Zoonotic Yersinia enterocolitica in Swine: One Health Implications for Food Safety and Public Health
Simple Summary
Abstract
1. Introduction
2. Taxonomy and General Characteristics of Yersinia
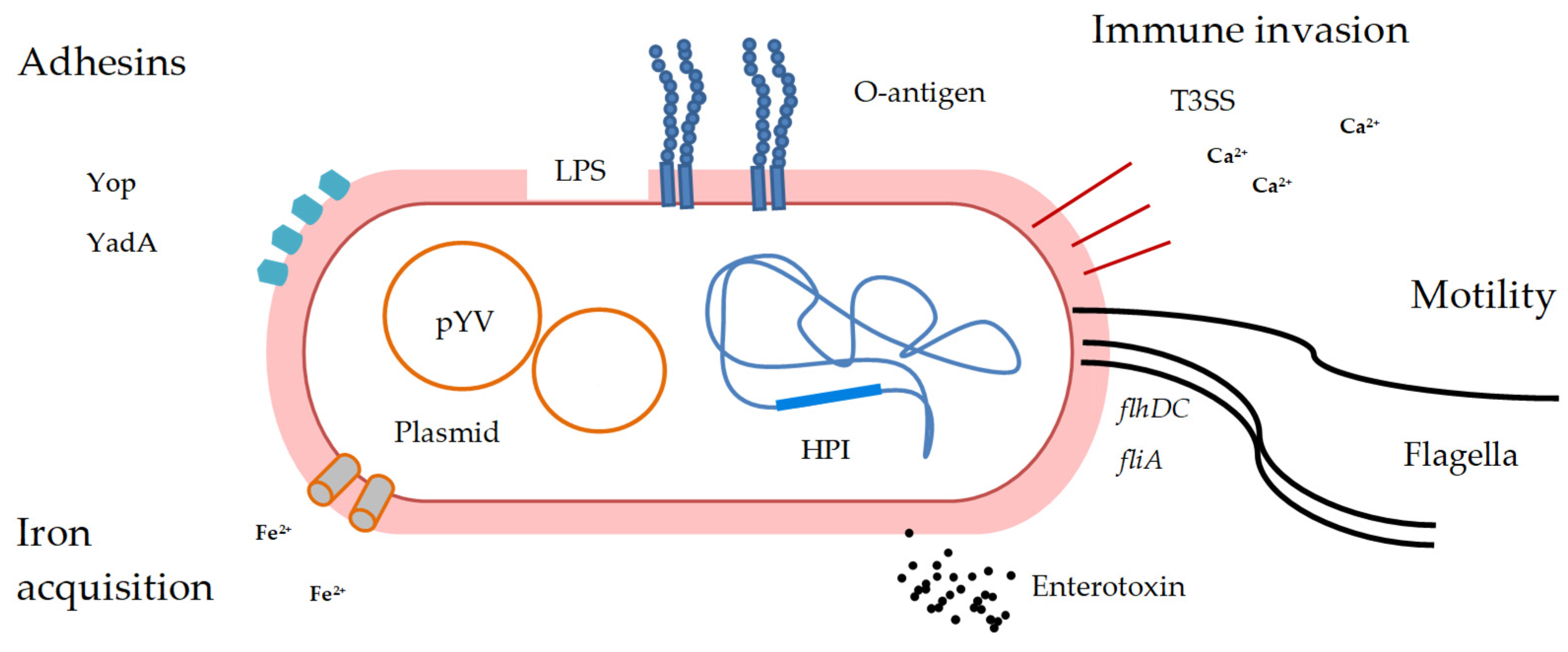
3. Virulence Factors
Role of Key Virulence Genes
4. Prevalence of Y. enterocolitica in Domestic and Wild Pigs
Geographic and Seasonal Trends of Y. enterocolitica
5. One Health Relevance
Foodborne Outbreaks (FBO)
6. Farm-to-Fork Transmission
6.1. Routes of Transmission of Yersinia enterocolitica in Pigs
6.1.1. Slaughter of Pigs
6.1.2. Meat Consumption
6.1.3. Swine Meat Products
6.2. Zoonotic Potential and Foodborne Transmission to Humans
7. Prevention and Control of Y. enterocolitica
7.1. Prevention and Biosecurity on Pig Farms
7.2. Prevention of Y. enterocolitica Infection in Humans
8. Knowledge Gaps and Future Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CFU | Colony Forming Unit |
| DNA | Deoxyribonucleic Acid |
| ECDA | European Centre for Disease Prevention and Control |
| EFSA | European Food Safety Authority |
| EU | European Union |
| FBO | Foodborne outbreak |
| HPI | High-Pathogenicity Island |
| IL-8 | Interleukin-8 |
| ISO | International Organization for Standardization |
| MAPKs | Mitogen-activated protein kinases |
| PCR | Polymerase Chain Reaction |
| PFGE | Pulsed-field gel electrophoresis |
| pYV | Yersinia virulence plasmid |
| RTE | Ready-to-eat |
| STEC | Shiga toxin-producing Escherichia coli |
| T3SS | Type III Secretion System |
| YadA | Yersinia adhesin A protein |
| YopA | Yersinia outer membrane protein |
| Yst | Yersinia stable toxin |
| MLST | Multilocus sequence typing |
References
- Adeolu, M.; Alnajar, S.; Naushad, S.; Gupta, R.S. Genome-Based Phylogeny and Taxonomy of the ‘Enterobacteriales’: Proposal for Enterobacterales Ord. Nov. Divided into the Families Enterobacteriaceae, Erwiniaceae Fam. Nov., Pectobacteriaceae Fam. Nov., Yersiniaceae Fam. Nov., Hafniaceae Fam. Nov., Morganellaceae Fam. Nov., and Budviciaceae Fam. Nov. Int. J. Syst. Evol. Microbiol. 2016, 66, 5575–5599. [Google Scholar] [CrossRef] [PubMed]
- Robins-Browne, R.M. Yersinia enterocolitica. In Food Microbiology Fundamentals and Frontiers; Doyle, M.P., Beauchat, L.R., Montville, T.J., Eds.; ASM Press: Washington, DC, USA, 1997; pp. 192–215. [Google Scholar]
- Moxley, R.A. Family Yersiniaceae. In Veterinary Microbiology; McVey, D.S., Kennedy, M., Chengappa, M.M., Wilkes, R., Eds.; Wiley Blackwell: Hoboken, NJ, USA, 2022; pp. 88–99. [Google Scholar]
- EFSA; ECDC. The European Union One Health 2023 Zoonoses Report. EFSA J. 2024, 22, e9106. [Google Scholar] [CrossRef] [PubMed]
- ECDC European Centre for Disease Prevention and Control. Yersiniosis—Annual Epidemiological Report for 2015. 2018. Available online: https://www.ecdc.europa.eu/en/publications-data/yersiniosis-annual-epidemiological-report-2015 (accessed on 30 June 2025).
