Colony-PCR Is a Rapid Method for DNA Amplification of Hyphomycetes
Abstract
:1. Introduction
2. Materials and Methods
2.1. Media and Cultivation
2.2. Morphological Identification of Isolates
2.3. Direct PCR of Fungal Cultures
2.4. Sequence Analysis and Data Handling
3. Results
PCR Success from Fungal Pure Cultures
4. Discussion
4.1. The Advantages of Direct Fungal Colony PCR
4.2. Factors Affecting Direct Colony PCR Success
4.3. Potential Applications for Direct Fungal Colony PCR
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| BSA | bovine serum albumin |
| CYA | Czapek yeast extract agar |
| DNA | deoxyribonucleic acid |
| G25N | 25% glycerol nitrate agar |
| ITS | internal transcribed spacer |
| JMRC | Jena Microbial Resource Collection |
| MEA | malt extract agar |
| MOTU | molecular operational taxonomic unit |
| PCR | polymerase chain reaction |
Appendix
| MOTU ID | MOTU Description | GenBank ACCN | JMRC:SF:Nr |
|---|---|---|---|
| MK_42 | Aureobasidium sp. | KP714635 | JMRC:SF:12047 |
| GW_52 | Bjerkandera adusta | KP714580 | JMRC:SF:12006 |
| GW_54 | Botrytis sp. | KP714582 | JMRC:SF:12008 |
| GW_07 | Cladosporium sp. 1 | KP714536 | JMRC:SF:11967 |
| GW_17 | Cladosporium sp. 2 | KP714546 | JMRC:SF:11977 |
| GW_43 | Cladosporium sp. 3 | KP714571 | JMRC:SF:12000 |
| GW_59 | Cladosporium sp. 4 | KP714587 | JMRC:SF:12012 |
| MK_40 | Cladosporium sp. 5 | KP714642 | JMRC:SF:12052 |
| GW_40 | Cryptococcus aff. albidosimilis | KP714568 | JMRC:SF:11998 |
| GW_58 | Cryptococcus aff. victoriae | KP714586 | - |
| MK_35 | Cryptococcus friedmannii | KP714628 | JMRC:SF:12041 |
| GW_18 | Cryptococcus sp. 1 | KP714547 | JMRC:SF:11978 |
| GW_24 | Cryptococcus sp. 2 | KP714553 | JMRC:SF:11984 |
| GW_33 | Cryptococcus sp. 3 | KP714562 | JMRC:SF:11992 |
| GW_36 | Cryptococcus sp. 4 | KP714565 | JMRC:SF:11995 |
| MK_45 | Cryptococcus sp. 5 | KP714638 | JMRC:SF:12048 |
| MK_72 | Cryptococcus sp. 6 | KP714662 | JMRC:SF:12074 |
| GW_19 | Cryptococcus terricola | KP714548 | JMRC:SF:11979 |
| MK_53 | Cryptococcus victoriae | KP714646 | JMRC:SF:12056 |
| MK_75 | Cystofilobasidium infirmominiatum | KP714665 | JMRC:SF:12077 |
| MK_14 | Davidiella sp. 1 | KP714607 | JMRC:SF:12027 |
| MK_70 | Davidiella sp. 2 | KP714660 | JMRC:SF:12072 |
| MK_10 | Dioszegia sp. 1 | KP714603 | JMRC:SF:12024 |
| MK_57 | Dioszegia sp. 2 | KP714649 | JMRC:SF:12060 |
| GW_48 | Dothideomycetes unknown | KP714576 | JMRC:SF:12003 |
