A Network of Sporogenesis-Responsive Genes Regulates the Growth, Asexual Sporogenesis, Pathogenesis and Fusaric Acid Production of Fusarium oxysporum f. sp. cubense
Abstract
:1. Introduction
2. Materials and Methods
2.1. Fungal Strain and Media
2.2. Gene Identification
2.3. Mutant Generation
2.4. Vegetative Growth and Hydrophobicity Assays
2.5. In Vitro and in Planta Sporulation Assay
2.6. Microscopy
2.7. Quantitative Real-Time PCR (qRT-PCR)
2.8. Yeast One-Hybrid (Y1H) Assay
2.9. Chromatin Immunoprecipitation (ChIP)-qPCR Assay
2.10. Pathogenicity Assays in Banana Plants
2.11. Cellophane Membrane Penetration Assays
2.12. Fusaric Acid Production Assays
2.13. Statistical Analyses
3. Results
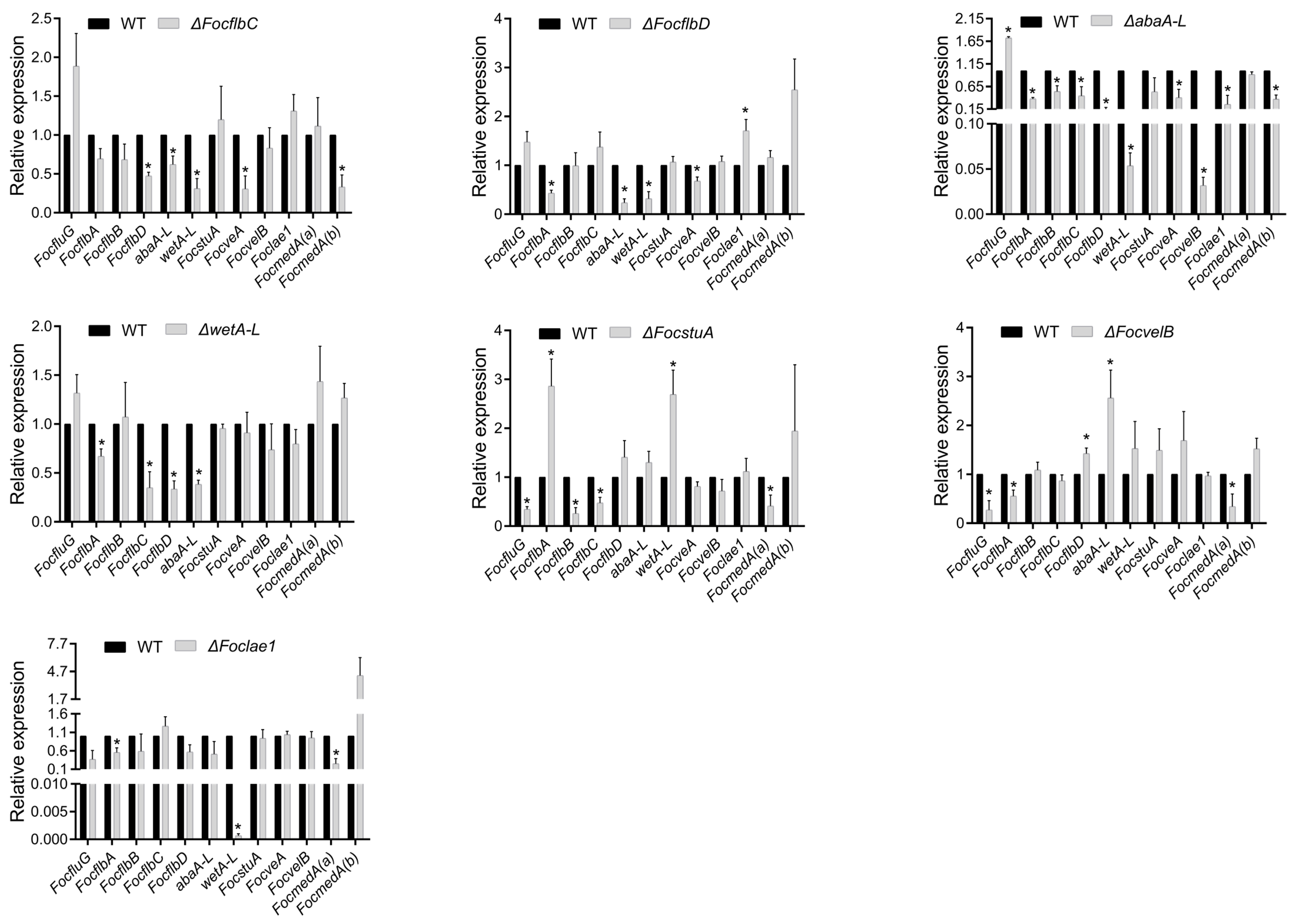
3.1. Putative Sporulation-Responsive Gene Identification and Expression
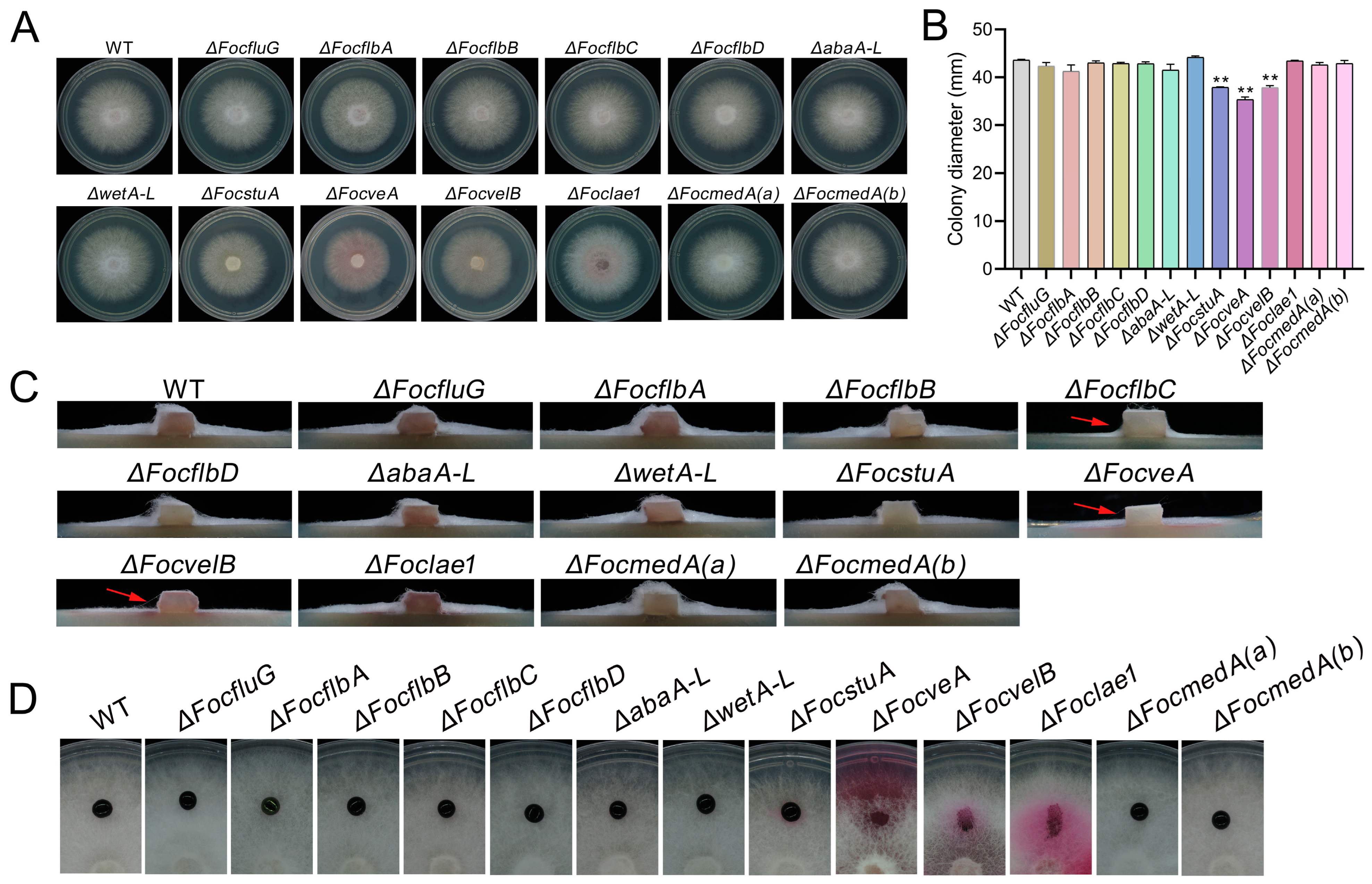
3.2. FocstuA, FocveA and FocvelB Are Required for Vegetative Growth in Foc
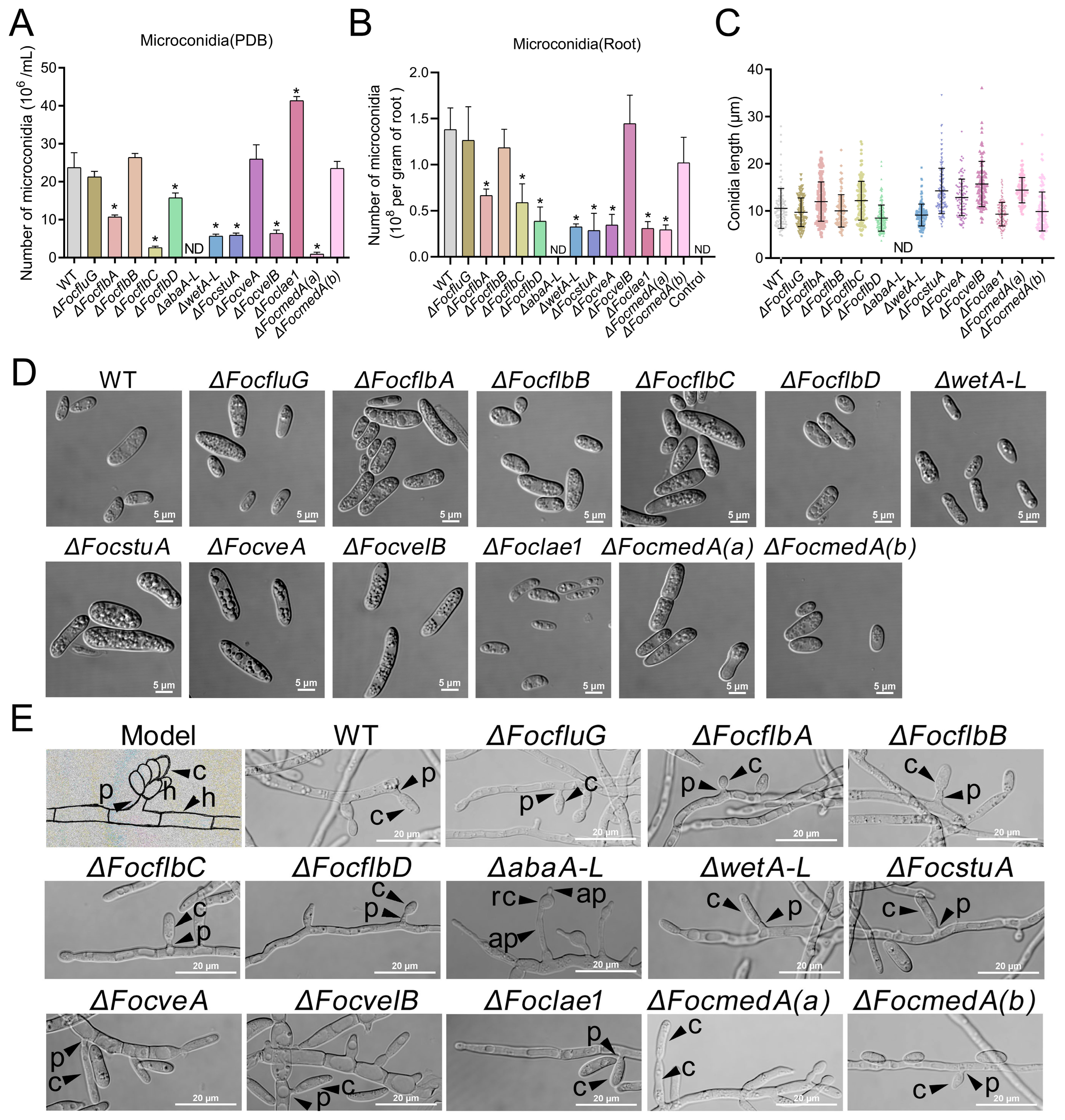
3.3. abaA-L Is Essential for Microconidia and Phialides Production in Foc
3.4. FocflbA, FocflbD, abaA-L, wetA-L, FocstuA, FocveA, Foclae1 and FocmedA(a) Are Required for Macroconidia Production and Conidiophores Formation in Foc
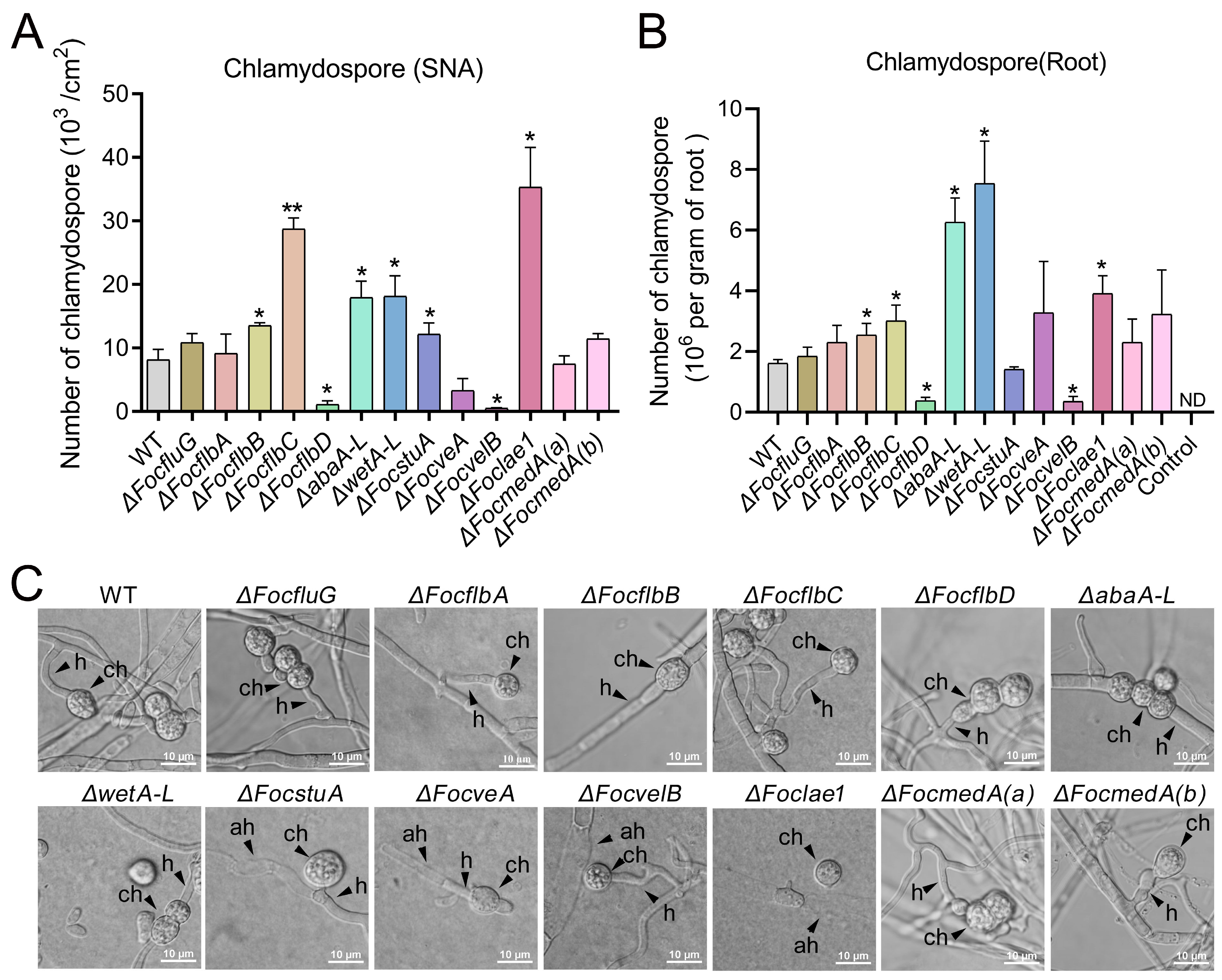
3.5. FocflbB, FocflbC, FocflbD, abaA-L, wetA-L, FocvelB and Foclae1 Control Chlamydosporulation In Vitro and in Planta in Foc
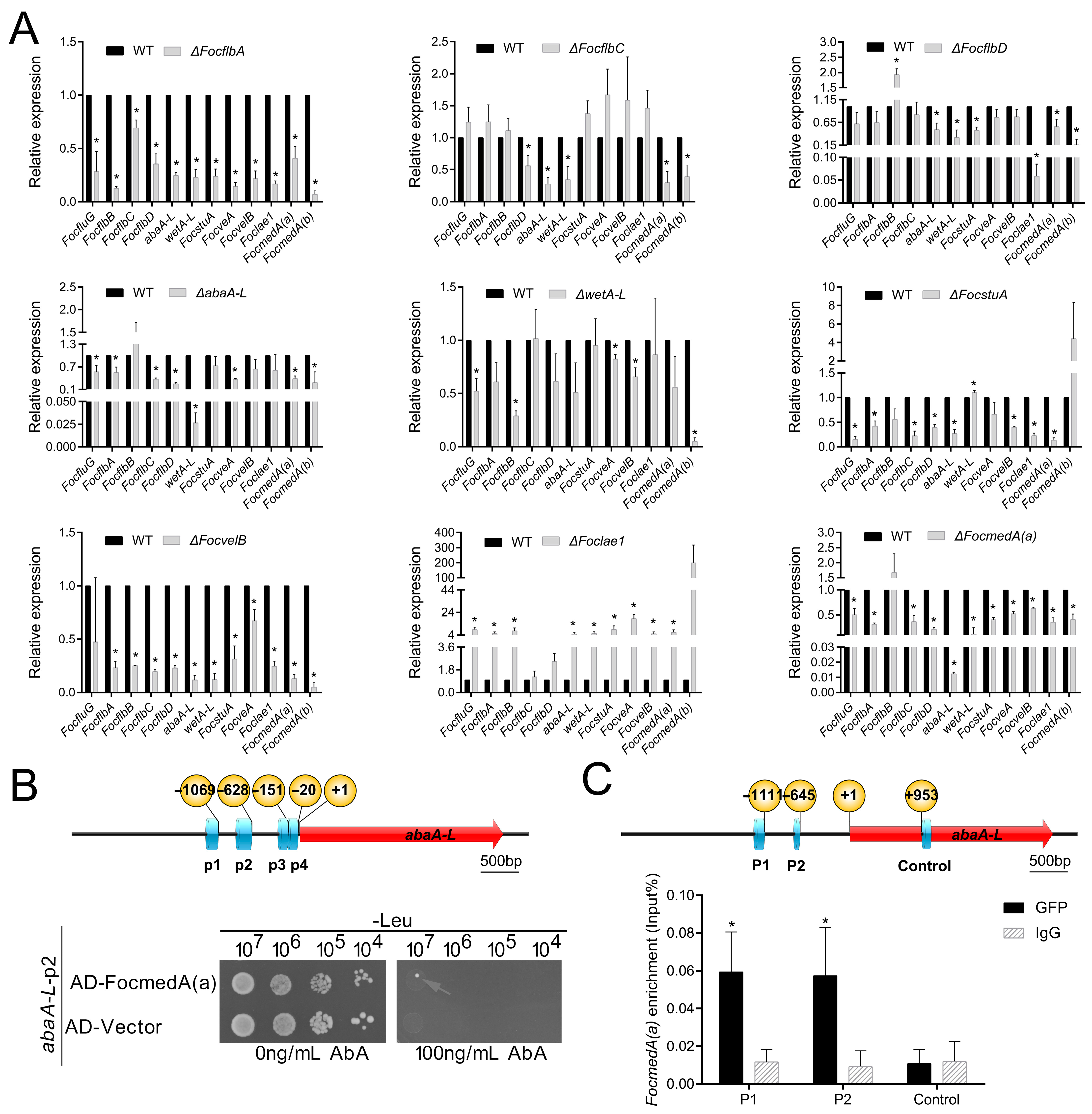
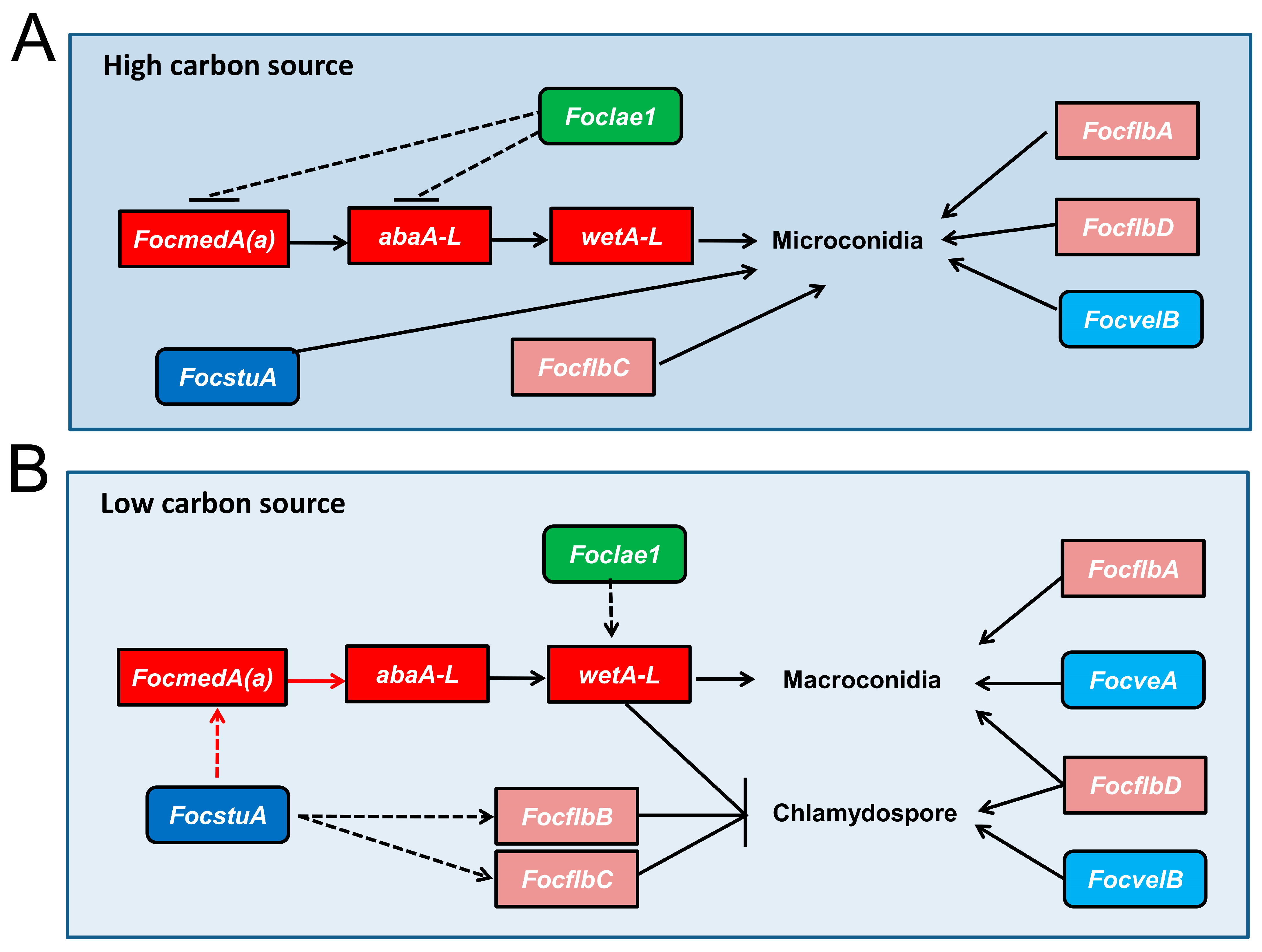
3.6. Functional Relationship of the Sporulation-Responsive Genes Regulating Microconidia Production in Foc
3.7. Functional Relationship of the Sporulation-Responsive Genes Regulating Macroconidia and Chlamydospores Production in Foc
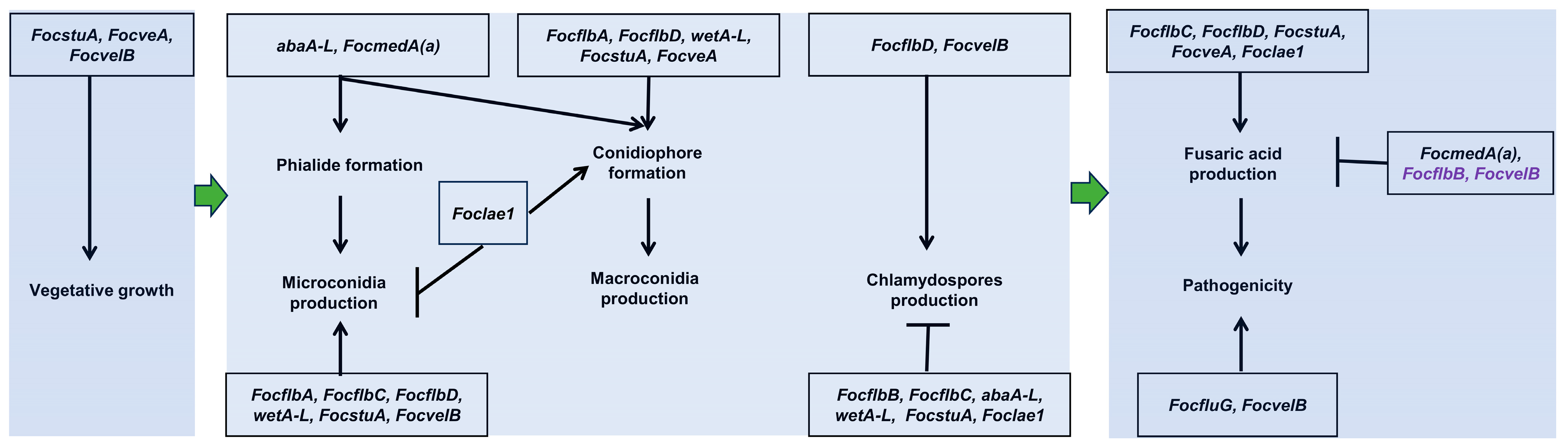
3.8. FocfluG, FocflbA, FocflbC, FocflbD, FocstuA, FocveA, FocvelB and Foclae1 Are Required for the Pathogenicity of Foc
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- EFSA Panel on Plant Health (PLH); Bragard, C.; Baptista, P.; Chatzivassiliou, E.K.; Di Serio, F.; Gonthier, P.; Jaques Miret, J.A.; Justesen, A.F.; MacLeod, A.; Magnusson, C.S.; et al. Pest categorisation of Fusarium oxysporum f. sp. cubense Tropical Race 4. EFSA J. 2022, 20, e07092. [Google Scholar] [PubMed]
- Bubici, G.; Kaushal, M.; Prigigallo, M.I.; Gómez-Lama Cabanás, C.; Mercado-Blanco, J. Biological control agents against Fusarium wilt of banana. Front. Microbiol. 2019, 10, 616. [Google Scholar] [CrossRef]
- Waite, B.H.; Stover, R.H. Studies on Fusarium wilt of bananas. VI. Variability and the cultivar concept in Fusarium oxysporum f. cubense. Can. J. Bot. 1960, 38, 985–994. [Google Scholar] [CrossRef]
- Stover, R.H.; Waite, B.H. Studies on Fusarium wilt of bananas. V. Pathogenicity and distribution of F. oxysporum f. cubense races 1 and 2. Can. J. Bot. 1960, 38, 51–61. [Google Scholar] [CrossRef]