- Fredriksson-Ahomaa, M.; Stolle, A.; Siitonen, A.; Korkeala, H. Sporadic Human Yersinia enterocolitica Infections Caused by Bioserotype 4/O:3 Originate Mainly from Pigs. J. Med. Microbiol. 2006, 55, 747–749. [Google Scholar] [CrossRef]
- Stevens, M.J.A.; Barmettler, K.; Kelbert, L.; Stephan, R.; Nüesch-Inderbinen, M. Genome Based Characterization of Yersinia enterocolitica from Different Food Matrices in Switzerland in 2024. Infect. Genet. Evol. 2025, 128, 105719. [Google Scholar] [CrossRef]
- Terentjeva, M.; Ķibilds, J.; Gradovska, S.; Alksne, L.; Streikiša, M.; Meistere, I.; Valciņa, O. Prevalence, Virulence Determinants, and Genetic Diversity in Yersinia enterocolitica Isolated from Slaughtered Pigs and Pig Carcasses. Int. J. Food Microbiol. 2022, 376, 109756. [Google Scholar] [CrossRef]
- Angelovska, M.; Zaharieva, M.M.; Dimitrova, L.L.; Dimova, T.; Gotova, I.; Urshev, Z.; Ilieva, Y.; Kaleva, M.D.; Kim, T.C.; Naydenska, S.; et al. Prevalence, Genetic Homogeneity, and Antibiotic Resistance of Pathogenic Yersinia enterocolitica Strains Isolated from Slaughtered Pigs in Bulgaria. Antibiotics 2023, 12, 716. [Google Scholar] [CrossRef]
- Colbran, C.; May, F.; Alexander, K.; Hunter, I.; Stafford, R.; Bell, R.; Cowdry, A.; Vosti, F.; Jurd, S.; Graham, T.; et al. Yersiniosis Outbreaks in Gold Coast Residential Aged Care Facilities Linked to Nutritionally-Supplemented Milkshakes, January–April 2023. Commun. Dis. Intell. 2024, 48, 1–8. [Google Scholar] [CrossRef]
- Isobe, J.; Kimata, K.; Shimizu, M.; Kanatani, J.I.; Sata, T.; Watahiki, M. Water-Borne Outbreak of Yersinia enterocolitica O8 Due to a Small Scale Water System. Kansenshogaku Zasshi 2014, 88, 827–832. [Google Scholar] [CrossRef]
- Lee, T.S.; Lee, S.W.; Seok, W.S.; Yoo, M.Y.; Yoon, J.W.; Park, B.K.; Moon, K.D.; Oh, D.H. Prevalence, Antibiotic Susceptibility, and Virulence Factors of Yersinia enterocolitica and Related Species from Ready-to-Eat Vegetables Available in Korea. J. Food Prot. 2004, 67, 1123–1127. [Google Scholar] [CrossRef] [PubMed]
- Syczyło, K.; Platt-Samoraj, A.; Bancerz-Kisiel, A.; Szczerba-Turek, A.; Pajdak-Czaus, J.; Łabuć, S.; Procajło, Z.; Socha, P.; Chuzhebayeva, G.; Szweda, W. The Prevalence of Yersinia enterocolitica in Game Animals in Poland. PLoS ONE 2018, 13, e0195136. [Google Scholar] [CrossRef]
- Fredriksson-Ahomaa, M.; Grönthal, T.; Heljanko, V.; Johansson, V.; Rantala, M.; Heikinheimo, A.; Laukkanen-Ninios, R. Enteropathogenic Yersinia with Public Health Relevance Found in Dogs and Cats in Finland. Pathogens 2024, 13, 54. [Google Scholar] [CrossRef]
- Sannö, A.; Rosendal, T.; Aspán, A.; Backhans, A.; Jacobson, M. Comparison of Multiple-Locus Variable-Number Tandem Repeat Analysis Profiles of Enteropathogenic Yersinia Spp. Obtained from Humans, Domestic Pigs, Wild Boars, Rodents, Pork and Dog Food. Animals 2023, 13, 3055. [Google Scholar] [CrossRef]
- Liang, J.; Duan, R.; Xia, S.; Hao, Q.; Yang, J.; Xiao, Y.; Qiu, H.; Shi, G.; Wang, S.; Gu, W.; et al. Ecology and Geographic Distribution of Yersinia enterocolitica among Livestock and Wildlife in China. Vet. Microbiol. 2015, 178, 125–131. [Google Scholar] [CrossRef]
- McIver, M.A.; Pike, R.M. Chronic Glanders-like Infection of Face Caused by an Organism Resembling Flavobacterium Pseudomallei Whitmore. Clin. Misc. 1934, 1, 16–21. [Google Scholar]
- Bottone, E.J. Yersinia enterocolitica: The Charisma Continues. Clin. Microbiol. Rev. 1997, 10, 257–276. [Google Scholar] [CrossRef]
- Hordofa, D.L.; Nuguse, A. Review on Yersiniosis and Its Public Health Importance. Int. J. Clin. Biol. Biochem. 2023, 5, 37–41. [Google Scholar] [CrossRef]
- Arsić, M.; Vićić, I.; Petrović, M.; Dmitrić, M.; Karabasil, N. Yersinia enterocolitica and Control Measures for Reducing Risks in the Pork Production Chain. Meat Technol. 2023, 64, 237–241. [Google Scholar] [CrossRef]
- Lobato, M.J.; Landeras, E.; González-Hevia, M.A.; Mendoza, M.C. Genetic Heterogeneity of Clinical Strains of Yersinia enterocolitica Traced by Ribotyping and Relationships between Ribotypes, Serotypes, and Biotypes. J. Clin. Microbiol. 1998, 36, 3297–3302. [Google Scholar] [CrossRef]
- Bancerz-Kisiel, A.; Pieczywek, M.; Łada, P.; Szweda, W. The Most Important Virulence Markers of Yersinia enterocolitica and Their Role during Infection. Genes 2018, 9, 235. [Google Scholar] [CrossRef]
- Fàbrega, A.; Vila, J. Yersinia enterocolitica: Pathogenesis, Virulence and Antimicrobial Resistance. Enferm. Infecc. Microbiol. Clin. 2012, 30, 24–32. [Google Scholar] [CrossRef]
- Kiani, P.; Bakhshi, B.; Soltan-Dallal, M.M.; Najar-Peerayeh, S. Heterogeneity of Highly Susceptible Yersinia enterocolitica Isolates of Clinical and Environmental Origin: A 5-Year Survey from Iran (2011–2016). Microb. Drug Resist. 2020, 26, 46–53. [Google Scholar] [CrossRef]
- Miller, V.L.; Beer, K.B.; Heusipp, G.; Young, B.M.; Wachtel, M.R. Identification of Regions of Ail Required for the Invasion and Serum Resistance Phenotypes. Mol. Microbiol. 2001, 41, 1053–1062. [Google Scholar] [CrossRef]
- Ventola, E.; Lovari, S.; Farneti, S.; Finazzi, G.; Bilei, S.; Owczarek, S.; Delibato, E. Molecular Characterization of Yersinia enterocolitica Strains to Evaluate Virulence Associated Genes. Ann. Ist. Super. Sanita. 2024, 59, 280–285. [Google Scholar] [CrossRef]
- Imori, P.F.M.; Passaglia, J.; Souza, R.A.; Rocha, L.B.; Falcão, J.P. Virulence-Related Genes, Adhesion and Invasion of Some Yersinia enterocolitica-like Strains Suggests Its Pathogenic Potential. Microb. Pathog. 2017, 104, 72–77. [Google Scholar] [CrossRef]
- Marggraf, M.; Barac, A.; Hammerl, J.A.; Hertwig, S. Improvement of the EN ISO 10273:2017 Method for the Cultural Detection of Yersinia enterocolitica in Meat. Food Microbiol. 2024, 117, 104384. [Google Scholar] [CrossRef]
