Chem. Pharm. 2023, 31, 100934. [Google Scholar] [CrossRef]
- Mattila, H.; Österman-Udd, J.; Mali, T.; Lundell, T. Basidiomycota fungi and ROS: Genomic perspective on key enzymes involved in generation and mitigation of reactive oxygen species. Front. Fungal Biol. 2022, 3, 837605. [Google Scholar] [CrossRef]
- Fonseca, M.I.; Zapata, P.D.; Villalba, L.L.; Fariña, J.I. Characterization of the oxidative enzyme potential in wild white rot fungi of misiones (Argentina). Acta Biol. Colomb. 2015, 20, 47–56. [Google Scholar] [CrossRef]
- Sharma, A.; Aggarwal, N.K.; Yadav, A. Isolation and screening of lignolytic fungi from various ecological niches. Univers. J. Microbiol. Res. 2017, 5, 25–34. [Google Scholar] [CrossRef]
- Tsiklauri, N.; Jobava, M.; Khvedelidze, R.; Kutateladze, L.Y.; Khokhashvili, I.; Kharatishvili, B. Lignocellulosic ezymes of basidial fungi-isolated from different ecological niches of georgia. EPSTEM 2021, 12, 113–118. [Google Scholar] [CrossRef]
- Zmitrovich, I.V.; Psurtseva, N.V.; Belova, N.V. Evolutionary and taxonomical aspects of search and study of lignin-degrading fungi—Active producers of oxidative enzymes. Mikol. Fitopatol. 2007, 41, 57–78. [Google Scholar]
- Binder, M.; Justo, A.; Riley, R.; Salamov, A.; Lopez-Giraldez, F.; Sjokvist, E.; Copeland, A.; Foster, B.; Sun, H.; Larsson, E.; et al. Phylogenetic and phylogenomic overview of the Polyporales. Mycologia 2013, 105, 1350–1373. [Google Scholar] [CrossRef] [PubMed]
- Myasoedova, N.M.; Chernykh, A.M.; Psurtseva, N.V.; Belova, N.V.; Golovleva, L.A. New efficient producers of fungal laccases. Appl. Biochem. Microbiol. 2008, 44, 73–77. [Google Scholar] [CrossRef]
- Fedorova, T.V.; Shakhova, N.V.; Klyain, O.I.; Glazunova, O.A.; Maloshenok, L.G.; Kulikova, N.A.; Psurtseva, N.V.; Koroleva, O.V. Comparative analysis of the ligninolytic potential of basidiomycetes belonging to different taxonomic and ecological groups. Appl. Biochem. Microbiol. 2013, 49, 570–580. [Google Scholar] [CrossRef] [PubMed]
- Glazunova, O.A.; Shakhova, N.V.; Psurtseva, N.V.; Moiseenko, K.V.; Kleimenov, S.Y.; Fedorova, T.V. White-rot basidiomycetes Junghuhnia nitida and Steccherinum bourdotii: Oxidative potential and laccase properties in comparison with Trametes hirsuta and Coriolopsis caperata. PLoS ONE 2018, 13, e0197667. [Google Scholar] [CrossRef] [PubMed]
- Chernykh, A.M.; Kolomytseva, M.P.; Myasoedova, N.M.; Golovleva, L.A.; Psurtseva, N.V.; Belova, N.V. Thermostable Laccase III of Fungus Steccherinum ochraceum LE (BIN) 1833 D and Method for Preparing Laccases. Patent RU 2385930, 10 April 2010. Available online: https://patents.google.com/patent/RU2385930C1/ru (accessed on 10 October 2023).
