Attention-Based UNet Deep Learning Model for Plaque Segmentation in Carotid Ultrasound for Stroke Risk Stratification: An Artificial Intelligence Paradigm
Abstract
1. Introduction
2. Database Selection, Preparation, and Baseline Characteristics
2.1. DB1: UK ICA Database
2.2. DB2: Japanese Diabetic CCA Database
2.3. DB3: Hong Kong Post-Menopausal Women CCA Database
2.4. Data Preparation and Augmentation Technique
2.5. Binary Mask Preparation for Supervised Learning
3. UNet Architectures
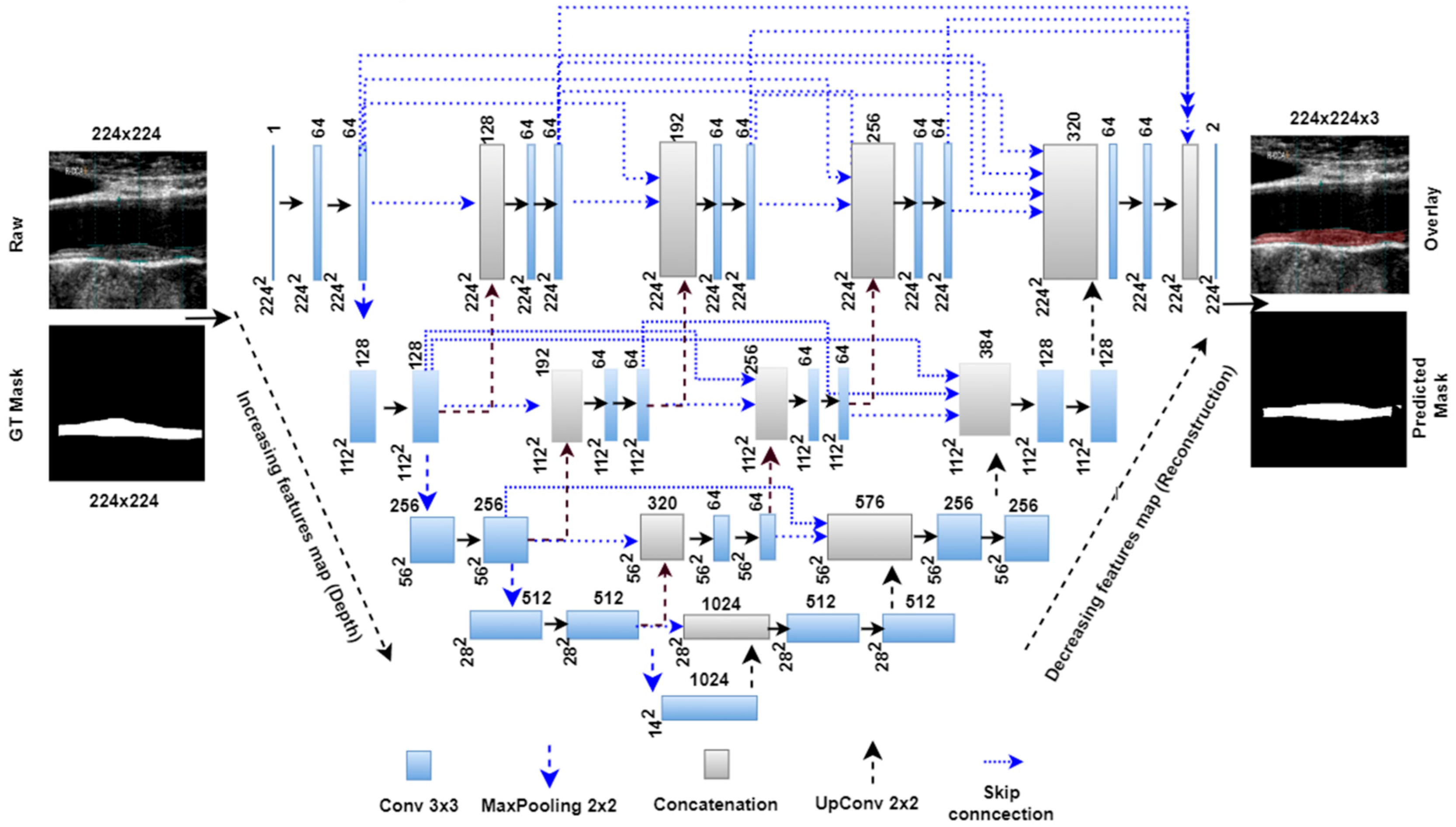
3.1. Basic UNet Model
3.2. Unet++ Architecture
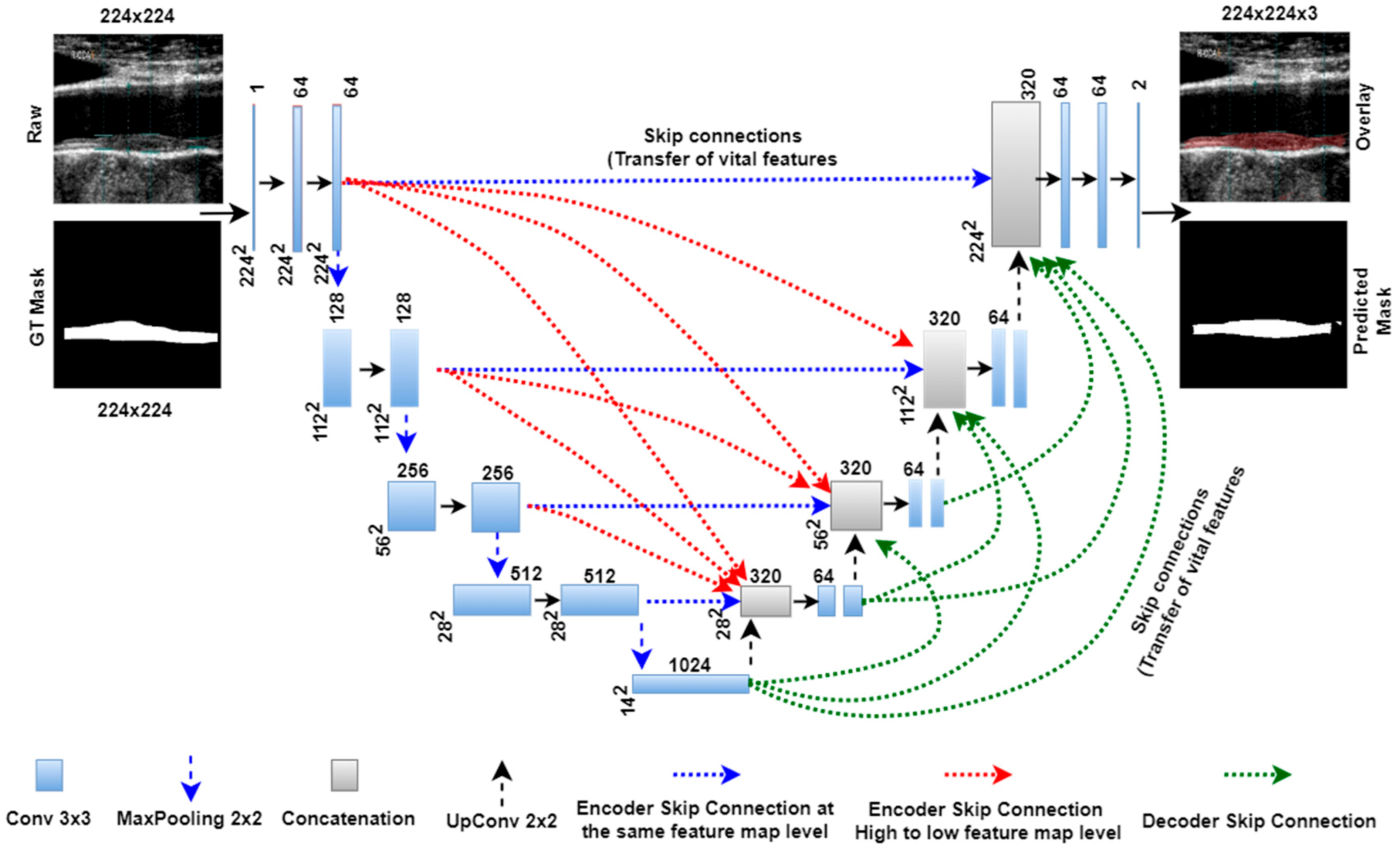
3.3. Unet3P
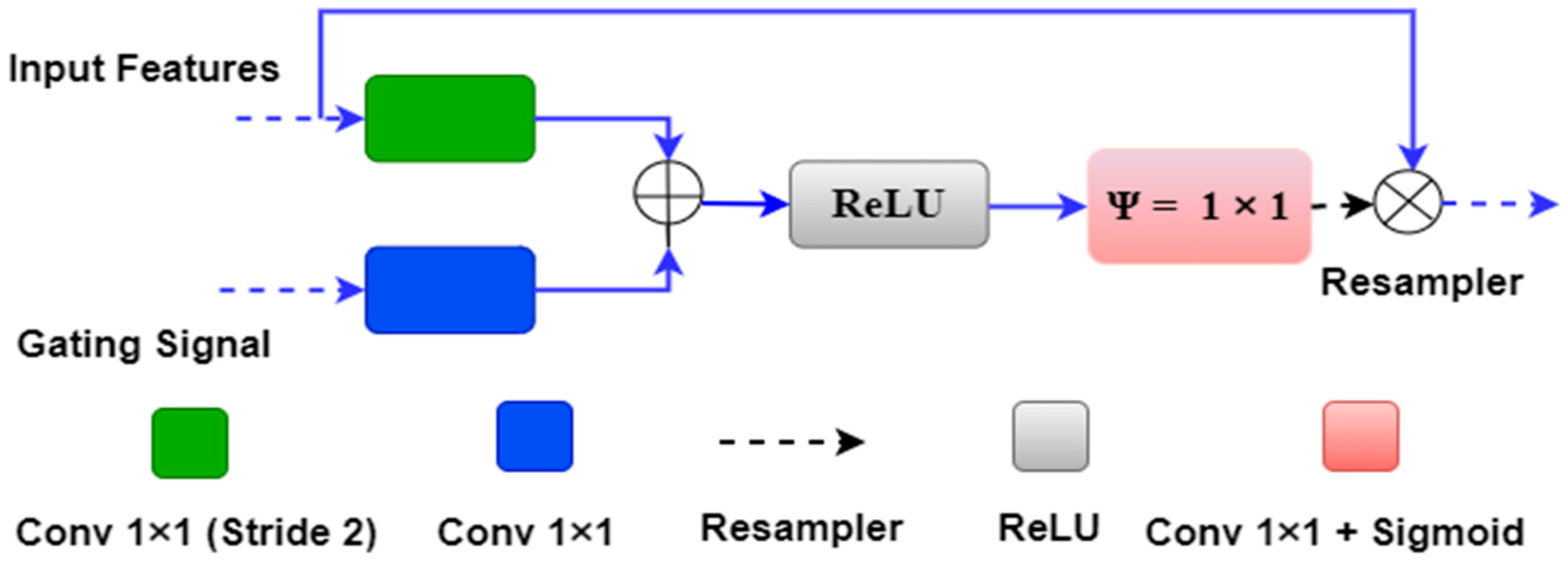
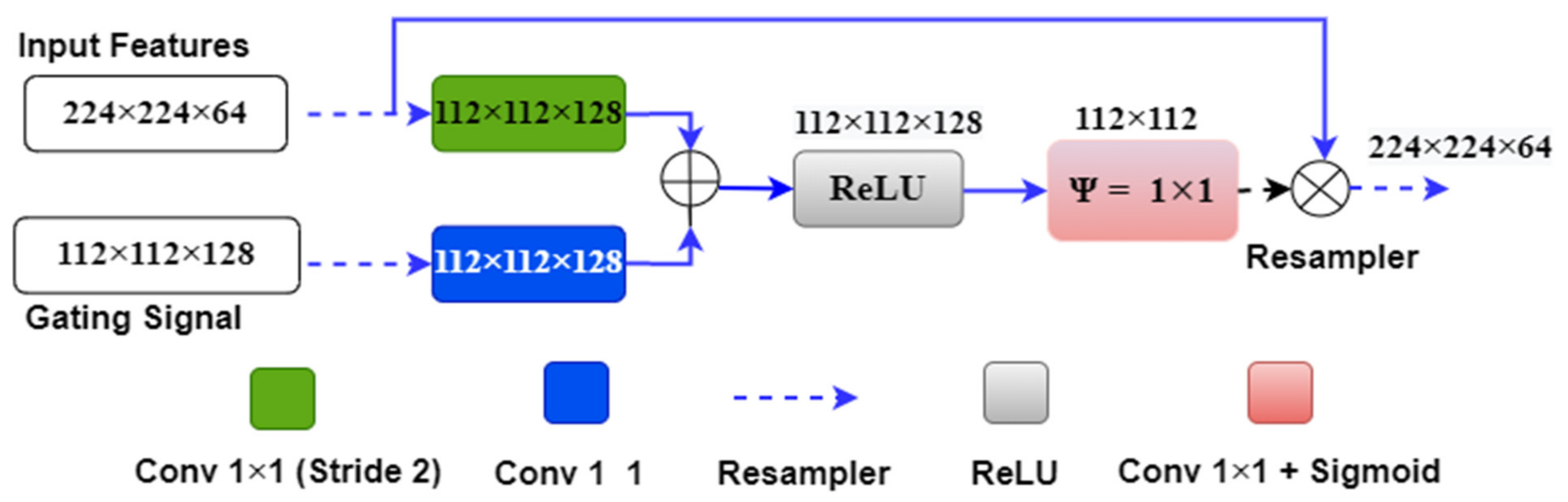
3.4. Attention-based Unet Model
4. Methodology and Experiments
4.1. Hyperparameter Selection and Optimization
4.2. Sparse Categorical Cross-Entropy Loss Function
4.3. K5 Cross-Validation
5. Results
6. Performance Evaluation
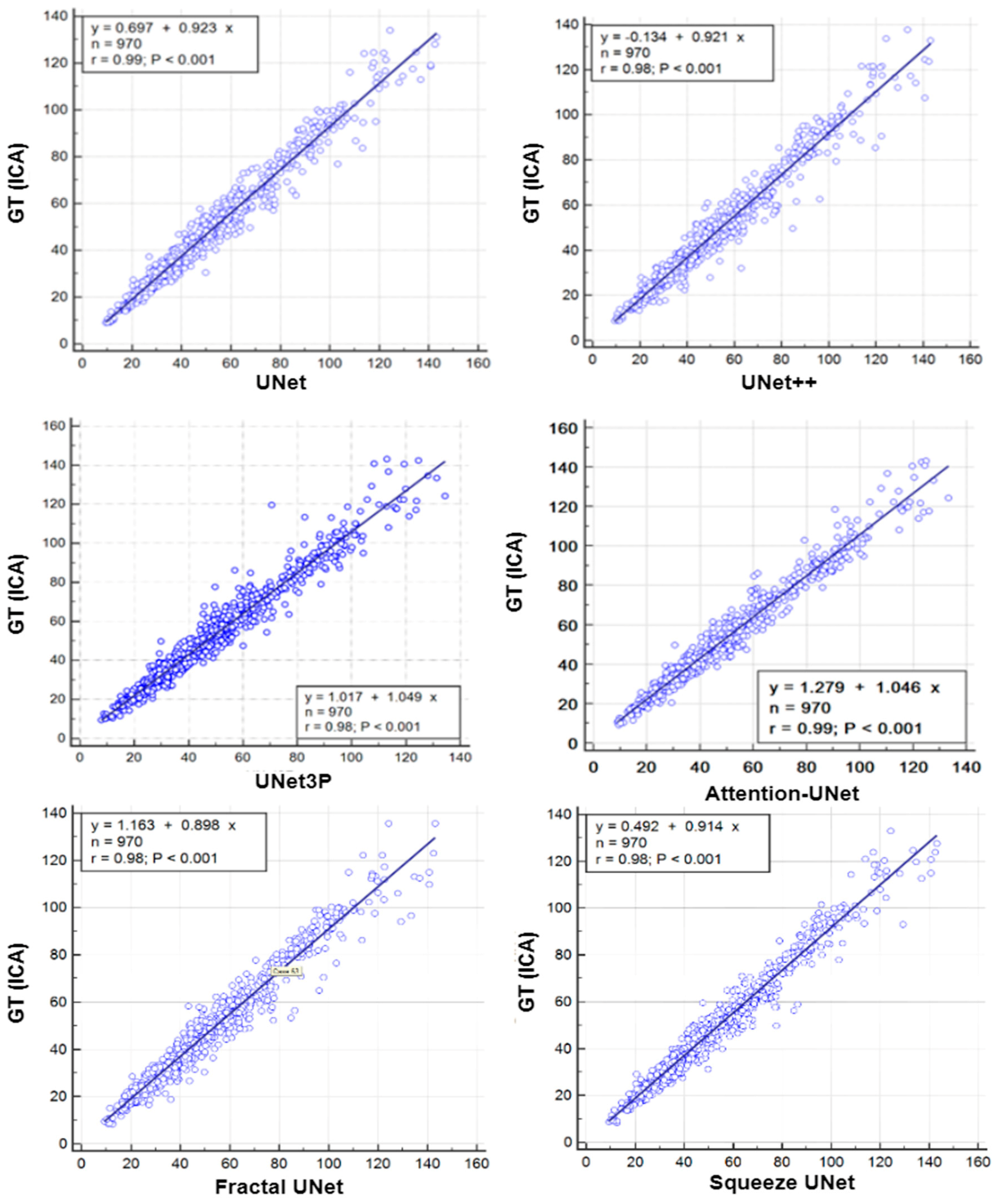
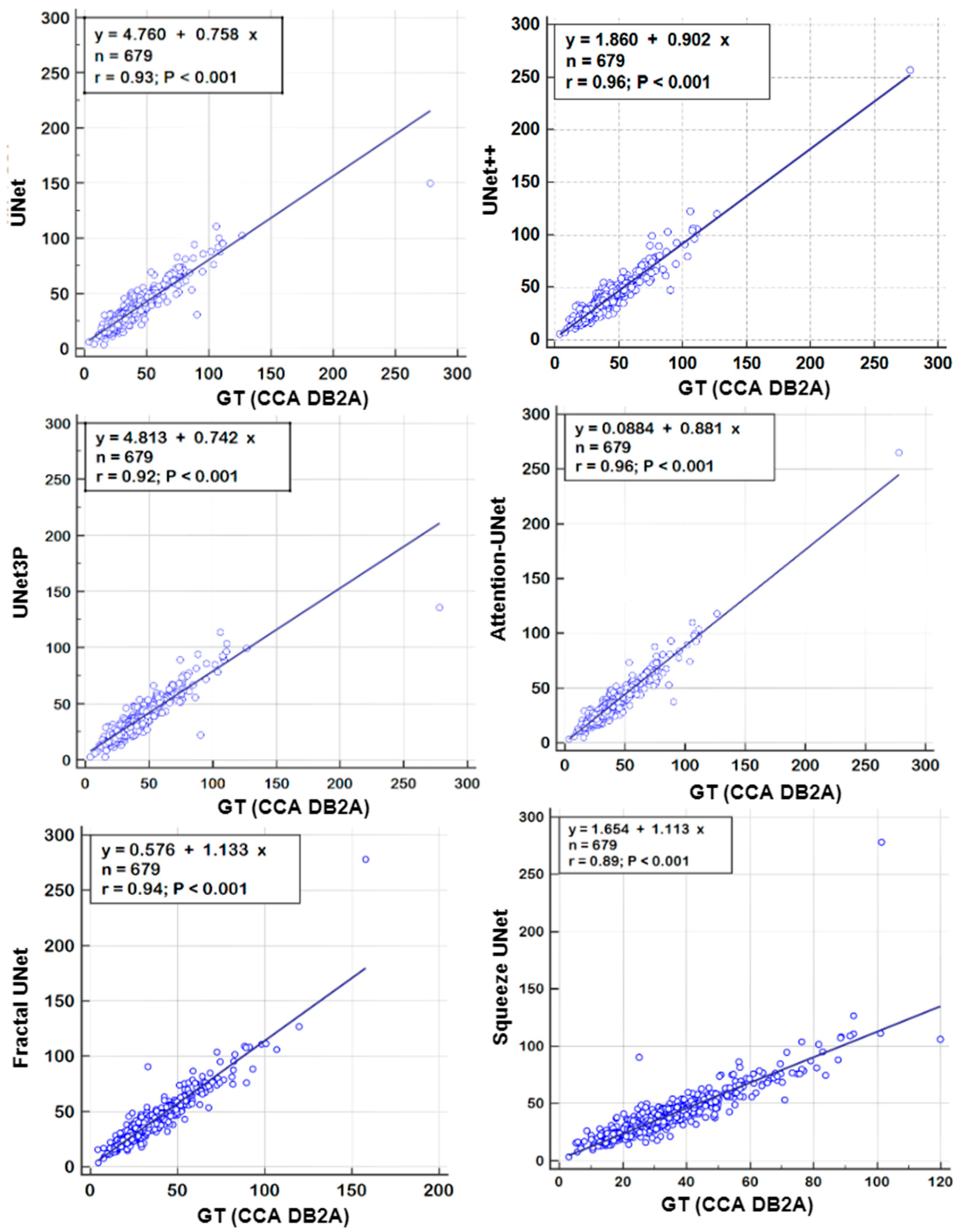
6.1. Regression Analysis
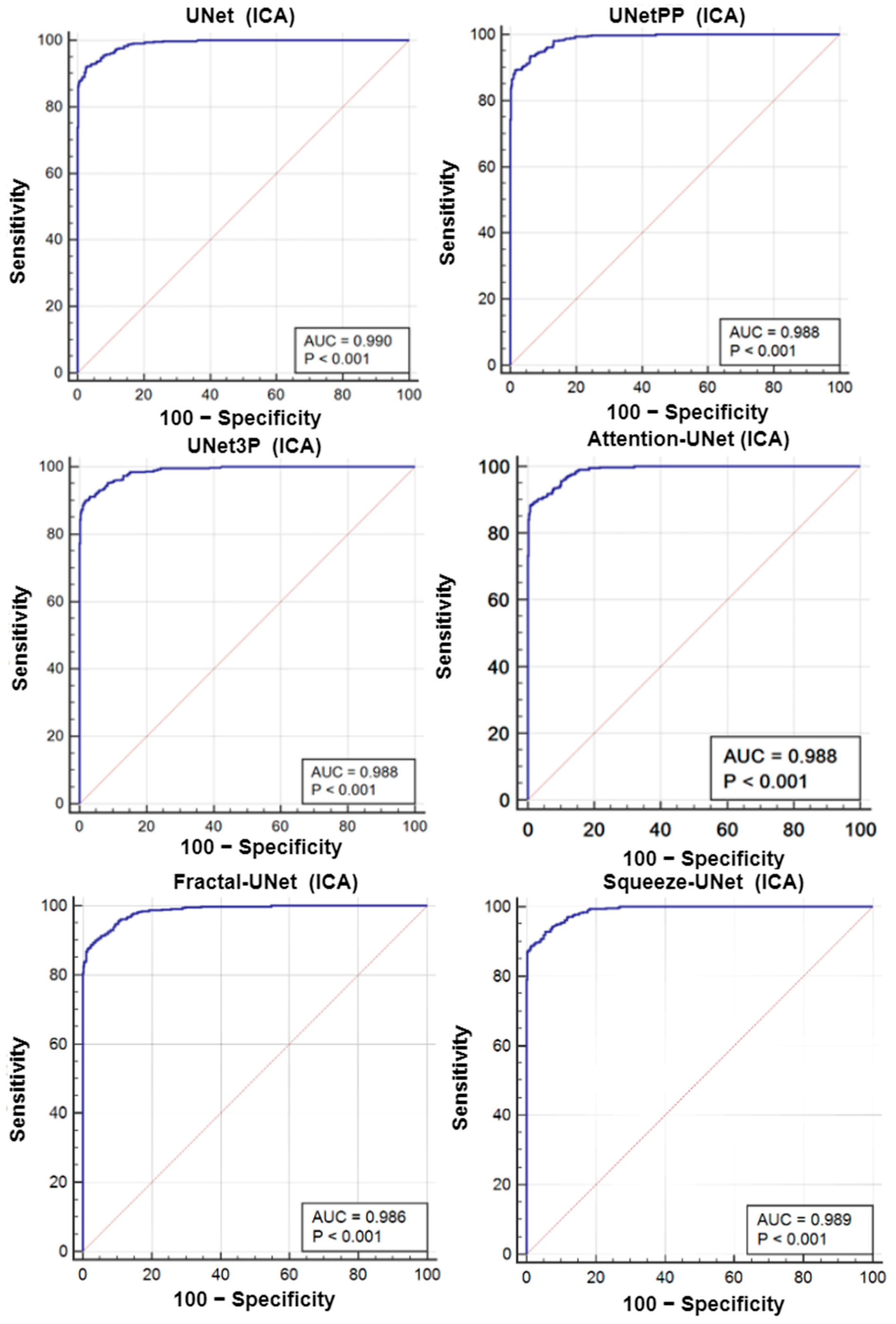
6.2. Receiver Operating Characteristics
6.3. Paired-t-Test Analysis
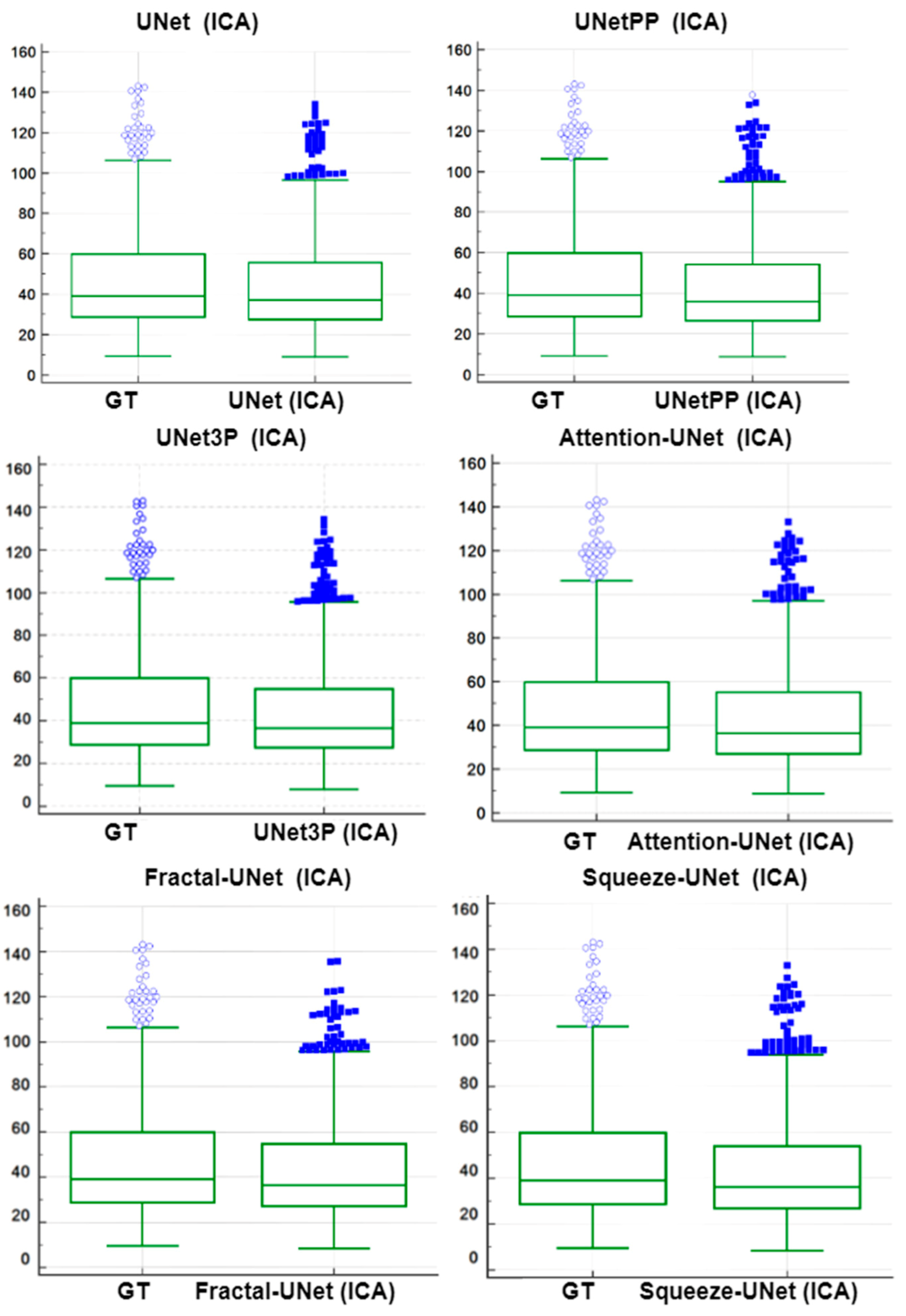
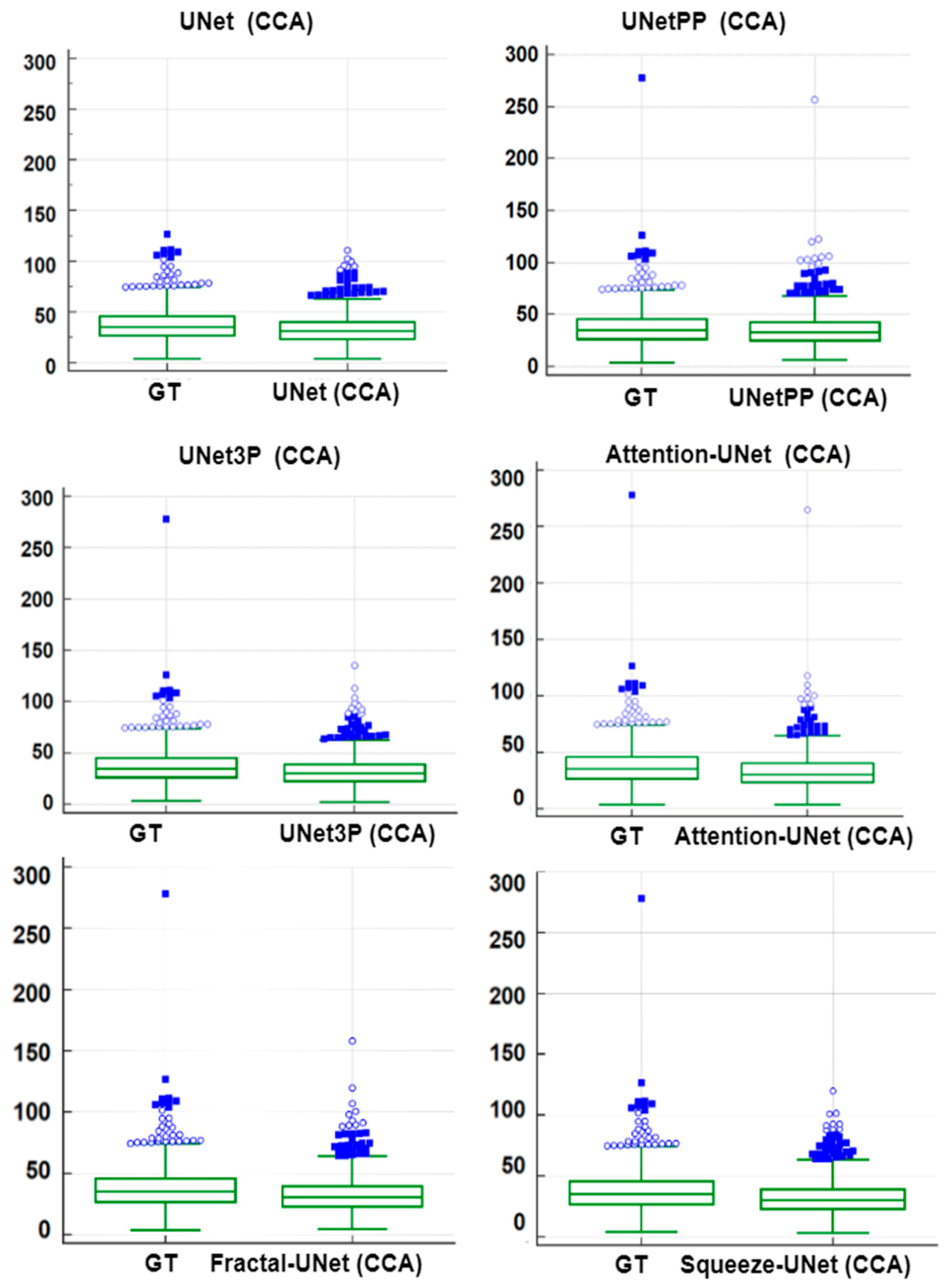
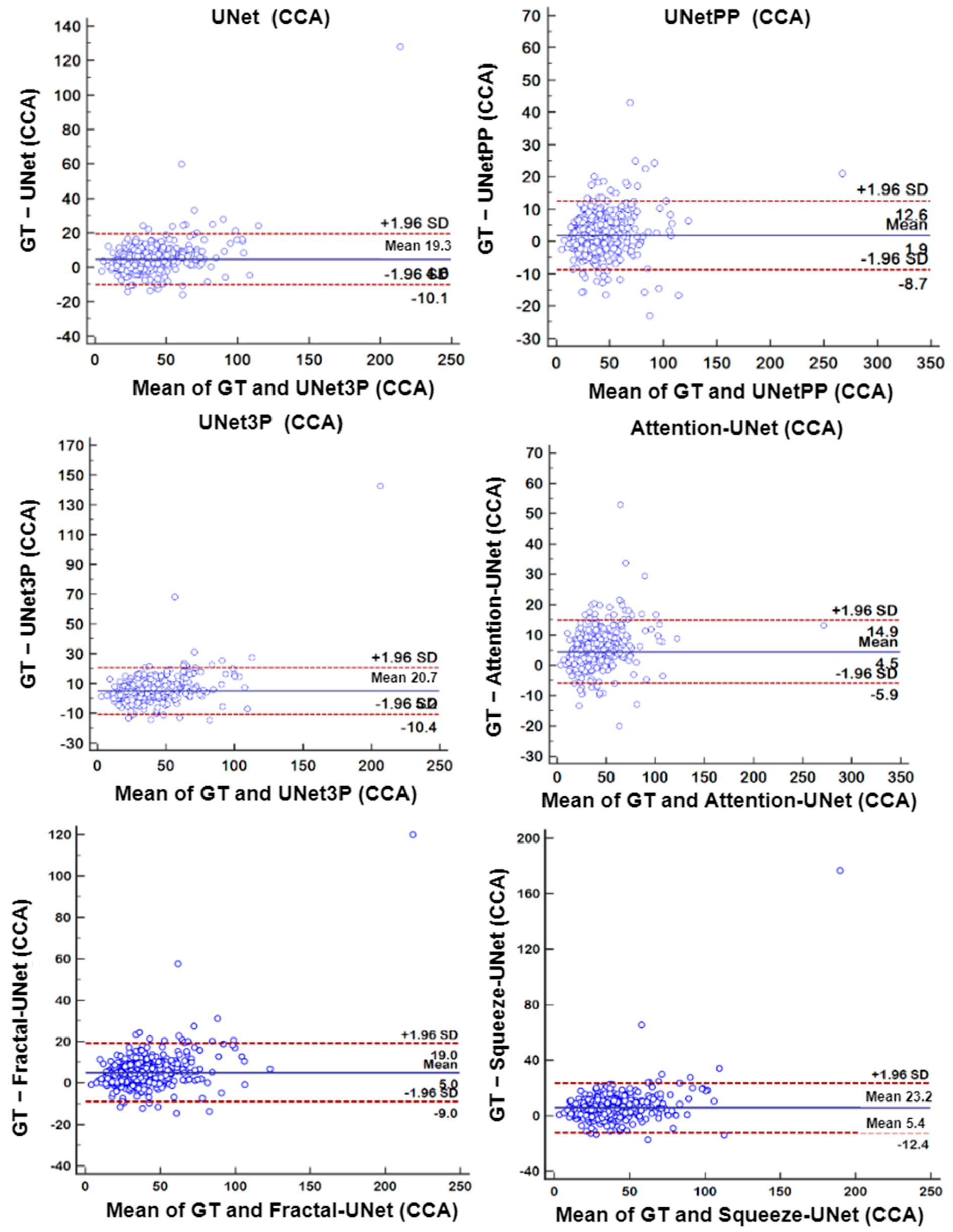
6.4. Bland-Altman’s Plot
7. Discussion
7.1. Bias in Medical Imaging Models
7.2. Supervised and Unsupervised Learning Based DL Models
7.3. Benchmarking
7.4. Strength, Weakness, and Extension
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chauhan, S.; Aeri, B.T. The rising incidence of cardiovascular diseases in India: Assessing its economic impact. J. Prev. Cardiol. 2015, 4, 735–740. [Google Scholar]
- Jones, S.P.; Baqai, K.; Clegg, A.; Georgiou, R.; Harris, C.; Holland, E.-J.; Kalkonde, Y.; Lightbody, C.E.; Maulik, P.K.; Srivastava, P.M.; et al. Stroke in India: A systematic review of the incidence, prevalence, and case fatality. Int. J. Stroke. 2022, 17, 132–140. [Google Scholar] [CrossRef] [PubMed]
- Dorairaj, P.; Jeemon, P.; Roy, A. Cardiovascular Diseases in India: Current Epidemiology and Future Directions. Circulation 2016, 133, 1605–1620. [Google Scholar] [CrossRef]
- Molinari, F.; Meiburger, K.M.; Saba, L.; Acharya, U.R.; Famiglietti, L.; Georgiou, N.; Nicolaides, A.; Mamidi, R.S.; Kuper, H.; Suri, J.S. Automated carotid IMT measurement and its validation in low contrast ultrasound database of 885 patient Indian population epidemiological study: Results of AtheroEdge® software. In Multi-Modality Atherosclerosis Imaging and Diagnosis; Springer: Berlin/Heidelberg, Germany, 2014; pp. 209–219. [Google Scholar]
- Kamalakannan, S.; Gudlavalleti, A.S.V.; Gudlavalleti, V.S.M.; Goenka, S.; Kuper, H. Incidence & prevalence of stroke in India: A systematic review. Ind. J. Med. Res. 2017, 146, 175–185. [Google Scholar] [CrossRef] [PubMed]
- Kleindorfer, D.O.; Towfighi, A.; Chaturvedi, S.; Cockroft, K.M.; Gutierrez, J.; Lombardi-Hill, D.; Kamel, H.; Kernan, W.N.; Kittner, S.J.; Leira, E.C.; et al. 2021 Guideline for the Prevention of Stroke in Patients With Stroke and Transient Ischemic Attack: A Guideline From the American Heart Association/American Stroke Association. Stroke 2021, 52. [Google Scholar] [CrossRef]
- Viswanathan, V.; Puvvula, A.; Jamthikar, A.D.; Saba, L.; Johri, A.M.; Kotsis, V.; Khanna, N.N.; Dhanjil, S.K.; Majhail, M.; Misra, D.P.; et al. Bidirectional link between diabetes mellitus and coronavirus disease 2019 leading to cardiovascular disease: A narrative review. World J. Diabetes 2021, 12, 215–237. [Google Scholar] [CrossRef]
- Puvvula, A.; Jamthikar, A.D.; Gupta, D.; Khanna, N.N.; Porcu, M.; Saba, L.; Viskovic, K.; Ajuluchukwu, J.N.A.; Gupta, A.; Mavrogeni, S.; et al. Morphological Carotid Plaque Area Is Associated With Glomerular Filtration Rate: A Study of South Asian Indian Patients With Diabetes and Chronic Kidney Disease. Angiology 2020, 71, 520–535. [Google Scholar] [CrossRef] [PubMed]
- Virani, S.S.; Alonso, A.; Aparicio, H.J.; Benjamin, E.J.; Bittencourt, M.S.; Callaway, C.W.; Carson, A.P.; Chamberlain, A.M.; Cheng, S.; Delling, F.N.; et al. Heart Disease and Stroke Statistics-2021 Update. Circulation 2021, 143, e254–e743. [Google Scholar] [CrossRef] [PubMed]
- Benjamin, E.J.; Virani, S.S.; Callaway, C.W.; Chamberlain, A.M.; Chang, A.R.; Cheng, S.; Chiuve, S.E.; Cushman, M.; Delling, F.N.; Deo, R. Heart disease and stroke statistics—2018 update: A report from the American Heart Association. Circulation 2018, 137, e67–e492. [Google Scholar] [CrossRef] [PubMed]
- Benjamin, E.J.; Muntner, P.; Alonso, A.; Bittencourt, M.S.; Callaway, C.W.; Carson, A.P.; Chamberlain, A.M.; Chang, A.R.; Cheng, S.; Das, S.R.; et al. Heart disease and stroke statistics—2019 update: A report from the American heart association. Circulation 2019, 139, e56–e528. [Google Scholar] [CrossRef]
- Suri, J.S.; Kathuria, C.; Molinari, F. Atherosclerosis Disease Management; Springer: New York, NY, USA, 2010. [Google Scholar]
