Integrating Dynamic Representation and Multi-Priors for Transnasal Intubation via Visual Foundation Model
Abstract
1. Introduction
- Dynamic representation and adaptation of multi-scale glottal image features to precisely parse local details and global semantic priors, enhancing robustness to deformation and motion blur.
- A high-efficiency fine-tuning strategy that extends low-rank adaptation for hierarchical feature adaptation, preserving geometric priors while reducing computational overhead.
- A feature aggregator that dynamically calibrates feature responses and constructs a dual-path dynamic feature pyramid by incorporating attention-based adaptive fusion of multi-scale features, which improves generalization under complex internal cavity anatomies.
2. Related Work
2.1. Glottal Detection and Segmentation
2.2. Foundation Model Adaptation for Medical Image Segmentation
3. Methods
3.1. Multi-Scale Contextual Feature Encoding
3.2. Hierarchical Low-Rank Adaptation for Efficient Fine-Tuning
3.3. Dynamic Semantic Feature Aggregation
3.3.1. Feature Aggregation
3.3.2. Dual-Path Dynamic Feature Pyramid
3.3.3. Multi-Head Cooperative Feature Decoupling
4. Experiments
4.1. Hardware System
4.2. Experiments Setup
4.2.1. Dataset
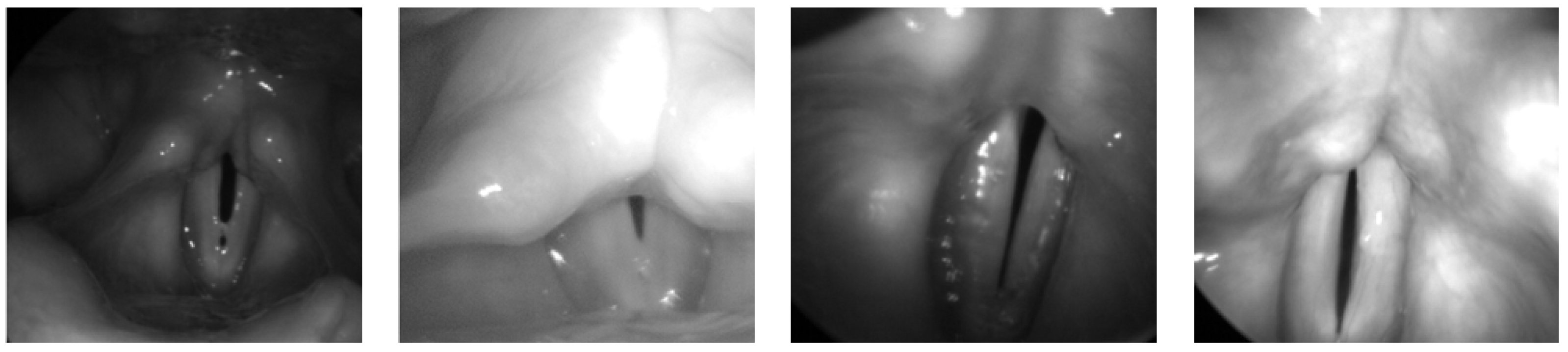
- Glottis: The Benchmark for Automatic Glottis Segmentation (BAGLS) [35] dataset comprises high-speed videoendoscopic recordings from 640 subjects, including both healthy individuals and patients with various laryngeal disorders. The data are collected by multiple medical professionals using diverse endoscopic systems, introducing substantial heterogeneity in image resolution, lighting conditions, and anatomical variations, as illustrated in Figure 5. To adapt this dataset for object detection tasks, we derive bounding boxes that tightly enclose the original segmentation masks and convert the annotations into the COCO [36] format. The resulting dataset includes 55,750 images for training and 3500 for testing. This large-scale and diverse dataset serves as a critical benchmark for evaluating the generalization capability and real-world applicability of our model across different patient groups and imaging devices.
- Phantom 2025: To mitigate ethical constraints during the initial development stage, we construct a synthetic dataset using our robotic NTI system. The Phantom dataset [37] is collected in a controlled laboratory setting using a fiberoptic bronchoscope equipped with real-time navigation and feedback control, with representative samples shown in Figure 6. The dataset includes 2267 training images and 479 test images. Bounding boxes are used to label general structures, such as the nose, channel, glottis, and trachea. Masks are employed to annotate structures with more clearly defined boundaries or specific shapes, including the right nostril, left nostril, glottic slit, right glottal valve, and left glottal valve, providing pixel-level accurate representations of their form and extent. These samples provide diverse anatomical representations that support early-stage model training and validation without involving human subjects.