| GW_35 | Drechslera sp. | KP714564 | JMRC:SF:11994 |
| GW_63 | Epicoccum sp. | KP714591 | JMRC:SF:12016 |
| GW_09 | Fusarium sp. 1 | KP714538 | JMRC:SF:11969 |
| MK_24 | Fusarium sp. 2 | KP714617 | JMRC:SF:12032 |
| MK_06 | Geomyces aff. vinaceus | KP714599 | JMRC:SF:12022 |
| MK_05 | Geomyces pannorum 1 | KP714598 | JMRC:SF:12021 |
| MK_20 | Geomyces pannorum 2 | KP714613 | JMRC:SF:12030 |
| GW_02 | Geomyces sp. 1 | KP714531 | JMRC:SF:11962 |
| GW_03 | Geomyces sp. 2 | KP714532 | JMRC:SF:11963 |
| GW_53 | Geomyces sp. 3 | KP714581 | JMRC:SF:12007 |
| MK_61 | Geomyces sp. 4 | KP714653 | JMRC:SF:12064 |
| MK_38 | Guehomyces pullulans | KP714631 | JMRC:SF:12043 |
| GW_46 | Helgardia sp. | KP714574 | JMRC:SF:12001 |
| MK_09 | Helotiales unknown 1 | KP714602 | JMRC:SF:12023 |
| MK_39 | Helotiales unknown 2 | KP714632 | JMRC:SF:12044 |
| GW_42 | Herpotrichia juniperi 1 | KP714570 | - |
| MK_32 | Herpotrichia juniperi 2 | KP714625 | JMRC:SF:12038 |
| MK_46 | Herpotrichia juniperi 3 | KP714639 | JMRC:SF:12049 |
| GW_39 | Hormonema sp. | KP714567 | JMRC:SF:11997 |
| GW_65 | Ilyonectria sp. | KP714593 | JMRC:SF:12018 |
| MK_63 | Leptodontidium orchidicola | KP714654 | JMRC:SF:12066 |
| MK_01 | Leucosporidiella sp. | KP714594 | JMRC:SF:12019 |
| GW_34 | Leucosporidium sp. | KP714563 | JMRC:SF:11993 |
| GW_12 | Monographella aff. lycopodina | KP714541 | JMRC:SF:11972 |
| GW_50 | Monographella sp. | KP714578 | JMRC:SF:12005 |
| GW_13 | Mortierella aff. gamsii | KP714542 | JMRC:SF:11973 |
| MK_29 | Mortierella alpina 1 | KP714622 | JMRC:SF:12035 |
| MK_34 | Mortierella alpina 2 | KP714627 | JMRC:SF:12040 |
| MK_77 | Mortierella alpina 3 | KP714667 | JMRC:SF:12079 |
| MK_52 | Mortierella antarctica | KP714645 | JMRC:SF:12055 |
| MK_50 | Mortierella globulifera 1 | KP714643 | JMRC:SF:12053 |
| MK_54 | Mortierella globulifera 2 | KP714647 | JMRC:SF:12057 |
| GW_08 | Mortierella humilis | KP714537 | JMRC:SF:11968 |
| GW_29 | Mortierella macrocystis | KP714558 | JMRC:SF:11988 |
| GW_01 | Mortierella sp. 1 | KP714530 | JMRC:SF:11961 |
| GW_16 | Mortierella sp. 2 | KP714545 | JMRC:SF:11976 |
| GW_20 | Mortierella sp. 3 | KP714549 | JMRC:SF:11980 |
| GW_27 | Mortierella sp. 4 | KP714556 | - |
| MK_31 | Mrakia blollopsis | KP714624 | JMRC:SF:12037 |
| MK_41 | Mrakia sp. | KP714634 | JMRC:SF:12046 |
| MK_25 | Mrakiella aquatica | KP714618 | JMRC:SF:12033 |
| GW_44 | Mucor aff. abundans | KP714572 | - |
