Scedosporium and Lomentospora Infections: Contemporary Microbiological Tools for the Diagnosis of Invasive Disease
Abstract
1. Introduction
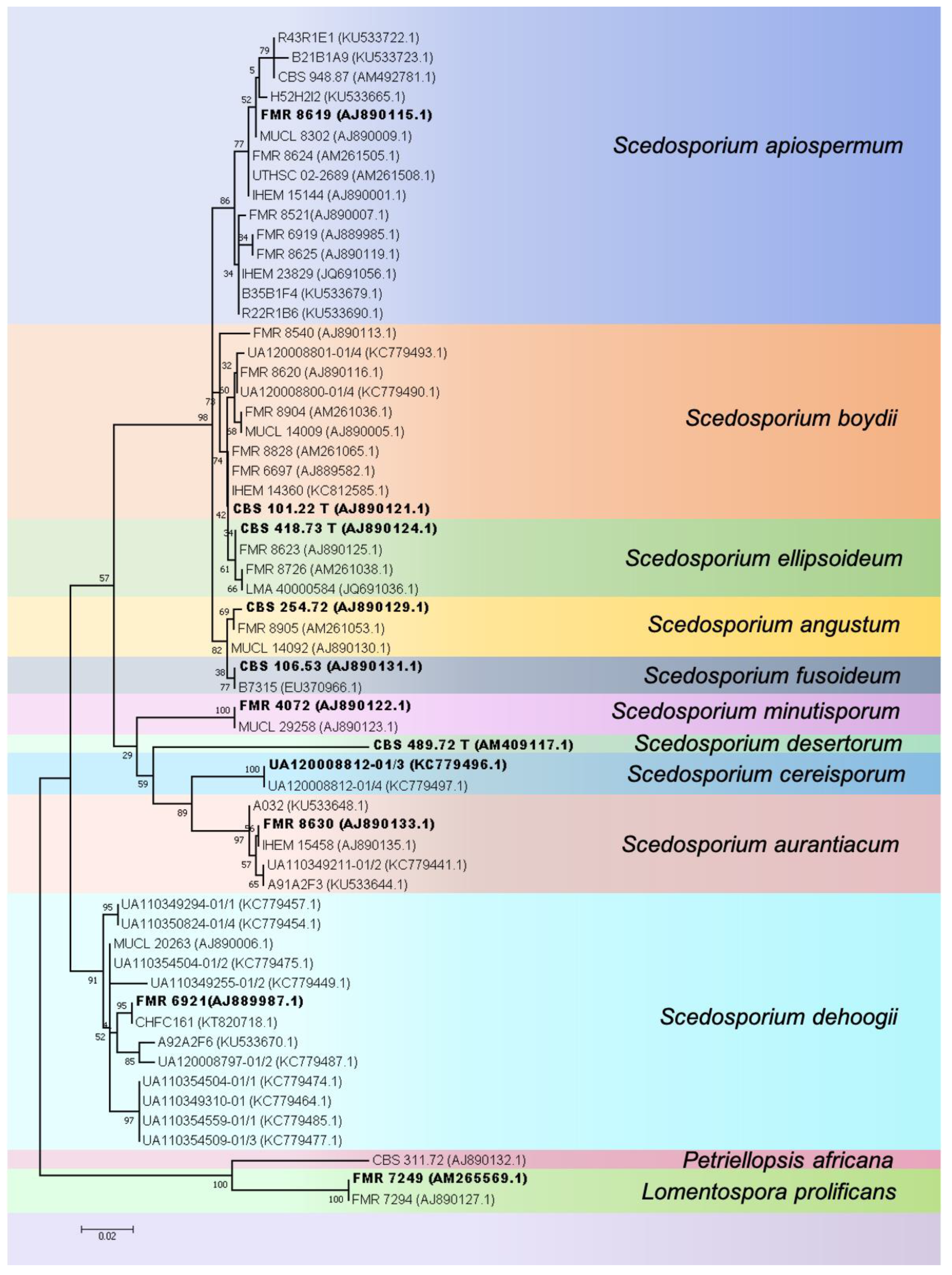
2. Taxonomy and Species
3. Microbiological Diagnosis
3.1. Histopathology
3.2. Microscopy and Culture
3.2.1. Culture
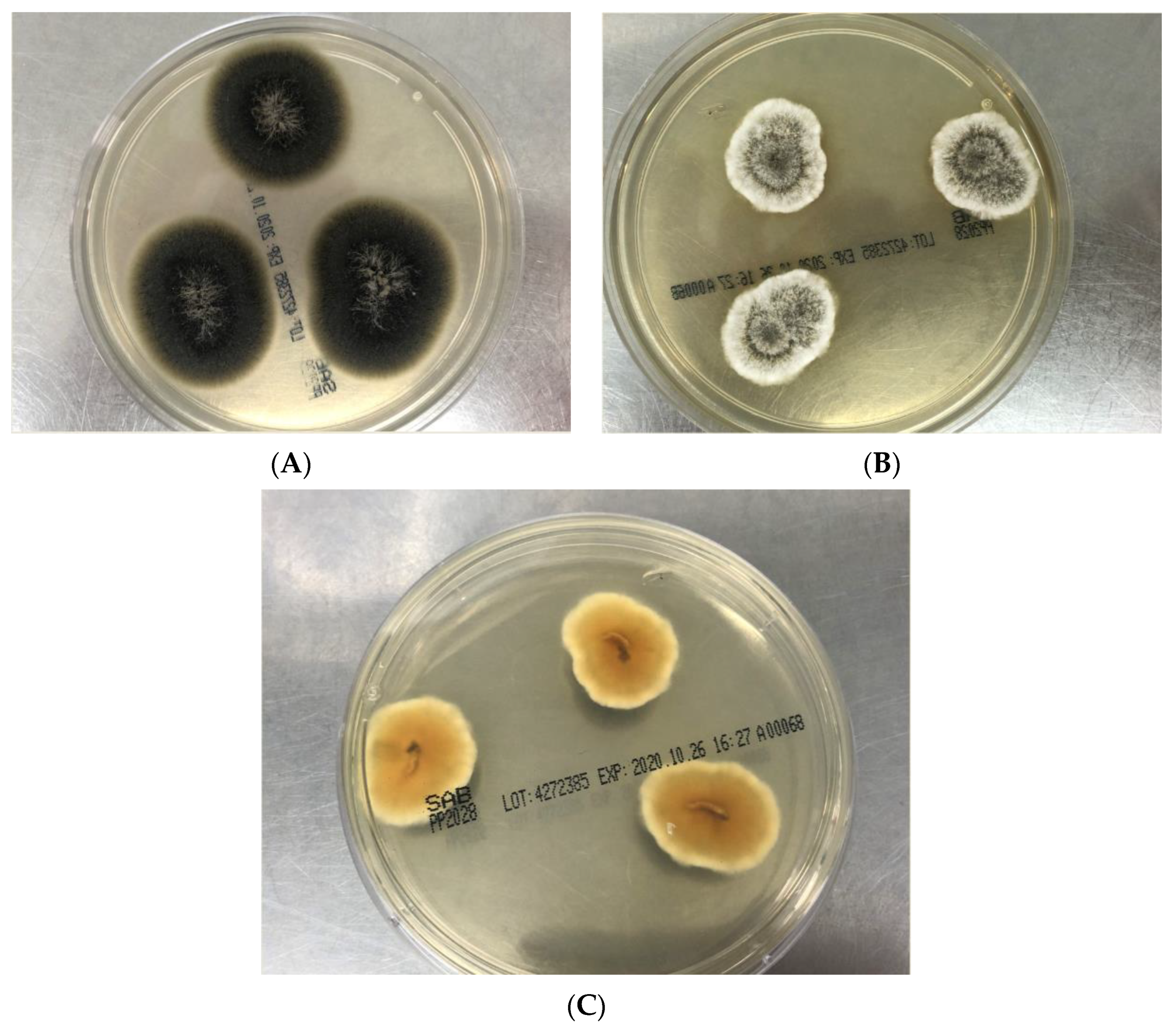
3.2.1.1. Macroscopic Features
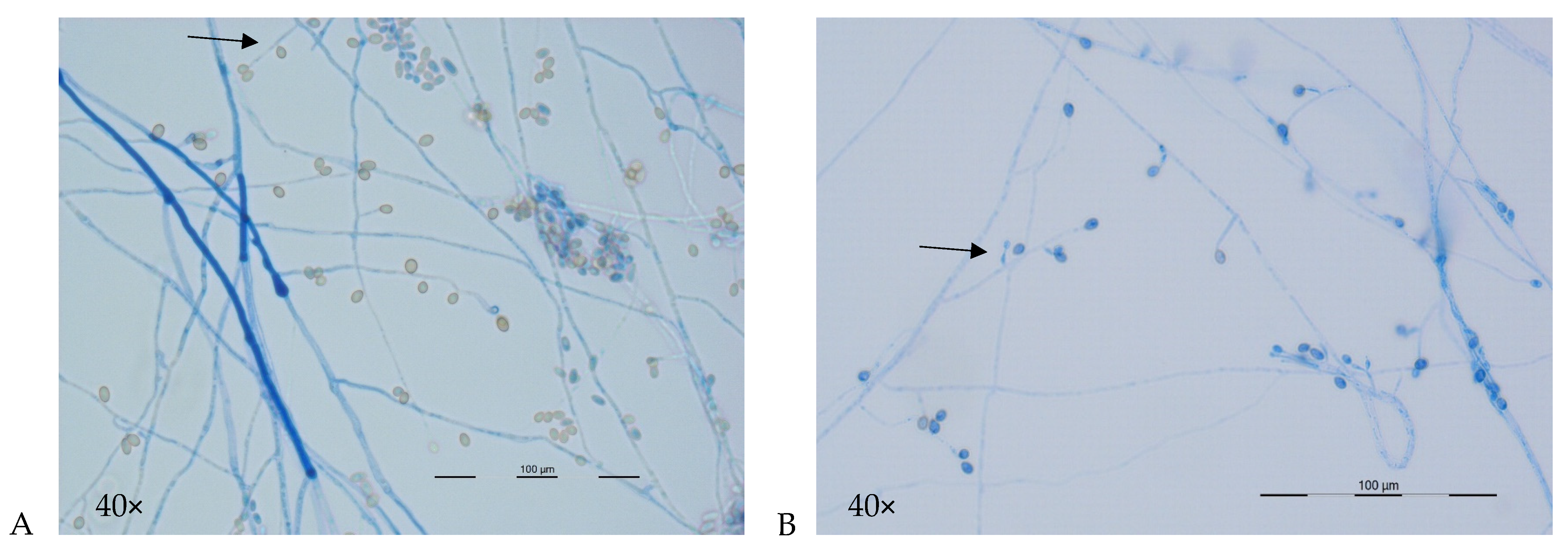
3.2.1.2. Microscopic Features
3.3. Species Identification
3.3.1. Phenotypic Methods
3.3.2. MALDI-TOF MS
3.3.3. Identification by Molecular Methods (Isolates)
3.4. Direct Detection in Clinical Specimens by Molecular Methods
3.4.1. Broad Range PCR
3.4.2. Targeted Multiplex PCR
3.4.3. Oligonucleotide Array Assay
3.5. Serology
3.6. In Vitro Susceptibility to Antifungal Agents
New Antifungal Drugs in the Pipeline
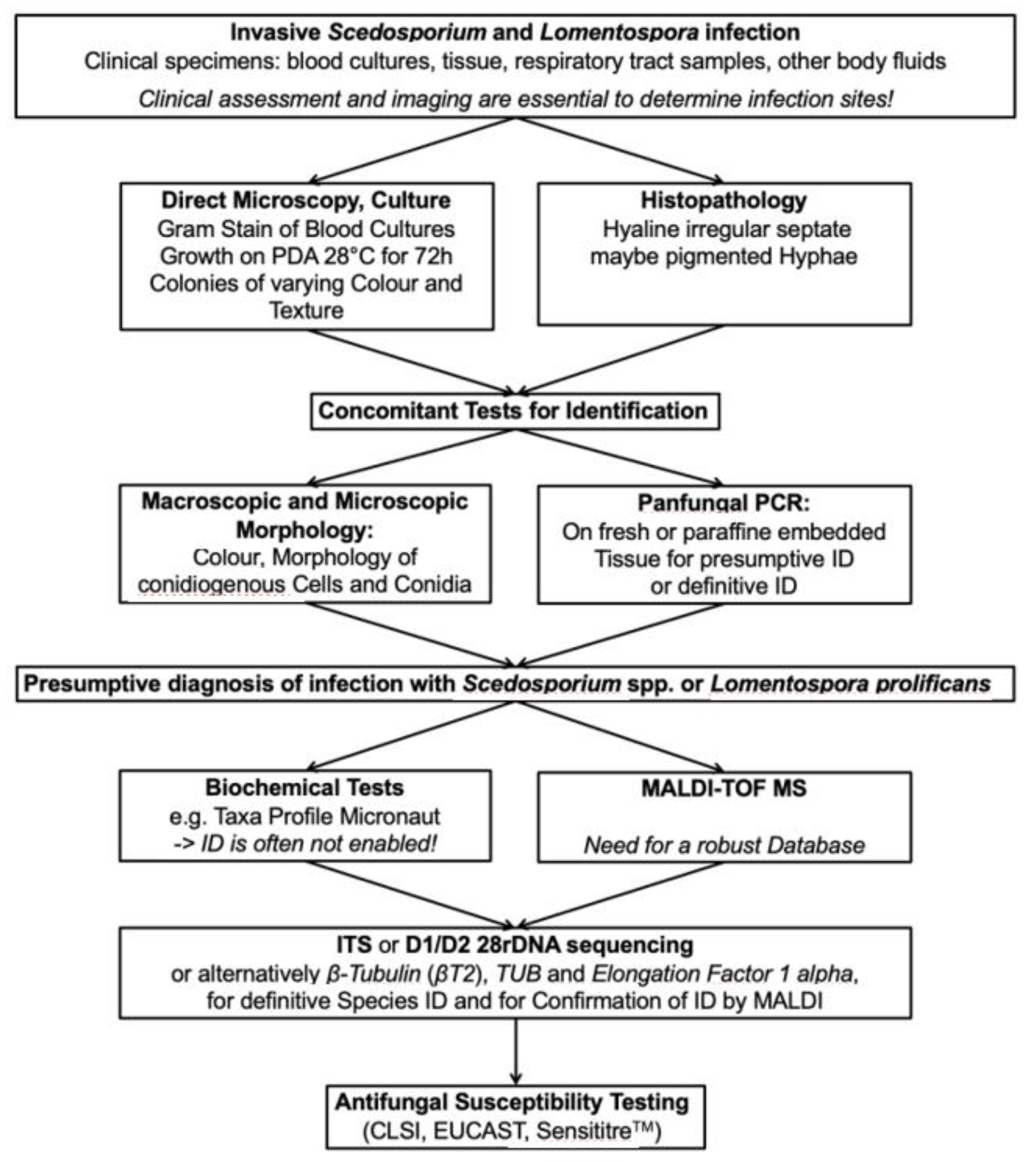
3.7. Diagnostic Algorithms in the Microbiology and Pathology Laboratories
4. Next Generation Sequencing Approaches
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lackner, M.; De Hoog, S.; Yang, L.; Moreno, L.F.; Ahmed, S.A.; Andreas, F.; Kaltseis, J.; Nagl, M.; Lass-Flörl, C.; Risslegger, B.; et al. Proposed nomenclature for Pseudallescheria, Scedosporium and related genera. Fungal Divers. 2014, 67, 1–10. [Google Scholar] [CrossRef]
- Cortez, K.J.; Roilides, E.; Quiroz-Telles, F.; Meletiadis, J.; Antachopoulos, C.; Knudsen, T.; Buchanan, W.; Milanovich, J.; Sutton, D.A.; Fothergill, A.; et al. Infections caused by Scedosporium spp. Clin. Microbiol. Rev. 2008, 21, 157–197. [Google Scholar] [CrossRef] [PubMed]
- Ramirez-Garcia, A.; Pellon, A.; Rementeria, A.; Buldain, I.; Barreto-Bergter, E.; Rolilin-Pinheiro, R.; de Meirelles, J.V.; Xisto, M.; Ranque, S.; Havlicek, H.; et al. Scedosporium and Lomentospora: An updated overview of underrated opportunists. Med. Mycol. 2018, 56, S102–S125. [Google Scholar] [CrossRef] [PubMed]
- Slavin, M.A.; van Hal, S.; Sorrell, T.C.; Lee, A.; Marriott, D.; Daveson, K.; Kennedy, K.; Hajkowicz, K.; Halliday, C.; Athan, E.; et al. Invasive infections due to filamentous fungi other than Aspergillus: Epidemiology and determinants of mortality. Clin. Microbiol. Infect. 2015, 21, 1e.1–1e.10. [Google Scholar] [CrossRef]