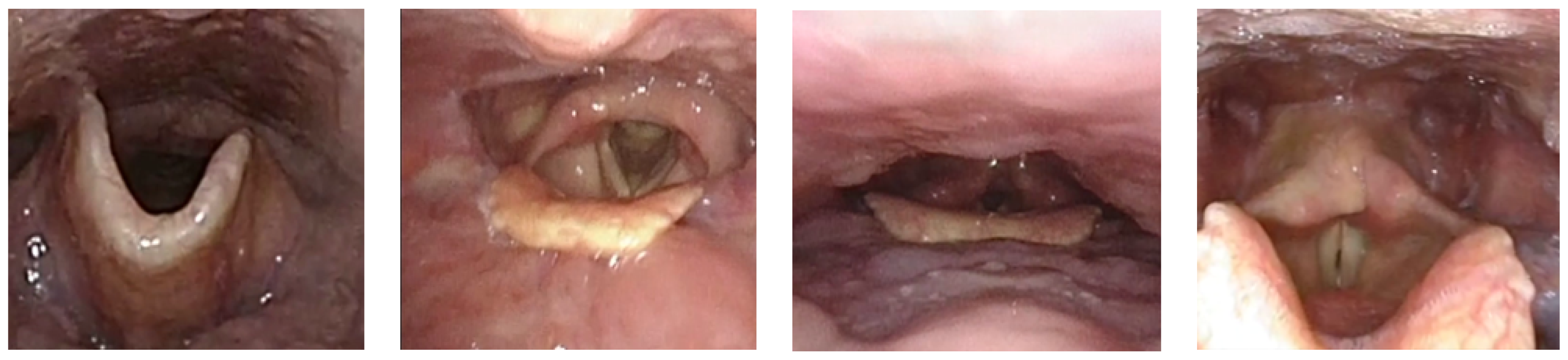
- Clinical 2025: The Clinical dataset [37] is obtained from nasopharyngoscopy procedures conducted at the Singapore General Hospital. It comprises 82 high-definition video recordings, each obtained from a unique patient, reflecting a broad spectrum of anatomical diversity and clinically realistic imaging conditions, as exemplified in Figure 7. All procedures are conducted by board-certified otolaryngologists using commercial flexible nasopharyngoscopes to ensure procedural consistency and clinical realism. From these recordings, we extract 2683 training images with 7030 annotated structures and 1131 validation images with 3295 annotations. These images capture key anatomical landmarks, such as the nose, nostrils, epiglottis, glottic valves, and surrounding tissues, as the endoscope passes through the glottis into the trachea.
4.2.2. Implementation Details
4.2.3. Evaluation Metrics
4.3. Comparison with State-of-the-Art Methods
4.3.1. Glottis
4.3.2. Phantom
4.3.3. Clinical
4.4. Ablation Study
4.4.1. Low-Rank Aaptation
4.4.2. Rank and Scaling Factor
4.4.3. Channels in FPN
4.4.4. Weight of the Loss Function
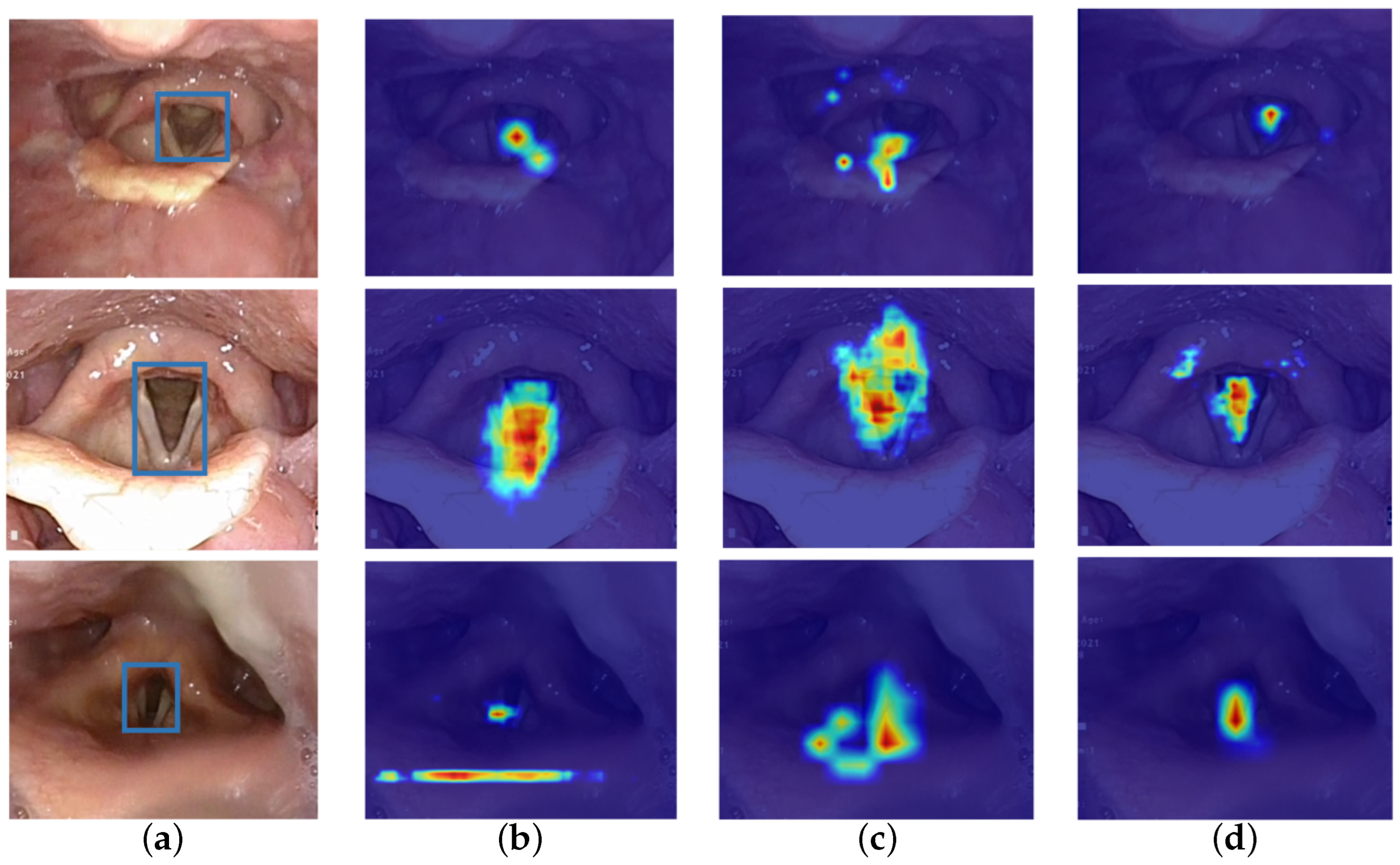
4.4.5. Qualitative Results
5. Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhu, Z.; Wang, Z.; Qi, G.; Mazur, N.; Yang, P.; Liu, Y. Brain tumor segmentation in MRI with multi-modality spatial information enhancement and boundary shape correction. Pattern Recognit. 2024, 153, 110553. [Google Scholar] [CrossRef]
- Liu, H.; Liu, S.Q.; Xie, X.L.; Li, Y.; Zhou, X.H.; Feng, Z.Q.; Li, G.T.; Ma, X.Y.; Hou, Z.G.; Yuan, Y.Y.; et al. An Original Design of Robotic-Assisted Flexible Nasotracheal Intubation System. In 2023 IEEE International Conference on Robotics and Biomimetics (ROBIO); IEEE: New York, NY, USA, 2023; pp. 1–6. [Google Scholar] [CrossRef]
- Zheng, L.; Wu, H.; Yang, L.; Lao, Y.; Lin, Q.; Yang, R. A Novel Respiratory Follow-up Robotic System for Thoracic-Abdominal Puncture. IEEE Trans. Ind. Electron. 2020, 68, 2368–2378. [Google Scholar] [CrossRef]
- Gloger, O.; Lehnert, B.; Schrade, A.; Völzke, H. Fully automated glottis segmentation in endoscopic videos using local color and shape features of glottal regions. IEEE Trans. Biomed. Eng. 2014, 62, 795–806. [Google Scholar] [CrossRef] [PubMed]