- Barrasa, J.M.; Blanco, M.N.; Esteve-Raventós, F.; Altés, A.; Checa, J.; Martínez, A.T.; Ruiz-Dueñas, F.J. Wood and humus decay strategies by white-rot basidiomycetes correlate with two different dye decolorization and enzyme secretion patterns on agar plates. Fungal Genet. Biol. 2014, 72, 106–114. [Google Scholar] [CrossRef]
- Martínková, L.; Kotik, M.; Marková, E.; Homolka, L. Biodegradation of phenolic compounds by Basidiomycota and its phenol oxidases: A review. Chemosphere 2016, 149, 373–382. [Google Scholar] [CrossRef]
- Mäkelä, M.R.; Hildén, K.; Kowalczyk, J.E.; Hatakka, A. Progress and research needs of plant biomass degradation by basidiomycete fungi. In Grand Challenges in Biology and Biotechnology; Springer: Cham, Switzerland, 2020; pp. 405–438. [Google Scholar] [CrossRef]
- Okal, E.J.; Aslam, M.M.; Karanja, J.K.; Nyimbo, W.J. Mini review: Advances in understanding regulation of cellulase enzyme in white-rot basidiomycetes. Microb. Pathog. 2020, 147, 104410. [Google Scholar] [CrossRef]
- Peralta, R.M.; da Silva, B.P.; Côrrea, R.C.G.; Kato, C.G.; Seixa, F.A.V.; Bracht, A. Chapter 5—Enzymes from basidiomycetes—Peculiar and efficient tools for biotechnology. In Biotechnology of Microbial Enzymes; Academic Press: Cambridge, MA, USA, 2017; pp. 119–149. [Google Scholar] [CrossRef]
- Shakhova, N.; Volobuev, S. Revealing new active and biotechnologically perspective producers of oxidative and cellulolytic enzymes among pure cultures of xylotrophic Agaricomycetes from the Southern Non-Chernozem zone of the European part of Russia. Curr. Res. Environ. Appl. Mycol. 2020, 10, 113–119. [Google Scholar] [CrossRef]
- Volobuev, S.; Shakhova, N. Towards the discovery of active lignocellulolytic enzyme producers: A screening study of xylotrophic macrofungi from the Central Russian Upland. Iran. J. Sci. Technol. Trans. A Sci. 2022, 46, 91–100. [Google Scholar] [CrossRef]
- Moiseenko, K.V.; Glazunova, O.A.; Shakhova, N.V.; Savinova, O.S.; Vasina, D.V.; Tyazhelova, T.V.; Psurtseva, N.V.; Fedorova, T.V. Fungal adaptation to the advanced stages of wood decomposition: Insights from the Steccherinum ochraceum. Microorganisms 2019, 7, 527. [Google Scholar] [CrossRef] [PubMed]
- Glazunova, O.A.; Moiseenko, K.V.; Shakhova, N.V.; Psurtseva, N.V.; Fedorova, T.V. Biodegradation potential of Steccherinum ochraceum: Growth on different wood types and preliminary evaluation of enzymatic activities. KnE Life Sci. 2022, 7, 207–215. [Google Scholar] [CrossRef]
- Schmidt, O.; Huckfeldt, T. Characterization and identification of indoor wood-decaying basidiomycetes. In Fundamentals of Mold Growth in Indoor Environments and Strategies for Healthy Living; Adan, D.G., Samson, R.A., Eds.; Academic Publication: Wageningen, The Netherlands, 2011; pp. 117–180. [Google Scholar] [CrossRef]
- Vlasov, D.Y.; Psurtseva, N.V.; Zmitrovich, I.V.; Sazanova, K.V.; Ezhov, O.N.; Bondartseva, M.A. Occurrence and adaptive potential of indoor macrofungi. In Bioprospects of Macrofungi: Recent Developments; CRS Press: Boca Raton, FL, USA, 2023; pp. 280–307. [Google Scholar]
- Denisova, N.P.; Alekhina, I.A. Fibrinolytic activity of basidiomycetes culture. Mikol. Fitopatol. 1987, 21, 471–477. [Google Scholar]
- Denisova, N.P.; Semenova, I.R.; Sukharevich, V.I. Biosynthesis of fibrinolytic proteinases by higher basidiomycetes in submerged culture. Mikol. Fitopatol. 1989, 23, 378–381. [Google Scholar]
- Romanovets, E.S.; Kapich, A.N.; Denisova, N.P.; Psurtseva, N.V. Submerged cultivation of higher basidiomycete from genus Coprinus. Biotekhnologiya 1990, 6, 41–43. [Google Scholar]
- Psurtseva, N.V.; Mnukhina, A.Y. Morphological, physiological and enzyme variability of Flammulina, P. Karst. Cultures. Mikol. Fitopatol. 1998, 32, 49–54. [Google Scholar]