- Hallanvuo, S.; Herranen, M.; Jaakkonen, A.; Nummela, M.; Ranta, J.; Botteldoorn, N.; De Zutter, L.; Fredriksson-Ahomaa, M.; Hertwig, S.; Johannessen, G.S.; et al. Validation of EN ISO Method 10273—Detection of Pathogenic Yersinia enterocolitica in Foods. Int. J. Food Microbiol. 2019, 288, 66–74. [Google Scholar] [CrossRef]
- Mancusi, A.; Delibato, E.; Francesca Peruzy, M.; Girardi, S.; Di Maro, O.; Cristiano, D.; Ventola, E.; Dini, I.; Thérèse Rose Proroga, Y. Evaluation of Two Real-Time PCR Methods to Detect Yersinia enterocolitica in Bivalve Molluscs Collected in Campania Region. Food Res. Int. 2024, 192, 114789. [Google Scholar] [CrossRef]
- Duan, R.; Liang, J.; Shi, G.; Cui, Z.; Hai, R.; Wang, P.; Xiao, Y.; Li, K.; Qiu, H.; Gu, W.; et al. Homology Analysis of Pathogenic Yersinia Species Yersinia enterocolitica, Yersinia I, and Yersinia pestis Based on Multilocus Sequence Typing. J. Clin. Microbiol. 2014, 52, 20–29. [Google Scholar] [CrossRef]
- Savin, C.; Criscuolo, A.; Guglielmini, J.; Le Guern, A.S.; Carniel, E.; Pizarro-Cerdá, J.; Brisse, S. Genus-Wide Yersinia Core-Genome Multilocus Sequence Typing for Species Identification and Strain Characterization. Microb. Genom 2019, 5, e000301. [Google Scholar] [CrossRef]
- Forsythe, S.; Sutherland, J.; Varnam, A. Yersinia, Shigella, Vibrio, Aeromonas, Plesiomonas, Cronobacter, Enterobacter, Klebsiella and Citrobacter. In Foodborne Pathogens; Elsevier: Amsterdam, The Netherlands, 2009; pp. 763–801. [Google Scholar]
- Wauters, G.; Kandolo, K.; Janssens, M. Revised Biogrouping Scheme of Yersinia enterocolitica. Contrib. Microbiol. Immunol. 1987, 9, 14–21. [Google Scholar]
- Liang, J.; Kou, Z.; Qin, S.; Chen, Y.; Li, Z.; Li, C.; Duan, R.; Hao, H.; Zha, T.; Gu, W.; et al. Novel Yersinia enterocolitica Prophages and a Comparative Analysis of Genomic Diversity. Front. Microbiol. 2019, 10, 1184. [Google Scholar] [CrossRef] [PubMed]
- Bottone, E.J. Yersinia enterocolitica: Revisitation of an Enduring Human Pathogen. Clin. Microbiol. Newsl. 2015, 37, 1–8. [Google Scholar] [CrossRef]
- Platt-Samoraj, A. Toxigenic Properties of Yersinia enterocolitica Biotype 1A. Toxins 2022, 14, 118. [Google Scholar] [CrossRef] [PubMed]
- Rivas, L.; Strydom, H.; Paine, S.; Wang, J.; Wright, J. Yersiniosis in New Zealand. Pathogens 2021, 10, 191. [Google Scholar] [CrossRef]
- Bengoechea, J.A.; Najdenski, H.; Skurnik, M. Lipopolysaccharide O Antigen Status of Yersinia enterocolitica O:8 Is Essential for Virulence and Absence of O Antigen Affects the Expression of Other Yersinia Virulence Factors. Mol. Microbiol. 2004, 52, 451–469. [Google Scholar] [CrossRef] [PubMed]
- Skurnik, M.; Bengoechea, J.A. The Biosynthesis and Biological Role of Lipopolysaccharide O-Antigens of Pathogenic Yersiniae. Carbohydr. Res. 2003, 338, 2521–2529. [Google Scholar] [CrossRef]
- Longenberger, A.H.; Gronostaj, M.P.; Yee, G.Y.; Johnson, L.M.; Lando, J.F.; Voorhees, R.E.; Waller, K.; Weltman, A.C.; Moll, M.; Lyss, S.B.; et al. Yersinia enterocolitica Infections Associated with Improperly Pasteurized Milk Products: Southwest Pennsylvania, March–August, 2011. Epidemiol. Infect. 2014, 142, 1640–1650. [Google Scholar] [CrossRef]
- Espenhain, L.; Riess, M.; Müller, L.; Colombe, S.; Ethelberg, S.; Litrup, E.; Jernberg, C.; Kühlmann-Berenzon, S.; Lindblad, M.; Hove, N.K.; et al. Cross-Border Outbreak of Yersinia enterocolitica O3 Associated with Imported Fresh Spinach, Sweden and Denmark, March 2019. Eurosurveillance 2019, 24, 1900368. [Google Scholar] [CrossRef]
- Singh, I.; Virdi, J.S. Production of Yersinia Stable Toxin (YST) and Distribution of Yst Genes in Biotype 1A Strains of Yersinia enterocolitica. J. Med. Microbiol. 2004, 53, 1065–1068. [Google Scholar] [CrossRef]
- Mühlenkamp, M.; Oberhettinger, P.; Leo, J.C.; Linke, D.; Schütz, M.S. Yersinia Adhesin A (YadA)—Beauty & Beast. Int. J. Med. Microbiol. 2015, 305, 252–258. [Google Scholar]
- Heise, T.; Dersch, P. Identification of a Domain in Yersinia Virulence Factor YadA That Is Crucial for Extracellular Matrix-Specific Cell Adhesion and Uptake. Proc. Natl. Acad. Sci. USA 2006, 103, 3375–3380. [Google Scholar] [CrossRef]
- Bhagat, N.; Virdi, J.S. The Enigma of Yersinia enterocolitica Biovar 1A. Crit. Rev. Microbiol. 2011, 37, 25–39. [Google Scholar] [CrossRef]
- Atkinson, S.; Williams, P. Yersinia Virulence Factors—A Sophisticated Arsenal for Combating Host Defences. F1000Research 2016, 5, F1000 Faculty Rev-1370. [Google Scholar] [CrossRef]
- von Tils, D.; Blädel, I.; Schmidt, M.A.; Heusipp, G. Type II Secretion in Yersinia—A Secretion System for Pathogenicity and Environmental Fitness. Front. Cell. Infect. Microbiol. 2012, 2, 160. [Google Scholar] [CrossRef]
- Gkouletsos, T.; Patas, K.; Lambrinidis, G.; Neubauer, H.; Sprague, L.D.; Ioannidis, A.; Chatzipanagiotou, S. Antimicrobial Resistance of Yersinia enterocolitica and Presence of Plasmid PYV Virulence Genes in Human and Animal Isolates. New Microbes New Infect. 2019, 32, 100604. [Google Scholar] [CrossRef] [PubMed]
- Peruzy, M.F.; Murru, N.; Perugini, A.G.; Capuano, F.; Delibato, E.; Mercogliano, R.; Korkeala, H.; Proroga, Y.T.R. Evaluation of Virulence Genes in Yersinia enterocolitica Strains Using SYBR Green Real-Time PCR. Food Microbiol. 2017, 65, 231–235. [Google Scholar] [CrossRef]
- Yue, Y.; Zheng, J.; Sheng, M.; Liu, X.; Hao, Q.; Zhang, S.; Xu, S.; Liu, Z.; Hou, X.; Jing, H.; et al. Public Health Implications of Yersinia enterocolitica Investigation: An Ecological Modeling and Molecular Epidemiology Study. Infect. Dis. Poverty 2023, 12, 41. [Google Scholar] [CrossRef]
- Morka, K.; Bystroń, J.; Bania, J.; Korzeniowska-Kowal, A.; Korzekwa, K.; Guz-Regner, K.; Bugla-Płoskońska, G. Identification of Yersinia enterocolitica Isolates from Humans, Pigs and Wild Boars by MALDI TOF MS. BMC Microbiol. 2018, 18, 86. [Google Scholar] [CrossRef] [PubMed]