- Schwartz, C.; Brandt, C.; Antwieler, E.; Karanich, A.; Staab, D.; Schmitt-Grobe, S.; Fischer, R.; Hartl, D.; Thronicke, A.; Tintelnot, K. Prospective multicenter German study on pulmonary colonization with Scedosporium/Lomentopsora species in cystic fibrosis: Epidemiology and new association factors. PLoS ONE 2017, 12, e00171485. [Google Scholar]
- Kell, M.; Stevens, R.; Konecny, P. Lomentospora prolificans endocarditis—case report and literature review. BMC Infect. Dis. 2016, 16, 36. [Google Scholar] [CrossRef] [PubMed]
- Lackner, M.; Hagen, F.; Meis, J.F.; Ende, A.H.G.G.V.D.; Vu, D.; Robert, V.; Fritz, J.; Moussa, T.A.A.; De Hoog, G.S. Susceptibility and diversity in the therapy-refractory genus Scedosporium. Antimicrob. Agents. Chemother. 2014, 58, 5877–5885. [Google Scholar] [CrossRef]
- Biswas, C.; Law, D.; Birch, M.; Halliday, C.; Sorrell, T.C.; Rex, J.; Slavin, M.; Chen, S.C.-A. In vitro activity of the novel antifungal compound F901318 against Australian Scedosporium and Lomentospora fungi. Med. Mycol. 2018, 56, 1050–1054. [Google Scholar] [CrossRef]
- Jenks, J.D.; Seidel, D.; Cornely, O.A.; Chen, S.; van Hal, S.; Kauffman, C.; Miceli, M.H.; Heinemann, M.; Christner, M.; Saenz, A.; et al. Clinical characteristics and outcomes of invasive Lomentospora prolificans: Analysis of patients in the FungiScopeR registry. Mycoses 2020. [Google Scholar] [CrossRef]
- Gilgado, F.; Cano, J.; Gené, J.; Guarro, J. Molecular Phylogeny of the Pseudallescheria boydii species complex: Proposal or two new species. J. Clin. Microbiol. 2005, 43, 4930–4942. [Google Scholar] [CrossRef]
- Gilgado, F.; Cano, J.; Gene, J.; Sutton, D.A.; Guarro, J. Molecular and phenotypic data supporting distinct species statues for Scedosporium apiospermum and Pseuadlescheria boydii and the proposed new species Scedosporium dehoogii. J. Clin. Microbiol. 2008, 46, 766–771. [Google Scholar] [CrossRef] [PubMed]
- Rainer, J.; Kaltseis, J. Diversity in Scedosporium dehoogii (Microascaceae): S. deficiens sp. nov. Sydowia 2010, 62, 137–147. [Google Scholar]