- Morozova, E.N.; Falina, N.N.; Denisova, N.P.; Barkova, L.V.; Psurtseva, N.V.; Samartsev, M.A.; Shitova, V.I. Analysis of the component composition and substrate specificity of a fibrinolytic preparation from the fungus Flammulina velutipes. Biochemistry 1982, 47, 1181–1185. [Google Scholar]
- Shaginyan, K.A.; Alekhina, I.A.; Denisova, N.P. Serine proteinase from higher basidiomycete of the genus Coprinus. Biochemistry 1990, 55, 1387–1395. [Google Scholar]
- Denisova, N.P.; Mikhailov, V.N.; Petrishchev, N.N. Thrombolytic activity of mushroom proteinases: Thrombolytic activity of proteinases from Coprinus domesticus, C. cinereus, and Cerrena unicolor. Int. J. Med. Mushrooms 1999, 1, 187–190. [Google Scholar] [CrossRef]
- Abugri, D.A.; McElhenney, W.H.; Willian, K.R. Fatty acid profiling in selected cultivated edible and wild medicinal mushrooms in southern United States. J. Exp. Food Chem. 2016, 2, 1000108. [Google Scholar] [CrossRef]
- Sande, D.; Oliveira, G.P.; Moura, M.A.F.E.; Martins, B.A.; Lima, M.T.N.S.; Takahashi, J.A. Edible mushrooms as a ubiquitous source of essential fatty acids. Food Res. Int. 2019, 125, 108524. [Google Scholar] [CrossRef] [PubMed]
- Nagy, K.; Tiuca, I.-D. Importance of fatty acids in physiopathology of human body. In Fatty acids; IntechOpen: London, UK, 2017; pp. 3–22. [Google Scholar] [CrossRef]
- Balić, A.; Vlašić, D.; Žužul, K.; Marinović, B.; Bukvić-Mokos, Z. Omega-3 versus omega-6 polyunsaturated fatty acids in the prevention and treatment of inflammatory skin diseases. Int. J. Mol. Sci. 2020, 21, 741. [Google Scholar] [CrossRef] [PubMed]
- Saini, R.K.; Rauf, A.; Khalil, A.A.; Ko, E.-Y.; Keum, Y.-S.; Anwar, S.; Alamri, A.; Rengasamy, K.R.R. Edible mushrooms show significant differences in sterols and fatty acid compositions. S. Afr. J. Bot. 2021, 141, 344–356. [Google Scholar] [CrossRef]
- Saenge, C.; Cheirsilp, B.; Suksaroge, T.T.; Bourtoom, T. Potential use of oleaginous red yeast Rhodotorula glutinis for the bioconversion of crude glycerol from biodiesel plant to lipids and carotenoids. Process Biochem. 2011, 46, 210–218. [Google Scholar] [CrossRef]
- Negri, R. Polyacetylenes from terrestrial plants and fungi: Recent phytochemical and biological advances. Fitoterapia 2015, 106, 92–109. [Google Scholar] [CrossRef]
- León, A.; Del-Ángel, M.; Ávila, J.L.; Delgado, G. Phthalides: Distribution in nature, chemical reactivity, synthesis and biological activity. Prog. Chem. Org. Nat. 2017, 140, 127–244. [Google Scholar]
- Whaley, A.K.; Ponkratova, A.O.; Orlova, A.A.; Volobuev, S.V.; Shakhova, N.V.; Serebryakov, E.B.; Smirnov, S.N.; Pustovit, N.V.; Kraeva, L.A.; Luzhanin, V.G.; et al. New benzoquinone pigments from the hydnoid fungus Sarcodontia setosa and their biosynthetic relationship. Nat. Prod. Res. 2023, 73, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Zhao, P.; Guan, M.; Tang, W.; Walayat, N.; Ding, Y.; Liu, J. Structural diversity, fermentation production, bioactivities and applications of triterpenoids from several common medicinal fungi: Recent advances and future perspectives. Fitoterapia 2023, 166, 105470. [Google Scholar] [CrossRef] [PubMed]
- León, F.; Quintana, J.; Rivera, A.; Estévez, F.; Bermejo, J. Lanostanoid triterpenes from Laetiporus sulphureus and apoptosis induction on HL-60 human myeloid leukemia cells. J. Nat. Prod. 2004, 67, 2008–2011. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Zhou, R.; Xie, J.; Ye, H.; Liang, X.; Zhong, C.; Shen, B.; Qin, Y.; Zhang, S.; Huang, L. Insights into triterpene acids in fermented mycelia of edible fungus Poria cocos by a comparative study. Molecules 2019, 24, 1331. [Google Scholar] [CrossRef]
