Impact of Glycerol as Carbon Source onto Specific Sugar and Inducer Uptake Rates and Inclusion Body Productivity in E. coli BL21(DE3)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bioreactor Cultivations
2.2. Cultivation Scheme and qs Screening Procedure
2.3. Process Analytics
3. Results and Discussion
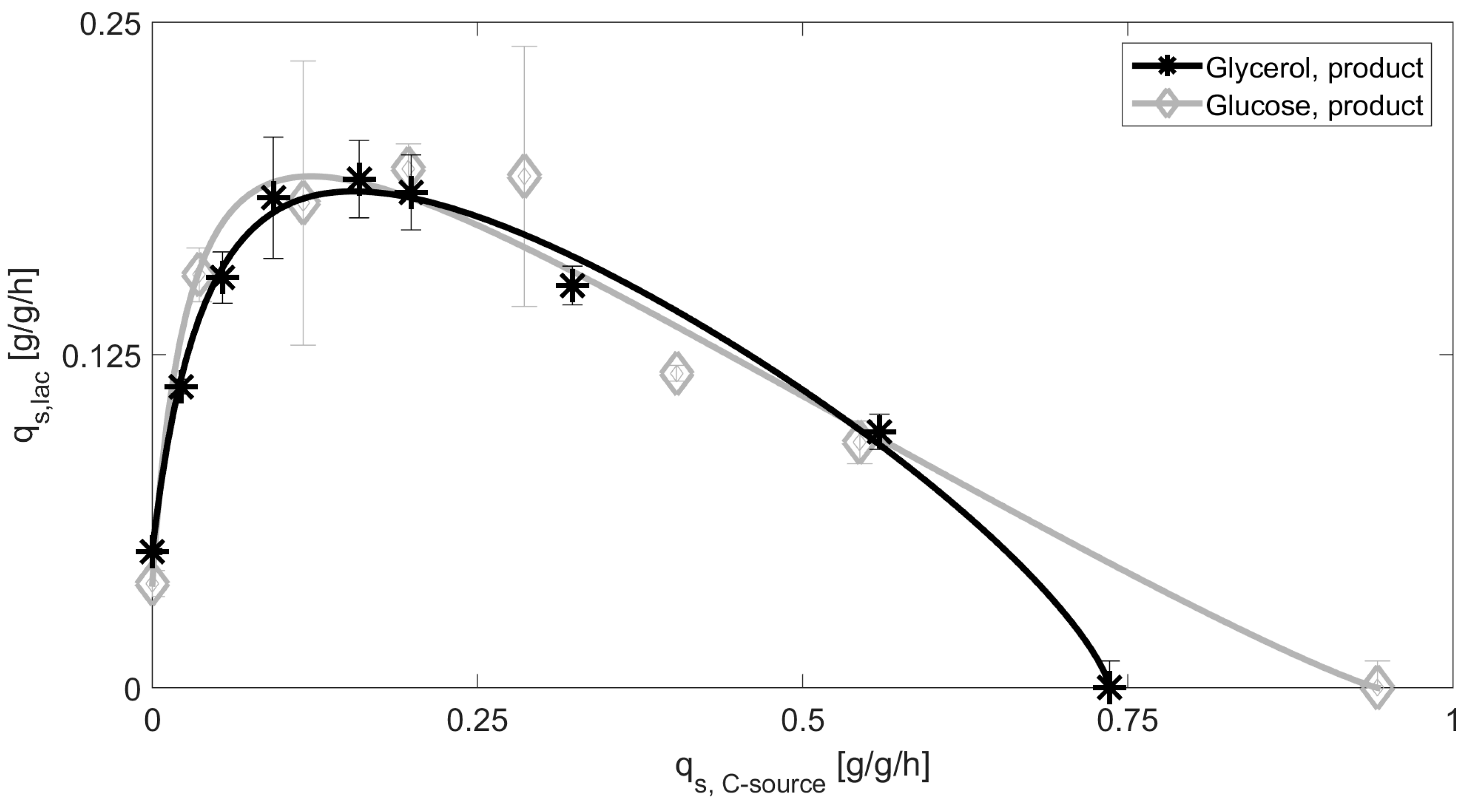
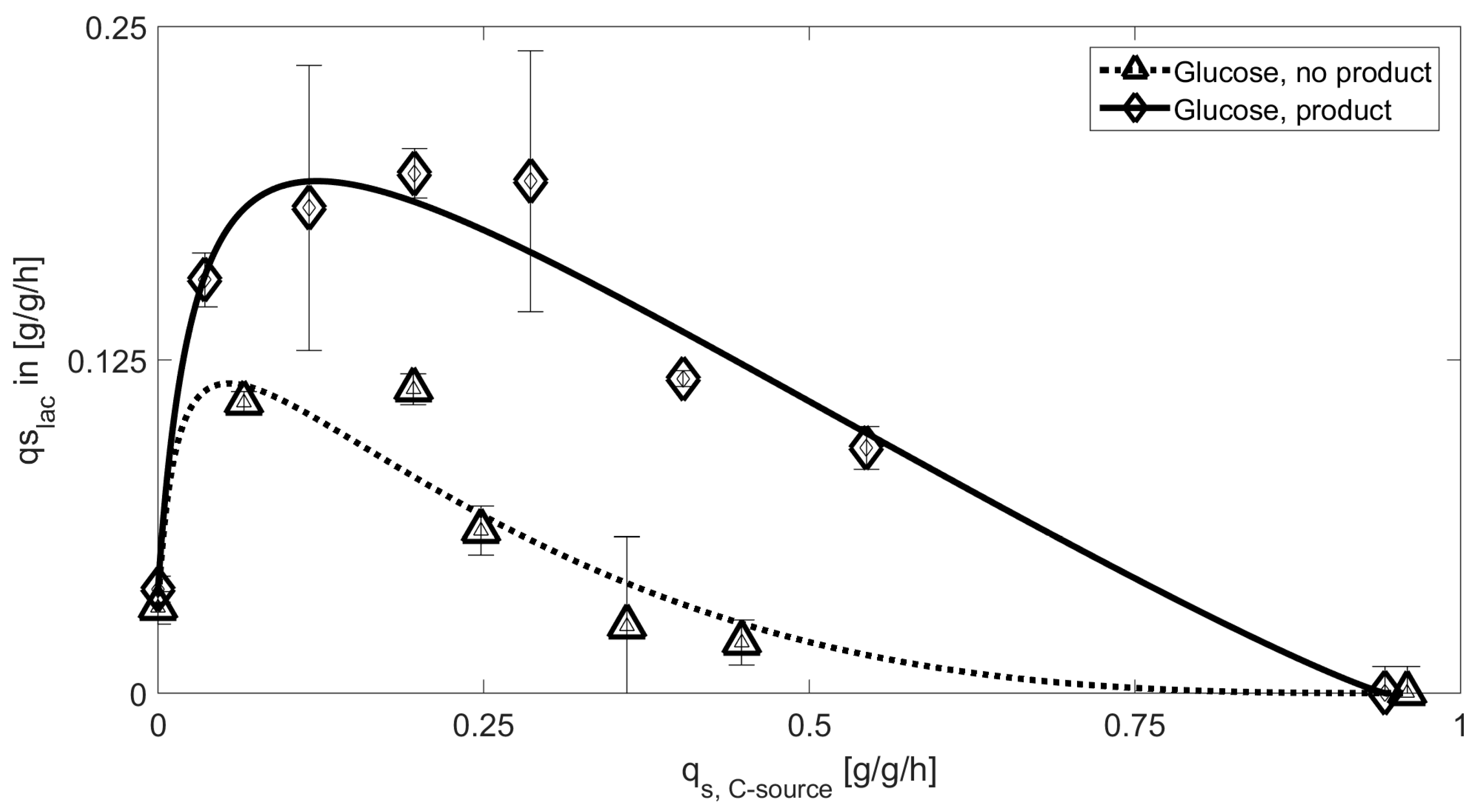
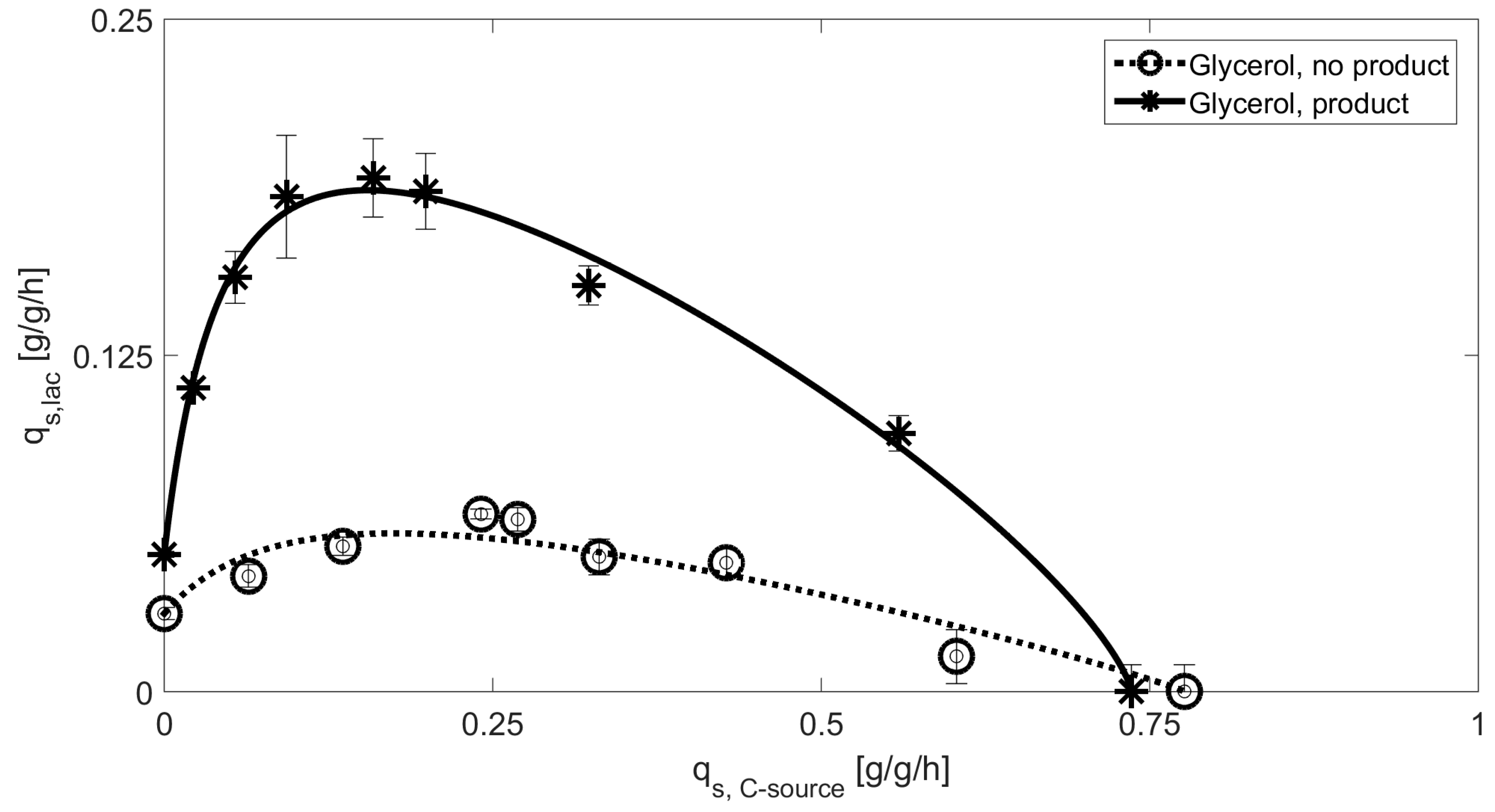