- Moore, N.Y.; Pegg, K.G.; Allen, R.N.; Irwin, J.A.G. Vegetative compatibility and distribution of Fusarium oxysporum f.sp. cubense in Australia. Aust. J. Exp. Arg. 1993, 33, 797–802. [Google Scholar] [CrossRef]
- Pegg, K.G.; Moore, N.Y.; Bentley, S. Fusarium wilt of banana in Australia: A review. Aust. J. Agric. Res. 1996, 47, 637–650. [Google Scholar] [CrossRef]
- Ploetz, R.C.; Freeman, S.; Konkol, J.L.; Al-Abed, A.; Naser, Z.; Shalan, K.; Barakat, R.M.; Israeli, Y. Tropical race 4 of Panama disease in the Middle East. Phytoparasitica 2015, 43, 283–293. [Google Scholar] [CrossRef]
- Butler, D. Fungus threatens top banana. Nature 2013, 504, 195–196. [Google Scholar] [CrossRef]
- Ploetz, R.C. Fusarium wilt of banana is caused by several pathogens referred to as Fusarium oxysporum f. sp. cubense. Phytopathology 2006, 96, 653–656. [Google Scholar] [CrossRef]
- Ma, L.J.; Geiser, D.M.; Proctor, R.H.; Rooney, A.P.; O’Donnell, K.; Trail, F.; Gardiner, D.M.; Manners, J.M.; Kazan, K. Fusarium pathogenomics. Annu. Rev. Microbiol. 2013, 67, 399–416. [Google Scholar] [CrossRef]
- Maryani, N.; Lombard, L.; Poerba, Y.S.; Subandiyah, S.; Crous, P.W.; Kema, G.H.J. Phylogeny and genetic diversity of the banana Fusarium wilt pathogen Fusarium oxysporum f. sp. cubense in the Indonesian centre of origin. Stud. Mycol. 2019, 92, 155–194. [Google Scholar] [CrossRef] [PubMed]
- Ohara, T.; Inoue, I.; Namiki, F.; Kunoh, H.; Tsuge, T. REN1 is required for development of microconidia and macroconidia, but not of chlamydospores, in the plant pathogenic fungus Fusarium oxysporum. Genetics 2004, 166, 113–124. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Chen, S.; Zuo, C.; Sun, Q.; Ye, Q.; Yi, G.; Huang, B. The use of GFP-transformed isolates to study infection of banana with Fusarium oxysporum f. sp. cubense race 4. Eur. J. Plant Pathol. 2011, 131, 327–340. [Google Scholar] [CrossRef]
- Rowe, R.C.; Farely, J.D.; Coplin, D.C. Airborne spore dispersal and recolonization of steamed soil by Fusarium oxysporum in tomato greenhouses. Phytopathology 1977, 67, 1513–1517. [Google Scholar] [CrossRef]
- Couteaudier, Y.; Alabouvette, C. Survival and inoculum potential of conidia and chlamydospores of Fusarium oxysporum f.sp. lini in soil. Can. J. Microbiol. 1990, 36, 551–556. [Google Scholar] [CrossRef]
- Katan, T.; Shlevin, E.; Katan, J. Sporulation of Fusarium oxysporum f. sp. lycopersici on stem surfaces of tomato plants and aerial dissemination of inoculum. Phytopathology 1997, 87, 712–719. [Google Scholar] [CrossRef] [PubMed]
- Rekah, Y.; Shtienberg, D.; Katan, J. Disease development following infection of tomato and basil foliage by airborne conidia of the soilborne pathogens Fusarium oxysporum f. sp. radicis-lycopersici and F. oxysporum f. sp. basilici. Phytopathology 2000, 90, 1322–1329. [Google Scholar] [CrossRef]