- Crous, P.W.; Wingfield, M.J.; Burgess, T.I.; Hardy, G.E.; Crane, C.; Barratt, S.; Cano-Lina, J.F.; Le roux, J.; Thangevel, R.; Guarro, J.; et al. Fungal planet description sheets: 469–557. Personia 2016, 37, 218–403. [Google Scholar] [CrossRef]
- Hawksworth, D.L.; Crous, P.W.; Redhead, S.A.; Reynolds, D.R.; Samson, R.A.; Seifert, K.A.; Taylor, J.W.; Wingfield, M.J.; Abaci, Ö.; Aime, C.; et al. The Amsterdam Declaration on Fungal Nomenclature. IMA Fungus 2011, 2, 105–112. [Google Scholar] [CrossRef]
- Heath, C.H.; Slavin, M.A.; Sorrell, T.C.; Handke, R.; Harun, A.; Phillips, M.; Nguyen, Q.; Delhaes, L.; Ellis, D.; Meyer, W.; et al. Population-based surveillance for scedopsporiosis in Australia: Epidemiology, disease manifestations and emergence of Scedosporium aurantiacum infection. Clin. Infect. Microbiol. 2009, 15, 689–693. [Google Scholar] [CrossRef] [PubMed]
- Johnson, L.S.; Shields, R.K.; Clancy, C.J. Epidemiology, clinical manifestations, and outcomes of Scedosporium infections among solid organ transplant recipients. Transpl. Infect. Dis. 2014, 16, 578–587. [Google Scholar] [CrossRef] [PubMed]
- Bronnimann, D.; Garcia-Hermoso, D.; Dromer, F.; Lanternier, F.; The French Mycoses Study Group. Scedosporiosis/lomentosporiosis observational study (SOS): Clinical significance of Scedosporium species identification. Med. Mycol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Donnelly, J.P.; Chen, S.C.-A.; Kauffman, C.A.; Stienbach, W.J.; Baddley, J.W.; Verweij, P.E.; Clancy, C.J.; Wingard, J.R.; Lockhart, S.R.; Groll, A.H.; et al. Revision and update of the consensus definitions for invasive fungal disease from the European Organization for Research and Treatment of Cancer and the Mycoses Study Group Education and Research Consortium. Clin. Infect. Dis. 2020, 71, 1367–1376. [Google Scholar]
- Hoenigl, M.; Slamanton-Garcia, J.; Walsh, T.J.; Nucci, M.; Neoh, C.; Jenks, J.D.; Lackner, M.; Sprute, R.; Alhatmi, A.; Bassetti, M.; et al. Global guideline for the diagnosis and management of rare mold infections: An initiative of the ECMM in cooperation with ISHAM and ASM. Lancet. Infect. Dis. 2020. (Accepted; in Press). [Google Scholar]