- Saba, L.; Sanches, J.M.; Pedro, L.M.; Suri, J.S. Multi-Modality Atherosclerosis Imaging and Diagnosis; Springer: Berlin/Heidelberg, Germany, 2014. [Google Scholar]
- Saba, L.; Antignani, P.L.; Gupta, A.; Cau, R.; Paraskevas, K.I.; Poredos, P.; Wasserman, B.A.; Kamel, H.; Avgerinos, E.D.; Salgado, R.; et al. International Union of Angiology (IUA) consensus paper on imaging strategies in atherosclerotic carotid artery imaging: From basic strategies to advanced approaches. Atherosclerosis 2022, 354, 23–40. [Google Scholar] [CrossRef] [PubMed]
- Patel, A.K.; Suri, H.S.; Singh, J.; Kumar, D.; Shafique, S.; Nicolaides, A.; Jain, S.K.; Saba, L.; Gupta, A.; Laird, J.R.; et al. A Review on Atherosclerotic Biology, Wall Stiffness, Physics of Elasticity, and Its Ultrasound-Based Measurement. Curr. Atheroscler. Rep. 2016, 18, 83. [Google Scholar] [CrossRef] [PubMed]
- Saba, L.; Nardi, V.; Cau, R.; Gupta, A.; Kamel, H.; Suri, J.S.; Balestrieri, A.; Congiu, T.; Butler, A.P.; Gieseg, S.; et al. Carotid Artery Plaque Calcifications: Lessons From Histopathology to Diagnostic Imaging. Stroke 2022, 53, 290–297. [Google Scholar] [CrossRef] [PubMed]
- Kotsis, V.; Jamthikar, A.; Araki, T.; Gupta, D.; Laird, J.R.; Giannopoulos, A.A.; Saba, L.; Suri, H.S.; Mavrogeni, S.; Kitas, G.D.; et al. Echolucency-based phenotype in carotid atherosclerosis disease for risk stratification of diabetes patients. Diabetes Res. Clin. Pract. 2018, 143, 322–331. [Google Scholar] [CrossRef]
- Placidi, G. MRI: Essentials for Innovative Technologies; CRC Press: Boca Raton, FL, USA, 2012. [Google Scholar]
- Saba, L.; Suri, J.S. Multi-Detector CT Imaging: Principles, Head, Neck, and Vascular Systems; CRC Press: Florida, FL, USA, 2013; Volume 1. [Google Scholar]
- Trivedi, R.; Saba, L.; Suri, J.S. 3D Imaging Technologies in Atherosclerosis; Springer: Berlin/Heidelberg, Germany, 2015. [Google Scholar]
- Suri, J.S. Advances in Diagnostic and Therapeutic Ultrasound Imaging; Artech House: London, UK, 2008. [Google Scholar]
- Laine, A.; Sanches, J.M.; Suri, J.S. Ultrasound Imaging: Advances and Applications; Springer: New York, NY, USA, 2012. [Google Scholar]
- Sharma, A.M.; Gupta, A.; Kumar, P.K.; Rajan, J.; Saba, L.; Nobutaka, I.; Laird, J.R.; Nicolades, A.; Suri, J.S. A Review on Carotid Ultrasound Atherosclerotic Tissue Characterization and Stroke Risk Stratification in Machine Learning Framework. Curr. Atheroscler. Rep. 2015, 17. [Google Scholar] [CrossRef]
- Kumar, P.K.; Araki, T.; Rajan, J.; Laird, J.R.; Nicolaides, A.; Suri, J.S. State-of-the-art review on automated lumen and adventitial border delineation and its measurements in carotid ultrasound. Comput. Methods Programs Biomed. 2018, 163, 155–168. [Google Scholar] [CrossRef]
- Saba, L.; Jamthikar, A.; Gupta, D.; Khanna, N.N.; Viskovic, K.; Suri, H.S.; Gupta, A.; Mavrogeni, S.; Turk, M.; Laird, J.R. Global perspective on carotid intima-media thickness and plaque: Should the current measurement guidelines be revisited? Int. Angiol. 2019, 38, 451–465. [Google Scholar] [CrossRef]
- Molinari, F.; Meiburger, K.M.; Acharya, U.R.; Zeng, G.; Rodrigues, P.S.; Saba, L.; Nicolaides, A.; Suri, J.S. CARES 3.0: A two stage system combining feature-based recognition and edge-based segmentation for CIMT measurement on a multi-institutional ultrasound database of 300 images. In Proceedings of the 2011 Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Boston, MA, USA, 30 August–3 September 2011; pp. 5149–5152. [Google Scholar] [CrossRef]
- Molinari, F.; Meiburger, K.M.; Zeng, G.; Nicolaides, A.; Suri, J.S. CAUDLES-EF: Carotid Automated Ultrasound Double Line Extraction System Using Edge Flow. In Ultrasound Imaging; Springer: Boston, MA, USA, 2012; pp. 129–162. [Google Scholar] [CrossRef]
- Molinari, F.; Liboni, W.; Pantziaris, M.; Suri, J. CALSFOAM-completed automated local statistics based first order absolute moment for carotid wall recognition, segmentation and IMT measurement: Validation and benchmarking on a 300 patient database. Int. J. Angiol. 2011, 30, 227–241. [Google Scholar]
- Molinari, F.; Meiburger, K.M.; Saba, L.; Acharya, U.R.; Ledda, G.; Zeng, G.; Ho, S.Y.S.; Ahuja, A.T.; Ho, S.C.; Nicolaides, A.; et al. Ultrasound IMT measurement on a multi-ethnic and multi-institutional database: Our review and experience using four fully automated and one semi-automated methods. Comput. Methods Programs Biomed. 2012, 108, 946–960. [Google Scholar] [CrossRef]
- Faita, F.; Gemignani, V.; Bianchini, E.; Giannarelli, C.; Ghiadoni, L.; Demi, M. Real-time Measurement System for Evaluation of the Carotid Intima-Media Thickness With a Robust Edge Operator. J. Ultrasound Med. 2008, 27, 1353–1361. [Google Scholar] [CrossRef]
- Molinari, F.; Pattichis, C.S.; Zeng, G.; Saba, L.; Acharya, U.R.; Sanfilippo, R.; Nicolaides, A.; Suri, J.S. Completely Automated Multiresolution Edge Snapper—A New Technique for an Accurate Carotid Ultrasound IMT Measurement: Clinical Validation and Benchmarking on a Multi-Institutional Database. IEEE Trans. Image Process. 2012, 21, 1211–1222. [Google Scholar] [CrossRef] [PubMed]