- Park, H.S.; Yu, J.H. Genetic control of asexual sporulation in filamentous fungi. Curr. Opin. Microbiol. 2012, 15, 669–677. [Google Scholar] [CrossRef]
- Krijgsheld, P.; Bleichrodt, R.J.; van Veluw, G.J.; Wang, F.W.; Müller, W.H.; Dijksterhuis, J.; Wösten, H.A.B. Development in Aspergillus. Stud. Mycol. 2012, 74, 1–29. [Google Scholar] [CrossRef]
- Etxebeste, O.; Garzia, A.; Espeso, E.A.; Ugalde, U.O. Aspergillus nidulans asexual development: Making the most of cellular modules. Trends Microbiol. 2010, 18, 569–576. [Google Scholar] [CrossRef]
- Adams, T.H.; Wieser, J.K.; Yu, J.H. Asexual sporulation in Aspergillus nidulans. Microbiol. Mol. Biol. Rev. 1998, 62, 35–54. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.H. Regulation of development in Aspergillus nidulans and Aspergillus fumigatus. Mycobiology 2010, 38, 229–237. [Google Scholar] [CrossRef] [PubMed]
- Park, H.S.; Yu, J.H. Developmental regulators in Aspergillus fumigatus. J. Microbiol. 2016, 54, 223–231. [Google Scholar] [CrossRef] [PubMed]
- Etxebeste, O.; Ni, M.; Garzia, A.; Kwon, N.-J.; Fischer, R.; Yu, J.H.; Espeso, E.A.; Ugalde, U.O. Basic-zipper-type transcription factor FlbB controls asexual development in Aspergillus nidulans. Eukaryot. Cell 2008, 7, 38–48. [Google Scholar] [CrossRef]
- Mah, J.H.; Yu, J.H. Upstream and downstream regulation of asexual development in Aspergillus fumigatus. Eukaryot. Cell 2006, 5, 1585–1595. [Google Scholar] [CrossRef] [PubMed]
- Mirabito, P.M.; Adams, T.H.; Timberlake, W.E. Interactions of three sequentially expressed genes control temporal and spatial specificity in aspergillus development. Cell 1989, 57, 859–868. [Google Scholar] [CrossRef] [PubMed]
- Ojeda-López, M.; Chen, W.X.; Eagle, C.E.; Gutiérrez, G.; Jia, W.; Swilaiman, S.S.; Huang, Z.; Park, H.S.; Yu, J.-H.; Cánovas, D.; et al. Evolution of asexual and sexual reproduction in the Aspergilli. Stud. Mycol. 2018, 91, 37–59. [Google Scholar] [CrossRef]
- Tao, L.; Yu, J.H. AbaA and WetA govern distinct stages of Aspergillus fumigatus development. Microbiology 2011, 157, 313–326. [Google Scholar] [CrossRef]
- Yu, J.H.; Mah, J.H.; Seo, J.A. Growth and developmental control in the model and pathogenic Aspergilli. Eukaryot. Cell 2006, 5, 1577–1584. [Google Scholar] [CrossRef]
- Adams, T.H.; Boylan, M.T.; Timberlake, W.E. brlA is necessary and sufficient to direct conidiophore development in Aspergillus nidulans. Cell 1988, 54, 353–362. [Google Scholar] [CrossRef]
- Wang, M.; Sun, X.; Zhu, C.; Xu, Q.; Ruan, R.; Yu, D.; Li, H. PdbrlA, PdabaA and PdwetA control distinct stages of conidiogenesis in Penicillium digitatum. Res. Microbiol. 2015, 166, 56–65. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.A.; Guan, Y.; Yu, J.H. FluG-dependent asexual development in Aspergillus nidulans occurs via derepression. Genetics 2006, 172, 1535–1544. [Google Scholar] [CrossRef] [PubMed]