- Guarner, J.; Brandt, M.E. Histopathologic diagnosis of fungal infections in the 21st century. Clin. Micrbiol. Rev. 2011, 24, 247–280. [Google Scholar] [CrossRef]
- Kimura, M.; Maenishi, O.; Ito, H.; Ohkusu, K. Unique histological characteristics of Scedosporium that could aid in its identification. Pathol. Int. 2010, 60, 131–136. [Google Scholar] [CrossRef] [PubMed]
- Hayden, R.T.; Isotalo, P.A.; Parrett, T.; Wolk, D.M.; Qian, X.; Roberts, G.D.; Lloyd, R.V. In situ hybridization for the differentiation of Aspergillus, Fusarium and Pseudallescheria in tissue section. Diagn. Mol. Pathol. 2003, 12, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Albernaz, V.; Huston, B.; Castillo, M.; Mukerji, S.; Bouldin, T.W. Pseudallescheria boydii infection of the brain: Imaging with pathologic confirmation. Am. J. Neuroradiol. 1996, 17, 589–592. [Google Scholar] [PubMed]
- Rainer, J.; Kaltseis, J.; de Hoog, S.G.; Summerbell, R.C. Efficacy of a selective isolation procedure for members of the Pseudallescheria boydii complex. Ant. Leeuwenhoek. 2008, 93, 315–322. [Google Scholar] [CrossRef] [PubMed]
- Pham, T.; Giraud, S.; Schuliar, G.; Rougeron, A.; Bouchara, J.P. Scedo-Select III: A new semi-selective culture medium for detection of the Scedosporium apiospermum species complex. Med. Mycol. 2015, 53, 512–519. [Google Scholar] [CrossRef]
- King, A.D., Jr.; Hocking, A.D.; Pitt, J.I. Dichloran-rose bengal medium for enumeration and isolation of molds from foods. Appl. Environ. Microbiol. 1979, 37, 959–964. [Google Scholar] [CrossRef]
- Blyth, C.C.; Harun, A.; Middleton, P.G.; Sleiman, S.; Lee, O.; Sorrell, T.; Meyer, W.; Chen, S.C. Detection of occult Scedosporium species in respiratory tract specimens form patients with cystic fibrosis by use of selective media. J. Clin. Microbiol. 2010, 48, 314–316. [Google Scholar] [CrossRef]
- De Hoog, G.S.; Guarro, J.; Gene, J.; Ahmed, S.; Al-Hatmi, A.M.S.; Figueras, J.; Vitale, R.G. Atlas of Clinical Fungi, 3rd ed. Utrecht/Reus. 2019. Available online: http://www.clinicalfunig.org/ (accessed on 31 October 2020).