- Kirillov, A.; Mintun, E.; Ravi, N.; Mao, H.; Rolland, C.; Gustafson, L.; Xiao, T.; Whitehead, S.; Berg, A.C.; Lo, W.Y.; et al. Segment Anything. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Paris, France, 2–6 October 2023; pp. 4015–4026. [Google Scholar] [CrossRef]
- Huang, Y.; Yang, X.; Liu, L.; Zhou, H.; Chang, A.; Zhou, X.; Chen, R.; Yu, J.; Chen, J.; Chen, C.; et al. Segment Anything Model for Medical Images. Med. Image Anal. 2024, 92, 103061. [Google Scholar] [CrossRef] [PubMed]
- Cerrolaza, J.J.; Osma-Ruiz, V.; Sáenz-Lechón, N.; Villanueva, A.; Gutiérrez-Arriola, J.M.; Godino-Llorente, J.I.; Cabeza, R. Fully-automatic glottis segmentation with active shape models. In MAVEBA; Firenze University Press: Firenze, Italy, 2011; pp. 35–38. [Google Scholar]
- Karakozoglou, S.; Henrich, N.; d’Alessandro, C.; Stylianou, Y. Automatic Glottal Segmentation Using Local-Based Active Contours and Application to Glottovibrography. Speech Commun. 2012, 54, 641–654. [Google Scholar] [CrossRef]
- Fehling, M.K.; Grosch, F.; Schuster, M.E.; Schick, B.; Lohscheller, J. Fully Automatic Segmentation of Glottis and Vocal Folds in Endoscopic Laryngeal High-Speed Videos Using a Deep Convolutional LSTM Network. PLoS ONE 2020, 15, e0227791. [Google Scholar] [CrossRef] [PubMed]
- Deng, K.; Tao, B.; Hu, J. Unpaired Medical Image Enhancement Based on Generative Adversarial Networks. In Fifteenth International Conference on Information Optics and Photonics; SPIE: Philadelphia, PA, USA, 2024; Volume 13418, pp. 331–339. [Google Scholar] [CrossRef]
- He, K.; Gkioxari, G.; Dollár, P.; Girshick, R. Mask R-CNN. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 2980–2988. [Google Scholar] [CrossRef]
- Kirillov, A.; Wu, Y.; He, K.; Girshick, R. PointRend: Image Segmentation as Rendering. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Seattle, WA, USA, 13–19 June 2020; pp. 9796–9805. [Google Scholar] [CrossRef]
- Huang, Z.; Huang, L.; Gong, Y.; Huang, C.; Wang, X. Mask Scoring R-CNN. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Long Beach, CA, USA, 15–20 June 2019; Volume 2019, pp. 6402–6411. [Google Scholar] [CrossRef]
- Fang, Y.; Yang, S.; Wang, X.; Li, Y.; Fang, C.; Shan, Y.; Feng, B.; Liu, W. Instances as Queries. In Proceedings of the IEEE International Conference on Computer Vision, Montreal, BC, Canada, 11–17 October 2021; pp. 6910–6919. [Google Scholar]