| GW_45 | Mucor flavus | KP714573 | - |
| GW_15 | Mucor hiemalis 1 | KP714544 | JMRC:SF:11975 |
| MK_15 | Mucor hiemalis 2 | KP714608 | JMRC:SF:12028 |
| MK_69 | Mucor hiemalis 3 | KP714659 | JMRC:SF:12071 |
| GW_47 | Mucor strictus | KP714575 | JMRC:SF:12002 |
| MK_27 | Nectriaceae unknown | KP714620 | JMRC:SF:12034 |
| GW_55 | Penicillium aff. brevicompactum | KP714583 | JMRC:SF:12009 |
| GW_14 | Penicillium aff. lividum | KP714543 | JMRC:SF:11974 |
| GW_31 | Penicillium aff. melinii | KP714560 | JMRC:SF:11990 |
| GW_10 | Penicillium aff. spinulosum | KP714539 | JMRC:SF:11970 |
| GW_25 | Penicillium aff. ubiquetum | KP714554 | JMRC:SF:11985 |
| GW_04 | Penicillium sp. 1 | KP714533 | JMRC:SF:11964 |
| GW_23 | Penicillium sp. 2 | KP714552 | JMRC:SF:11983 |
| GW_32 | Penicillium sp. 3 | KP714561 | JMRC:SF:11991 |
| GW_49 | Penicillium sp. 4 | KP714577 | JMRC:SF:12004 |
| GW_64 | Penicillium sp. 5 | KP714592 | JMRC:SF:12017 |
| MK_60 | Penicillium sp. 6 | KP714652 | JMRC:SF:12063 |
| GW_06 | Phacidium aff. pseudophacidioides | KP714535 | JMRC:SF:11966 |
| GW_05 | Phacidium aff. trichophori | KP714534 | JMRC:SF:11965 |
| GW_41 | Phaeosphaeria sp. | KP714569 | JMRC:SF:11999 |
| GW_56 | Pleosporales unknown 1 | KP714584 | JMRC:SF:12010 |
| MK_13 | Pleosporales unknown 2 | KP714606 | JMRC:SF:12026 |
| MK_36 | Pleosporales unknown 3 | KP714629 | JMRC:SF:12042 |
| MK_47 | Pleosporales unknown 4 | KP714640 | JMRC:SF:12050 |
| MK_30 | Pseudeurotiaceae sp. | KP714623 | JMRC:SF:12036 |
| MK_40 | Pseudogymnoascus destructans 1 | KP714633 | JMRC:SF:12045 |
| MK_51 | Pseudogymnoascus destructans 2 | KP714644 | JMRC:SF:12054 |
| MK_56 | Pseudogymnoascus destructans 3 | KP714648 | JMRC:SF:12059 |
| MK_33 | Rhodotorula colostri | KP714626 | JMRC:SF:12039 |
| GW_26 | Rhodotorula sp. | KP714555 | JMRC:SF:11986 |
| GW_51 | Stemphylium sp. | KP714579 | - |
| GW_57 | Stereum sanguinolentum | KP714585 | JMRC:SF:12011 |
| MK_21 | Sterile Mycelium (Ascomycete) 1 | KP714614 | JMRC:SF:12031 |
| MK_59 | Sterile Mycelium (Ascomycete) 2 | KP714651 | JMRC:SF:12062 |
| MK_65 | Sterile Mycelium (Ascomycete) 3 | KP714655 | JMRC:SF:12067 |
| MK_66 | Sterile Mycelium (Ascomycete) 4 | KP714656 | JMRC:SF:12068 |
| MK_73 | Sterile Mycelium (Ascomycete) 5 | KP714663 | JMRC:SF:12075 |
| MK_74 | Sterile Mycelium (Ascomycete) 6 | KP714664 | JMRC:SF:12076 |
| MK_76 | Sterile Mycelium (Ascomycete) 7 | KP714666 | JMRC:SF:12078 |