- Sannö, A.; Rosendal, T.; Aspán, A.; Backhans, A.; Jacobson, M. Distribution of Enteropathogenic Yersinia Spp. and Salmonella Spp. in the Swedish Wild Boar Population, and Assessment of Risk Factors That May Affect Their Prevalence. Acta Vet. Scand. 2018, 60, 40. [Google Scholar] [CrossRef] [PubMed]
- Gürtler, M.; Alter, T.; Kasimir, S.; Linnebur, M.; Fehlhaber, K. Prevalence of Yersinia enterocolitica in Fattening Pigs. J. Food Prot. 2005, 68, 850–854. [Google Scholar] [CrossRef]
- Zdolec, N.; Kiš, M.; Jankuloski, D.; Blagoevska, K.; Kazazić, S.; Pavlak, M.; Blagojević, B.; Antić, D.; Fredriksson-Ahomaa, M.; Pažin, V. Prevalence and Persistence of Multidrug-Resistant Yersinia enterocolitica 4/O:3 in Tonsils of Slaughter Pigs from Different Housing Systems in Croatia. Foods 2022, 11, 1459. [Google Scholar] [CrossRef]
- Fredriksson-Ahomaa, M.; Wacheck, S.; Bonke, R.; Stephan, R. Different Enteropathogenic Yersinia Strains Found in Wild Boars and Domestic Pigs. Foodborne Pathog. Dis. 2011, 8, 733–737. [Google Scholar] [CrossRef]
- De Boer, E.; Zwartkruis-Nahuis, J.T.M.; Lesuis, R. Prevalentie Humaanpathogene Yersinia enterocolitica Bij Varkens. Tijdschr Diergeneeskd 2008, 133, 158–161. [Google Scholar]
- Bancerz-Kisiel, A.; Platt-Samoraj, A.; Szczerba-Turek, A.; SyczyŁo, K.; Szweda, W. The First Pathogenic Yersinia enterocolitica Bioserotype 4/O: 3 Strain Isolated from a Hunted Wild Boar (Sus scrofa) in Poland. Epidemiol. Infect. 2015, 143, 2758–2765. [Google Scholar] [CrossRef] [PubMed]
- von Altrock, A.; Seinige, D.; Kehrenberg, C. Yersinia enterocolitica Isolates from Wild Boars Hunted in Lower Saxony, Germany. Appl. Environ. Microbiol. 2015, 81, 4835–4840. [Google Scholar] [CrossRef] [PubMed]
- Lorencova, A.; Babak, V.; Lamka, J. Serological Prevalence of Enteropathogenic Yersinia Spp. in Pigs and Wild Boars from Different Production Systems in the Moravian Region, Czech Republic. Foodborne Pathog. Dis. 2016, 13, 275–279. [Google Scholar] [CrossRef]
- Bhaduri, S.; Wesley, I.V.; Bush, E.J. Prevalence of Pathogenic Yersinia enterocolitica Strains in Pigs in the United States. Appl. Environ. Microbiol. 2005, 71, 7117–7121. [Google Scholar] [CrossRef] [PubMed]
- Bowman, A.S.; Glendening, C.; Wittum, T.E.; LeJeune, J.T.; Stich, R.W.; Funk, J.A. Prevalence of Yersinia enterocolitica in Different Phases of Production on Swine Farms. J. Food Prot. 2007, 70, 11–16. [Google Scholar] [CrossRef]
- Bancerz-Kisiel, A.; Socha, P.; Szweda, W. Detection and Characterisation of Yersinia enterocolitica Strains in Cold-Stored Carcasses of Large Game Animals in Poland. Vet. J. 2016, 208, 102–103. [Google Scholar] [CrossRef]
- Arrausi-Subiza, M.; Gerrikagoitia, X.; Alvarez, V.; Ibabe, J.C.; Barral, M. Prevalence of Yersinia enterocolitica and Yersinia pseudotuberculosis in Wild Boars in the Basque Country, Northern Spain. Acta Vet. Scand. 2016, 58, 4. [Google Scholar] [CrossRef]
- Bonardi, S.; Brémont, S.; Vismarra, A.; Poli, I.; Diegoli, G.; Bolzoni, L.; Corradi, M.; Gilioli, S.; Le Guern, A.S. Is Yersinia Bercovieri Surpassing Yersinia enterocolitica in Wild Boars (Sus scrofa)? Ecohealth 2020, 17, 388–392. [Google Scholar] [CrossRef] [PubMed]
- Modesto, P.; De Ciucis, C.G.; Vencia, W.; Pugliano, M.C.; Mignone, W.; Berio, E.; Masotti, C.; Ercolini, C.; Serracca, L.; Andreoli, T.; et al. Evidence of Antimicrobial Resistance and Presence of Pathogenicity Genes in Yersinia enterocolitica Isolate from Wild Boars. Pathogens 2021, 10, 398. [Google Scholar] [CrossRef] [PubMed]
- Fredriksson-Ahomaa, M.; London, L.; Skrzypczak, T.; Kantala, T.; Laamanen, I.; Biström, M.; Maunula, L.; Gadd, T. Foodborne Zoonoses Common in Hunted Wild Boars. Ecohealth 2020, 17, 512–522. [Google Scholar] [CrossRef] [PubMed]
- Pilon, J.; Higgins, R.; Quessy, S. Epidemiological Study of Yersinia enterocolitica in Swine Herds in Québec. Can. Vet. J. 2000, 41, 383–387. [Google Scholar]
- Liang, J.; Wang, X.; Xiao, Y.; Cui, Z.; Xia, S.; Hao, Q.; Yang, J.; Luo, L.; Wang, S.; Li, K.; et al. Prevalence of Yersinia enterocolitica in Pigs Slaughtered in Chinese Abattoirs. Appl. Environ. Microbiol. 2012, 78, 2949–2956. [Google Scholar] [CrossRef]
- Fondrevez, M.; Minvielle, B.; Labbé, A.; Houdayer, C.; Rose, N.; Esnault, E.; Denis, M. Prevalence of Pathogenic Yersinia enterocolitica in Slaughter-Aged Pigs during a One-Year Survey, 2010–2011, France. Int. J. Food Microbiol. 2014, 174, 56–62. [Google Scholar] [CrossRef]
- Ibanez, T.R.; Laukkanen-Ninios, R.; Hakkinen, M.; Johansson, T.; Vilar, M.; Korkeala, H. Prevalence of Pathogenic Yersinia enterocolitica in Finnish Slaughter Pigs. J. Food Prot. 2016, 79, 677–681. [Google Scholar] [CrossRef]
- Pugazhenthi, T.R.; Elango, A.; Naresh Kumar, C.; Dhanalakshmi, B. Seasonwise Prevalence of Yersinia Species Isolated from Milk and Milk Products in Chennai. Indian Vet. J. 2013, 90, 29–31. [Google Scholar]
- Pal, M. Yersinia enterocolitica as an Important Pathogen of Food Safety Concern. J. Exp. Food Chem. 2018, 4, e111. [Google Scholar] [CrossRef]
- Galvis, J.C.; Beltrán, C.; Sierra, E.; Montaño, A.M.; Ramírez, F.; Villa, P.; Diaz, A.; Robledo, C.; Robledo, J. Outbreak of Gastrointestinal Infections Caused by Yersinia enterocolitica in Children, Medellín, Colombia. J. Pediatr. Infect. Dis. 2023, 18, 289–294. [Google Scholar] [CrossRef]
- Alnabwani, D.; Durrani, M.; Prasad, A.; Pandya, S.; Ghodasara, K.; Hasan, B.I.; Greenberg, A.; Cheriyath, P. Yersinia enterocolitica Sepsis in an Elderly Male with No Iron Overload: A Case Report From the Northeastern United States. Cureus 2022, 14, e26431. [Google Scholar] [CrossRef]
- Aziz, M.; Yelamanchili, V.S. Yersinia enterocolitica; StatPearls Publishing: Treasure Island, FL, USA, 2023. Available online: https://www.ncbi.nlm.nih.gov/books/NBK499837/ (accessed on 30 June 2025).