- Zahnd, G.; Kapellas, K.; van Hattem, M.; van Dijk, A.; Sérusclat, A.; Moulin, P.; van der Lugt, A.; Skilton, M.; Orkisz, M. A Fully-Automatic Method to Segment the Carotid Artery Layers in Ultrasound Imaging: Application to Quantify the Compression-Decompression Pattern of the Intima-Media Complex During the Cardiac Cycle. Ultrasound Med. Biol. 2016, 43, 239–257. [Google Scholar] [CrossRef] [PubMed]
- Rouco, J.; Carvalho, C.; Domingues, A.; Azevedo, E.; Campilho, A. A robust anisotropic edge detection method for carotid ultrasound image processing. Procedia Comput. Sci. 2018, 126, 723–732. [Google Scholar] [CrossRef]
- Loizou, C.P.; Pattichis, C.; Pantziaris, M.; Tyllis, T.; Nicolaides, A. Snakes based segmentation of the common carotid artery intima media. Med. Biol. Eng. Comput. 2007, 45, 35–49. [Google Scholar] [CrossRef] [PubMed]
- Molinari, F.; Meiburger, K.M.; Saba, L.; Rajendra Acharya, U.; Ledda, M.; Nicolaides, A.; Suri, J.S. Constrained snake vs. conventional snake for carotid ultrasound automated IMT measurements on multi-center data sets. Ultrasonics 2012, 52, 949–961. [Google Scholar] [CrossRef]
- Molinari, F.; Meiburger, K.M.; Saba, L.; Zeng, G.; Acharya, U.R.; Ledda, M.; Nicolaides, A.; Suri, J.S. Fully Automated Dual-Snake Formulation for Carotid Intima-Media Thickness Measurement. J. Ultrasound Med. 2012, 31, 1123–1136. [Google Scholar] [CrossRef]
- Kumar, P.K.; Araki, T.; Rajan, J.; Saba, L.; Lavra, F.; Ikeda, N.; Sharma, A.M.; Shafique, S.; Nicolaides, A.; Laird, J.R.; et al. Accurate lumen diameter measurement in curved vessels in carotid ultrasound: An iterative scale-space and spatial transformation approach. Med. Biol. Eng. Comput. 2016, 55, 1415–1434. [Google Scholar] [CrossRef]
- Suri, J.S.; Liu, K.; Reden, L.; Laxminarayan, S. A review on MR vascular image processing algorithms: Acquisition and prefiltering: Part I. IEEE Trans. Inf. Technol. Biomed. 2002, 6, 324–337. [Google Scholar] [CrossRef]
- Molinari, F.; Zeng, G.; Suri, J.S. Intima-media thickness: Setting a standard for a completely automated method of ultrasound measurement. IEEE Trans. Ultrason. Ferroelectr. Freq. Control 2010, 57, 1112–1124. [Google Scholar] [CrossRef]
- Araki, T.; Ikeda, N.; Dey, N.; Acharjee, S.; Molinari, F.; Saba, L.; Godia, E.C.; Nicolaides, A.; Suri, J.S. Shape-Based Approach for Coronary Calcium Lesion Volume Measurement on Intravascular Ultrasound Imaging and Its Association With Carotid Intima-Media Thickness. J. Ultrasound Med. 2015, 34, 469–482. [Google Scholar] [CrossRef]
- Yang, X.; Jin, J.; Xu, M.; Wu, H.; He, W.; Yuchi, M.; Ding, M. Ultrasound Common Carotid Artery Segmentation Based on Active Shape Model. Comput. Math. Methods Med. 2013, 2013, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Molinari, F.; Krishnamurthi, G.; Acharya, U.R.; Subbhuraam, V.S.; Saba, L.; Nicolaides, A.; Suri, J. Hypothesis Validation of Far-Wall Brightness in Carotid-Artery Ultrasound for Feature-Based IMT Measurement Using a Combination of Level-Set Segmentation and Registration. IEEE Trans. Instrum. Meas. 2012, 61, 1054–1063. [Google Scholar] [CrossRef]
- El-Baz, A.; Suri, J.S. Level Set Method in Medical Imaging Segmentation; CRC Press: Boca Raton, FL, USA, 2019. [Google Scholar] [CrossRef]
- Li, C.; Xu, C.; Gui, C.; Fox, M.D. Level set evolution without re-initialization: A new variational formulation. In Proceedings of the 2005 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR’05), San Diego, CA, USA, 20–25 June 2005; pp. 430–436. [Google Scholar]
- Araki, T.; Kumar, P.K.; Suri, H.S.; Ikeda, N.; Gupta, A.; Saba, L.; Rajan, J.; Lavra, F.; Sharma, A.M.; Shafique, S. Two automated techniques for carotid lumen diameter measurement: Regional versus boundary approaches. J. Med. Syst. 2016, 40, 1–19. [Google Scholar] [CrossRef]
- Suri, J.; Liu, K.; Singh, S.; Laxminarayan, S.; Zeng, X.; Reden, L. Shape recovery algorithms using level sets in 2-D/3-D medical imagery: A state-of-the-art review. IEEE Trans. Inf. Technol. Biomed. 2002, 6, 8–28. [Google Scholar] [CrossRef]
- Giraldi, G.A.; Rodrigues, P.S.; Suri, J.S. Implicit dual snakes for medical imaging. In Proceedings of the 2006 International Conference of the IEEE Engineering in Medicine and Biology Society, New York, NY, USA, 30 August–3 September 2006; pp. 3025–3028. [Google Scholar]
- Roy, S.; Bhattacharyya, D.; Bandyopadhyay, S.K.; Kim, T.-H. An iterative implementation of level set for precise segmentation of brain tissues and abnormality detection from MR images. IETE J. Res. 2017, 63, 769–783. [Google Scholar] [CrossRef]