- Ni, M.; Yu, J.-H. A novel regulator couples sporogenesis and trehalose biogenesis in Aspergillus nidulans. PLoS ONE 2007, 2, e970. [Google Scholar] [CrossRef] [PubMed]
- Bayram, O.; Braus, G.H. Coordination of secondary metabolism and development in fungi: The velvet family of regulatory proteins. FEMS Microbiol. Rev. 2012, 36, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Hoff, B.; Kamerewerd, J.; Sigl, C.; Mitterbauer, R.; Zadra, I.; Kürnsteiner, H.; Kück, U. Two components of a velvet-like complex control hyphal morphogenesis, conidiophore development, and penicillin biosynthesis in Penicillium chrysogenum. Eukaryot. Cell 2010, 9, 1236–1250. [Google Scholar] [CrossRef] [PubMed]
- Wiemann, P.; Brown, D.W.; Kleigrewe, K.; Bok, J.W.; Keller, N.P.; Humpf, H.U.; Tudzynski, B. FfVel1 and FfLae1, components of a velvet-like complex in Fusarium fujikuroi, affect differentiation, secondary metabolism and virulence. Mol. Microbiol. 2010, 77, 972–994. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.L.; Oide, S.; Zhang, N.; Choi, M.Y.; Turgeon, B.G. ChLae1 and ChVel1 regulate T-toxin production, virulence, oxidative stress response, and development of the maize pathogen Cochliobolus heterostrophus. PLoS Pathog. 2012, 8, e1002542. [Google Scholar] [CrossRef]
- Sarikaya Bayram, Ö.; Bayram, Ö.; Valerius, O.; Park, H.S.; Irniger, S.; Gerke, J.; Ni, M.; Han, K.H.; Yu, J.H.; Braus, G.H. LaeA control of velvet family regulatory proteins for light-dependent development and fungal cell-type specificity. PLoS Genet. 2010, 6, e1001226. [Google Scholar] [CrossRef]
- Bayram, O.; Krappmann, S.; Ni, M.; Bok, J.W.; Helmstaedt, K.; Valerius, O.; Braus-Stromeyer, S.A.; Kwon, N.J.; Keller, N.P.; Yu, J.H.; et al. VelB/VeA/LaeA complex coordinates light signal with fungal development and secondary metabolism. Science 2008, 320, 1504–1506. [Google Scholar] [CrossRef]
- Clutterbuck, A.J. A mutational analysis of conidial development in Aspergillus nidulans. Genetics 1969, 63, 317–327. [Google Scholar] [CrossRef]
- Busby, T.M.; Miller, K.Y.; Miller, B.L. Suppression and enhancement of the Aspergillus nidulans medusa mutation by altered dosage of the bristle and stunted genes. Genetics 1996, 143, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Miller, K.Y.; Wu, J.; Miller, B.L. StuA is required for cell pattern formation in Aspergillus. Genes Dev. 1992, 6, 1770–1782. [Google Scholar] [CrossRef] [PubMed]
- Miller, K.Y.; Toennis, T.M.; Adams, T.H.; Miller, B.L. Isolation and transcriptional characterization of a morphological modifier: The Aspergillus nidulans stunted (stuA) gene. Mol. Gen. Genet. 1991, 227, 285–292. [Google Scholar] [CrossRef] [PubMed]