- Horre, R.; Schaal, K.P.; Marklein, G.; de Hoog, G.S.; Reiffert, S.M. Physiological typing of Psedualleascheria and Scedosporium strains using Taxa Profile, a semi-automated, 384-well microtiter system. Mycoses 2011, 54, 56–65. [Google Scholar] [CrossRef]
- Kaur, J.; Duan, S.Y.; Vaas, L.A.I.; Penesyan, A.; Meyer, W.; Paulsen, I.; Nevalainen, H. Phenotypic profiling of Scedosporium aurantiacum, an opportunistic pathogen colonizing human lungs. PLoS ONE 2015, 10, e0122354. [Google Scholar] [CrossRef]
- Wilkendorf, L.S.; Bowles, E.; Buil, J.B.; van der Lee, H.A.L.; Posteraro, B.; Sanguientti, M.; Verweij, P.E. Update on MALDI-TOF MS identification of filamentous fungi. J. Clin. Microbiol. 2020. [Google Scholar] [CrossRef]
- Patel, R. A moldy application of MALDI: MALDI-TOF mass spectrometry for fungal identification. J. Fungi 2019, 5, 4. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, D.; Monteiro, C.; Faria, M.A.; Pinto, E. VitekR MS v3.0 system in the identification of filamentous fungi. Mycopathologia 2019, 184, 645–651. [Google Scholar] [CrossRef] [PubMed]
- Sleiman, S.; Halliday, C.; Chapman, B.; Brown, M.; Nitschke, J.; Lau, A.F.; Chen, S.C.-A. Performance of matrix-assisted laser desorption ionization-time of flight mass spectrometry for identification of Aspergillus, Scedosporium and Fusarium spp. in the Australian clinical setting. J. Clin. Microbiol. 2016, 54, 2182–2186. [Google Scholar] [CrossRef] [PubMed]
- Bernhard, M.; Zautner, A.E.; Steinmann, J.; Weig, M.; GroB, U.; Bader, O. Towards proteomic species barcoding of fungi—An example using Scedosporium/Pseudallescheria complex isolates. Fungal. Biol. 2016, 120, 162–165. [Google Scholar] [CrossRef]
- Sitterle, E.; Giraud, S.; Leto, J.; Bouchara, J.P.; Rougeron, A.; Morio, F.; Bougnoux, M.E. matrix-assisted laser desorption ionization-time of flight mass spectrometry for fast and accurate identification of Pseudallescheria/Scedosporium species. Clin. Microbiol. Infect. 2014, 20, 929–935. [Google Scholar] [CrossRef]
- Lau, A.F.; Walchak, R.C.; Miller, H.B.; Slechta, E.S.; Kamboj, K.; Rieebe, K.; Frank, K.M. Multicenter study demonstrates standardization requirements for mold identification by MALDI-TOF MS. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
- Zvezdanova, M.E.; Escribano, P.; Riuz, A.; Martinez-Jimenez, M.C.; Pepaez, T.; Ccollazos, A.; Rodriguez-Sanchez, B. Increased species assignment of filamentous fungi using MALDI-TOF MS coupled with simplified sample processing and an in-house library. Med. Mycol. 2018, 57, 63–70. [Google Scholar] [CrossRef]
- Irinyi, L.; Serena, C.; Garcia-Hermoso, D.; Arabatzis, M.; Desnos-Ollivier, M.; Vu, D.; Cardindi, G.; Arthur, I.; Normand, A.-C.; Giraldo, A.; et al. International Society of Human and Animal Mycology (ISHAM)-ITS reference DNA barcoding database: The quality controlled standard tool for routine identification of human and animal pathogenic fungi. Med. Mycol. 2015, 53, 313–337. [Google Scholar] [CrossRef]
- Matray, O.; Mouhajir, A.; Giraud, S.; Gordon, C.; Gargala, G.; Labbe, F.; Rougeron, A.; Ballet, J.J.; Zouhair, R.; Bouchara, J.-P. Semi-automated repetitive sequence-based PCR amplification for species of the Scedosporium apiospermum complex. Med. Mycol. 2016, 54, 409–419. [Google Scholar] [CrossRef]
- Bernhardt, A.; Sedlacek, L.; Wagner, S.; Schwarz, C.; Wurstl, B.; Tintelnot, K. Multilocus sequence typing of Scedosporium apiospermum and Pseudallescheria boydii isolates from cystic fibrosis patients. J. Cyst. Fibros. 2013, 12, 592–598. [Google Scholar] [CrossRef]
- Demidov, V.V. Rolling circle amplification in DNA diagnostics: The power of simplicity. Expert. Rev. Mol. Diagn. 2002, 2, 542–548. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Kong, F.; Sorrell, T.C.; Wang, H.; Daun, Y.; Chen, S.C. Practical method for detection and identification of Candida, Aspergillus and Scedosporium spp. by use of rolling circle amplification. J. Clin. Microbiol. 2008, 46, 2423–2427. [Google Scholar] [CrossRef] [PubMed]
- Kidd, S.; Chen, S.C.-A.; Meyer, W.; Halliday, C. A new age in molecular diagnostics for invasive fungal diseases: Are we ready? Front. Microbiol. 2020. [Google Scholar] [CrossRef]
- Sabino, R.; Simoes, H.; Verissimo, C. Detection of deep fungal infections: A polyphasic approach. J. Med. Microbiol. 2019, 68, 81–86. [Google Scholar] [CrossRef]