- Bolya, D.; Zhou, C.; Xiao, F.; Lee, Y.J. Yolact: Real-time instance segmentation. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Seoul, Republic of Korea, 27 October–2 November 2019; pp. 9157–9166. [Google Scholar]
- Tian, Z.; Shen, C.; Wang, X.; Chen, H. BoxInst: High-Performance Instance Segmentation with Box Annotations. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Nashville, TN, USA, 19–25 June 2021; pp. 5439–5448. [Google Scholar] [CrossRef]
- Tian, Z.; Shen, C.; Chen, H. Conditional Convolutions for Instance Segmentation. In European Conference on Computer Vision (ECCV); Springer: Cham, Switzerland, 2020; Volume 12346, pp. 282–298. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, R.; Kong, T.; Li, L.; Shen, C. SOLOv2: Dynamic and fast instance segmentation. Adv. Neural Inf. Process. Syst. 2020, 33, 17721–17732. [Google Scholar]
- Lyu, C.; Zhang, W.; Huang, H.; Zhou, Y.; Wang, Y.; Liu, Y.; Zhang, S.; Chen, K. RTMDet: An Empirical Study of Designing Real-Time Object Detectors. arXiv 2022, arXiv:2212.07784. [Google Scholar] [CrossRef]
- Cheng, T.; Wang, X.; Chen, S.; Zhang, W.; Zhang, Q.; Huang, C.; Zhang, Z.; Liu, W. Sparse Instance Activation for Real-Time Instance Segmentation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), New Orleans, LA, USA, 18–24 June 2022; pp. 4423–4432. [Google Scholar] [CrossRef]
- Zhou, Z.; Rahman Siddiquee, M.M.; Tajbakhsh, N.; Liang, J. Unet++: A nested u-net architecture for medical image segmentation. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support: 4th International Workshop, DLMIA 2018, and 8th International Workshop, ML-CDS 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, September 20, 2018, Proceedings 4; Springer: Cham, Switzerland, 2018; pp. 3–11. [Google Scholar] [CrossRef]
- Huang, H.; Lin, L.; Tong, R.; Hu, H.; Zhang, Q.; Iwamoto, Y.; Han, X.; Chen, Y.W.; Wu, J. UNet 3+: A Full-Scale Connected UNet for Medical Image Segmentation. In Proceedings of the IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), Barcelona, Spain, 4–8 May 2020. [Google Scholar] [CrossRef]
- Derdiman, Y.S.; Koc, T. Deep learning model development with U-net architecture for glottis segmentation. In 2021 29th Signal Processing and Communications Applications Conference (SIU); IEEE: New York, NY, USA, 2021; pp. 1–4. [Google Scholar] [CrossRef]