- Deng, J.S.; Huang, S.S.; Lin, T.H.; Lee, M.M.; Kuo, C.C.; Sung, P.J.; Hou, W.C.; Huang, G.J.; Kuo, Y.H. Analgesic and anti-inflammatory bioactivities of eburicoic acid and dehydroeburicoic acid isolated from Antrodia camphorata on the inflammatory mediator expression in mice. J. Agric. Food Chem. 2013, 61, 5064–5071. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.J.; Deng, J.S.; Huang, S.S.; Lee, C.Y.; Hou, W.C.; Wang, S.Y.; Sung, P.J.; Kuo, Y.H. Hepatoprotective effects of eburicoic acid and dehydroeburicoic acid from Antrodia camphorata in a mouse model of acute hepatic injury. Food Chem. 2013, 141, 3020–3027. [Google Scholar] [CrossRef]
- Kuo, Y.H.; Lin, C.H.; Shih, C.C. Dehydroeburicoic acid from Antrodia camphorata prevents the diabetic and dyslipidemic state via modulation of glucose transporter 4, peroxisome proliferator-activated receptor α expression and AMP-activated protein kinase phosphorylation in high-fat-fed mice. Int. J. Mol. Sci. 2016, 17, 872. [Google Scholar] [CrossRef]
- Kim, Y.J.; Park, J.; Min, B.S.; Shim, S.H. Chemical constituents from the sclerotia of Inonotus obliquus. J. Korean Soc. Appl. Biol. Chem. 2011, 54, 287–294. [Google Scholar] [CrossRef]
- Kang, H.M.; Lee, S.K.; Shin, D.S.; Lee, M.-Y.; Han, D.C.; Baek, N.-I.; Son, K.-H.; Kwon, B.-M. Dehydrotrametenolic acid selectively inhibits the growth of H-ras transformed rat2 cells and induces apoptosis through caspase-3 pathway. Life Sci. 2006, 78, 607–613. [Google Scholar] [CrossRef]
- Novikova, S.P.; Denisova, N.P.; Il’ina, M.B.; Dobrova, N.B.; Psurtseva, N.V.; Falina, N.N. The Method of Obtaining of Polymer Materials with Thromboresistant Features. SU Patent 1182709, 1 June 1985. [Google Scholar]
- Denisova, N.P.; Psurtseva, N.V.; Alekhina, I.A.; Petrishchev, N.N.; Mihailov, V.N.; Sashek, V.; Musilek, V. Somatic Culture Strain Cerrena unicolor (Bull. Fr.) Murr.—Producer of Fibrino- and Thrombolytic Enzymes. SU Patent 1459232, 15 October 1988. [Google Scholar]
- Denisova, N.P.; Psurtseva, N.V.; Falina, N.N.; Petrishchev, N.N.; Kiyanov, V.I. Strain of Somatic Cultures of the Basidiomycete Fungus Coprinus domesticus (Fr.) Gray—Producer of Fibrino- and Thrombolytic Enzymes. SU Patent 1137762, 7 August 1991. [Google Scholar]
- Diwan, D.; Usmani, Z.; Sharma, M.; Nelson, J.W.; Thakur, V.K.; Christie, G.; Molina, G.; Gupta, V.K. Thrombolytic Enzymes of Microbial Origin: A Review. Int. J. Mol. Sci. 2021, 22, 10468. [Google Scholar] [CrossRef] [PubMed]
- Takehara, M.; Kuida, K.; Mori, K. Antiviral activity of virus-like particles from Lentinus edodes (Shiitake). Brief report. Arch. Virol. 1979, 59, 269–274. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, G.; Ling, J. Medicinal fungi with antiviral effect. Molecules 2022, 27, 4457. [Google Scholar] [CrossRef] [PubMed]
- Teplyakova, T.V.; Psurtseva, N.V.; Kosogova, T.A.; Mazurkova, N.A.; Khanin, V.A.; Vlasenko, V.A. Antiviral activity of polyporoid mushrooms from Altai Mountains. Int. J. Med. Mushrooms 2012, 14, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Seo, D.J.; Choi, C. Antiviral bioactive compounds of mushrooms and their antiviral mechanisms: A review. Viruses 2021, 13, 350. [Google Scholar] [CrossRef] [PubMed]
- Arunachalam, K.; Sasidharan, S.P.; Yang, X. A concise review of mushrooms antiviral and immunomodulatory properties that may combat against COVID-19. Food Chem. Adv. 2022, 1, 100023. [Google Scholar] [CrossRef]
- Sen, D.; Debnath, B.; Debnath, P.; Debnath, S.; Zaki, M.E.A.; Masand, V.H. Identification of potential edible mushroom as SARS-CoV-2 main protease inhibitor using rational drug designing approach. Sci. Rep. 2022, 12, 1503. [Google Scholar] [CrossRef]
- Chkhenkeli, V.A.; Belova, N.V.; Shkil’, N.A.; Chkhenkeli, G.D. A Drug for the Treatment of Gastrointestinal Diseases of Calves and a Method of Its Application. RU Patent 2429871, 27 September 2011. [Google Scholar]
- List of cultures of basidiomycetes present in the collection at Komarov Botanical Institute (Academy of sciences of the U.S.S.R.). Mikol. Fitopatol. 1980, 14, 465–469.