- Gomez, C.A.; Budvytuene, I.; Zenek, A.J.; Banal, N. Performance of targeted fungal sequencing for culture-independent diagnosis of invasive fungal disease. Clin. Infect. Dis. 2017, 65, 2035–2041. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J.W. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Lau, A.; Chen, S.; Sorrell, T.; Martin, P.; Malik, R.; Carter, D.; Halliday, C. Development and clinical application of a panfungal PCR assay to detect and identify fungal DNA in tissue specimens. J. Clin. Microbiol. 2007, 45, 380–385. [Google Scholar] [CrossRef]
- Valero, C.; de la Cruz-Villar, L.; Zaragoza, O.; Buitrago, M.J. New panfungal real-time PCR assay for diagnosis of invasive fungal infections. J. Clin. Microbiol. 2016, 54, 2910–2918. [Google Scholar] [CrossRef]
- Salehi, E.; Hedayati, M.T.; Zoll, J.; Rafati, H.; Ghasemi, M.; Doroudinia, A.; Abastabar, M.; Tolooe, A.; Snelders, E.; van der Lee, H.A.; et al. Discrimination of aspergillosis, mucormycosis, fusariosis and scedosporiosis in formalin-fixed paraffin-embedded tissue specimens by use of multiple real-time quantitative PCR assays. J. Clin. Microbiol. 2016, 11, 2798–2903. [Google Scholar] [CrossRef]
- Zeller, I.; Schabereiter-Gurtner, C.; Mihaltis, V.; Selitsch, B.; Barousch, W.; Hirschi, A.M.; Makristathis, A.; Willinger, B. Detection of fungal pathogens by a new broad range real time PCR assay targeting the fungal ITS region. J. Med. Microbiol. 2017, 16, 1383–1392. [Google Scholar] [CrossRef]
- Trubiano, J.A.; Dennison, A.M.; Morrissey, C.O.; Chua, K.Y.; Halliday, C.L.; Chen, S.C.-A. Clinical utility of panfungal polymerase chain reaction for the diagnosis of invasive fungal disease: A single centre experience. Med. Mycol. 2016, 54, 138–146. [Google Scholar] [CrossRef]
- Harun, A.; Blyth, C.C.; Gilgado, F.; Middleton, P.; Chen, S.C.; Meyer, W. Development and validation of a multiplex PCR for detection of Scedosporium spp. in respiratory tract specimens form patients with cystic fibrosis. J. Clin. Microbiol. 2011, 49, 1508–1512. [Google Scholar] [CrossRef] [PubMed]
- Castelli, M.V.; Buitrago, M.J.; Bernal-Martinez, L.; Gomez-Lopez, A.; Rodriguez-Tudela, J.L.; Cuenca-Estrella, M. Development and validation of a quantitative PCR assay for diagnosis of scedosporiosis. J. Clin. Microbiol. 2008, 46, 3412–3416. [Google Scholar] [CrossRef] [PubMed]
- Bouchara, J.P.; Hsieh, H.Y.; Croquefer, S.; Barton, R.; Marchais, V.; Pihet, M.; Chang, T.C. Development of an oligonucleotide assay for direct detection of fungi in sputum samples from patients with cystic fibrosis. J. Clin. Microbiol. 2009, 47, 142–152. [Google Scholar] [CrossRef] [PubMed]
- He, S.; Hang, J.P.; Zhang, L.; Wang, F.; Zhang, D.C.; Gong, F.H. A systematic review and meta-analysis of diagnostic accuracy of serum 1,3-β-D-glucan for invasive fungal infection: Focus on cutoff levels. J. Microbiol. Immunol. Infect 2015, 48, 351–356. [Google Scholar] [CrossRef] [PubMed]
- Karageorgopoulos, D.E.; Voulouman, E.K.; Nziora, F.; Michalpoulos, A.; Rafailidos, P.I.; Falagas, M.E. β-D-glucan assay for the diagnosis of invasive fungal infections: A meta-analysis. Clin. Infect. Dis. 2011, 52, 750–770. [Google Scholar] [CrossRef]
- Cuerara, M.S.; Alhambra, A.; Moragues, M.; Gonzalez-Elorza, E.; Ponton, J.; de Palacio, A. Detection of (1,2)-β-D-glucan as an adjunct to diagnosis in a mixed population with uncommon proven invasive fungal diseases or with an unusual clinical presentation. Clin. Vaccine. Immunol. 2009, 16, 423–426. [Google Scholar]
- Levesque, E.; Rizk, F.; Noorah, Z.; Ait-Ammar, N.; Cordonnier-Jourdin, C.; El-Anbassi, S.; Bonnal, C.; Azoulay, D.; Merle, J.-C.; Botterel, F. Detection of (1,3)-β-D-glucan for the diagnosis of invasive fungal infection in liver transplant recipients. Int. J. Mol. Sci. 2017, 18, 862. [Google Scholar] [CrossRef]
- Nishimori, M.; Takahashi, T.; Suzuki, E.; Kodaka, T.; Hiramoto, N.; Itoh, K.; Tsunemine, H.; Yarita, K.; Kamei, K.; Takegawa, H.; et al. Fatal fungemia with Scedosporium prolificans in a patient with acute myeloid leukemia. Med. Mycol. J. 2014, 55E, E63–E70. [Google Scholar] [CrossRef]
- Thornton, C.R.; Ryder, L.S.; Le Cocq, K.; Soanes, D.M. Identifying the emerging human pathogen Scedosporium prolificans by using a species-specific monoclonal antibody that bind to the melanin biosynthetic enzyme tetrahydroxynaphthalene reductase. Environ. Microbiol. 2015, 17, 1023–1039. [Google Scholar] [CrossRef]