3.1. Mechanistic Correlations of Glycerol onto Specific Lactose Uptake Rate
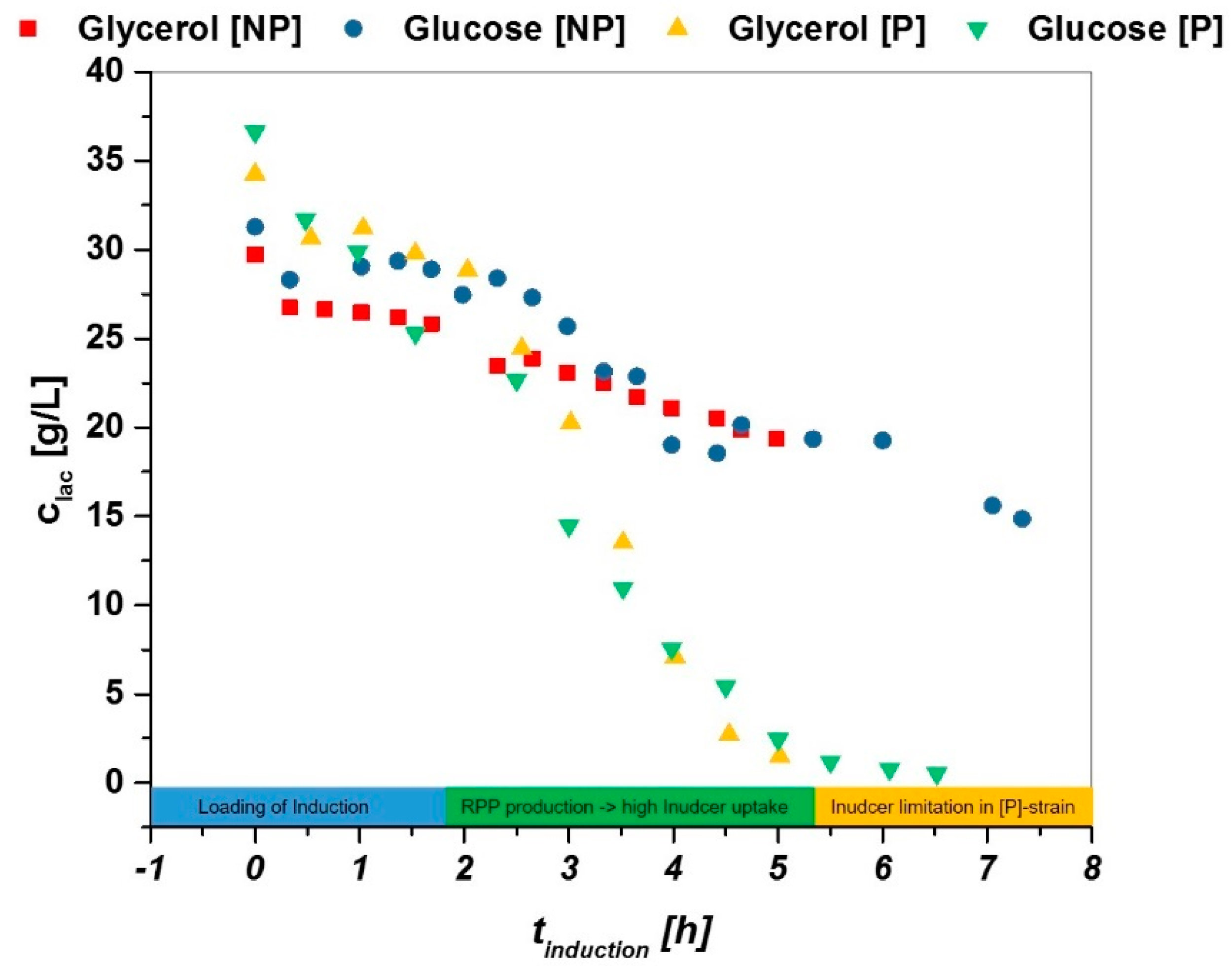
- (i)
- Adaption phase: lactose gets transferred to alloactose and loads the induction (0–2 h in induction phase).