- Ikeda, N.; Dey, N.; Sharma, A.; Gupta, A.; Bose, S.; Acharjee, S.; Shafique, S.; Cuadrado-Godia, E.; Araki, T.; Saba, L.; et al. Automated segmental-IMT measurement in thin/thick plaque with bulb presence in carotid ultrasound from multiple scanners: Stroke risk assessment. Comput. Methods Programs Biomed. 2017, 141, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Saba, L.; Banchhor, S.K.; Suri, H.S.; Londhe, N.D.; Araki, T.; Ikeda, N.; Viskovic, K.; Shafique, S.; Laird, J.R.; Gupta, A. Accurate cloud-based smart IMT measurement, its validation and stroke risk stratification in carotid ultrasound: A web-based point-of-care tool for multicenter clinical trial. Comput. Biol. Med. 2016, 75, 217–234. [Google Scholar] [CrossRef]
- Meiburger, K.M.; Marzola, F.; Zahnd, G.; Faita, F.; Loizou, C.P.; Lainé, N.; Carvalho, C.; Steinman, D.A.; Gibello, L.; Bruno, R.M. Carotid Ultrasound Boundary Study (CUBS): Technical considerations on an open multi-center analysis of computerized measurement systems for intima-media thickness measurement on common carotid artery longitudinal B-mode ultrasound scans. Comput. Biol. Med. 2022, 144, 105333. [Google Scholar] [CrossRef]
- Menchón-Lara, R.-M.; Bastida-Jumilla, M.-C.; Larrey-Ruiz, J.; Verdú-Monedero, R.; Morales-Sánchez, J.; Sancho-Gómez, J.-L. Measurement of Carotid Intima-Media Thickness in ultrasound images by means of an automatic segmentation process based on machine learning. Proceedings of Eurocon 2013, Zagreb, Croatia, 1–4 July 2013; pp. 2086–2093. [Google Scholar]
- Biswas, M.; Saba, L.; Omerzu, T.; Johri, A.M.; Khanna, N.N.; Viskovic, K.; Mavrogeni, S.; Laird, J.R.; Pareek, G.; Miner, M. A review on joint carotid intima-media thickness and plaque area measurement in ultrasound for cardiovascular/stroke risk monitoring: Artificial intelligence framework. J. Digit. Imaging. 2021, 34, 581–604. [Google Scholar] [CrossRef]
- Kuppili, V.; Biswas, M.; Sreekumar, A.; Suri, H.S.; Saba, L.; Edla, D.R.; Marinhoe, R.T.; Sanches, J.; Suri, J.S. Extreme Learning Machine Framework for Risk Stratification of Fatty Liver Disease Using Ultrasound Tissue Characterization. J. Med Syst. 2017, 41, 152. [Google Scholar] [CrossRef]
- Shrivastava, V.K.; Londhe, N.D.; Sonawane, R.S.; Suri, J.S. Reliable and accurate psoriasis disease classification in dermatology images using comprehensive feature space in machine learning paradigm. Expert Syst. Appl. 2015, 42, 6184–6195. [Google Scholar] [CrossRef]
- Araki, T.; Ikeda, N.; Shukla, D.; Jain, P.K.; Londhe, N.D.; Shrivastava, V.K.; Banchhor, S.K.; Saba, L.; Nicolaides, A.; Shafique, S.; et al. PCA-based polling strategy in machine learning framework for coronary artery disease risk assessment in intravascular ultrasound: A link between carotid and coronary grayscale plaque morphology. Comput. Methods Programs Biomed. 2016, 128, 137–158. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Faust, O.S.; Faust, O.S.V.S.; Krishnamurthi, A.; Faust, G.; Seabra, J.; Sanches, J.; Suri, J.S. Understanding symptomatology of atherosclerotic plaque by image-based tissue characterization. Comput. Methods Programs Biomed. 2013, 110, 66–75. [Google Scholar] [CrossRef] [PubMed]
- Boi, A.; Jamthikar, A.D.; Saba, L.; Gupta, D.; Sharma, A.; Loi, B.; Laird, J.R.; Khanna, N.N.; Suri, J.S. A survey on coronary atherosclerotic plaque tissue characterization in intravascular optical coherence tomography. Curr. Atheroscler. Rep. 2018, 20, 1–17. [Google Scholar] [CrossRef]
- Saba, L.; Biswas, M.; Kuppili, V.; Godia, E.C.; Suri, H.S.; Edla, D.R.; Omerzu, T.; Laird, J.R.; Khanna, N.N.; Mavrogeni, S.; et al. The present and future of deep learning in radiology. Eur. J. Radiol. 2019, 114, 14–24. [Google Scholar] [CrossRef]
- Mainak, B.; Venkatanareshbabu, K.; Luca, S.; Damodar, R.E.; Elisa, C.-G.; Tato, M.R.; Andrew, N. State-of-the-art review on deep learning in medical imaging. Front. Bioscience-Landmark 2019, 24, 380–406. [Google Scholar]
- Saba, L.; Jain, P.K.; Suri, H.S.; Ikeda, N.; Araki, T.; Singh, B.K.; Nicolaides, A.; Shafique, S.; Gupta, A.; Laird, J.R.; et al. Plaque Tissue Morphology-Based Stroke Risk Stratification Using Carotid Ultrasound: A Polling-Based PCA Learning Paradigm. J. Med Syst. 2017, 41, 98. [Google Scholar] [CrossRef]
- Araki, T.; Jain, P.K.; Suri, H.S.; Londhe, N.D.; Ikeda, N.; El-Baz, A.; Shrivastava, V.K.; Saba, L.; Nicolaides, A.; Shafique, S.; et al. Stroke Risk Stratification and its Validation using Ultrasonic Echolucent Carotid Wall Plaque Morphology: A Machine Learning Paradigm. Comput. Biol. Med. 2017, 80, 77–96. [Google Scholar] [CrossRef]
- Menchón-Lara, R.-M.; Sancho-Gómez, J.-L. Fully automatic segmentation of ultrasound common carotid artery images based on machine learning. Neurocomputing 2015, 151, 161–167. [Google Scholar] [CrossRef]
- Cuadrado-Godia, E.; Srivastava, S.K.; Saba, L.; Araki, T.; Suri, H.S.; Giannopolulos, A.; Omerzu, T.; Laird, J.; Khanna, N.N.; Mavrogeni, S. Geometric total plaque area is an equally powerful phenotype compared with carotid intima-media thickness for stroke risk assessment: A deep learning approach. J. Vasc. Ultrasound 2018, 42, 162–188. [Google Scholar] [CrossRef]
- Zhou, R.; Azarpazhooh, M.R.; Spence, J.D.; Hashemi, S.; Ma, W.; Cheng, X.; Gan, H.; Ding, M.; Fenster, A. Deep Learning-Based Carotid Plaque Segmentation from B-Mode Ultrasound Images. Ultrasound Med. Biol. 2021, 47, 2723–2733. [Google Scholar] [CrossRef] [PubMed]