- Honda, K.; Iwanaga, N.; Izumi, Y.; Tsuji, Y.; Kawahara, C.; Michitsuji, T.; Higashi, S.; Kawakami, A.; Migita, K. Reactive Arthritis Caused by Yersinia enterocolitica Enteritis. Intern. Med. 2017, 56, 1239–1242. [Google Scholar] [CrossRef]
- EFSA; ECDC. The European Union One Health 2022 Zoonoses Report. EFSA J. 2023, 21, e8442. [Google Scholar] [CrossRef]
- EFSA; ECDC. The European Union One Health 2021 Zoonoses Report. EFSA J. 2022, 20, e05926. [Google Scholar] [CrossRef]
- Black, R.E.; Jackson, R.J.; Tsai, T.; Medvesky, M.; Shayegani, M.; Feeley, J.C.; MacLeod, K.I.E.; Wakelee, A.M. Epidemic Yersinia enterocolitica Infection Due to Contaminated Chocolate Milk. N. Engl. J. Med. 1978, 298, 76–79. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, T. Yersinia enterocolitica Infection in Humans and Isolation of the Microorganism from Pigs in Japan. Contrib. Microbiol. Immunol. 1987, 9, 48–55. [Google Scholar] [PubMed]
- Fenwick, S.G.; McCarthy, M.D. Yersinia enterocolitica Is a Common Cause of Gastroenteritis in Auckland. N. Z. Med. J. 1995, 108, 269–271. [Google Scholar]
- Lee, L.A.; Gerber, A.R.; Lonsway, D.R.; Smith, J.D.; Carter, G.P.; Puhr, N.D.; Parrish, C.M.; Sikes, R.K.; Finton, R.J.; Tauxe, R.V. Yersinia enterocolitica O:3 Infections in Infants and Children, Associated with the Household Preparation of Chitterlings. N. Engl. J. Med. 1990, 322, 984–987. [Google Scholar] [CrossRef] [PubMed]
- Ackers, M.L.; Schoenfeld, S.; Markman, J.; Smith, M.G.; Nicholson, M.A.; DeWitt, W.; Cameron, D.N.; Griffin, P.M.; Slutsker, L. An Outbreak of Yersinia enterocolitica O:8 Infections Associated with Pasteurized Milk. J. Infect. Dis. 2000, 181, 1834–1837. [Google Scholar] [CrossRef]
- Sakai, T.; Nakayama, A.; Hashida, M.; Yamamoto, Y.; Takebe, H.; Imai, S. Outbreak of Food Poisoning by Yersinia enterocolitica Serotype O8 in Nara Prefecture: The First Case Report in Japan. Jpn. J. Infect. Dis. 2005, 58, 257–258. [Google Scholar] [CrossRef]
- Grahek-Ogden, D.; Schimmer, B.; Cudjoe, K.S.; Nygård, K.; Kapperud, G. Outbreak of Yersinia enterocolitica Serogroup O:9 Infection and Processed Pork, Norway. Emerg. Infect. Dis. 2007, 13, 754. [Google Scholar] [CrossRef] [PubMed]
- Moriki, S.; Nobata, A.; Shibata, H.; Nagai, A.; Minami, N.; Taketani, T.; Fukushima, H. Familial Outbreak of Yersinia enterocolitica Serotype O9 Biotype 2. J. Infect. Chemother. 2010, 16, 56–58. [Google Scholar] [CrossRef] [PubMed]
- Rosner, B.M.; Stark, K.; Höhle, M.; Werber, D. Risk Factors for Sporadic Yersinia enterocolitica Infections, Germany 2009–2010. Epidemiol. Infect. 2012, 140, 1738–1747. [Google Scholar] [CrossRef] [PubMed]
- Duan, R.; Liang, J.; Zhang, J.; Chen, Y.; Wang, J.; Tong, J.; Guo, B.; Hu, W.; Wang, M.; Zhao, J.; et al. Prevalence of Yersinia enterocolitica Bioserotype 3/O:3 among Children with Diarrhea, China, 2010–2015. Emerg. Infect. Dis. 2017, 23, 1502. [Google Scholar] [CrossRef]
- MacDonald, E.; Heier, B.T.; Nygård, K.; Stalheim, T.; Cudjoe, K.S.; Skjerdal, T.; Wester, A.L.; Lindstedt, B.A.; Stavnes, T.L.; Vold, L. Yersinia enterocolitica Outbreak Associated with Ready-to-Eat Salad Mix, Norway, 2011. Emerg. Infect. Dis. 2012, 18, 1496. [Google Scholar] [CrossRef]
- MacDonald, E.; Einöder-Moreno, M.; Borgen, K.; Brandal, L.T.; Diab, L.; Fossli, Ø.; Herrador, B.G.; Hassan, A.A.; Johannessen, G.S.; Johansen, E.J.; et al. National Outbreak of Yersinia enterocolitica Infections in Military and Civilian Populations Associated with Consumption of Mixed Salad, Norway, 2014. Eurosurveillance 2016, 21, 30321. [Google Scholar] [CrossRef]
- Konishi, N.; Ishitsuka, R.; Yokoyama, K.; Saiki, D.; Akase, S.; Monma, C.; Hirai, A.; Sadamasu, K.; Kai, A. Two Outbreaks of Yersinia enterocolitica O:8 Infections in Tokyo and the Characterization of Isolates. Kansenshogaku Zasshi 2016, 90, 66–72. [Google Scholar] [CrossRef]
- Le Guern, A.S.; Martin, L.; Savin, C.; Carniel, E. Yersiniosis in France: Overview and Potential Sources of Infection. Int. J. Infect. Dis. 2016, 46, 1–7. [Google Scholar] [CrossRef]
- Špačková, M.; Daniel, O.; Klimešová, P.; Ileninová, Z. Overview of Basic Epidemiological Characteristics and Descriptive Analysis of the Incidence of Human Yersiniosis in the Czech Republic in 2018–2020. Epidemiol. Mikrobiol. Imunol. 2022, 71, 32–39. [Google Scholar]
- Karlsson, P.A.; Tano, E.; Jernberg, C.; Hickman, R.A.; Guy, L.; Järhult, J.D.; Wang, H. Molecular Characterization of Multidrug-Resistant Yersinia enterocolitica From Foodborne Outbreaks in Sweden. Front. Microbiol. 2021, 12, 664665. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA). EU One Health Zoonoses Report 2020. EFSA J. 2021, 19, e06971. [Google Scholar]
- Virtanen, S.; Laukkanen-Ninios, R.; Martínez, P.O.; Siitonen, A.; Fredriksson-Ahomaa, M.; Korkeala, H. Multiple-Locus Variable-Number Tandem-Repeat Analysis in Genotyping Yersinia enterocolitica Strains from Human and Porcine Origins. J. Clin. Microbiol. 2013, 51, 2154–2159. [Google Scholar] [CrossRef]
- Vilar, M.J.; Virtanen, S.; Laukkanen-Ninios, R.; Korkeala, H. Bayesian Modelling to Identify the Risk Factors for Yersinia enterocolitica Contamination of Pork Carcasses and Pluck Sets in Slaughterhouses. Int. J. Food. Microbiol. 2015, 197, 53–57. [Google Scholar] [CrossRef]