- (ii)
- Linear decrease of lactose as the system needs inducer for recombinant protein expression (2–5 h).
- (iii)
- Limitation of lactose in P strain: not sufficient inducer present, need for mixed feed system (5–7 h), no inducer limitation seen in NP strain, further decrease of inducer analogue to phase 2.
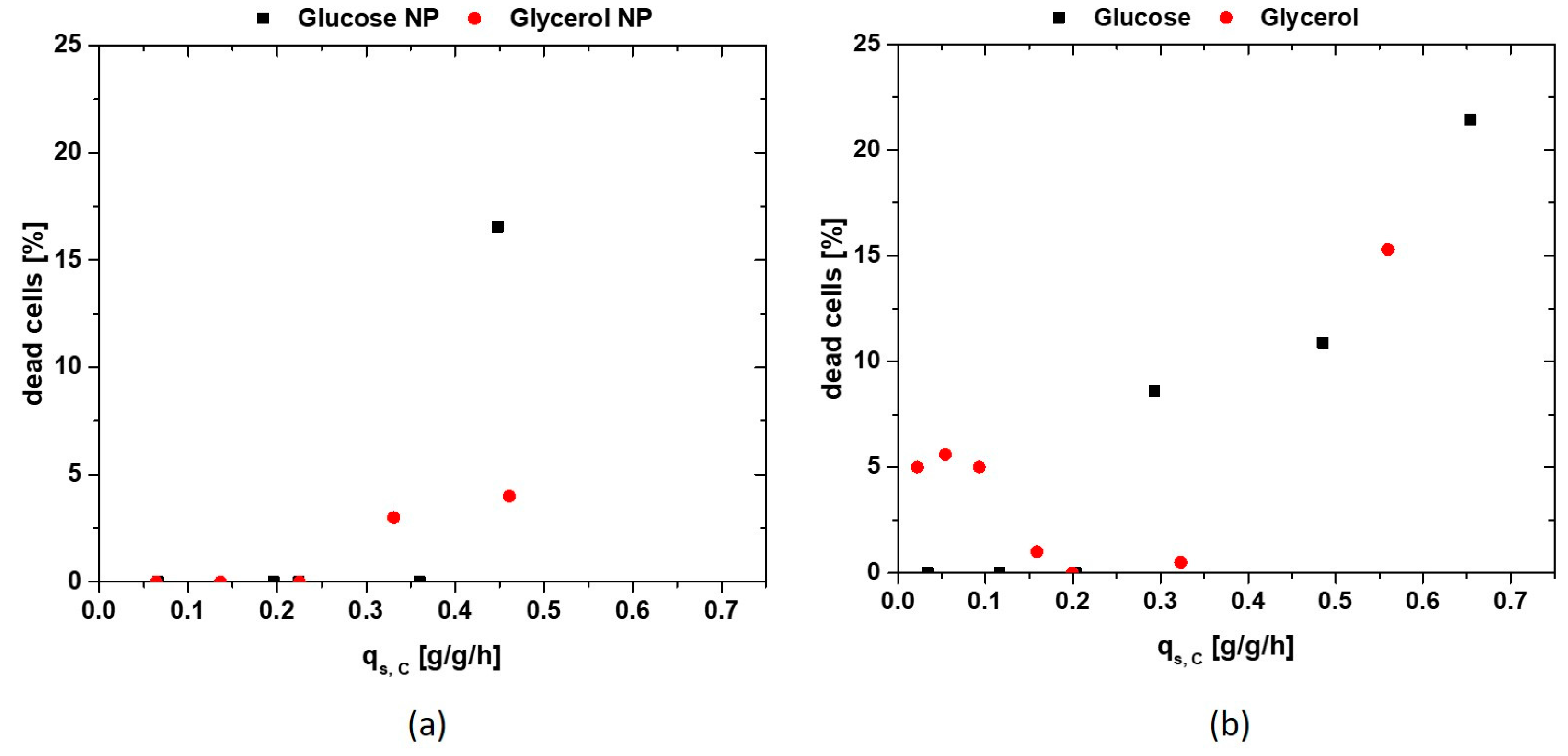
3.2. Productivity and Physiology Using Glycerol as Primary Carbon Source
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gupta, S.K.; Shukla, P. Microbial platform technology for recombinant antibody fragment production: A review. Crit. Rev. Microbiol. 2017, 43, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Walsh, G. Biopharmaceutical benchmarks 2010. Nat. Biotechnol. 2010, 28, 917–924. [Google Scholar] [CrossRef] [PubMed]
- Wurm, D.J.; Veiter, L.; Ulonska, S.; Eggenreich, B.; Herwig, C.; Spadiut, O. The E. coli pET expression system revisited-mechanistic correlation between glucose and lactose uptake. Appl. Microbiol. Biotechnol. 2016, 100, 8721–8729. [Google Scholar] [CrossRef] [PubMed]
- Meuris, L.; Santens, F.; Elson, G.; Festjens, N.; Boone, M.; Dos Santos, A.; Devos, S.; Rousseau, F.; Plets, E.; Houthuys, E.; et al. GlycoDelete engineering of mammalian cells simplifies N-glycosylation of recombinant proteins. Nat. Biotechnol. 2014, 32, 485–489. [Google Scholar] [CrossRef] [PubMed]
- DeLisa, M.P.; Li, J.; Rao, G.; Weigand, W.A.; Bentley, W.E. Monitoring GFP-operon fusion protein expression during high cell density cultivation of Escherichia coli using an on-line optical sensor. Biotechnol. Bioeng. 1999, 65, 54–64. [Google Scholar] [CrossRef]
- Berlec, A.; Strukelj, B. Current state and recent advances in biopharmaceutical production in Escherichia coli, yeasts and mammalian cells. J. Ind. Microbiol. Biotechnol. 2013, 40, 257–274. [Google Scholar] [CrossRef] [PubMed]
- Baeshen, M.N.; Al-Hejin, A.M.; Bora, R.S.; Ahmed, M.M.; Ramadan, H.A.; Saini, K.S.; Baeshen, N.A.; Redwan, E.M. Production of Biopharmaceuticals in E. coli: Current Scenario and Future Perspectives. J. Microbiol. Biotechnol. 2015, 25, 953–962. [Google Scholar] [CrossRef] [PubMed]
- Spadiut, O.; Capone, S.; Krainer, F.; Glieder, A.; Herwig, C. Microbials for the production of monoclonal antibodies and antibody fragments. Trends Biotechnol. 2014, 32, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Studier, F.W.; Moffatt, B.A. Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J. Mol. Biol. 1986, 189, 113–130. [Google Scholar] [CrossRef]
- Steen, R.; Dahlberg, A.E.; Lade, B.N.; Studier, F.W.; Dunn, J.J. T7 RNA polymerase directed expression of the Escherichia coli rrnB operon. EMBO J. 1986, 5, 1099–1103. [Google Scholar] [PubMed]
- Studier, F.W.; Rosenberg, A.H.; Dunn, J.J.; Dubendorff, J.W. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol. 1990, 185, 60–89. [Google Scholar] [PubMed]
- Dubendorff, J.W.; Studier, F.W. Controlling basal expression in an inducible T7 expression system by blocking the target T7 promoter with lac repressor. J. Mol. Biol. 1991, 219, 45–59. [Google Scholar] [CrossRef]
- Neubauer, P.; Hofmann, K. Efficient use of lactose for the lac promoter-controlled overexpression of the main antigenic protein of the foot and mouth disease virus in Escherichia coli under fed-batch fermentation conditions. FEMS Microbiol. Rev. 1994, 14, 99–102. [Google Scholar] [CrossRef] [PubMed]