- Ohara, T.; Tsuge, T. FoSTUA, encoding a basic helix-loop-helix protein, differentially regulates development of three kinds of asexual spores, macroconidia, microconidia, and chlamydospores, in the fungal plant pathogen Fusarium oxysporum. Eukaryot. Cell 2004, 3, 1412–1422. [Google Scholar] [CrossRef] [PubMed]
- López-Berges, M.S.; Hera, C.; Sulyok, M.; Schäfer, K.; Capilla, J.; Guarro, J.; Di Pietro, A. The velvet complex governs mycotoxin production and virulence of Fusarium oxysporum on plant and mammalian hosts. Mol. Microbiol. 2013, 87, 49–65. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.-l.; Chen, W.; Yang, S.; Wen, Y.; Zheng, Y.; Anjago, W.M.; Yun, Y.; Wang, Z. Isolation and identification of Fusarium oxysporum f. sp. cubense in Fujian Province, China. J. Integr. Agr. 2019, 18, 1905–1913. [Google Scholar] [CrossRef]
- Yun, Y.; Song, A.; Bao, J.; Chen, S.; Lu, S.; Cheng, C.; Zheng, W.; Wang, Z.; Zhang, L. Genome data of Fusarium oxysporum f. sp. cubense race 1 and tropical race 4 isolates using long-read sequencing. Mol. Plant Microbe Interact. 2019, 32, 1270–1272. [Google Scholar] [CrossRef]
- Yang, S.; Zhou, X.; Guo, P.; Lin, Y.; Fan, Q.; Zuriegat, Q.; Lu, S.; Yang, J.; Yu, W.; Liu, H.; et al. The exocyst regulates hydrolytic enzyme secretion at hyphal tips and septa in the banana Fusarium wilt fungus Fusarium odoratissimum. Appl. Environ. Microbiol. 2021, 87, e0308820. [Google Scholar] [CrossRef]
- Yun, Y.; Liu, Z.; Yin, Y.; Jiang, J.; Chen, Y.; Xu, J.R.; Ma, Z. Functional analysis of the Fusarium graminearum phosphatome. N. Phytol. 2015, 207, 119–134. [Google Scholar] [CrossRef]
- Leslie, J.F.; Summerell, B.A. The Fusarium Laboratory Manual, 1st ed.; Blackwell Publishing: Oxford, UK, 2006. [Google Scholar]
- Dai, Y.; Cao, Z.; Huang, L.; Liu, S.; Shen, Z.; Wang, Y.; Wang, H.; Zhang, H.; Li, D.; Song, F. CCR4-Not complex subunit Not2 plays critical roles in vegetative growth, conidiation and virulence in watermelon Fusarium wilt pathogen Fusarium oxysporum f. sp. niveum. Front. Microbiol. 2016, 7, 1449. [Google Scholar] [CrossRef]
- Yun, Y.; Zhou, X.; Yang, S.; Wen, Y.; You, H.; Zheng, Y.; Norvienyeku, J.; Shim, W.B.; Wang, Z. Fusarium oxysporum f. sp. lycopersici C2H2 transcription factor FolCzf1 is required for conidiation, fusaric acid production, and early host infection. Curr. Genet. 2019, 65, 773–783. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.H.; Hamari, Z.; Han, K.H.; Seo, J.A.; Reyes-Domínguez, Y.; Scazzocchio, C. Double-joint PCR: A PCR-based molecular tool for gene manipulations in filamentous fungi. Fungal Genet. Biol. 2004, 41, 973–981. [Google Scholar] [CrossRef] [PubMed]
- Di Pietro, A.; Roncero, M.I.G. Cloning, expression, and role in pathogenicity of pg1 encoding the major extracellular endopolygalacturonase of the vascular wilt pathogen Fusarium oxysporum. Mol. Plant Microbe Interact. 1998, 11, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Jian, Y.; Chen, Y.; Kistler, H.C.; He, P.; Ma, Z.; Yin, Y. A phosphorylated transcription factor regulates sterol biosynthesis in Fusarium graminearum. Nat. Commun. 2019, 10, 1228. [Google Scholar] [CrossRef] [PubMed]