- Esnault, E.; Rouaud, A.; Labbé, A.; Houdayer, C.; Bailly, Y.; Houard, E.; Bougeard, S.; Paboeuf, F.; Eterradossi, N.; Chemaly, M.; et al. Controlled Experimental Infection in Pigs with a Strain of Yersinia enterocolitica Harboring Genetic Markers for Human Pathogenicity: Colonization and Stability. Infect. Immun. 2023, 91, e00157-23. [Google Scholar] [CrossRef]
- Arsić, M.; Vićić, I.; Galić, N.; Dmitrić, M.; Kureljušić, J.; Dimitrijević, M.; Petrović, M.; Šarić, L.; Karabasil, N. Risk Factors and the Overall Characterization of Yersinia enterocolitica as an Initial Model of Pathogen Surveillance in the Pig Production System in Serbia. Res. Vet. Sci. 2022, 152, 167–174. [Google Scholar] [CrossRef]
- Kruse, H.; Kapperud, G.; Lassen, J.; Lunestad, B.T.; Nesbakken, T.; Rimstad, E.; Robertson, L.; Skjerve, E.; Wasteson, Y. A Preliminary Risk Assessment of Yersinia enterocolitica in the Food Chain: Some Aspects Related to Human Health in Norway. Eur. J. Nutr. Food Saf. 2021, 13, 1–2. [Google Scholar] [CrossRef]
- Virtanen, S.E.; Salonen, L.K.; Laukkanen, R.; Hakkinen, M.; Korkeala, H. Factors Related to the Prevalence of Pathogenic Yersinia enterocolitica on Pig Farms. Epidemiol. Infect. 2011, 139, 1919–1927. [Google Scholar] [CrossRef]
- Laukkanen-Ninios, R.; Fredriksson-Ahomaa, M.; Korkeala, H. Enteropathogenic Yersinia in the Pork Production Chain: Challenges for Control. Compr. Rev. Food Sci. Food Saf. 2014, 13, 1165–1191. [Google Scholar] [CrossRef]
- Buder, C.; Meemken, D.; Fürstenberg, R.; Langforth, S.; Kirse, A.; Langkabel, N. Drinking Pipes and Nipple Drinkers in Pig Abattoir Lairage Pens—A Source of Zoonotic Pathogens as a Hazard to Meat Safety. Microorganisms 2023, 11, 2554. [Google Scholar] [CrossRef]
- Laukkanen, R.; Martínez, P.O.; Siekkinen, K.M.; Ranta, J.; Maijala, R.; Korkeala, H. Contamination of Carcasses with Human Pathogenic Yersinia enterocolitica 4/O:3 Originates from Pigs Infected on Farms. Foodborne Pathog. Dis. 2009, 6, 681–688. [Google Scholar] [CrossRef]
- Vanantwerpen, G.; Berkvens, D.; De Zutter, L.; Houf, K. Association between Microbiological and Serological Prevalence of Human Pathogenic Yersinia Spp. in Pigs and Pig Batches. Vet. Microbiol. 2015, 178, 114–118. [Google Scholar] [CrossRef]
- Wesley, I.V.; Bhaduri, S.; Bush, E. Prevalence of Yersinia enterocolitica in Market Weight Hogs in the United States. J. Food Prot. 2008, 71, 1162–1168. [Google Scholar] [CrossRef]
- Guillier, L.; Fravalo, P.; Leclercq, A.; Thébault, A.; Kooh, P.; Cadavez, V.; Gonzales-Barron, U. Risk Factors for Sporadic Yersinia enterocolitica Infections: A Systematic Review and Meta-Analysis. Microb. Risk Anal. 2021, 17, 100141. [Google Scholar] [CrossRef]
- Farzan, A.; Friendship, R.M.; Cook, A.; Pollari, F. Monitoring of Salmonella, Campylobacter, Yersinia enterocolitica, E. coli 0157, and Listeria monocytogenes on a Subset of Canadian Swine Farms. In Proceedings of the Eighth International Symposium on the Epidemiology and Control of Foodborne Pathogens in Pork, Québec, QC, Canada, 30 September–2 October 2009. [Google Scholar]
- Vanantwerpen, G.; Berkvens, D.; Van Damme, I.; De Zutter, L.; Houf, K. Assessment of Risk Factors for a High Within-Batch Prevalence of Yersinia enterocolitica in Pigs Based on Microbiological Analysis at Slaughter. Foodborne Pathog. Dis. 2015, 12, 571–575. [Google Scholar] [CrossRef]
- Schaake, J.; Kronshage, M.; Uliczka, F.; Rohde, M.; Knuuti, T.; Strauch, E.; Fruth, A.; Wos-Oxley, M.; Dersch, P. Human and Animal Isolates of Yersinia enterocolitica Show Significant Serotype-Specific Colonization and Host-Specific Immune Defense Properties. Infect. Immun. 2013, 81, 4013–4025. [Google Scholar] [CrossRef] [PubMed]
- Robins-Browne, R.M.; Tzipori, S.; Gonis, G.; Hayes, J.; Withers, M.; Prpic, J.K. The Pathogenesis of Yersinia enterocolitica Infection in Gnotobiotic Piglets. J. Med. Microbiol. 1985, 19, 297–308. [Google Scholar] [CrossRef]
- Joutsen, S.; Johansson, P.; Laukkanen-Ninios, R.; Björkroth, J.; Fredriksson-Ahomaa, M. Two Copies of the Ail Gene Found in Yersinia enterocolitica and Yersinia Kristensenii. Vet. Microbiol. 2020, 247, 108798. [Google Scholar] [CrossRef] [PubMed]
- Niwa, T.; Ikeuchi, S.; Bui, T.H.; Okumura, M.; Taniguchi, T.; Hayashidani, H. Seroepidemiological Survey of Pathogenic Yersinia in Domestic Pigs in Chiba Prefecture, Japan. J. Vet. Med. Sci. 2024, 86, 322–324. [Google Scholar] [CrossRef] [PubMed]
- Borch, E.; Nesbakken, T.; Christensen, H. Hazard Identification in Swine Slaughter with Respect to Foodborne Bacteria. Int. J. Food Microbiol. 1996, 30, 9–25. [Google Scholar] [CrossRef]
- Fredriksson-Ahomaa, M.; Hallanvuo, S.; Korte, T.; Siitonen, A.; Korkeala, H. Correspondence of Genotypes of Sporadic Yersinia enterocolitica Bioserotype 4/O:3 Strains from Human and Porcine Sources. Epidemiol. Infect. 2001, 127, 37–47. [Google Scholar] [CrossRef]
- Van Damme, I.; Habib, I.; De Zutter, L. Yersinia enterocolitica in Slaughter Pig Tonsils: Enumeration and Detection by Enrichment versus Direct Plating Culture. Food Microbiol. 2010, 27, 158–161. [Google Scholar] [CrossRef] [PubMed]
- Funk, J.A.; Troutt, H.F.; Isaacson, R.E.; Fossler, C.P. Prevalence of Pathogenic Yersinia enterocolitica in Groups of Swine at Slaughter. J. Food Prot. 1998, 61, 677–682. [Google Scholar] [CrossRef] [PubMed]