- Thornton, C.R. Tracking the emerging human pathogen Pseudallescheria boydii by using high specific monoclonal antibodies. Clin. Vaccine. Immunol. 2009, 16, 756–7645. [Google Scholar] [CrossRef]
- Mina, S.; Marot-Leblond, A.; Cimon, B.; Fleury, M.J.J.; Larcher, G.; Bouchara, J.-P. Purification and characterization of a mycelial catalase from Scedosporium boydii, a useful tool for specific antibody detection in patients with cystic fibrosis. Clin. Vaccine. Immunol. 2015, 22, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Pluhacek, T.; Lemr, K.; Ghosh, D.; Milde, D.; Novak, J.; Havlicek, V. Characterization of microbial siderophores by mass spectrometry. Mass. Spectrom. Rev. 2016, 35, 35–47. [Google Scholar] [CrossRef] [PubMed]
- Halliday, C.L.; Chen, S.C.; Kidd, S.E.; van Hal, S.; Chapman, B.; Heath, C.H.; Lee, A.; Kennedy, K.; Daveson, K.; Sorrell, T.C.; et al. Antifungal susceptibilities of non-Aspergillus filamentous fungi causing invasive infection in Australia: Support for current antifungal guideline recommendations. Int. J. Antimicrob. Agents. 2016, 48, 453–458. [Google Scholar] [CrossRef] [PubMed]
- Lackner, M.; De Hoog, G.S.; Verweij, P.E.; Najafzadeh, M.; Curfs-Breuker, I.; Klassen, C.; Meis, J. Species-specific antifungal susceptibility patterns of Scedosporium and Pseudallescheria species. Antimicrob. Agents Chemother. 2012, 56, 2635–2642. [Google Scholar] [CrossRef] [PubMed]
- Alastruey-Izuierdo, A.; Alcazar-Fuoli, L.; Rivero-Menendez, O.; Ayats, J.; Castro, C.; Garcia-Rodriguez, J.; Goterris-Bonet, L.; Ibanez-Martinez, E.; Linares-Sicilia, M.J.; Martin-Gomez, M.T.; et al. Molecular identification and susceptibility testing of molds in a prospective surveillance of triazole resistance in Spain (FILPOP2 Study). Antimicrob. Agents. Chemother. 2018, 62, e00358-18. [Google Scholar] [CrossRef]
- Pellon, A.; Ramirez-Garcia, A.; Buldain, I.; Antoran, A.; Martin-Souto, L.; Rementeria, A.; Hernando, F. Pathobiology of Lomentospora prolificans: Could this species serve as a model of primary antifungal resistance? Int. J. Antimicrob. Agents 2018, 51, 10–15. [Google Scholar] [CrossRef]
- Sedlacek, L.; Graf, B.; Schwarz, C.; Albert, F.; Peter, S.; Wurstl, B.; Wagner, S.; Klotz, M.; Becker, A.; Haase, G.; et al. Prevalence of Scedosporium species and Lomentospora prolificans in patients with cystic fibrosis in a multicentre trial by use of a selective medium. J. Cyst. Fibros. 2015, 14, 237–241. [Google Scholar] [CrossRef]
- Trovato, L.; Scalia, G.; Palermo, C.I.; Costanzo, C.M.; Oliveri, S. Evaluation of isavuconazole MICs trips for susceptibility testing of Aspergillus and Scedosporium species. Med. Mycol. 2019, 57, 429–433. [Google Scholar] [CrossRef]
- Jenks, J.D.; Seidel, D.; Cornely, O.A.; Chen, S.; van Hal, S.; Kauffman, C.; Miceli, M.; Heinemann, M.; Christner, M.; Jover-Saenz, A.; et al. Voricoanzole plus terbinafine combination antifungal therapy for invasive Lomentospora prolificans infections; analysis of 41 patients from the FungiScope R registry, 2008–2019. Clin Microbiol Infect 2020, 26, 784.e1–784.e5. [Google Scholar] [CrossRef]
- Blyth, C.C.; Gilroy, N.M.; Guy, S.D.; Chambers, S.T.; Cheong, E.Y.; Gottlieb, T.; McGuiness, S.L.; Thursky, K.A. Consensus guidelines for the treatment of invasive mold infections in hematological malignancy and hematopoeitic stem cell transplantation, 2014. Int. Med. J. 2014, 44, 1333–1345. [Google Scholar] [CrossRef]
- Shaw, K.J.; Ibrahim, A.S. Fosmanogepix: A review of the first-in-class broad spectrum agent for the treatment of invasive fungal infections. J. Fungi 2020, 6, 239. [Google Scholar] [CrossRef] [PubMed]
- Castaheira, M.; Duncanson, F.P.; Diekema, D.J.; Guarro, J.; Jones, R.N.; Pfaller, M.A. Activities of E1210 and comparator agents tested by CLSI and EUCAST broth microdilution methods against Fusarium and Scedosporium species identified using molecular methods. Antimicrob. Agents Chemother. 2012, 56, 352–357. [Google Scholar] [CrossRef] [PubMed]
- Oliver, J.D.; Sibley, G.E.; Beckmann, N.; Dobb, K.S.; Slater, M.J.; McEntee, L.; du Pre, S.; Livermore, J.; Bronley, M.J.; Wiederhold, N.P.; et al. F901318 represents a novel class of antifungal drug that inhibits dihydro-ororate dehydrogenase. Proc. Natl. Acad. Sci. USA 2016, 113, 12809–12814. [Google Scholar] [CrossRef] [PubMed]
- Wiederhold, N.; Law, D.; Birch, M. Dihydroorotate dehydrogenase inhibitor F901318 has potent in vitro activity against Scedosporium species and Lomentospora prolificans. J. Antimicrob. Chemother. 2017, 27, 1977–1980. [Google Scholar] [CrossRef] [PubMed]