| MK_48 | Sterile Mycelium (Basidiomycete) 1 | KP714641 | JMRC:SF:12051 |
| MK_67 | Sterile Mycelium (Basidiomycete) 2 | KP714657 | JMRC:SF:12069 |
| MK_68 | Sterile Mycelium (Basidiomycete) 3 | KP714658 | JMRC:SF:12070 |
| MK_03 | Tetracladium sp. 1 | KP714596 | JMRC:SF:12020 |
| MK_17 | Tetracladium sp. 2 | KP714610 | JMRC:SF:12029 |
| MK_71 | Tetracladium sp. 3 | KP714661 | JMRC:SF:12073 |
| MK_58 | Thelebolus sp. | KP714650 | JMRC:SF:12061 |
| GW_22 | Trichoderma sp. 1 | KP714551 | JMRC:SF:11982 |
| GW_62 | Trichoderma sp. 2 | KP714590 | JMRC:SF:11996 |
| GW_38 | Trichoderma sp. 3 | KP714566 | JMRC:SF:12015 |
| MK_12 | Truncatella angustata | KP714605 | JMRC:SF:12025 |
| GW_11 | Umbelopsis sp. 1 | KP714540 | JMRC:SF:11971 |
| GW_21 | Umbelopsis sp. 2 | KP714550 | JMRC:SF:11981 |
| GW_28 | Umbelopsis sp. 3 | KP714557 | JMRC:SF:11987 |
| GW_30 | Umbelopsis sp. 4 | KP714559 | JMRC:SF:11989 |
| GW_60 | Umbelopsis sp. 5 | KP714588 | JMRC:SF:12013 |
| GW_61 | Umbelopsis sp. 6 | KP714589 | JMRC:SF:12014 |
References
- Schadt, C.W.; Martin, A.P.; Lipson, D.A.; Schmidt, S.K. Seasonal dynamics of previously unknown fungal lineages in tundra soils. Science 2003, 301, 1359–1361. [Google Scholar] [CrossRef] [PubMed]
- Ganley, R.J.; Brunsfeld, S.J.; Newcombe, G. A community of unknown, endophytic fungi in western white pine. Proc. Natl. Acad. Sci. USA 2004, 101, 10107–10112. [Google Scholar] [CrossRef] [PubMed]
- Abarenkov, K.; Henrik Nilsson, R.; Larsson, K.-H.; Alexander, I.J.; Eberhardt, U.; Erland, S.; Høiland, K.; Kjøller, R.; Larsson, E.; Pennanen, T.; et al. The UNITE database for molecular identification of fungi—Recent updates and future perspectives. New Phytol. 2010, 186, 281–285. [Google Scholar] [CrossRef] [PubMed]
- Tedersoo, L.; Bahram, M.; Põlme, S.; Kõljalg, U.; Yorou, N.S.; Wijesundera, R.; Ruiz, L.V.; Vasco-Palacios, A.M.; Thu, P.Q.; Suija, A.; et al. Global diversity and geography of soil fungi. Science 2014, 346. [Google Scholar] [CrossRef] [PubMed]
- El-Sayed, N.M.; Younis, M.S.; Elhamshary, A.M.; Abd-Elmaboud, A.I.; Kishik, S.M. Acanthamoeba DNA can be directly amplified from corneal scrapings. Parasitol. Res. 2014, 113, 3267–3272. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Zhai, H.; Yuan, Q.; Sun, S.; Liu, T.; Xie, L. Rapid and sensitive diagnosis of fungal keratitis with direct PCR without template DNA extraction. Clin. Microbiol. Infect. 2014, 20, O776–O782. [Google Scholar] [CrossRef] [PubMed]