- Chen, X.; Liu, Q.; Deng, H.H.; Kuang, T.; Lin, H.H.Y.; Xiao, D.; Gateno, J.; Xia, J.J.; Yap, P.T. Improving image segmentation with contextual and structural similarity. Pattern Recognit. 2024, 152, 110489. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Cheng, S.; Wang, L. Medical image segmentation based on dynamic positioning and region-aware attention. Pattern Recognit. 2024, 151, 110375. [Google Scholar] [CrossRef]
- Cao, G.; Sun, Z.; Wang, C.; Geng, H.; Fu, H.; Yin, Z.; Pan, M. RASNet: Renal automatic segmentation using an improved U-Net with multi-scale perception and attention unit. Pattern Recognit. 2024, 150, 110336. [Google Scholar] [CrossRef]
- Ma, J.; He, Y.; Li, F.; Han, L.; You, C.; Wang, B. Segment Anything in Medical Images. Nat. Commun. 2024, 15, 654. [Google Scholar] [CrossRef] [PubMed]
- Silva-Rodríguez, J.; Dolz, J.; Ayed, I.B. Towards foundation models and few-shot parameter-efficient fine-tuning for volumetric organ segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Cham, Switzerland, 2023; pp. 213–224. [Google Scholar] [CrossRef]
- Wen, J.; Qin, F.; Du, J.; Fang, M.; Wei, X.; Chen, C.L.P.; Li, P. Msgfusion: Medical semantic guided two-branch network for multimodal brain image fusion. IEEE Trans. Multimed. 2023, 26, 944–957. [Google Scholar] [CrossRef]
- Dai, Z.; Yi, J.; Yan, L.; Xu, Q.; Hu, L.; Zhang, Q.; Li, J.; Wang, G. PFEMed: Few-shot medical image classification using prior guided feature enhancement. Pattern Recognit. 2023, 134, 109108. [Google Scholar] [CrossRef]
- Zhao, X.; Zhang, P.; Song, F.; Ma, C.; Fan, G.; Sun, Y.; Feng, Y.; Zhang, G. Prior attention network for multi-lesion segmentation in medical images. IEEE Trans. Med. Imaging 2022, 41, 3812–3823. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Han, S.; Huang, S.; Liao, Y.; Li, G.; Kong, X.; Liu, S. Mask-enhanced segment anything model for tumor lesion semantic segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Cham, Switzerland, 2024; pp. 403–413. [Google Scholar] [CrossRef]
- You, X.; He, J.; Yang, J.; Gu, Y. Learning with explicit shape priors for medical image segmentation. IEEE Trans. Med. Imaging 2024, 44, 927–940. [Google Scholar] [CrossRef] [PubMed]
- Hu, E.J.; Shen, Y.; Wallis, P.; Allen-Zhu, Z.; Li, Y.; Wang, S.; Wang, L.; Chen, W. LoRA: Low-Rank Adaptation of Large Language Models. Int. Conf. Learn. Represent. 2022, 1, 3. [Google Scholar] [CrossRef]
- Gómez, P.; Kist, A.M.; Schlegel, P.; Berry, D.A.; Chhetri, D.K.; Dürr, S.; Echternach, M.; Johnson, A.M.; Kniesburges, S.; Kunduk, M.; et al. BAGLS, a Multihospital Benchmark for Automatic Glottis Segmentation. Sci. Data 2020, 7, 186. [Google Scholar] [CrossRef] [PubMed]