- Zhou, R.; Guo, F.; Azarpazhooh, M.R.; Hashemi, S.; Cheng, X.; Spence, J.D.; Ding, M.; Fenster, A. Deep Learning-Based Measurement of Total Plaque Area in B-Mode Ultrasound Images. IEEE J. Biomed. Heal. Informatics 2021, 25, 2967–2977. [Google Scholar] [CrossRef] [PubMed]
- Meshram, N.H.; Mitchell, C.C.; Wilbrand, S.; Dempsey, R.J.; Varghese, T. Deep Learning for Carotid Plaque Segmentation using a Dilated U-Net Architecture. Ultrason. Imaging 2020, 42, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Jain, P.K.; Sharma, N.; Giannopoulos, A.A.; Saba, L.; Nicolaides, A.; Suri, J.S. Hybrid deep learning segmentation models for atherosclerotic plaque in internal carotid artery B-mode ultrasound. Comput. Biol. Med. 2021, 136, 104721. [Google Scholar] [CrossRef]
- Jena, B.; Saxena, S.; Nayak, G.K.; Saba, L.; Sharma, N.; Suri, J.S. Artificial intelligence-based hybrid deep learning models for image classification: The first narrative review. Comput. Biol. Med. 2021, 137, 104803. [Google Scholar] [CrossRef]
- Suri, J.S.; Agarwal, S.; Elavarthi, P.; Pathak, R.; Ketireddy, V.; Columbu, M.; Saba, L.; Gupta, S.K.; Faa, G.; Singh, I.M.; et al. Inter-Variability Study of COVLIAS 1.0: Hybrid Deep Learning Models for COVID-19 Lung Segmentation in Computed Tomography. Diagnostics 2021, 11, 2025. [Google Scholar] [CrossRef]
- Skandha, S.S.; Nicolaides, A.; Gupta, S.K.; Koppula, V.K.; Saba, L.; Johri, A.M.; Kalra, M.S.; Suri, J.S. A hybrid deep learning paradigm for carotid plaque tissue characterization and its validation in multicenter cohorts using a supercomputer framework. Comput. Biol. Med. 2022, 141, 105131. [Google Scholar] [CrossRef]
- Suri, J.S.; Agarwal, S.; Chabert, G.L.; Carriero, A.; Paschè, A.; Danna, P.S.C.; Saba, L.; Mehmedović, A.; Faa, G.; Singh, I.M.; et al. COVLIAS 2.0-cXAI: Cloud-Based Explainable Deep Learning System for COVID-19 Lesion Localization in Computed Tomography Scans. Diagnostics 2022, 12, 1482. [Google Scholar] [CrossRef]
- Jain, P.K.; Sharma, N.; Saba, L.; Paraskevas, K.I.; Kalra, M.K.; Johri, A.; Nicolaides, A.N.; Suri, J.S. Automated deep learning-based paradigm for high-risk plaque detection in B-mode common carotid ultrasound scans: An asymptomatic Japanese cohort study. Int. Angiol. 2021, 41, 9–23. [Google Scholar] [CrossRef]
- Jain, P.K.; Sharma, N.; Kalra, M.K.; Johri, A.; Saba, L.; Suri, J.S. Far wall plaque segmentation and area measurement in common and internal carotid artery ultrasound using U-series architectures: An unseen Artificial Intelligence paradigm for stroke risk assessment. Comput. Biol. Med. 2022, 149, 106017. [Google Scholar] [CrossRef]
- Oktay, O.; Schlemper, J.; Folgoc, L.; Lee, M.; Heinrich, M.; Misawa, K.; Mori, K.; McDonagh, S.; Hammerla, N.; Kainz, B.; et al. Attention U-Net: Learning Where to Look for the Pancreas. arXiv 2018, arXiv:1804.03999v3. [Google Scholar]
- Pal, D.; Reddy, P.B.; Roy, S. Attention UW-Net: A fully connected model for automatic segmentation and annotation of chest X-ray. Comput. Biol. Med. 2022, 106083. [Google Scholar] [CrossRef] [PubMed]
- Molinari, F.; Liboni, W.; Giustetto, P.; Badalamenti, S.; Suri, J.S. AUTOMATIC COMPUTER-BASED TRACINGS (ACT) IN LONGITUDINAL 2-D ULTRASOUND IMAGES USING DIFFERENT SCANNERS. J. Mech. Med. Biol. 2009, 9, 481–505. [Google Scholar] [CrossRef]
- Jain, P.K.; Sharma, N.; Saba, L.; Paraskevas, K.I.; Kalra, M.K.; Johri, A.; Laird, J.R.; Nicolaides, A.N.; Suri, J.S. Unseen Artificial Intelligence—Deep Learning Paradigm for Segmentation of Low Atherosclerotic Plaque in Carotid Ultrasound: A Multicenter Cardiovascular Study. Diagnostics 2021, 11, 2257. [Google Scholar] [CrossRef]
- Suri, J.S.; Agarwal, S.; Chabert, G.L.; Carriero, A.; Paschè, A.; Danna, P.S.C.; Saba, L.; Mehmedović, A.; Faa, G.; Singh, I.M.; et al. COVLIAS 1.0Lesion vs. MedSeg: An Artificial Intelligence Framework for Automated Lesion Segmentation in COVID-19 Lung Computed Tomography Scans. Diagnostics 2022, 12, 1283. [Google Scholar] [CrossRef]
- Suri, J.; Agarwal, S.; Pathak, R.; Ketireddy, V.; Columbu, M.; Saba, L.; Gupta, S.; Faa, G.; Singh, I.; Turk, M.; et al. COVLIAS 1.0: Lung Segmentation in COVID-19 Computed Tomography Scans Using Hybrid Deep Learning Artificial Intelligence Models. Diagnostics 2021, 11, 1405. [Google Scholar] [CrossRef]
- Saba, L.; Banchhor, S.K.; Araki, T.; Viskovic, K.; Londhe, N.D.; Laird, J.R.; Suri, H.S.; Suri, J.S. Intra- and inter-operator reproducibility of automated cloud-based carotid lumen diameter ultrasound measurement. Ind. Heart J. 2018, 70, 649–664. [Google Scholar] [CrossRef]
- Than, J.C.; Saba, L.; Noor, N.M.; Rijal, O.M.; Kassim, R.M.; Yunus, A.; Suri, H.S.; Porcu, M.; Suri, J.S. Lung disease stratification using amalgamation of Riesz and Gabor transforms in machine learning framework. Comput. Biol. Med. 2017, 89, 197–211. [Google Scholar] [CrossRef]