- Wong, W.H.; Tay, Y.C.; Puniamoorthy, J.; Balke, M.; Cranston, P.S.; Meier, R. “Direct PCR” optimization yields a rapid, cost-effective, nondestructive and efficient method for obtaining DNA barcodes without DNA extraction. Mol. Ecol. Resour. 2014, 14, 1271–1280. [Google Scholar] [CrossRef] [PubMed]
- Calmin, G.; Belbahri, L.; Lefort, F. Direct PCR for DNA barcoding in the genera Phytophopthora and Pythium. Biotechnol. Biotechnol. Equip. 2007, 21, 40–42. [Google Scholar] [CrossRef]
- Urayama, S.-I.; Katoh, Y.; Fukuhara, T.; Arie, T.; Moriyama, H.; Teraoka, T. Rapid detection of Magnaporthe oryzae chrysovirus 1-A from fungal colonies on agar plates and lesions of rice blast. J. Gen. Plant Pathol. 2015, 81, 97–102. [Google Scholar] [CrossRef]
- Kuusisto, P.; Chum, P.Y. Thermo Scientific Phire Plant Direct PCR Kit: Plant genotyping and transgene detection without DNA purification. In Application Note; Thermo Fisher Scientific: Vantaa, Finland, 2013. [Google Scholar]
- Dudaite, N.; Navickaite, M.; Dinarina, A. Direct PCR from yeast cells. In Application Note; Thermo Fisher Scientific: Vantaa, Finland, 2015. [Google Scholar]
- Lin, X.; Yang, F.; Zhou, Y.; Zhu, Z.; Jin, G.; Zhang, S.; Zhao, Z.K. Highly-efficient colony PCR method for red yeasts and its application to identify mutations within two leucine auxotroph mutants. Yeast 2012, 29, 467–474. [Google Scholar] [CrossRef] [PubMed]
- Luo, G.; Mitchell, T.G. Rapid identification of pathogenic fungi directly from cultures by using multiplex PCR. Clin. Microbiol. 2002, 40, 2860–2865. [Google Scholar] [CrossRef]
- Iotti, M.; Zambonelli, A. A quick and precise technique for identifying ectomycorrhizas by PCR. Mycol. Res. 2006, 110, 60–65. [Google Scholar] [CrossRef] [PubMed]
- Bonito, G. Fast DNA-based identification of the black truffle Tuber melanosporum with direct PCR and species-specific primers. FEMS Microbiol. Lett. 2009, 301, 171–175. [Google Scholar] [CrossRef] [PubMed]
- Pancher, M.; Ceol, M.; Corneo, P.E.; Longa, C.M.O.; Yousaf, S.; Pertot, I.; Campisano, A. Fungal endophytic communities in grapevines (Vitis vinifera L.) respond to crop management. Appl. Environ. Microbiol. 2012, 78, 4308–4317. [Google Scholar] [CrossRef] [PubMed]
- Rainer, G.; Kuhnert, R.; Unterholzer, M.; Dresch, P.; Gruber, A.; Peintner, U. Host-specialist dominated ectomycorrhizal communities of Pinus cembra are not affected by temperature manipulation. J. Fungi 2015, 1, 55–75. [Google Scholar] [CrossRef]