- Sivochub, O.A. Catalogue of Badidiomycetes Cultures of the Komarov Botanical Institute Collection; SPb: Nauka, Russia, 1992. [Google Scholar]
- Psurtseva, N.V.; Kiyashko, A.A.; Gachkova, E.Y.; Belova, N.V. Basidiomycetes Culture Collection LE (BIN): Catalogue of Strains, 2nd ed.; M.: KMK Scientific Press Ltd.: St. Petersburg, Russia, 2007. [Google Scholar]



| Number of Strains | Survival (%) | Storage t (°C) | Storage Time (Years) | Reference |
|---|---|---|---|---|
| 151 | 94 | 5 | 7 | [50] |
| 34 | 88 | 5 | 20 | [51] |
| 393 | 26 | 20 | 10 | [53] |
| 159 | 32 | 22–25 | 5–10 | LE-BIN strains |
| Class | Order | Family | Genus |
|---|---|---|---|
| Agaricomycetes | |||
| Agaricales | Agaricaceae | Agaricus 8 (36), Chlorophyllum 2 (3), Coprinus 2 (5), Disciseda 1 (1), Lepiota 1 (2), Leucoagaricus 3 (6), Macrolepiota 2 (8), Micropsalliota 1 (1), Montagnea 1 (3), Mycenastrum 1 (3) | |
| Amanitaceae | Amanita 1 (1) | ||
| Battarreaceae | Tulostoma 3 (5) | ||
| Bolbitiaceae | Bolbitius 1 (1), Conocybe 1 (3) | ||
| Callistosporiaceae | Callistosporium 1 (2) | ||
| Clitocybaceae | Clitocybe 8 (29), Lepista 1 (2) | ||
| Crepidotaceae | Crepidotus 5 (15), Simocybe 1 (1) | ||
| Cyphellaceae | Atheniella 2 (5), Baeospora 1 (2), Chondrostereum 1 (6), Gloeostereum 1 (2), Granulobasidium 1 (1), Hemimycena 1 (4), Henningsomyces 1 (1), Mycopan 1 (1) | ||
| Entolomataceae | Clitopilopsis 1 (1), Clitopilus 3 (5), Entoloma 1 (2) | ||
| Fayodiaceae | Conchomyces 1 (2) | ||
| Galeropsidaceae | Panaeolina 1 (1), Panaeolus 5 (16) | ||
| Hygrophoraceae | Arrhenia 2 (4), Lignomphalia 1 (1), Pseudoarmillariella 1 (1) | ||
| Hymenogastraceae | Flammula 2 (8), Galerina 9 (27), Gymnopilus 8 (37), Hemistropharia 1 (1), Psilocybe 5 (24) | ||
| Lycoperdaceae | Apioperdon 1 (13), Bovista 4 (15), Bovistella 1 (4), Calvatia 3 (3), Lycoperdon 6 (38) | ||
| Lyophyllaceae | Asterophora 1 (1), Calocybe 2 (3), Hypsizygus 3 (19), Lyophyllum 5 (10), Ossicaulis 2 (13), Tephrocybe 1 (2), Termitomyces 1 (1) | ||
| Marasmiaceae | Campanella 1 (2), Chaetocalathus 1 (1), Crinipellis 1 (2), Marasmius 11 (70), Paramarasmius 1 (1), Tetrapyrgos 1 (4) | ||
| Mycenaceae | Cruentomycena 1 (2), Dictyopanus 1 (5), Favolaschia 3 (9), Filoboletus 1 (3), Heimiomyces 1 (1), Mycena 38 (157), Panellus 5 (29), Roridomyces 1 (1), Xeromphalina 6 (27) | ||
| Nidulariaceae | Crucibulum 1 (8), Cyathus 6 (31), Nidula 1 (1), Nidularia 1 (5) | ||
| Omphalinaceae | Infundibulicybe 2 (9), Omphalina 1 (1) | ||
| Omphalotaceae | Anthracophyllum 1 (1), Collybiopsis 10 (46), Connopus 1 (1), Gymnopus 11 (56), Lentinula 2 (29), Marasmiellus 2 (5), Mycetinis 3 (22), Neonothopanus 1 (9), Omphalotus 2 (10), Paragymnopus 1 (4), Pseudomarasmius 1 (1), Rhodocollybia 2 (14) | ||
| Phyllotopsidaceae | Phyllotopsis 1 (7) | ||
| Physalacriaceae | Armillaria 5 (52), Cylindrobasidium 1 (2), Flammulina 9 (57), Hymenopellis 2 (19), Mucidula 2 (30), Oudemansiella 1 (3), Rhizomarasmius 2 (3), Rhodotus 1 (5), Xerula 2 (7) | ||