- Saba, L.; Than, J.C.M.; Noor, N.M.; Rijal, O.M.; Kassim, R.M.; Yunus, A.; Ng, C.R.; Suri, J.S. Inter-observer Variability Analysis of Automatic Lung Delineation in Normal and Disease Patients. J. Med. Syst. 2016, 40, 1–18. [Google Scholar] [CrossRef]
- Lucatelli, P.; Raz, E.; Saba, L.; Argiolas, G.M.; Montisci, R.; Wintermark, M.; King, K.S.; Molinari, F.; Ikeda, N.; Siotto, P.; et al. Relationship between leukoaraiosis, carotid intima-media thickness and intima-media thickness variability: Preliminary results. Eur. Radiol. 2016, 26, 4423–4431. [Google Scholar] [CrossRef]
- Ikeda, N.; Araki, T.; Sugi, K.; Nakamura, M.; Deidda, M.; Molinari, F.; Meiburger, K.M.; Acharya, U.R.; Saba, L.; Bassareo, P.P.; et al. Ankle–Brachial Index and Its Link to Automated Carotid Ultrasound Measurement of Intima–Media Thickness Variability in 500 Japanese Coronary Artery Disease Patients. Curr. Atheroscler. Rep. 2014, 16, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Saba, L.; Molinari, F.; Meiburger, K.M.; Acharya, U.R.; Nicolaides, A.; Suri, J.S. Inter- and intra-observer variability analysis of completely automated cIMT measurement software (AtheroEdge™) and its benchmarking against commercial ultrasound scanner and expert Readers. Comput. Biol. Med. 2013, 43, 1261–1272. [Google Scholar] [CrossRef] [PubMed]
- Rajendra Acharya, U.; Paul Joseph, K.; Kannathal, N.; Lim, C.M.; Suri, J.S. Heart rate variability: A review. Med. Biol. Eng. Comput. 2006, 44, 1031–1051. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Suri, J.S.; Salvado, O.; Chen, Y.; Wacker, F.K.; Wilson, D.L.; Duerk, J.L.; Lewin, J.S. Inter- and Intra-Observer Variability Assessment of in Vivo Carotid Plaque Burden Quantification Using Multi-Contrast Dark Blood MR Images. Stud. Heal. Technol. Inform. 2005, 113, 384–393. [Google Scholar]
- Bahdanau, D.; Cho, K.; Bengio, Y. Neural machine translation by jointly learning to align and translate. arXiv preprint 2014, arXiv:1409.0473. [Google Scholar]
- Luong, M.-T.; Pham, H.; Manning, C.D. Effective approaches to attention-based neural machine translation. arXiv Preprint 2015, arXiv:1508.04025. [Google Scholar]
- Jain, P.K.; Gupta, S.; Bhavsar, A.; Nigam, A.; Sharma, N. Localization of common carotid artery transverse section in B-mode ultrasound images using faster RCNN: A deep learning approach. Med. Biol. Eng. Comput. 2020, 58, 471–482. [Google Scholar] [CrossRef]
- Suri, J.S.; Agarwal, S.; Gupta, S.K.; Puvvula, A.; Viskovic, K.; Suri, N.; Alizad, A.; El-Baz, A.; Saba, L.; Fatemi, M.; et al. Systematic Review of Artificial Intelligence in Acute Respiratory Distress Syndrome for COVID-19 Lung Patients: A Biomedical Imaging Perspective. IEEE J. Biomed. Health Inform. 2021, 25, 4128–4139. [Google Scholar] [CrossRef]
- Suri, J.S.; Agarwal, S.; Jena, B.; Saxena, S.; El-Baz, A.; Agarwal, V.; Kalra, M.K.; Saba, L.; Viskovic, K.; Fatemi, M.; et al. Five Strategies for Bias Estimation in Artificial Intelligence-based Hybrid Deep Learning for Acute Respiratory Distress Syndrome COVID-19 Lung Infected Patients using AP(ai)Bias 2.0: A Systematic Review. IEEE Trans. Instrum. Meas 2022, 1. [Google Scholar] [CrossRef]
- Acharya, U.R.; Joseph, K.P.; Kannathal, N.; Min, L.C.; Suri, J.S. Heart rate variability. In Advances in Cardiac Signal Processing; Springer: Berlin/Heidelberg, Germany, 2007; pp. 121–165. [Google Scholar]
- Suri, J.S.; Bhagawati, M.; Paul, S.; Protogerou, A.D.; Sfikakis, P.P.; Kitas, G.D.; Khanna, N.N.; Ruzsa, Z.; Sharma, A.M.; Saxena, S.; et al. A Powerful Paradigm for Cardiovascular Risk Stratification Using Multiclass, Multi-Label, and Ensemble-Based Machine Learning Paradigms: A Narrative Review. Diagnostics 2022, 12, 722. [Google Scholar] [CrossRef]
- Paul, S.; Maindarkar, M.; Saxena, S.; Saba, L.; Turk, M.; Kalra, M.; Krishnan, P.R.; Suri, J.S. Bias Investigation in Artificial Intelligence Systems for Early Detection of Parkinson’s Disease: A Narrative Review. Diagnostics 2022, 12, 166. [Google Scholar] [CrossRef] [PubMed]
- Suri, J.S.; Bhagawati, M.; Paul, S.; Protogeron, A.; Sfikakis, P.P.; Kitas, G.D.; Khanna, N.N.; Ruzsa, Z.; Sharma, A.M.; Saxena, S.; et al. Understanding the bias in machine learning systems for cardiovascular disease risk assessment: The first of its kind review. Comput. Biol. Med. 2022, 142, 105204. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Nayak, G.; Saba, L.; Kalra, M.; Suri, J.S.; Saxena, S. An artificial intelligence framework and its bias for brain tumor segmentation: A narrative review. Comput. Biol. Med. 2022, 143, 105273. [Google Scholar] [CrossRef] [PubMed]
- El-Baz, A.; Gimel’Farb, G.; Suri, J.S. Stochastic Modeling for Medical Image Analysis; CRC Press: Boca Raton, FL, USA, 2015. [Google Scholar] [CrossRef]
- Acharya, U.R.; Mookiah, M.R.K.; Subbhuraam, V.S.; Yanti, R.; Martis, R.; Saba, L.; Molinari, F.; Guerriero, S.; Suri, J. Evolutionary Algorithm-Based Classifier Parameter Tuning for Automatic Ovarian Cancer Tissue Characterization and Classification. Ultraschall Der Med. Eur. J. Ultrasound 2012, 35, 237–245. [Google Scholar] [CrossRef]