- Knapp, M. Fungal Communities in snow-covered alpine soil: A comparison between molecular and cultivation-based techniques. Master’s Thesis, University of Innsbruck, Innsbruck, Austria, 2015. [Google Scholar]
- Kuhnert, R.; Oberkofler, I.; Peintner, U. Fungal growth and biomass development is boosted by plants in snow-covered soil. Microbial. Ecol. 2012, 64, 79–90. [Google Scholar] [CrossRef] [PubMed]
- Dresch, P.; D’Aguanno, M.; Rosam, K.; Grienke, U.; Rollinger, J.; Peintner, U. Fungal strain matters: Colony growth and bioactivity of the European medicinal polypores Fomes fomentarius, Fomitopsis pinicola and Piptoporus betulinus. AMB Express 2015, 5, 4. [Google Scholar] [CrossRef] [PubMed]
- Pitt, J.I. A Laboratory Guide to Common Penicillium Species; Commonwealth Scientific and Industrial Research Organization, Division of Food Research: North Ryde, NSW, Australia, 1985. [Google Scholar]
- Domsch, K.H.; Gams, W.; Anderson, T.H. Compendium of Soil Fungi; IHW-Verlag: Eching, Germany, 2007. [Google Scholar]
- Seifert, K.; Morgan-Jones, G.; Gams, W.; Kendrick, B. The Genera of Hyphomycetes; CBS-KNAW Fungal Biodiversity Centre: Utrecht, The Netherlands, 2011. [Google Scholar]
- Bensch, K.; Braun, U.; Groenewald, J.Z.; Crous, P.W. The genus Cladosporium. Stud. Mycol. 2012, 72, 1–401. [Google Scholar] [CrossRef] [PubMed]
- Gardes, M.; Bruns, T.D. ITS primers with enhanced specificity for basidiomycetes—Application to the identification of mycorrhizae and rusts. Mol. Ecol. 1993, 2, 113–118. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Michael, A.I., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press, Inc.: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Xu, C.; Dong, W.; Shi, S.; Cheng, T.; Li, C.; Liu, Y.; Wu, P.; Wu, H.; Gao, P.; Zhou, S. Accelerating plant DNA barcode reference library construction using herbarium specimens: Improved experimental techniques. Mol. Ecol. Resour. 2015, 15, 1366–1374. [Google Scholar] [CrossRef] [PubMed]
- Vazquez-Angulo, J.C.; Mendez-Trujillo, V.; Gonzalez-Mendoza, D.; Morales-Trejo, A.; Grimaldo-Juarez, O.; Cervantes-Diaz, L. A rapid and inexpensive method for isolation of total DNA from Trichoderma spp. (Hypocreaceae). Genet. Mol. Res. 2012, 11, 1379–1384. [Google Scholar] [CrossRef] [PubMed]