- Merhej, J.; Urbán, M.; Dufresne, M.; Hammond-Kosack, K.E.; Richard-Forget, F.; Barreau, C. The velvet gene, FgVe1, affects fungal development and positively regulates trichothecene biosynthesis and pathogenicity in Fusarium graminearum. Mol. Plant Pathol. 2012, 13, 363–374. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Kaufmann, K.B.; Muiño, J.M.; Østerås, M.; Farinelli, L.; Krajewski, P.; Angenent, G.C. Chromatin immunoprecipitation (ChIP) of plant transcription factors followed by sequencing (ChIP-SEQ) or hybridization to whole genome arrays (ChIP-CHIP). Nat. Protoc. 2010, 5, 457–472. [Google Scholar] [CrossRef]
- Jian, Y.; Liu, Z.; Wang, H.; Chen, Y.; Yin, Y.; Zhao, Y.; Ma, Z. Interplay of two transcription factors for recruitment of the chromatin remodeling complex modulates fungal nitrosative stress response. Nat. Commun. 2021, 12, 2576. [Google Scholar] [CrossRef]
- Orr, R.; Pattison, A.B.; East, D.J.; Warman, N.; O’Neill, W.T.; Czislowski, E.; Nelson, P.N. Image-based quantification of Fusarium wilt severity in banana. Australas. Plant Dis. 2019, 14, 14. [Google Scholar] [CrossRef]
- Gu, Q.; Zhang, C.; Liu, X.; Ma, Z. A transcription factor FgSte12 is required for pathogenicity in Fusarium graminearum. Mol. Plant Pathol. 2015, 16, 1–13. [Google Scholar] [CrossRef]
- Qin, H.; Yang, L.; Li, S.; Xie, Y.; Huang, J. Effect of nutrients in medium on growth of Fusarium oxysporum f.sp. cubense causing fusarial wilt on banana. Chin. J. Trop. Crops 2009, 30, 1852–1857. [Google Scholar]
- Nelson, P.E.; Toussoun, T.A.; Marasas, W.F.O. Fusarium Species: An Illustrated Manual for Identification, 1st ed.; Pennsylvania State University Press: University Park, PA, USA, 1983. [Google Scholar]
- Li, C.; Zuo, C.; Deng, G.; Kuang, R.B.; Yang, Q.; Hu, C.H.; Sheng, O.; Zhang, S.; Ma, L.J.; Wei, Y.R.; et al. Contamination of bananas with beauvericin and fusaric acid produced by Fusarium oxysporum f. sp. cubense. PLoS ONE 2013, 8, e70226. [Google Scholar] [CrossRef] [PubMed]
- Aguirre, J. Spatial and temporal controls of the Aspergillus brlA developmental regulatory gene. Mol. Microbiol. 1993, 8, 211–218. [Google Scholar] [CrossRef] [PubMed]
- Fan, G.; Zhang, K.; Zhang, J.; Yang, J.; Yang, X.; Hu, Y.; Huang, J.; Zhu, Y.; Yu, W.; Hu, H.; et al. The transcription factor FgMed1 is involved in early conidiogenesis and DON biosynthesis in the plant pathogenic fungus Fusarium graminearum. Appl. Microbiol. Biotechnol. 2019, 103, 5851–5865. [Google Scholar] [CrossRef]
- Gravelat, F.N.; Ejzykowicz, D.E.; Chiang, L.Y.; Chabot, J.; Urb, M.; Macdonald, K.; al-Bader, N.; Filler, S.G.; Sheppard, D.C. Aspergillus fumigatus MedA governs adherence, host cell interactions and virulence. Cell Microbiol. 2010, 12, 473–488. [Google Scholar] [CrossRef]