- Lyakhov, D.L.; He, B.; Zhang, X.; Studier, F.W.; Dunn, J.J.; McAllister, W.T. Pausing and termination by bacteriophage T7 RNA polymerase. J. Mol. Biol. 1998, 280, 201–213. [Google Scholar] [CrossRef] [PubMed]
- Jeong, H.; Barbe, V.; Lee, C.H.; Vallenet, D.; Yu, D.S.; Choi, S.H.; Couloux, A.; Lee, S.W.; Yoon, S.H.; Cattolico, L.; et al. Genome sequences of Escherichia coli B strains REL606 and BL21(DE3). J. Mol. Biol. 2009, 394, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Jeong, H.; Kim, H.J.; Lee, S.J. Complete Genome Sequence of Escherichia coli Strain BL21. Genome Announc. 2015, 3, e00134-15. [Google Scholar] [CrossRef] [PubMed]
- Tseng, T.T.; Tyler, B.M.; Setubal, J.C. Protein secretion systems in bacterial-host associations, and their description in the Gene Ontology. BMC Microbiol. 2009, 9 (Suppl. 1), S2. [Google Scholar] [CrossRef] [PubMed]
- Marbach, A.; Bettenbrock, K. Lac operon induction in Escherichia coli: Systematic comparison of IPTG and TMG induction and influence of the transacetylase LacA. J. Biotechnol. 2012, 157, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Keiler, K.C. Biology of trans-translation. Annu. Rev. Microbiol. 2008, 62, 133–151. [Google Scholar] [CrossRef] [PubMed]
- Viitanen, M.I.; Vasala, A.; Neubauer, P.; Alatossava, T. Cheese whey-induced high-cell-density production of recombinant proteins in Escherichia coli. Microb. Cell Fact. 2003, 2, 2. [Google Scholar] [CrossRef] [PubMed]
- Marisch, K.; Bayer, K.; Cserjan-Puschmann, M.; Luchner, M.; Striedner, G. Evaluation of three industrial Escherichia coli strains in fed-batch cultivations during high-level SOD protein production. Microb. Cell Fact. 2013, 12, 58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ukkonen, K.; Mayer, S.; Vasala, A.; Neubauer, P. Use of slow glucose feeding as supporting carbon source in lactose autoinduction medium improves the robustness of protein expression at different aeration conditions. Protein Expr. Purif. 2013, 91, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Neubauer, P.; Hofmann, K.; Holst, O.; Mattiasson, B.; Kruschke, P. Maximizing the expression of a recombinant gene in Escherichia coli by manipulation of induction time using lactose as inducer. Appl. Microbiol. Biotechnol. 1992, 36, 739–744. [Google Scholar] [CrossRef] [PubMed]
- Marschall, L.; Sagmeister, P.; Herwig, C. Tunable recombinant protein expression in E. coli: Enabler for continuous processing? Appl. Microbiol. Biotechnol. 2016, 100, 5719–5728. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blommel, P.G.; Becker, K.J.; Duvnjak, P.; Fox, B.G. Enhanced bacterial protein expression during auto-induction obtained by alteration of lac repressor dosage and medium composition. Biotechnol. Prog. 2007, 23, 585–598. [Google Scholar] [CrossRef] [PubMed]
- Dvorak, P.; Chrast, L.; Nikel, P.I.; Fedr, R.; Soucek, K.; Sedlackova, M.; Chaloupkova, R.; de Lorenzo, V.; Prokop, Z.; Damborsky, J. Exacerbation of substrate toxicity by IPTG in Escherichia coli BL21(DE3) carrying a synthetic metabolic pathway. Microb. Cell Fact. 2015, 14, 201. [Google Scholar] [CrossRef] [PubMed]
- Wurm, D.J.; Herwig, C.; Spadiut, O. How to Determine Interdependencies of Glucose and Lactose Uptake Rates for Heterologous Protein Production with E. coli. Methods Mol. Biol. 2017, 1586, 397–408. [Google Scholar] [PubMed]
- García-Fruitós, E.; Vázquez, E.; Díez-Gil, C.; Corchero, J.L.; Seras-Franzoso, J.; Ratera, I.; Veciana, J.; Villaverde, A. Bacterial inclusion bodies: Making gold from waste. Trends Biotechnol. 2012, 30, 65–70. [Google Scholar] [CrossRef] [PubMed]