- Bokulich, N.A.; Mills, D.A. Improved selection of internal transcribed spacer-specific primers enables quantitative, ultra-high-throughput profiling of fungal communities. Appl. Environ. Microb. 2013, 79, 2519–2526. [Google Scholar] [CrossRef] [PubMed]
- Bellemain, E.; Carlsen, T.; Brochmann, C.; Coissac, E.; Taberlet, P.; Kauserud, H. ITS as an environmental DNA barcode for fungi: An in silico approach reveals potential PCR biases. BMC Microbiol. 2010, 10, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Wang, X.-R.; Blomquist, G. Evaluation of PCR primers and PCR conditions for specific detection of common airborne fungi. J. Environ. Monit. 2002, 4, 377–382. [Google Scholar] [CrossRef] [PubMed]
- Nikolcheva, L.G.; Bärlocher, F. Taxon-specific fungal primers reveal unexpectedly high diversity during leaf decomposition in a stream. Mycol. Prog. 2004, 3, 41–49. [Google Scholar] [CrossRef]
- Hagn, A.; Wallisch, S.; Radl, V.; Charles Munch, J.; Schloter, M. A new cultivation independent approach to detect and monitor common Trichoderma species in soils. J. Microbiol. Meth. 2007, 69, 86–92. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-N.; Liu, X.-Y.; Zheng, R.-Y. Umbelopsis changbaiensis sp. nov. from China and the typification of Mortierella vinacea. Mycol. Prog. 2014, 13, 657–669. [Google Scholar] [CrossRef]
- Voigt, K.; Cigelnik, E.; O’Donnell, K. Phylogeny and PCR identification of clinically important zygomycetes based on nuclear ribosomal-DNA sequence data. J. Clin. Microbiol. 1999, 37, 3957–3964. [Google Scholar] [PubMed]
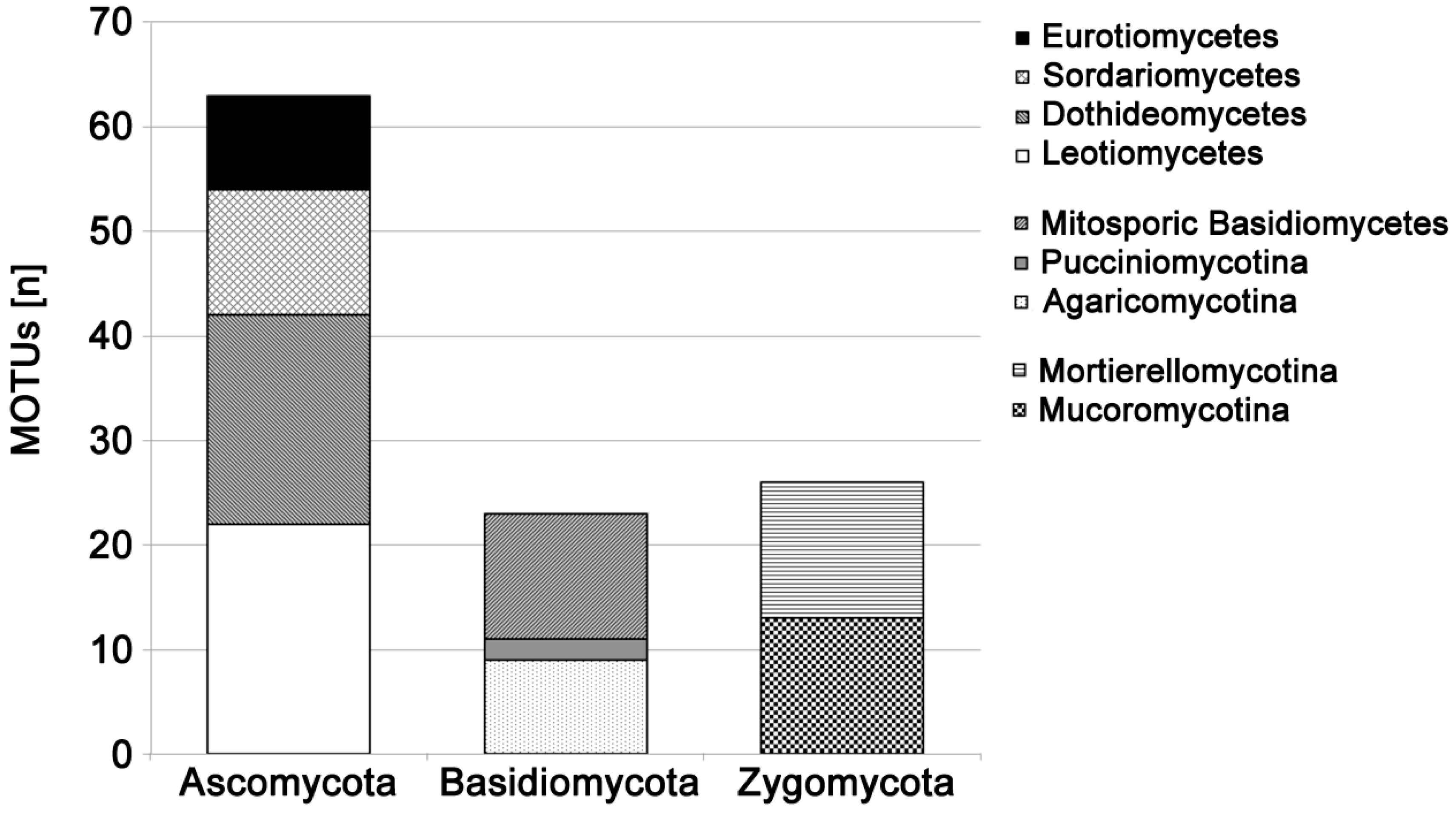
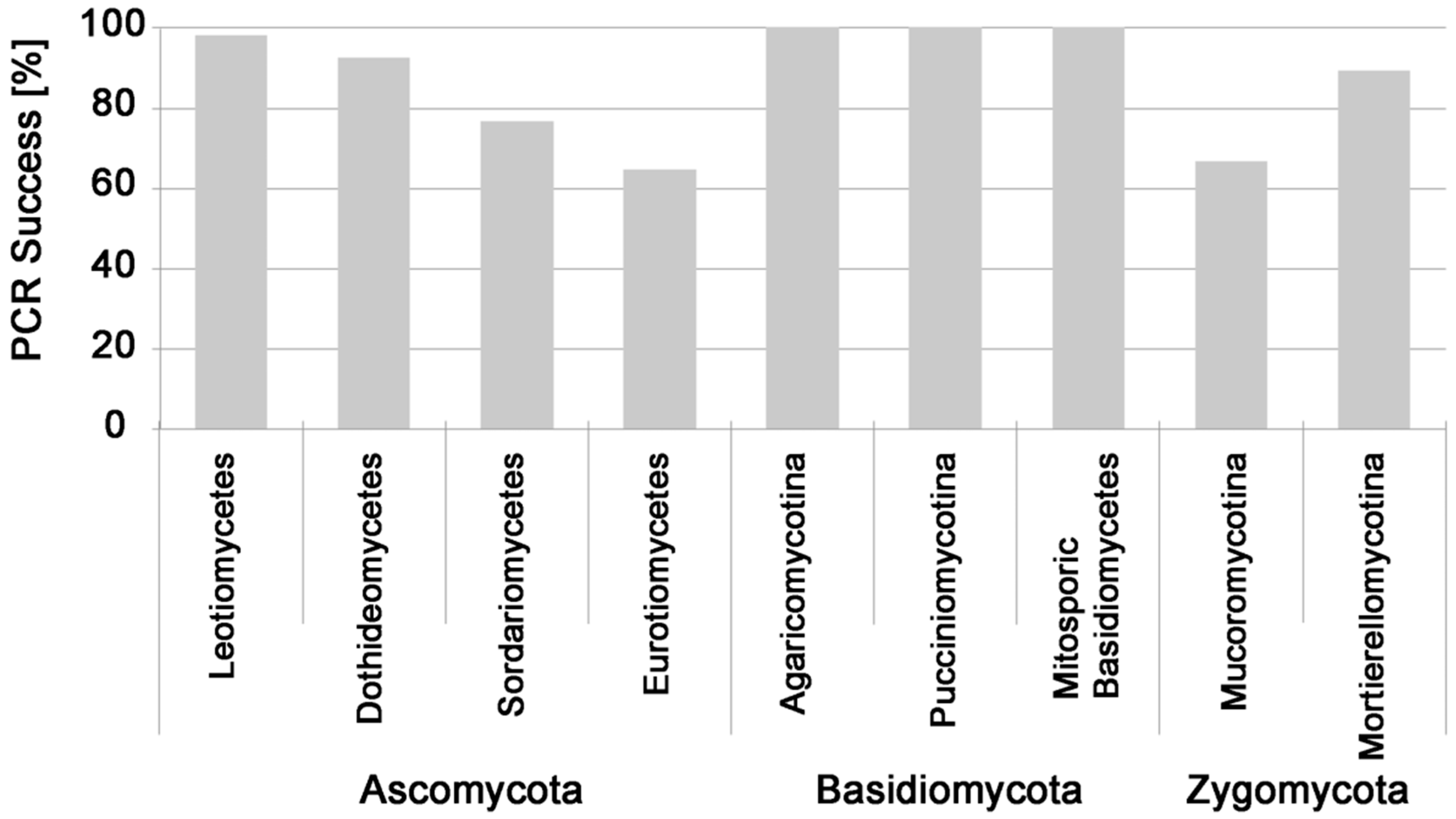
| Genus/Name | Taxonomic Affiliation | MOTUs | Tested Isolates | PCR-Success (%) |
|---|---|---|---|---|
| Absidia | Mucoromycotina | 0 | 7 | 0 |
| Aureobasidium | Dothideomycetes | 1 | 2 | 100 |
| Bjerkandera | Agaricomycotina | 1 | 1 | 100 |
| Botrytis/Sclerotinia | Leotiomycetes | 1 | 2 | 100 |
| Cadophora | Leotiomycetes | 1 | 1 | 100 |
| Chaetosphaeronema | Dothideomycetes | 1 | 1 | 100 |
| Cladosporium/Davidiella | Dothideomycetes | 5 | 63 | 89 |
| Cryptococcus | Mitosporic Basidiomycetes | 10 | 67 | 100 |
| Cystodendron | Leotiomycetes | 1 | 1 | 100 |
| Cystofilobasidium | Agaricomycotina | 1 | 1 | 100 |
| Didymella | Dothideomycetes | 1 | 6 | 100 |
| Dioszegia | Agaricomycotina | 2 | 3 | 100 |
| Drechslera | Dothideomycetes | 1 | 1 | 100 |
| Epicoccum | Dothideomycetes | 1 | 1 | 100 |
| Fusarium/Gibberella | Sordariomycetes | 1 | 33 | 85 |
| Geomyces/Pseudogymnoascus | Leotiomycetes | 8 | 147 | 99 |