- Agarwal, M.; Agarwal, S.; Saba, L.; Chabert, G.L.; Gupta, S.; Carriero, A.; Pasche, A.; Danna, P.; Mehmedovic, A.; Faa, G.; et al. Eight pruning deep learning models for low storage and high-speed COVID-19 computed tomography lung segmentation and heatmap-based lesion localization: A multicenter study using COVLIAS 2. Comput. Biol. Med. 2022, 146, 105571. [Google Scholar] [CrossRef]
- Saba, L.; Sanagala, S.S.; Gupta, S.K.; Koppula, V.K.; Johri, A.M.; Khanna, N.N.; Mavrogeni, S.; Laird, J.R.; Pareek, G.; Miner, M.; et al. Multimodality carotid plaque tissue characterization and classification in the artificial intelligence paradigm: A narrative review for stroke application. Ann. Transl. Med. 2021, 9, 1206. [Google Scholar] [CrossRef]
- Sanagala, S.S.; Nicolaides, A.; Gupta, S.K.; Koppula, V.K.; Saba, L.; Agarwal, S.; Johri, A.M.; Kalra, M.S.; Suri, J.S. Ten Fast Transfer Learning Models for Carotid Ultrasound Plaque Tissue Characterization in Augmentation Framework Embedded with Heatmaps for Stroke Risk Stratification. Diagnostics 2021, 11, 2109. [Google Scholar] [CrossRef]
- Ikeda, N.; Gupta, A.; Dey, N.; Bose, S.; Shafique, S.; Arak, T.; Godia, E.C.; Saba, L.; Laird, J.R.; Nicolaides, A.N.; et al. Improved correlation between carotid and coronary atherosclerosis SYNTAX score using automated ultrasound carotid bulb plaque IMT measurement. Ultrasound Med. Biol. 2015, 41, 1247–1262. [Google Scholar] [CrossRef]
- Khanna, N.N.; Jamthikar, A.D.; Gupta, D.; Piga, M.; Saba, L.; Carcassi, C.; Giannopoulos, A.A.; Nicolaides, A.; Laird, J.R.; Suri, H.S.; et al. Rheumatoid Arthritis: Atherosclerosis Imaging and Cardiovascular Risk Assessment Using Machine and Deep Learning–Based Tissue Characterization. Curr. Atheroscler. Rep. 2019, 21, 7. [Google Scholar] [CrossRef]
- Suri, J.S.; Puvvula, A.; Biswas, M.; Majhail, M.; Saba, L.; Faa, G.; Singh, I.M.; Oberleitner, R.; Turk, M.; Chadha, P.S.; et al. COVID-19 pathways for brain and heart injury in comorbidity patients: A role of medical imaging and artificial intelligence-based COVID severity classification: A review. Comput. Biol. Med. 2020, 124, 103960. [Google Scholar] [CrossRef]




| Risk Type | - | High-Risk | Low-Risk | - | |||
|---|---|---|---|---|---|---|---|
| PA Threshold | - | GTPA ≥ 40 mm2 | GTPA < 40 mm2 | - | |||
| N | Total (379) | 48 | 331 | - | |||
| Gender | 293(M) | 86(F) | 37(M) | 11(F) | 256(M) | 75(F) | |
| - | Mean | SD | Mean | SD | Mean | SD | p-values |
| Age | 68.78 | 10.88 | 69.69 | 10.71 | 68.64 | 10.91 | 0.54 |
| SBP | 134.62 | 8.88 | 137.1 | 7.133 | 134.3 | 9.061 | 0.04 |
| DBP | 87.31 | 4.44 | 88.54 | 3.567 | 87.13 | 4.531 | 0.04 |
| HbA1c | 6.25 | 1.06 | 6.442 | 1.132 | 6.221 | 1.052 | 0.18 |
| eGFR | 45.67 | 20.43 | 43.77 | 23.35 | 45.95 | 19.99 | 0.49 |
| LDL | 100.84 | 31.48 | 101.6 | 29.01 | 100.7 | 31.87 | 0.87 |
| HDL | 50.60 | 14.73 | 47.63 | 12.63 | 51.03 | 14.98 | 0.14 |
| TC | 174.27 | 36.55 | 174.9 | 36.01 | 174.2 | 36.68 | 0.9 |
| HT | 277 | - | 41 | - | 236 | - | <0.0001 |
| Smoking | 151 | - | 14 | - | 137 | - | <0.0001 |
| FH | 47 | - | 7 | - | 40 | - | <0.0001 |
| Model | Encoder Arm | Bottleneck Layer | Decoder Arm | Intermediate Stages | FC | Classification Layer | Total Parameters |
|---|---|---|---|---|---|---|---|
| UNet | 4685376 | 14157824 | 12188480 | None | 1154 | 6 | 31032840 |
| UNet++ | 4685376 | 14157824 | 12593984 | 1275008 | None | 130 | 32712322 |
| UNet3P | 4685376 | 14157824 | 1196800 | 1447040 | 1154 | 6 | 21488200 |
| Attention-UNet | 4685376 | 14157824 | 12535680 | 523684 | None | 130 | 31902694 |
| Model | Acc | Sens | Spec | Prec | MCC | Dice | Jaccard |
|---|---|---|---|---|---|---|---|
| UNet | 98.58 ± 0.61 | 87.43 ± 5.45 | 99.48 ± 0.40 | 93.11 ± 4.61 | 89.39 ± 3.63 | 90.02 ± 3.53 | 82.03 ± 5.66 |
| UNet++ | 98.51 ± 0.65 | 85.72 ± 6.56 | 99.53 ± 0.38 | 93.70 ± 4.55 | 88.74 ± 4.14 | 89.32 ± 4.16 | 80.94 ± 6.45 |
| UNet3P | 98.51 ± 0.66 | 86.73 ± 6.35 | 99.47 ± 0.41 | 92.90 ± 4.84 | 88.87 ± 3.96 | 89.49 ± 3.90 | 81.19 ± 6.16 |
| Fractal-UNet | 98.41 ± 0.74 | 85.68 ± 6.71 | 99.45 ± 0.43 | 92.61 ± 5.35 | 88.13 ± 4.58 | 88.78 ± 4.50 | 80.10 ± 6.92 |
| Squeeze-UNet | 98.53 ± 0.63 | 86.29 ± 6.01 | 99.51 ± 0.39 | 93.40 ± 4.95 | 88.91 ± 4.09 | 89.52 ± 4.01 | 81.25 ± 6.30 |
| Attention-UNet | 98.58 ± 0.59 | 86.86 ± 5.73 | 99.52 ± 0.38 | 93.54 ± 4.62 | 89.31 ± 3.76 | 89.90 ± 3.69 | 81.86 ± 5.90 |
| Model | Acc | Sens | Spec | Prec | MCC | Dice | Jaccard |
|---|---|---|---|---|---|---|---|
| UNet | 98.97 ± 0.53 | 81.58 ± 8.58 | 99.72 ± 0.27 | 92.20 ± 7.39 | 85.97 ± 5.77 | 86.07 ± 6.17 | 76.01 ± 8.59 |
| UNet++ | 99.05 ± 0.38 | 85.82 ± 7.21 | 99.62 ± 0.31 | 90.04 ± 8.09 | 87.23 ± 5.43 | 87.48 ± 5.57 | 78.14 ± 8.01 |
| UNet3P | 98.93 ± 0.56 | 80.39 ± 9.09 | 99.73 ± 0.26 | 92.42 ± 7.25 | 85.43 ± 6.24 | 85.49 ± 6.68 | 75.18 ± 9.00 |
| Fractal-UNet | 98.91 ± 0.54 | 80.08 ± 8.67 | 99.72 ± 0.27 | 85.17 ± 6.56 | 91.96 ± 7.61 | 85.06 ± 6.29 | 74.68 ± 8.92 |