- Fois, F.; Piras, F.; Torpdahl, M.; Mazza, R.; Ladu, D.; Consolati, S.G.; Spanu, C.; Scarano, C.; De Santis, E.P.L. Prevalence, Bioserotyping and Antibiotic Resistance of Pathogenic Yersinia enterocolitica Detected in Pigs at Slaughter in Sardinia. Int. J. Food Microbiol. 2018, 283, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Bhaduri, S.; Wesley, I.V. Prevalence, Serotype, Virulence Characteristics, Clonality, and Antibiotic Susceptibility of Pathogenic Yersinia enterocolitica from Swine Feces. Adv. Exp. Med. Biol. 2012, 954, 111–116. [Google Scholar] [PubMed]
- European Union. Commission Implementing Regulation (EU) 2019/627 Laying down Uniform Practical Arrangements for the Performance of Official Controls on Products of Animal Origin Intended for Human Consumption. Off. J. Eur. Union 2019, 131, 51–100. [Google Scholar]
- Valentin-Weigand, P.; Heesemann, J.; Dersch, P. Unique Virulence Properties of Yersinia enterocolitica O:3—An Emerging Zoonotic Pathogen Using Pigs as Preferred Reservoir Host. Int. J. Med. Microbiol. 2014, 304, 824–834. [Google Scholar] [CrossRef]
- Drummond, N.; Murphy, B.P.; Ringwood, T.; Prentice, M.B.; Buckley, J.F.; Fanning, S. Yersinia enterocolitica: A Brief Review of the Issues Relating to the Zoonotic Pathogen, Public Health Challenges, and the Pork Production Chain. Foodborne Pathog. Dis. 2012, 9, 179–189. [Google Scholar] [CrossRef]
- Primavilla, S.; Farneti, S.; Roila, R.; Branciari, R.; Altissimi, C.; Valiani, A.; Ranucci, D. Retrospective Study on the Prevalence of Yersinia enterocolitica in Food Collected in Umbria Region (Central Italy). Ital. J. Food Saf. 2023, 12, 10996. [Google Scholar] [CrossRef]
- Rivas, L.; Horn, B.; Armstrong, B.; Wright, J.; Strydom, H.; Wang, J.; Paine, S.; Thom, K.; Orton, A.; Robson, B.; et al. A Case-Control Study and Molecular Epidemiology of Yersiniosis in Aotearoa New Zealand. J. Clin. Microbiol. 2024, 62, e00754-24. [Google Scholar] [CrossRef]
- Yue, Y.; Shen, M.; Liu, X.; Hao, Q.; Kang, Y.; Che, Y.; Li, F.; Chen, S.; Xu, S.; Jing, H.; et al. Whole-Genome Sequencing-Based Prediction and Analysis of Antimicrobial Resistance in Yersinia enterocolitica from Ningxia, China. Front. Microbiol. 2022, 13, 936425. [Google Scholar] [CrossRef]
- Seakamela, E.M.; Diseko, L.; Malatji, D.; Makhado, L.; Motau, M.; Jambwa, K.; Magwedere, K.; Ntushelo, N.; Matle, I. Characterisation and Antibiotic Resistance of Yersinia enterocolitica from Various Meat Categories, South Africa. Onderstepoort J. Vet. Res. 2022, 89, 2006. [Google Scholar] [CrossRef]
- Fredriksson-Ahomaa, M.; Hielm, S.; Korkeala, H. High Prevalence of YadA-Positive Yersinia enterocolitica in Pig Tongues and Minced Meat at the Retail Level in Finland. J. Food Prot. 1999, 62, 123–127. [Google Scholar] [CrossRef]
- Thong, K.L.; Tan, L.K.; Ooi, P.T. Genetic Diversity, Virulotyping and Antimicrobial Resistance Susceptibility of Yersinia enterocolitica Isolated from Pigs and Porcine Products in Malaysia. J. Sci. Food Agric. 2018, 98, 87–95. [Google Scholar] [CrossRef]
- Klinzing, D.C.; Matias, R.R.; Skowronski, E.; Alvarez, M.; Liles, V.; Dimamay, M.P.; Natividad, F.F. Shotgun Genome Sequence of a Yersinia enterocolitica Isolate from the Philippines. J. Bacteriol. 2012, 194, 542–543. [Google Scholar] [CrossRef] [PubMed]
- Thomson, N.R.; Howard, S.; Wren, B.W.; Holden, M.T.G.; Crossman, L.; Challis, G.L.; Churcher, C.; Mungall, K.; Brooks, K.; Chillingworth, T.; et al. The Complete Genome Sequence and Comparative Genome Analysis of the High Pathogenicity Yersinia enterocolitica Strain 8081. PLoS Genet. 2006, 2, e206. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, Y.; Jing, H.; Ren, Y.; Zhou, Z.; Wang, S.; Kan, B.; Xu, J.; Wang, L. Complete Genome Sequence of a Yersinia enterocolitica “Old World” (3/o:9) Strain and Comparison with the “New World” (1B/O:8) Strain. J. Clin. Microbiol. 2011, 49, 1251–1259. [Google Scholar] [CrossRef] [PubMed]
- Batzilla, J.; Höper, D.; Antonenka, U.; Heesemann, J.; Rakin, A. Complete Genome Sequence of Yersinia enterocolitica Subsp. Palearctica Serogroup O:3. J. Bacteriol. 2011, 193, 2067. [Google Scholar] [CrossRef]
- Batzilla, J.; Antonenka, U.; Höper, D.; Heesemann, J.; Rakin, A. Yersinia enterocolitica Palearctica Serobiotype O:3/4—A Successful Group of Emerging Zoonotic Pathogens. BMC Genom. 2011, 12, 348. [Google Scholar] [CrossRef]
- Fuchs, T.M.; Brandt, K.; Starke, M.; Rattei, T. Shotgun Sequencing of Yersinia enterocolitica Strain W22703 (Biotype 2, Serotype O:9): Genomic Evidence for Oscillation between Invertebrates and Mammals. BMC Genom. 2011, 12, 168. [Google Scholar] [CrossRef]
- Martins, B.T.F.; de Meirelles, J.L.; Omori, W.P.; de Oliveira, R.R.; Yamatogi, R.S.; Call, D.R.; Nero, L.A. Comparative Genomics and Antibiotic Resistance of Yersinia enterocolitica Obtained from a Pork Production Chain and Human Clinical Cases in Brazil. Food Res. Int. 2022, 152, 110917. [Google Scholar] [CrossRef]
- Li, C.; Gölz, G.; Alter, T.; Barac, A.; Hertwig, S.; Riedel, C. Prevalence and Antimicrobial Resistance of Yersinia enterocolitica in Retail Seafood. J. Food Prot. 2018, 81, 497–501. [Google Scholar] [CrossRef]