- Son, H.; Kim, M.G.; Chae, S.K.; Lee, Y.W. FgFlbD regulates hyphal differentiation required for sexual and asexual reproduction in the ascomycete fungus Fusarium graminearum. J. Microbiol. 2014, 52, 930–939. [Google Scholar] [CrossRef]
- Dutton, J.R.; Johns, S.; Miller, B.L. StuAp is a sequence-specific transcription factor that regulates developmental complexity in Aspergillus nidulans. EMBO J. 1997, 16, 5710–5721. [Google Scholar] [CrossRef]
- Martinelli, S.D. Phenotypes of double conidiation mutants of Aspergillus nidulans. J. Gen. Microbiol. 1979, 114, 277–287. [Google Scholar] [CrossRef]
- Lysøe, E.; Pasquali, M.; Breakspear, A.; Kistler, H.C. The transcription factor FgStuAp influences spore development, pathogenicity, and secondary metabolism in Fusarium graminearum. Mol. Plant Microbe Interact. 2011, 24, 54–67. [Google Scholar] [CrossRef]
- Sheppard, D.C.; Doedt, T.; Chiang, L.Y.; Kim, H.S.; Chen, D.; Nierman, W.C.; Filler, S.G. The Aspergillus fumigatus StuA protein governs the up-regulation of a discrete transcriptional program during the acquisition of developmental competence. Mol. Biol. Cell 2005, 16, 5866–5879. [Google Scholar] [CrossRef]
- Borneman, A.R.; Hynes, M.J.; Andrianopoulos, A. A basic helix–loop–helix protein with similarity to the fungal morphological regulators, Phd1p, Efg1p and StuA, controls conidiation but not dimorphic growth in Penicillium marneffei. Mol. Microbiol. 2002, 44, 621–631. [Google Scholar] [CrossRef] [PubMed]
- Van Eck, W.H.; Schippers, B. Ultrastructure of developing chlamydospores of Fusarium solani f. cucurbitae in vitro. Soil Biol. Biochem. 1976, 8, 1–6. [Google Scholar] [CrossRef]



| Aspergillus nidulans | Fusarium oxysporum f. sp. cubense | |||||||
|---|---|---|---|---|---|---|---|---|
| Gene | Protein Accession Number a | Domain b | Number of Homologs c | Homologs c | Max Score (e-Value) c | Query Coverage c | Percent Identity c | Function |
| fluG | P38094 | Glutamine synthetase, catalytic domain; amidohydrolase | 2 | FOIG_07155 | 612 (0.0) | 99% | 41% | Present work (FocfluG) |
| FOIG_11529 | 182 (6 × 10 −50) | 50% | 29% | Unknown | ||||
| flbA | P38093 | Regulator of G protein signaling domain; domain found in Dishevelled, Egl-10 and Pleckstrin (DEP) | 3 | FOIG_08170 | 607 (0.0) | 72% | 60% | Present work (FocflbA) |
| FOIG_16270 | 454 (2 × 10 −153) | 67% | 49% | Unknown | ||||
| FOIG_13410 | 309 (3 × 10 −112) | 65% | 44% | Unknown | ||||
| flbB | C8VBM8 | bZIP transcription factor | 1 | FOIG_00530 | 205 (2 × 10 −62) | 90% | 36% | Present work (FocflbB) |
| flbC | G5EAS8 | Zinc finger, C2H2 type | 1 | FOIG_03036 | 217 (5 × 10 −68) | 66% | 56% | Present work (FocflbC) |
| flbD | G5EAY5 | Myb-like DNA-binding domain | 1 | FOIG_06682 | 142 (9 × 10 −41) | 33% | 65% | Present work (FocflbD) |