- Palmer, I.; Wingfield, P.T. Preparation and extraction of insoluble (inclusion-body) proteins from Escherichia coli. Curr. Protoc. Protein Sci. 2012. [Google Scholar] [CrossRef]
- Ramón, A.; Señorale-Pose, M.; Marín, M. Inclusion bodies: Not that bad…. Front. Microbiol. 2014, 5, 56. [Google Scholar] [CrossRef] [PubMed]
- Villaverde, A.; Corchero, J.L.; Seras-Franzoso, J.; Garcia-Fruitós, E. Functional protein aggregates: Just the tip of the iceberg. Nanomedicine (Lond.) 2015, 10, 2881–2891. [Google Scholar] [CrossRef] [PubMed]
- Wingfield, P.T.; Palmer, I.; Liang, S.M. Folding and Purification of Insoluble (Inclusion Body) Proteins from Escherichia coli. Curr. Protoc. Protein Sci. 2014. [Google Scholar] [CrossRef]
- Wingfield, P.T. Preparation of Soluble Proteins from Escherichia coli. Curr. Protoc. Protein Sci. 2014, 78, 6.2.1–6.2.22. [Google Scholar] [PubMed]
- Postma, P.W.; Lengeler, J.W.; Jacobson, G.R. Phosphoenolpyruvate:carbohydrate phosphotransferase systems of bacteria. Microbiol. Rev. 1993, 57, 543–594. [Google Scholar] [PubMed]
- Deutscher, J.; Francke, C.; Postma, P.W. How phosphotransferase system-related protein phosphorylation regulates carbohydrate metabolism in bacteria. Microbiol. Mol. Biol. Rev. 2006, 70, 939–1031. [Google Scholar] [CrossRef] [PubMed]
- Ronimus, R.S.; Morgan, H.W. Distribution and phylogenies of enzymes of the Embden-Meyerhof-Parnas pathway from archaea and hyperthermophilic bacteria support a gluconeogenic origin of metabolism. Archaea 2003, 1, 199–221. [Google Scholar] [CrossRef] [PubMed]
- Bettenbrock, K.; Fischer, S.; Kremling, A.; Jahreis, K.; Sauter, T.; Gilles, E.D. A quantitative approach to catabolite repression in Escherichia coli. J. Biol. Chem. 2006, 281, 2578–2584. [Google Scholar] [CrossRef] [PubMed]
- Kremling, A.; Bettenbrock, K.; Laube, B.; Jahreis, K.; Lengeler, J.W.; Gilles, E.D. The organization of metabolic reaction networks. III. Application for diauxic growth on glucose and lactose. Metab. Eng. 2001, 3, 362–379. [Google Scholar] [CrossRef] [PubMed]
- Stülke, J.; Hillen, W. Carbon catabolite repression in bacteria. Curr. Opin. Microbiol. 1999, 2, 195–201. [Google Scholar] [CrossRef]
- Wurm, D.J.; Hausjell, J.; Ulonska, S.; Herwig, C.; Spadiut, O. Mechanistic platform knowledge of concomitant sugar uptake in Escherichia coli BL21(DE3) strains. Sci. Rep. 2017, 7, 45072. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Gómez, K.; Flores, N.; Castañeda, H.M.; Martínez-Batallar, G.; Hernández-Chávez, G.; Ramírez, O.T.; Gosset, G.; Encarnación, S.; Bolivar, F. New insights into Escherichia coli metabolism: Carbon scavenging, acetate metabolism and carbon recycling responses during growth on glycerol. Microb. Cell Fact. 2012, 11, 46. [Google Scholar] [CrossRef] [PubMed]
- Lin, E.C. Glycerol dissimilation and its regulation in bacteria. Annu. Rev. Microbiol. 1976, 30, 535–578. [Google Scholar] [CrossRef] [PubMed]
- Achmüller, C.; Kaar, W.; Ahrer, K.; Wechner, P.; Hahn, R.; Werther, F.; Schmidinger, H.; Cserjan-Puschmann, M.; Clementschitsch, F.; Striedner, G.; et al. N(pro) fusion technology to produce proteins with authentic N termini in E. coli. Nat. Methods 2007, 4, 1037–1043. [Google Scholar] [CrossRef] [PubMed]