| Pleurotaceae | Hohenbuehelia 3 (5), Lignomyces 1 (7), Pleurotus 13 (152), Resupinatus 1 (1) | ||
| Pluteaceae | Volvariella 1 (1) | ||
| Porotheleaceae | Clitocybula 2 (7), Delicatula 1 (3), Gerronema 1 (3), Hydropus 3 (12), Megacollybia 3 (20), Trogia 1 (4) | ||
| Psathyrellaceae | Candolleomyces 2 (10), Coprinellus 7 (33), Coprinopsis 9 (21), Cystoagaricus 1 (1), Parasola 1 (1), Psathyrella 5 (27) | ||
| Pseudoclitocybaceae | Clitopaxillus 1 (1) | ||
| Pterulaceae | Pterulicium 1 (1) | ||
| Radulomycetaceae | Radulomyces 3 (6) | ||
| Sarcomyxaceae | Sarcomyxa 1 (4) | ||
| Schizophyllaceae | Schizophyllum 2 (24) | ||
| Squamanitaceae | Cystoderma 1 (5), Phaeolepiota 1 (2) | ||
| Strophariaceae | Agrocybe 6 (18), Deconica 3 (7), Hypholoma 6 (31), Kuehneromyces 2 (24), Leratiomyces 3 (5), Melanotus 1 (4), Pholiota 18 (92), Protostropharia 1 (6), Stropharia 3 (9) | ||
| Tricholomataceae | Leucopaxillus 2 (5) | ||
| Tubariaceae | Cyclocybe 2 (3), Tubaria 1 (4) | ||
| Incertae sedis | Aspropaxillus 1 (1), Collybia 3 (16), Cynema 1 (2), Cystodermella 3 (6), Fistulina 1 (7), Leucocortinarius 1 (1), Leucocybe 2 (3), Meottomyces 1 (1), Notholepista 1 (1), Paralepista 1 (1), Rhizocybe 1 (1), Ripartites 1 (1) | ||
| Unspecified | “Coprinus” (12) | ||
| Amylocorticiales | Amylocorticiaceae | Ceraceomyces 1 (1), Irpicodon 1 (2), Plicaturopsis 1 (2) | |
| Auriculariales | Auriculariaceae | Alloexidiopsis 1 (1), Auricularia 6 (21), Elmerina 2 (4), 1 (1) | |
| Incertae sedis | Guepinia 1 (1), Protomerulius 1 (1), Pseudohydnum 1 (2) | ||
| Boletales | Boletaceae | Boletus 1 (1) | |
| Coniophoraceae | Coniophora 1 (7), | ||
| Hygrophoropsidaceae | Hygrophoropsis 1 (2) | ||
| Serpulaceae | Serpula 2 (4) | ||
| Suillaceae | Boletinus 1 (3), Suillus 9 (11) | ||
| Tapinellaceae | Pseudomerulius 1 (1), Tapinella 1 (1) | ||
| Cantharellales | Hydnaceae | Rogersiomyces 1 (1), Sistotrema 1 (6) | |
| Corticiales | Punctulariaceae | Punctularia 1 (3) | |
| Vuilleminiaceae | Vuilleminia 1 (3) | ||
| Geastrales | Geastraceae | Geastrum 1 (3), Sphaerobolus 2 (4) | |
| Gloeophyllales | Gloeophyllaceae | Gloeophyllum 5 (24), Neolentinus 3 (21) | |
| Gomphales | Clavariadelphaceae | Clavariadelphus 1 (1) | |
| Gomphaceae | Ramaria 2 (4) | ||
| Lentariaceae | Hydnocristella 1 (1), Lentaria 1 (2) | ||
| Hymenochaetales | Hirschioporaceae | Hirschioporus 2 (9), Pallidohirschioporus 1 (16) | |
| Hymenochaetaceae | Fomitiporella 1 (1), Fomitiporia 2 (7), Fulvifomes 1 (2), Fuscoporia 3 (5), Hydnoporia 1 (1), Hymenochaete 5 (7), Hymenochaetopsis 1 (2), Inocutis 3 (8), Inonotus 4 (24), Phellinopsis 1 (2), Phellinus 8 (70), Phylloporia 1 (2), Porodaedalea 4 (8), Pyrrhoderma 1 (2), Sanghuangporus 1 5), Tropicoporus 1 (2), Xanthoporia 1 (3) | ||
| Hyphodontiaceae | Hyphodontia 3 (8) | ||
| Oxyporaceae | Oxyporus 3 (10) | ||
| Rickenellaceae | Peniophorella 1 (1) | ||