- Nagl, M.; Mair, C.; Dietrich, H. Tolerability of inhaled N-chlorotaurine versus sodium chloride in the mouse. J. Med. Res. Prac. 2013, 2, 163–170. [Google Scholar]
- Luo, R.; Zimin, A.; Workman, R.; Fan, Y.; Pertea, G.; Grossman, N.; Wear, W.; Jia, B.; Miller, H.; Casadevall, A.; et al. First draft genome sequence of the pathogenic fungus Lomentospora prolificans (formerly Scedosporium prolificans). G3 2017, 7, 3831–3836. [Google Scholar] [CrossRef]
- Duvaux, L.; Shiller, J.; Vanderputte, P.; De Bernonville, T.; Thornton, C.; Papon, N.; Le Cam, B.; Bouchara, J.P.; Gastebois, A. Draft genome sequence of the human pathogenic fungus Scedosporium boydii. Genome Announc. 2017, 5, e00871-17. [Google Scholar] [CrossRef]
- Vendeputte, P.; Ghamrawi, S.; Rechenmann, M.; Iltis, A.; Giraud, S.; Fleury, M.; Thornton, C.; Delhaes, L.; Meyer, W.; Papon, N.; et al. Draft genome sequence of the pathogenic fungi Scedosporium apiospermum. Genome. Announc. 2014, 2, 10–11. [Google Scholar]
- Perez-Bercoff, A.; Papanicolaou, A.; Ramsperger, M.; Kaur, J.; Patel, H.R.; Harun, A.; Duan, S.Y.; Elbourne, L.; Bouchara, J.P.; Paulsen, I.T.; et al. Draft genome of Australian environmental strain WM09.24 of the opportunistic human pathogen Scedosporium aurantiacum. Genome Announc. 2015, 3, e01526-14. [Google Scholar] [CrossRef]
- Boan, P.; Pang, S.; Gardam, D.J.; Darragh, H.; Wright, M.; Coombs, G.W. Investigation of a Lomentospora prolificans case cluster with whole genome sequencing. Med. Mycol. Case. Rep. 2020, 29, 1–4. [Google Scholar] [CrossRef]
| Species | Colony Morphology on PDA (25 °C) 1 | Growth Temperature | |||
|---|---|---|---|---|---|
| Color (Obverse) | Color (Reverse) | Texture | 40 °C | 45 °C | |
| Lomentospora prolificans | Olive-gray to black; white colored tufts (mycelium) | Dark brown or gray, almost black | Downy to cottony | Yes | Variable |
| Scedosporium apiospermum | White gray, becoming darker gray or brown | Dark brown or gray, almost black | Downy to cottony | Yes | No |
| Scedosporium boydii | White gray, becoming darker gray or brown | Dark brown or gray, almost black | Downy to cottony | Yes | No |
| Scedosporium aurantiacum | Yellow gray to brown gray; concentric growth pattern; white margin | Brown orange or brown but may be colorless | Dense, cottony to woolly | Yes | Yes |
| Species | Cleistothecia | Ascospores | Conidiogenous Cells | Conidia | ||
|---|---|---|---|---|---|---|
| Shape | Distribution | Shape | Size | |||
| Lomentospora prolificans | No Teleomorph | Flask-shaped with basal swelling | Solitary or on branched conidiophores | Globose to subglobose, thick-walled | 3 × 5 µm | |
| Scedosporium apiospermum | P. apiosperma (previously P. boydii) | Cylindrical | Solitary or on branched conidiophores | Globose to subglobose, thick- walled | 5–7 × 4–6 µm | |
| Scedosporium boydii | Globose to subglobose, 100–300 µm diameter | Broadly fusiform, pale yellow or brown—copper; 6–9 × 5–6 µm | Cylindrical | On branched coniodiophores or solitary | Globose to subglobse, thick-walled | 4–9 × 6–10 µm |
| Scedosporium aurantiacum | Teleomorph unknown | Cylindrical or slightly flask-shaped | On branched coniodiophores or solitary | Obvoid, thick-walled | 6–10 × 3–5 µm | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, S.C.-A.; Halliday, C.L.; Hoenigl, M.; Cornely, O.A.; Meyer, W. Scedosporium and Lomentospora Infections: Contemporary Microbiological Tools for the Diagnosis of Invasive Disease. J. Fungi 2021, 7, 23. https://doi.org/10.3390/jof7010023
Chen SC-A, Halliday CL, Hoenigl M, Cornely OA, Meyer W. Scedosporium and Lomentospora Infections: Contemporary Microbiological Tools for the Diagnosis of Invasive Disease. Journal of Fungi. 2021; 7(1):23. https://doi.org/10.3390/jof7010023
Chicago/Turabian StyleChen, Sharon C.-A., Catriona L. Halliday, Martin Hoenigl, Oliver A. Cornely, and Wieland Meyer. 2021. "Scedosporium and Lomentospora Infections: Contemporary Microbiological Tools for the Diagnosis of Invasive Disease" Journal of Fungi 7, no. 1: 23. https://doi.org/10.3390/jof7010023
APA StyleChen, S. C.-A., Halliday, C. L., Hoenigl, M., Cornely, O. A., & Meyer, W. (2021). Scedosporium and Lomentospora Infections: Contemporary Microbiological Tools for the Diagnosis of Invasive Disease. Journal of Fungi, 7(1), 23. https://doi.org/10.3390/jof7010023