| Squeeze-UNet | 98.90 ± 0.63 | 79.67 ± 10.45 | 99.73 ± 0.27 | 84.94 ± 7.56 | 92.33 ± 7.09 | 84.94 ± 7.60 | 74.48 ± 9.98 |
| Attention-UNet | 99.01 ± 0.42 | 81.75 ± 7.94 | 99.74 ± 0.26 | 92.71 ± 7.20 | 86.38 ± 5.70 | 86.50 ± 5.94 | 76.65 ± 8.36 |
| ICA | CCA | |||||
|---|---|---|---|---|---|---|
| Model | CC | AUC | p-Values | CC | AUC | p-Values |
| UNet | 0.99 | 0.99 | p < 0.001 | 0.93 | 0.964 | p < 0.001 |
| UNet++ | 0.98 | 0.988 | p < 0.001 | 0.96 | 0.966 | p < 0.001 |
| UNet3P | 0.98 | 0.988 | p < 0.001 | 0.92 | 0.965 | p < 0.001 |
| Fractal-UNet | 0.96 | 0.962 | p < 0.001 | 0.94 | 0.959 | p < 0.001 |
| Squeeze-UNet | 0.96 | 0.969 | p < 0.001 | 0.89 | 0.956 | p < 0.001 |
| Attention-UNet | 0.99 | 0.988 | p < 0.001 | 0.96 | 0.97 | p < 0.001 |
| Models | Mean ± SD | Std Error of Mean | Mean Difference | SD of Differences | Std Error of Mean Difference | 95% CI | Test Statistic-t | p-Value |
|---|---|---|---|---|---|---|---|---|
| UNet | 44.5496 ± 24.2130 | 0.7774 | −2.9449 | 4.5716 | 0.1468 | −3.2330 to −2.6568 | −20.063 | <0.0001 |
| UNet++ | 43.5908 ± 24.2386 | 0.7783 | −3.9037 | 5.0739 | 0.1629 | −4.2234 to −3.5840 | −23.962 | <0.0001 |
| UNet3P | 44.2960 ± 24.1313 | 0.7748 | −3.1985 | 5.3020 | 0.1702 | −3.5326 to −2.8644 | −18.789 | <0.0001 |
| Fractal-UNet | 43.8306 ± 23.6977 | 0.7609 | −3.6639 | 5.4349 | 0.1745 | −4.0063 to −3.3314 | −20.996 | <0.0001 |
| Squeeze-UNet | 43.8814 ± 23.9853 | 0.7701 | −36131 | 4.7915 | 0.1538 | −3.9150 to −3.3112 | −23.485 | <0.0001 |
| Attention-UNet | 44.1855 ± 24.3427 | 0.7816 | −3.3090 | 4.5570 | 0.1463 | −3.5961 to −3.0219 | −22.615 | <0.0001 |
| Mean ± SD | Std Error of Mean | Mean Difference | SD of Differences | Std Error of Mean Difference | 95% CI | Test Statistic-t | p-Value < | |
|---|---|---|---|---|---|---|---|---|
| UNet | 34.0097 ± 15.8775 | 0.6093 | −4.5858 | 7.5137 | 0.2883 | −5.1520 to −4.0197 | −15.904 | 0.0001 |
| UNet++ | 36.6595 ± 18.2781 | 0.7014 | −1.9361 | 5.4177 | 0.2079 | −2.3443 to −1.5278 | −9.312 | 0.0001 |
| UNet3P | 33.4363 ± 15.6883 | 0.6031 | −5.1592 | 7.9247 | 0.3041 | −5.7564 to −4.5621 | −16.964 | 0.0001 |
| Fractal-UNet | 33.5501 ± 16.1022 | 0.6179 | −5.0454 | 7.1421 | 0.2741 | −5.5836 to −4.5073 | −18.408 | 0.0001 |
| Squeeze-UNet | 33.1997 ± 15.5589 | 0.5971 | −5.3958 | 9.0953 | 0.349 | −6.0811 to −4.7105 | −15.459 | 0.0001 |
| Attention-UNet | 34.0731 ± 17.8005 | 0.6831 | −4.5224 | 5.3039 | 0.2035 | −4.9221 to −4.1228 | −22.218 | 0.0001 |
| Authors | Artery Segment | DL Model | #Patients/#Images | Results | Bias Identified |
|---|---|---|---|---|---|
| Zhou et al. [66] | ICA, CCA | UNet++ | N1 = 144/510 N2 = 497/638 | TPA error: 5.55 ± 4.34 mm2 | Data selection, model selection, validation bias |
| Jain et al. [68] | ICA | UNet, UNet+, SegNet, SegNet-UNet, SegNet-UNet+, | N = 97/970 | PA error 3.49 mm2 for SDL; 4.21 mm2 for HDL | Data selection bias |
| Jain et al. [78] | CCA | UNet | N1 = 379 N2 = 300 | FoM of 70.96 and 91.14 (unseen) against 97.57, 88.89, and 99.14 (seen) | Validation bias |
| Jain et al. [73] | CCA | UNet, SegNet-UNet, AtheroEdge 2.0 | N1 = 379 | PA error HDL = 8 mm2 SDL = 9.9 mm2 AtheroEdge 2.0 = 9.6 mm2 | Data selection bias, racial bias |
| Proposed method | ICA, CCA | UNet, UNet++, UNet3P, Attention-UNet | N1 = 970; N2 = 379; N3 = 300 | CC: 0.99 and 0.96 for ICA and CCA experiments | Free from data selection, racial, and validation biases |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jain, P.K.; Dubey, A.; Saba, L.; Khanna, N.N.; Laird, J.R.; Nicolaides, A.; Fouda, M.M.; Suri, J.S.; Sharma, N. Attention-Based UNet Deep Learning Model for Plaque Segmentation in Carotid Ultrasound for Stroke Risk Stratification: An Artificial Intelligence Paradigm. J. Cardiovasc. Dev. Dis. 2022, 9, 326. https://doi.org/10.3390/jcdd9100326
Jain PK, Dubey A, Saba L, Khanna NN, Laird JR, Nicolaides A, Fouda MM, Suri JS, Sharma N. Attention-Based UNet Deep Learning Model for Plaque Segmentation in Carotid Ultrasound for Stroke Risk Stratification: An Artificial Intelligence Paradigm. Journal of Cardiovascular Development and Disease. 2022; 9(10):326. https://doi.org/10.3390/jcdd9100326
Chicago/Turabian StyleJain, Pankaj K., Abhishek Dubey, Luca Saba, Narender N. Khanna, John R. Laird, Andrew Nicolaides, Mostafa M. Fouda, Jasjit S. Suri, and Neeraj Sharma. 2022. "Attention-Based UNet Deep Learning Model for Plaque Segmentation in Carotid Ultrasound for Stroke Risk Stratification: An Artificial Intelligence Paradigm" Journal of Cardiovascular Development and Disease 9, no. 10: 326. https://doi.org/10.3390/jcdd9100326
APA StyleJain, P. K., Dubey, A., Saba, L., Khanna, N. N., Laird, J. R., Nicolaides, A., Fouda, M. M., Suri, J. S., & Sharma, N. (2022). Attention-Based UNet Deep Learning Model for Plaque Segmentation in Carotid Ultrasound for Stroke Risk Stratification: An Artificial Intelligence Paradigm. Journal of Cardiovascular Development and Disease, 9(10), 326. https://doi.org/10.3390/jcdd9100326