| Schizoporaceae | Schizopora 1 (3), Xylodon 6 (13) | ||
| Trichaptaceae | Pseudotrichaptum 1 (2) | ||
| Unspecified | “Trichaptum” (4) | ||
| Phallales | Phallaceae | Clathrus 1 (1), Mutinus 2 (2), Phallus 4 (20) | |
| Polyporales | Adustoporiaceae | Adustoporia 1 (1), Amyloporia 1 (6), Rhodonia 1 (4) | |
| Auriporiaceae | Auriporia 1 (1) | ||
| Cerrenaceae | Cerrena 3 (22) | ||
| Climacocystaceae | Climacocystis 1 (2) | ||
| Fibroporiaceae | Fibroporia 1 (1) | ||
| Fomitopsidaceae | Antrodia 4 (10), Daedalea 2 (7), Fomitopsis 3 (41), Neoantrodia 2 (2), Ranadivia 1 (1), Rhodofomes 2 (16), Rhodofomitopsis 1 (3) | ||
| Ganodermataceae | Ganoderma 7 (77), Sanguinoderma 1 (1) | ||
| Grifolaceae | Grifola 1 (7) | ||
| Hyphodermataceae | Hyphoderma 3 (7), Mutatoderma 1 (5) | ||
| Incrustoporiaceae | Skeletocutis 3 (4), Tyromyces 3 (7) | ||
| Irpicaceae | Byssomerulius 1 (6), Ceriporia 2 (9), Efibula 1 (4), Flavodon 1 (3), Gloeoporus 1 (1), Irpex 3 (21), Leptoporus 1 (4), Resiniporus 1 (1), Trametopsis 1 (5), Vitreoporus 1 (8) | ||
| Ischnodermataceae | Ischnoderma 1 (10) | ||
| Laetiporaceae | Laetiporus 3 (20) | ||
| Laricifomitaceae | Laricifomes 1 (7) | ||
| Meripilaceae | Meripilus 1 (4), Physisporinus 1 (3), Rigidoporus 3 (5) | ||
| Meruliaceae | Aurantiporus 1 (3), Ceriporiopsis 1 (2), Climacodon 2 (10), Crustodontia 1 (1), Hermanssonia 1 (2), Hydnophlebia 1 (4), Lilaceophlebia 1 (1), Merulius 1 (2), Mycoacia 2 (4), Pappia 1 (5), Phlebia 4 (31), Phlebiodontia 1 (1), Phlebiopsis 2 (2), Sarcodontia 1 (32) | ||
| Panaceae | Cymatoderma 2 (2), Panus 3 (23) | ||
| Phaeolaceae | Phaeolus 1 (9) | ||
| Phanerochaetaceae | Bjerkandera 2 (26), Hapalopilus 2 (6), Phanerochaete 2 (15), Porostereum 1 (5), Rhizochaete 1 (3), Terana 1 (1) | ||
| Podoscyphaceae | Abortiporus 1 (1), Podoscypha 1 (1) | ||
| Polyporaceae | Abundisporus 1 (1), Cellulariella 2 (9), Cerioporus 4 (14), Cubamyces 3 (8), Daedaleopsis 4 38), Earliella 1 (1), Echinochaete 1 (1), Favolus 3 (8), Fomes 2 (13), Funalia 2 (2), Grammothele 1 (1), Haploporus 1 (1), Hexagonia 1 (1), Lentinus 8 (47), Lenzites 1 (9), Lopharia 1 (1), Microporellus 1 (1), Microporus 2 (21), Neofavolus 2 (8), Neofomitella 1 (1), Perenniporia 1 (2), Picipes 4 (15), Podofomes 2 (4), Polyporus 3 (20), Poronidulus 1 (3), Pseudofavolus 1 (2), Pyrofomes 1 (1), Szczepkamyces 1 (1), Trametes 16 (161) | ||
| Postiaceae | Amaropostia 1 (2), Calcipostia 1 (2), Cyanosporus 1 (2), Jahnoporus 1 (1), Oligoporus 1 (1), Osteina 2 (2), Postia 1 (7), Ptychogaster 1 (2), Spongiporus 1 (1) | ||
| Pycnoporellaceae | Crustoderma 2 (4), Pycnoporellus 2 (8) | ||
| Sparassidaceae | Sparassis 2 (5) | ||
| Steccherinaceae | Antrodiella 5 (13), Junghuhnia 1 (5), Metuloidea 2 (22), Mycorrhaphium 1 (4), Nigroporus 1 (4), Steccherinum 5 (39) | ||
| Russulales | Auriscalpiaceae | Artomyces 1 (10), Auriscalpium 1 (12), Lentinellus 7 (26) | |
| Bondarzewiaceae | Bondarzewia 3 (9), Gloiodon 1 (1), Heterobasidion 5 (7), Laurilia 1 (3) | ||