- Zwaig, N.; Kistler, W.S.; Lin, E.C. Glycerol kinase, the pacemaker for the dissimilation of glycerol in Escherichia coli. J. Bacteriol. 1970, 102, 753–759. [Google Scholar] [PubMed]
- Voegele, R.T.; Sweet, G.D.; Boos, W. Glycerol kinase of Escherichia coli is activated by interaction with the glycerol facilitator. J. Bacteriol. 1993, 175, 1087–1094. [Google Scholar] [CrossRef] [PubMed]
- Slouka, C.; Wurm, D.J.; Brunauer, G.; Welzl-Wachter, A.; Spadiut, O.; Fleig, J.; Herwig, C. A Novel Application for Low Frequency Electrochemical Impedance Spectroscopy as an Online Process Monitoring Tool for Viable Cell Concentrations. Sensors (Basel) 2016, 16, 1900. [Google Scholar] [CrossRef] [PubMed]
- Langemann, T.; Mayr, U.B.; Meitz, A.; Lubitz, W.; Herwig, C. Multi-parameter flow cytometry as a process analytical technology (PAT) approach for the assessment of bacterial ghost production. Appl. Microbiol. Biotechnol. 2016, 100, 409–418. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Banerjee, A.; Pan, S.H.; Li, Z.J. Galactose can be an inducer for production of therapeutic proteins by auto-induction using E. coli BL21 strains. Protein Expr. Purif. 2012, 83, 30–36. [Google Scholar] [CrossRef] [PubMed]
- Studier, F.W.; Daegelen, P.; Lenski, R.E.; Maslov, S.; Kim, J.F. Understanding the differences between genome sequences of Escherichia coli B strains REL606 and BL21(DE3) and comparison of the E. coli B and K-12 genomes. J. Mol. Biol. 2009, 394, 653–680. [Google Scholar] [CrossRef] [PubMed]
- Daegelen, P.; Studier, F.W.; Lenski, R.E.; Cure, S.; Kim, J.F. Tracing ancestors and relatives of Escherichia coli B, and the derivation of B strains REL606 and BL21(DE3). J. Mol. Biol. 2009, 394, 634–643. [Google Scholar] [CrossRef] [PubMed]
- Korz, D.J.; Rinas, U.; Hellmuth, K.; Sanders, E.A.; Deckwer, W.D. Simple fed-batch technique for high cell density cultivation of Escherichia coli. J. Biotechnol. 1995, 39, 59–65. [Google Scholar] [CrossRef]
- Inada, T.; Kimata, K.; Aiba, H. Mechanism responsible for glucose-lactose diauxie in Escherichia coli: Challenge to the cAMP model. Genes Cells 1996, 1, 293–301. [Google Scholar] [CrossRef] [PubMed]
- Larson, T.J.; Ye, S.Z.; Weissenborn, D.L.; Hoffmann, H.J.; Schweizer, H. Purification and characterization of the repressor for the sn-glycerol 3-phosphate regulon of Escherichia coli K12. J. Biol. Chem. 1987, 262, 15869–15874. [Google Scholar] [PubMed]
- Iuchi, S.; Cole, S.T.; Lin, E.C. Multiple regulatory elements for the glpA operon encoding anaerobic glycerol-3-phosphate dehydrogenase and the glpD operon encoding aerobic glycerol-3-phosphate dehydrogenase in Escherichia coli: Further characterization of respiratory control. J. Bacteriol. 1990, 172, 179–184. [Google Scholar] [CrossRef] [PubMed]
- Heyland, J.; Blank, L.M.; Schmid, A. Quantification of metabolic limitations during recombinant protein production in Escherichia coli. J. Biotechnol. 2011, 155, 178–184. [Google Scholar] [CrossRef] [PubMed]
- Glick, B.R. Metabolic load and heterologous gene expression. Biotechnol. Adv. 1995, 13, 247–261. [Google Scholar] [CrossRef]
- Weissenborn, D.L.; Wittekindt, N.; Larson, T.J. Structure and regulation of the glpFK operon encoding glycerol diffusion facilitator and glycerol kinase of Escherichia coli K-12. J. Biol. Chem. 1992, 267, 6122–6131. [Google Scholar] [PubMed]