- Lin, T.Y.; Maire, M.; Belongie, S.; Hays, J.; Perona, P.; Ramanan, D.; Dollár, P.; Zitnick, C.L. Microsoft COCO: Common Objects in Context. In European Conference on Computer Vision (ECCV); Springer: Cham, Switzerland, 2014; Volume 8693, pp. 740–755. [Google Scholar] [CrossRef]
- Hao, R.; Lai, J.; Zhong, W.; Xie, D.; Tian, Y.; Zhang, T.; Zhang, Y.; Chan, C.P.L.; Chan, J.Y.K.; Ren, H. Variable-stiffness nasotracheal intubation robot with passive buffering: A modular platform in mannequin studies. In 2025 IEEE International Conference on Robotics and Automation (ICRA); IEEE: New York, NY, USA, 2025; pp. 1–8. [Google Scholar]
- Cheng, B.; Misra, I.; Schwing, A.G.; Kirillov, A.; Girdhar, R. Masked-Attention Mask Transformer for Universal Image Segmentation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), New Orleans, LA, USA, 18–24 June 2022; pp. 1280–1289. [Google Scholar] [CrossRef]
- Chen, K.; Liu, C.; Chen, H.; Zhang, H.; Li, W.; Zou, Z.; Shi, Z. RSPrompter: Learning to Prompt for Remote Sensing Instance Segmentation Based on Visual Foundation Model. IEEE Trans. Geosci. Remote Sens. 2024, 62, 4701117. [Google Scholar] [CrossRef]
- Zhao, X.; Ding, W.; An, Y.; Du, Y.; Yu, T.; Li, M.; Tang, M.; Wang, J. Fast Segment Anything. arXiv 2023, arXiv:2306.12156. [Google Scholar] [CrossRef]






| Stage | Block | Configuration | Output Size |
|---|---|---|---|
| 1 | MBConv | × 2 | 56 × 56 |
| 2 | Transformer | × 2 | 28 × 28 |
| 3 | Transformer | × 6 | 14 × 14 |
| 4 | Transformer | × 2 | 7 × 7 |
| Models | Backbones | Glottis Dataset | Phantom Dataset | Clinical Dataset | Model Size (MB) | Infer (FPS) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| mAP | AP50 | mDice | mIoU | mAP | AP50 | mDice | mIoU | mAP | AP50 | mDice | mIoU | ||||
| BoxInst [16] | ResNet101 | 10.6 | 31.8 | 34.4 | 20.8 | 25.1 | 45.2 | 64.1 | 47.2 | 8.8 | 27.1 | 33.0 | 19.8 | 447.3 | 45.7 |
| CondInst [17] | ResNet50 | 13.5 | 39.1 | 43.9 | 28.1 | 31.0 | 55.4 | 77.1 | 62.7 | 13.2 | 36.5 | 43.3 | 27.6 | 279.2 | 46.8 |
| Mask-RCNN [11] | ResNet101 | 44.7 | 79.0 | 79.6 | 66.1 | 39.7 | 65.6 | 70.4 | 54.3 | 30.1 | 55.3 | 66.8 | 50.2 | 506.3 | 42.2 |
| Mask-RCNN [11] | Swin-T | 47.1 | 82.9 | 83.5 | 71.7 | 40.0 | 64.9 | 67.8 | 51.3 | 28.3 | 53.1 | 65.0 | 48.1 | 572.0 | 38.2 |