- Fosse, J.; Seegers, H.; Magras, C. Prevalence and Risk Factors for Bacterial Food-Borne Zoonotic Hazards in Slaughter Pigs: A Review. Zoonoses Public Health 2009, 56, 429–454. [Google Scholar] [CrossRef] [PubMed]
- Koskinen, J.; Keto-Timonen, R.; Virtanen, S.; Vilar, M.J.; Korkeala, H. Prevalence and Dynamics of Pathogenic Yersinia enterocolitica 4/O:3 among Finnish Piglets, Fattening Pigs, and Sows. Foodborne Pathog. Dis. 2019, 16, 831–839. [Google Scholar] [CrossRef]
- Arden, K.; Gedye, K.; Angelin-Bonnet, O.; Murphy, E.; Antic, D. Yersinia enterocolitica in wild and peridomestic rodents within Great Britain, a prevalence study. Zoonoses Public Health 2022, 69, 537–549. [Google Scholar] [CrossRef]
- Angelovska, M.; Zaharieva, M.M.; Najdenski, H. Yersinia enterocolitica—Isolation, Pathogenicity, and Prevalence in Farms for Slaughtered Pigs. Acta Microbiol. Bulg. 2023, 39, 118–129. [Google Scholar] [CrossRef]
- Virtanen, S.; Salonen, L.; Laukkanen-Ninios, R.; Fredriksson-Ahomaa, M.; Korkeala, H. Piglets Are a Source of Pathogenic Yersinia enterocolitica on Fattening-Pig Farms. Appl. Environ. Microbiol. 2012, 78, 3000–3003. [Google Scholar] [CrossRef] [PubMed]
- Nowak, B.; Mueffling, T.v.; Caspari, K.; Hartung, J. Validation of a Method for the Detection of Virulent Yersinia enterocolitica and Their Distribution in Slaughter Pigs from Conventional and Alternative Housing Systems. Vet. Microbiol. 2006, 117, 219–228. [Google Scholar] [CrossRef]
- Quion, N.B.; Torga, A.P. Yersinia enterocolitica. Pediatr. Rev. 2021, 42, 712–713. [Google Scholar] [CrossRef]
- Nesbakken, T. Yersinia. In Foodborne Infections and Intoxications; Elsevier: Amsterdam, The Netherlands, 2021; pp. 187–199. [Google Scholar]
- Xue, Y.; Zhai, S.; Wang, Z.; Ji, Y.; Wang, G.; Wang, T.; Wang, X.; Xi, H.; Cai, R.; Zhao, R.; et al. The Yersinia Phage X1 Administered Orally Efficiently Protects a Murine Chronic Enteritis Model Against Yersinia enterocolitica Infection. Front. Microbiol. 2020, 11, 351. [Google Scholar] [CrossRef]
| Year | Location | Month | Cases (No.) | Sevorar | Source | Reference |
|---|---|---|---|---|---|---|
| 1976 | New York | September | 38 | O:3 | Flavored milk | Black et al. [80] |
| 1980 | Japan | April | 1051 | O:3 | Pork | Maruyama et al. [81] |
| 1988–1993 | New Zealand | - | 918 | 4/O:3 | - | Fenwick et al. [82] |
| 1989 | Georgia | November | 15 | O:3 | Pork chitterlings (intestines) | Lee et al. [83] |
| 1995 | Vermont and New Hampshire | October | 10 (1 fatal) | O:8 | Bottles of pasteurized milk from a local dairy | Ackers et al. [84] |
| 2004 | Japan | July | 42 | O:8 | Salads with ham | Sakai et al. [85] |
| 2005–2006 | Norway | December–February | 11 | 2/O:9 | Processed pork | Grahek-Ogden et al. [86] |
| 2006 | Japan | July | 3 (family) | 2/O:9 | Contaminated food, such as pork (speculated) | Moriki et al. [87] |
| 2009–2010 | Germany | - | 563 | O:3 (93.6%), O:9 (5.1%) O:5,27 (0.4%) | Raw minced pork | Rosner et al. [88] |
| 2010–2015 | China | - | 7304 | 3/O:3 | Contaminated food | Duan et al. [89] |
| 2011 | Pennsylvania | July | 22 | 1B/O:8 | Improperly pasteurized milk | Longenberger et al. [41] |
| 2011–2014 | Norway | May | 21,133 | 2/O:9 | RTE salad mix | MacDonald et al. [90,91] |
| 2012 | Tokyo | August | 39 | O:8 | Fresh vegetable salad | Konishi et al. [92] |
| 2013 | Tokyo | April | 52 | O:9 | Fresh vegetable salad | Konishi et al. [92] |
| 2017–2021 | France | Summer | 7642 | 4 (87.2%); 2/3-9b (10.6%) | Barbecued sausages, chipolatas, ribs, and other pork meat products | Le Guern et al. [93] |
| 2018–2020 | Czech Republic | January | 1686 | - | Undercooked pork, less often vegetables, or water | Špačková et al. [94] |
| 2019 | Sweden | March–May (Spring) | 57 | 4/O:3 | Fresh spinach | Karlsson et al. [95] |
| 2023 | Austria | January-April | 11 | 1A | Nutritional milkshakes | Colbran et al. [10] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Saraiva, S.; García-Díez, J.; de Sousa, T.; Calouro, R.; Fernandes, D.; Mourão, A.V.; Saraiva, C.; Mesquita, J.R.; Coelho, A.C.; Poeta, P. Zoonotic Yersinia enterocolitica in Swine: One Health Implications for Food Safety and Public Health. Vet. Sci. 2025, 12, 795. https://doi.org/10.3390/vetsci12090795
Saraiva S, García-Díez J, de Sousa T, Calouro R, Fernandes D, Mourão AV, Saraiva C, Mesquita JR, Coelho AC, Poeta P. Zoonotic Yersinia enterocolitica in Swine: One Health Implications for Food Safety and Public Health. Veterinary Sciences. 2025; 12(9):795. https://doi.org/10.3390/vetsci12090795
Chicago/Turabian StyleSaraiva, Sónia, Juan García-Díez, Telma de Sousa, Rita Calouro, Diana Fernandes, Ana V. Mourão, Cristina Saraiva, João R. Mesquita, Ana C. Coelho, and Patrícia Poeta. 2025. "Zoonotic Yersinia enterocolitica in Swine: One Health Implications for Food Safety and Public Health" Veterinary Sciences 12, no. 9: 795. https://doi.org/10.3390/vetsci12090795
APA StyleSaraiva, S., García-Díez, J., de Sousa, T., Calouro, R., Fernandes, D., Mourão, A. V., Saraiva, C., Mesquita, J. R., Coelho, A. C., & Poeta, P. (2025). Zoonotic Yersinia enterocolitica in Swine: One Health Implications for Food Safety and Public Health. Veterinary Sciences, 12(9), 795. https://doi.org/10.3390/vetsci12090795