- Hogema, B.M.; Arents, J.C.; Bader, R.; Postma, P.W. Autoregulation of lactose uptake through the LacY permease by enzyme IIAGlc of the PTS in Escherichia coli K-12. Mol. Microbiol. 1999, 31, 1825–1833. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.M.; Tymoczko, J.L.; Stryer, L. Biochemistry, 5th ed.; W. H. Freeman: New York, NY, USA, 2002. [Google Scholar]
- Marr, A.G.; Ingraham, J.L.; Squires, C.L. Effect of the temperature of growth of Escherichia coli on the formation of beta-galactosidase. J. Bacteriol. 1964, 87, 356–362. [Google Scholar] [PubMed]
- Mayer, S.; Junne, S.; Ukkonen, K.; Glazyrina, J.; Glauche, F.; Neubauer, P.; Vasala, A. Lactose autoinduction with enzymatic glucose release: Characterization of the cultivation system in bioreactor. Protein Expr. Purif. 2014, 94, 67–72. [Google Scholar] [CrossRef] [PubMed]

| Amount of Glucose | Amount of Glycerol | |
|---|---|---|
| Preculture | 8.8 g/L | 8.9 g/L |
| Batch-Media | 22 g/L | 23 g/L |
| Feed | either 250 g/L or 300 g/L | |
| Cultivation System | qs,lac,max | KA | qs,C,crit | n | qs,lac,noglu | NRMSE |
|---|---|---|---|---|---|---|
| [g/g/h] | [g/g/h] | [g/g/h] | [-] | [g/g/h] | [%] | |
| Glucose | 0.23 | 0.032 | 0.94 | 1.14 | 0.039 | 6.5 |
| Glycerol | 0.23 | 0.053 | 0.74 | 0.74 | 0.051 | 2.6 |
| Cultivation System | qs,lac,max [g/g/h] | KA [g/g/h] | qs,glu,crit [g/g/h] | n [-] | qs,lac,noglu [g/g/h] | NRMSE [%] |
|---|---|---|---|---|---|---|
| Glucose [NP] | 0.14 | 0.016 | 0.96 | 2.92 | 0.032 | 12.7 |
| Glycerol [NP] | 0.10 | 0.13 | 0.78 | 0.90 | 0.029 | 9.7 |
| Glucose | Glucose NP | Glycerol | Glycerol NP | ||||
|---|---|---|---|---|---|---|---|
| qs,C [g/g/h] | qCO2 [g/g/h] | qs,C [g/g/h] | qCO2 [g/g/h] | qs,C [g/g/h] | qCO2 [g/g/h] | qs,C [g/g/h] | qCO2 [g/g/h] |
| 0.036 | 2.15 ± 0.33 | 0.066 | 1.69 ± 0.25 | 0.022 | 2.91 ± 0.46 | 0.064 | 0.82 ± 0.09 |
| 0.116 | 3.12 ± 0.46 | 0.196 | 3.75 ± 0.44 | 0.054 | 4.41 ± 0.78 | 0.136 | 1.85 ± 0.21 |
| 0.197 | 3.98 ± 0.55 | 0.224 | 3.35 ± 0.42 | 0.093 | 3.88 ± 0.64 | 0.225 | 2.86 ± 0.31 |
| 0.286 | 5.72 ± 0.41 | 0.36 | 5.96 ± 0.26 | 0.159 | 3.12 ± 0.43 | 0.331 | 3.31 ± 0.22 |
| 0.403 | 6.42 ± 1.48 | 0.448 | 5.64 ± 0.47 | 0.199 | 4.14 ± 0.64 | 0.428 | 4.07 ± 0.51 |
| 0.544 | 7.30 ± 1.64 | 0.323 | 5.13 ± 0.48 | 0.603 | 1.75 ± 1.58 | ||
| 0.559 | 7.18 ± 2.10 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kopp, J.; Slouka, C.; Ulonska, S.; Kager, J.; Fricke, J.; Spadiut, O.; Herwig, C. Impact of Glycerol as Carbon Source onto Specific Sugar and Inducer Uptake Rates and Inclusion Body Productivity in E. coli BL21(DE3). Bioengineering 2018, 5, 1. https://doi.org/10.3390/bioengineering5010001
Kopp J, Slouka C, Ulonska S, Kager J, Fricke J, Spadiut O, Herwig C. Impact of Glycerol as Carbon Source onto Specific Sugar and Inducer Uptake Rates and Inclusion Body Productivity in E. coli BL21(DE3). Bioengineering. 2018; 5(1):1. https://doi.org/10.3390/bioengineering5010001
Chicago/Turabian StyleKopp, Julian, Christoph Slouka, Sophia Ulonska, Julian Kager, Jens Fricke, Oliver Spadiut, and Christoph Herwig. 2018. "Impact of Glycerol as Carbon Source onto Specific Sugar and Inducer Uptake Rates and Inclusion Body Productivity in E. coli BL21(DE3)" Bioengineering 5, no. 1: 1. https://doi.org/10.3390/bioengineering5010001
APA StyleKopp, J., Slouka, C., Ulonska, S., Kager, J., Fricke, J., Spadiut, O., & Herwig, C. (2018). Impact of Glycerol as Carbon Source onto Specific Sugar and Inducer Uptake Rates and Inclusion Body Productivity in E. coli BL21(DE3). Bioengineering, 5(1), 1. https://doi.org/10.3390/bioengineering5010001