| MS-RCNN [13] | X101 | 45.5 | 78.1 | 79.0 | 65.3 | 31.3 | 57.0 | 62.9 | 45.9 | 25.5 | 50.0 | 60.2 | 43.1 | 634.4 | 43.0 |
| PointRend [12] | ResNet50 | 45.1 | 81.8 | 82.9 | 70.8 | 40.5 | 64.4 | 65.2 | 48.4 | 30.9 | 55.7 | 66.5 | 49.8 | 447.3 | 41.4 |
| Mask2Former [38] | ResNet101 | 37.6 | 79.7 | 83.0 | 70.9 | 36.7 | 56.9 | 72.9 | 57.4 | 23.4 | 52.1 | 59.8 | 42.7 | 787.1 | 19.8 |
| Mask2Former [38] | Swin-T | 34.5 | 76.5 | 80.1 | 66.8 | 34.0 | 53.3 | 61.9 | 44.8 | 28.3 | 53.1 | 65.0 | 48.1 | 599.6 | 18.8 |
| QueryInst [14] | ResNet50 | 42.2 | 84.9 | 89.2 | 80.5 | 18.7 | 33.8 | 45.5 | 29.4 | 15.6 | 32.6 | 44.5 | 28.6 | 208.7 | 30.8 |
| RTMDet [19] | CSPDarkNet | 42.4 | 83.2 | 86.5 | 76.2 | 23.0 | 39.6 | 42.3 | 26.8 | 19.3 | 40.6 | 50.7 | 34.0 | 977.7 | 37.2 |
| U-Net++ [21] | – | – | – | 78.3 | 64.3 | – | – | 69.3 | 53.0 | – | – | 49.6 | 33.0 | 32 | 55.6 |
| U-Net3+ [22] | – | – | – | 77.3 | 63.0 | – | – | 54.0 | 37.0 | – | – | 35.9 | 21.9 | 104 | 33.3 |
| SOLOv2 [18] | ResNet50 | 22.5 | 57.0 | 58.2 | 41.0 | 13.3 | 37.0 | 32.7 | 19.5 | 4.6 | 17.4 | 25.8 | 14.8 | 371.6 | 45.9 |
| SparseInst [20] | ResNet50 | 52.1 | 89.1 | 90.6 | 82.8 | 28.5 | 44.6 | 51.4 | 34.6 | 19.7 | 43.5 | 48.3 | 31.8 | 418.5 | 69.9 |
| YOLACT [15] | ResNet101 | 48.7 | 90.4 | 91.5 | 84.3 | 40.9 | 66.7 | 71.4 | 55.5 | 24.2 | 50.2 | 61.3 | 44.2 | 460.8 | 72.8 |
| RSPrompter [39] | ViT-B | 14.6 | 39.6 | 45.5 | 29.4 | 15.1 | 31.7 | 33.4 | 20.0 | 5.4 | 15.1 | 25.4 | 14.5 | 470.0 | 12.2 |
| SAM [5] | ViT-B | 52.7 | 90.4 | 92.7 | 86.4 | 40.7 | 64.3 | 67.2 | 50.6 | 28.2 | 50.5 | 64.6 | 47.7 | 441.5 | 12.8 |
| FastSAM-S [40] | CSPDarkNet | 56.9 | 92.7 | – | – | 39.4 | 59.8 | – | – | 27.5 | 46.2 | – | – | 148.5 | 25.9 |
| FastSAM-X [40] | CSPDarkNet | 57.2 | 93.5 | – | – | 35.1 | 55.5 | – | – | 26.5 | 43.4 | – | – | 868.0 | 7.4 |
| Glottis-SAM (Ours) | ViT-B | 53.8 | 92.0 | 93.0 | 86.9 | 44.7 | 71.3 | 74.4 | 59.2 | 28.8 | 55.0 | 65.7 | 48.9 | 392.6 | 13.2 |
| Glottis-SAM (Ours) | ViT-T | 55.8 | 93.6 | 94.7 | 89.9 | 43.8 | 66.8 | 73.1 | 57.6 | 34.4 | 63.2 | 72.6 | 57.0 | 55.2 | 44.3 |
| QKV | FC1 | FC2 | mAP | AP50 | AP75 | mDice | Model Size (MB) |
|---|---|---|---|---|---|---|---|
| × | × | × | 40.7 | 64.3 | 49.2 | 66.9 | 445.2 |
| ✓ | × | × | 43.3 | 66.1 | 49.7 | 69.6 | 417.1 |
| ✓ | ✓ | × | 42.3 | 65.1 | 49.0 | 67.8 | 417.5 |
| ✓ | × | ✓ | 42.9 | 65.4 | 50.9 | 68.2 | 414.6 |