| Guehomyces | Agaricomycotina | 1 | 2 | 100 |
| Helgardia | Leotiomycetes | 1 | 1 | 100 |
| Helotiales unknown | Leotiomycetes | 2 | 3 | 100 |
| Herpotrichia | Dothideomycetes | 4 | 5 | 100 |
| Holtermaniella | Agaricomycotina | 1 | 1 | 100 |
| Ilyonectria | Sordariomycetes | 1 | 1 | 100 |
| Leptodontidium | Leotiomycetes | 1 | 1 | 100 |
| Leuconeurospora | Leotiomycetes | 1 | 2 | 100 |
| Leucosporidiella/-ium | Pucciniomycotina | 2 | 5 | 100 |
| Monodictys | Sordariomycetes | 1 | 1 | 100 |
| Monographella/Microdochium | Sordariomycetes | 3 | 17 | 94 |
| Mortierella | Mortierellomycotina | 13 | 112 | 89 |
| Mrakia | Agaricomycotina | 2 | 3 | 100 |
| Mrakiella | Agaricomycotina | 1 | 2 | 100 |
| Mucor | Mucoromycotina | 6 | 48 | 58 |
| Neonectria | Sordariomycetes | 1 | 1 | 100 |
| Paraconiothyrium | Dothideomycetes | 1 | 1 | 100 |
| Penicillium | Eurotiomycetes | 9 | 105 | 65 |
| Phacidium | Leotiomycetes | 2 | 45 | 96 |
| Phaeosphaeria | Dothideomycetes | 1 | 2 | 100 |
| Phoma | Dothideomycetes | 2 | 9 | 100 |
| Rhodotorula | Mitosporic Basidiomycetes | 1 | 6 | 100 |
| Seimatosporium | Sordariomycetes | 1 | 2 | 100 |
| Stagonosporopsis | Dothideomycetes | 1 | 2 | 100 |
| Sydowia | Dothideomycetes | 1 | 2 | 100 |
| Tetracladium | Leotiomycetes | 3 | 3 | 100 |
| Thelebolus | Leotiomycetes | 1 | 1 | 100 |
| Trichoderma/Hypocrea | Sordariomycetes | 2 | 22 | 36 |
| Trichosporon | Mitosporic Basidiomycetes | 1 | 2 | 100 |
| Truncatella | Sordariomycetes | 2 | 9 | 100 |
| Umbelopsis | Mucoromycotina | 7 | 32 | 94 |
| Unknown sterile mycelia | Unknown | 3 | 5 | 60 |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Walch, G.; Knapp, M.; Rainer, G.; Peintner, U. Colony-PCR Is a Rapid Method for DNA Amplification of Hyphomycetes. J. Fungi 2016, 2, 12. https://doi.org/10.3390/jof2020012
Walch G, Knapp M, Rainer G, Peintner U. Colony-PCR Is a Rapid Method for DNA Amplification of Hyphomycetes. Journal of Fungi. 2016; 2(2):12. https://doi.org/10.3390/jof2020012
Chicago/Turabian StyleWalch, Georg, Maria Knapp, Georg Rainer, and Ursula Peintner. 2016. "Colony-PCR Is a Rapid Method for DNA Amplification of Hyphomycetes" Journal of Fungi 2, no. 2: 12. https://doi.org/10.3390/jof2020012
APA StyleWalch, G., Knapp, M., Rainer, G., & Peintner, U. (2016). Colony-PCR Is a Rapid Method for DNA Amplification of Hyphomycetes. Journal of Fungi, 2(2), 12. https://doi.org/10.3390/jof2020012