| flbE | Q5BFF9 | No | 0 | - | - | - | - | - |
| brlA | P10069 | Zinc-finger double domain | 0 | - | - | - | - | - |
| abaA | P20945 | TEA/ATTS domain | 1 | FOIG_01396 | 67.4 (2 × 10 −11) | 10% | 41% | Present work (abaA-L) |
| wetA | P22022 | ESC1/WetA-related domain | 1 | FOIG_07289 | 61.6 (3 × 10 −10) | 10% | 68% | Present work (wetA-L) |
| stuA | P36011 | KilA-N domain | 1 | FOIG_07247 | 249 (7 × 10 −75) | 33% | 61% | Present work (FocstuA) |
| veA | C8VTV4 | Velvet factor | 1 | FOIG_00370 | 221 (1 × 10 −76) | 49% | 48% | Present work (FocveA) |
| velB | C8VTS4 | Velvet factor | 1 | FOIG_00471 | 170 (2 × 10 −89) | 83% | 50% | Present work (FocvelB) |
| velC | Q5BBM1 | Velvet factor | 0 | - | - | - | - | - |
| vosA | Q5BBX1 | Velvet factor | 1 | FOIG_05756 | 111 (8 × 10 −27) | 38% | 37% | Unknown |
| laeA | C8VQG9 | Methyltransferase domain | 6 | FOIG_01530 | 180 (3 × 10 −53) | 75% | 35% | Present work (Foclae1) |
| FOIG_10149 | 188 (8 × 10 −57) | 77% | 37% | Unknown | ||||
| FOIG_15419 | 191 (9 × 10 −58) | 81% | 36% | Unknown | ||||
| FOIG_16515 | 181 (4 × 10 −54) | 77% | 36% | Unknown | ||||
| FOIG_10023 | 181 (1 × 10 −53) | 76% | 33% | Unknown | ||||
| FOIG_11569 | 179 (2 × 10 −53) | 77% | 34% | Unknown | ||||
| medA | O74251 | No | 2 | FOIG_10811 | 292 (2 × 10 −91) | 72% | 41% | Present work (FocmedA(a)) |
| FOIG_16330 | 254 (2 × 10 −80) | 58% | 42% | Present work (FocmedA(b)) | ||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, S.; Deng, H.; Lin, Y.; Huang, M.; You, H.; Zhang, Y.; Zhuang, W.; Lu, G.; Yun, Y. A Network of Sporogenesis-Responsive Genes Regulates the Growth, Asexual Sporogenesis, Pathogenesis and Fusaric Acid Production of Fusarium oxysporum f. sp. cubense. J. Fungi 2024, 10, 1. https://doi.org/10.3390/jof10010001
Lu S, Deng H, Lin Y, Huang M, You H, Zhang Y, Zhuang W, Lu G, Yun Y. A Network of Sporogenesis-Responsive Genes Regulates the Growth, Asexual Sporogenesis, Pathogenesis and Fusaric Acid Production of Fusarium oxysporum f. sp. cubense. Journal of Fungi. 2024; 10(1):1. https://doi.org/10.3390/jof10010001
Chicago/Turabian StyleLu, Songmao, Huobing Deng, Yaqi Lin, Meimei Huang, Haixia You, Yan Zhang, Weijian Zhuang, Guodong Lu, and Yingzi Yun. 2024. "A Network of Sporogenesis-Responsive Genes Regulates the Growth, Asexual Sporogenesis, Pathogenesis and Fusaric Acid Production of Fusarium oxysporum f. sp. cubense" Journal of Fungi 10, no. 1: 1. https://doi.org/10.3390/jof10010001
APA StyleLu, S., Deng, H., Lin, Y., Huang, M., You, H., Zhang, Y., Zhuang, W., Lu, G., & Yun, Y. (2024). A Network of Sporogenesis-Responsive Genes Regulates the Growth, Asexual Sporogenesis, Pathogenesis and Fusaric Acid Production of Fusarium oxysporum f. sp. cubense. Journal of Fungi, 10(1), 1. https://doi.org/10.3390/jof10010001