| ✓ | ✓ | ✓ | 43.0 | 67.4 | 51.0 | 70.5 | 414.1 |
| α | r | mAP | AP50 | AP75 | mDice | Trainable Params (M) |
|---|---|---|---|---|---|---|
| 64 | 2 | 38.8 | 63.1 | 44.8 | 66.7 | 0.07 |
| 4 | 40.1 | 63.9 | 44.5 | 68.1 | 0.15 | |
| 8 | 41.2 | 65.5 | 47.9 | 70.1 | 0.29 | |
| 16 | 41.8 | 66.6 | 48.1 | 69.8 | 0.59 | |
| 32 | 40.8 | 66.3 | 45.9 | 68.7 | 1.38 | |
| 64 | 42.6 | 65.6 | 49.5 | 69.6 | 2.36 | |
| 32 | 2 | 41.5 | 64.8 | 47.9 | 68.1 | 0.07 |
| 4 | 42.7 | 67.4 | 48.4 | 70.5 | 0.15 | |
| 8 | 41.3 | 65.6 | 44.2 | 69.1 | 0.29 | |
| 16 | 41.9 | 66.7 | 47.1 | 69.9 | 0.59 | |
| 32 | 41.6 | 66.3 | 47.1 | 69.6 | 1.38 | |
| 64 | 41.3 | 66.3 | 45.9 | 67.9 | 2.36 |
| Channels | mAP | AP50 | AP75 | mDice | Infer (FPS) |
|---|---|---|---|---|---|
| 256 | 41.8 | 66.8 | 46.6 | 70.3 | 11.40 |
| 128 | 40.1 | 64.3 | 45.0 | 67.5 | 11.47 |
| 64 | 42.5 | 67.2 | 48.6 | 70.7 | 12.16 |
| 32 | 41.7 | 65.2 | 46.1 | 68.7 | 12.52 |
| 16 | 38.6 | 60.2 | 42.4 | 67.1 | 12.54 |
| 8 | 16.5 | 36.5 | 11.9 | 44.9 | 12.87 |
| λ1 | λ2 | mAP | AP50 | AP75 | mDice | Model Size (MB) |
|---|---|---|---|---|---|---|
| 0.5 | 2 | 44.7 | 71.3 | 51.9 | 74.4 | 421.3 |
| 1 | 0.5 | 41.7 | 64.3 | 47.9 | 67.9 | 427.0 |
| 1 | 1 | 43.0 | 67.8 | 50.9 | 73.3 | 414.7 |
| 1 | 2 | 43.6 | 66.9 | 50.8 | 70.2 | 412.2 |
| 2 | 0.5 | 41.2 | 64.3 | 48.5 | 68.1 | 410.5 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Liu, J.; Zhou, Y.; Hao, R.; Li, M.; Zhang, Y.; Ren, H. Integrating Dynamic Representation and Multi-Priors for Transnasal Intubation via Visual Foundation Model. Bioengineering 2026, 13, 217. https://doi.org/10.3390/bioengineering13020217
Liu J, Zhou Y, Hao R, Li M, Zhang Y, Ren H. Integrating Dynamic Representation and Multi-Priors for Transnasal Intubation via Visual Foundation Model. Bioengineering. 2026; 13(2):217. https://doi.org/10.3390/bioengineering13020217
Chicago/Turabian StyleLiu, Jinyu, Yang Zhou, Ruoyi Hao, Mingying Li, Yang Zhang, and Hongliang Ren. 2026. "Integrating Dynamic Representation and Multi-Priors for Transnasal Intubation via Visual Foundation Model" Bioengineering 13, no. 2: 217. https://doi.org/10.3390/bioengineering13020217
APA StyleLiu, J., Zhou, Y., Hao, R., Li, M., Zhang, Y., & Ren, H. (2026). Integrating Dynamic Representation and Multi-Priors for Transnasal Intubation via Visual Foundation Model. Bioengineering, 13(2), 217. https://doi.org/10.3390/bioengineering13020217