| Hericiaceae | Dentipellis 1 (5), Hericium 5 (29), Laxitextum 1 (10) | ||
| Peniophoraceae | Baltazaria 1 (1), Gloiothele 1 (2), Peniophora 8 (26), Vararia 1 (1) | ||
| Stereaceae | Conferticium 1 (1), Stereum 6 (36), Xylobolus 4 (12) | ||
| Xenasmatellales | Xenasmatellaceae | Xenasmatella 1 (1) | |
| Dacrymycetes | Dacrymycetales | Dacrymycetaceae | Dacryopinax 1 (1) |
| Tremellomycetes | Tremellales | Phaeotremellaceae | Phaeotremella 1 (2) |
| Species | Strain Number | Content, mg/g, Dry Weight * | |||
|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | ||
| Daedalea xantha | LE-BIN 3823 | 5.99 | 0.12 | 3.81 | 0.00 |
| Gloeophyllum sepiarium | LE-BIN 3412 | 4.55 | 0.59 | 0.94 | 0.40 |
| Gloeophyllum trabeum | LE-BIN 0157 | 9.62 | 0.04 | 2.50 | tr |
| Inonotus obliquus | LE-BIN 3209 | 4.5 | tr | n/d | n/d |
| Laetiporus sulphureus | LE-BIN 3867 | 4.74 | 0.02 | 0.93 | 0.00 |
| Neolentinus lepideus | LE-BIN 4332 | 1.27 | 8.83 | 0.29 | 0.55 |
| Oxyporus populinus | LE-BIN 3818 | 0.44 | 0.06 | 0.03 | 0.00 |
| Pholiota aurivella | LE-BIN 3929 | 1.59 | 0.24 | 1.00 | 0.23 |
| Pseudofavolus tenuis | LE-BIN 4210 | 0.03 | n/d | 0.00 | n/d |
| Rhizomarasmius undadus | LE-BIN 3571 | 0.09 | tr | tr | n/d |
| Sparassis crispa | LE-BIN 2902 | 0.08 | n/d | tr | n/d |
| Stereum subtomentosum | LE-BIN 2841 | 0.09 | tr | tr | n/d |
| Tyromyces lacteus | LE-BIN 3990 | 8.19 | 1.40 | 0.07 | 0.01 |
| Vitreoporus dichrous | LE-BIN 3131 | 0.07 | 0.02 | 0.01 | 0.01 |
| Vuilleminia comedens | LE-BIN 3878 | 0.63 | 0.07 | 0.05 | 0.01 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Psurtseva, N.V.; Kiyashko, A.A.; Senik, S.V.; Shakhova, N.V.; Belova, N.V. The Conservation and Study of Macromycetes in the Komarov Botanical Institute Basidiomycetes Culture Collection—Their Taxonomical Diversity and Biotechnological Prospects. J. Fungi 2023, 9, 1196. https://doi.org/10.3390/jof9121196
Psurtseva NV, Kiyashko AA, Senik SV, Shakhova NV, Belova NV. The Conservation and Study of Macromycetes in the Komarov Botanical Institute Basidiomycetes Culture Collection—Their Taxonomical Diversity and Biotechnological Prospects. Journal of Fungi. 2023; 9(12):1196. https://doi.org/10.3390/jof9121196
Chicago/Turabian StylePsurtseva, Nadezhda V., Anna A. Kiyashko, Svetlana V. Senik, Natalya V. Shakhova, and Nina V. Belova. 2023. "The Conservation and Study of Macromycetes in the Komarov Botanical Institute Basidiomycetes Culture Collection—Their Taxonomical Diversity and Biotechnological Prospects" Journal of Fungi 9, no. 12: 1196. https://doi.org/10.3390/jof9121196
APA StylePsurtseva, N. V., Kiyashko, A. A., Senik, S. V., Shakhova, N. V., & Belova, N. V. (2023). The Conservation and Study of Macromycetes in the Komarov Botanical Institute Basidiomycetes Culture Collection—Their Taxonomical Diversity and Biotechnological Prospects. Journal of Fungi, 9(12), 1196. https://doi.org/10.3390/jof9121196





