Explainability of a Deep Learning Model for Mediastinal Lymph Node Station Classification in Endobronchial Ultrasound (EBUS)
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Preoperative Preparations
2.3. Intraoperative Workflow
2.4. Postoperative Image Processing
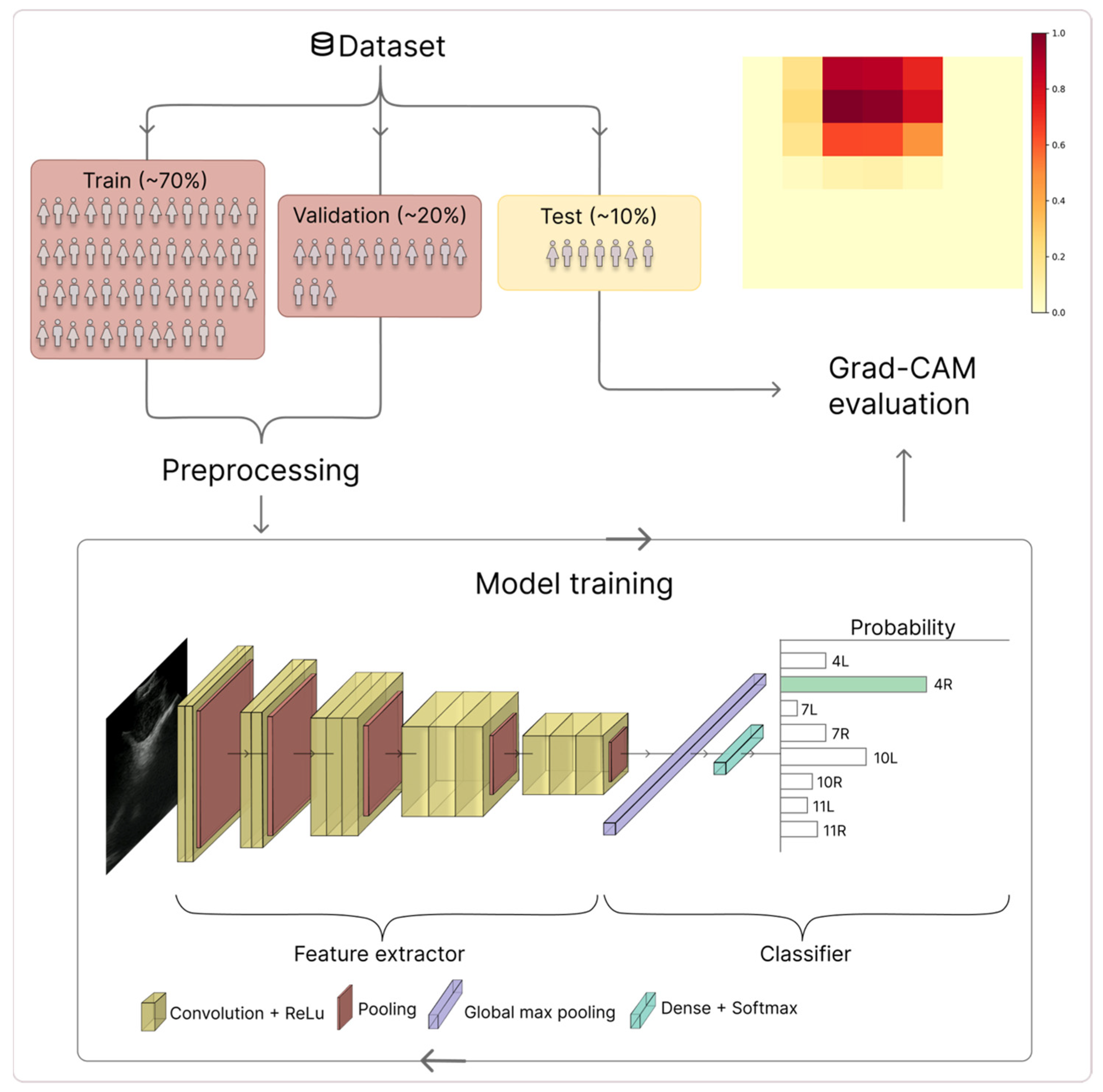
2.4.1. CNN
2.4.2. Grad-CAM Generation
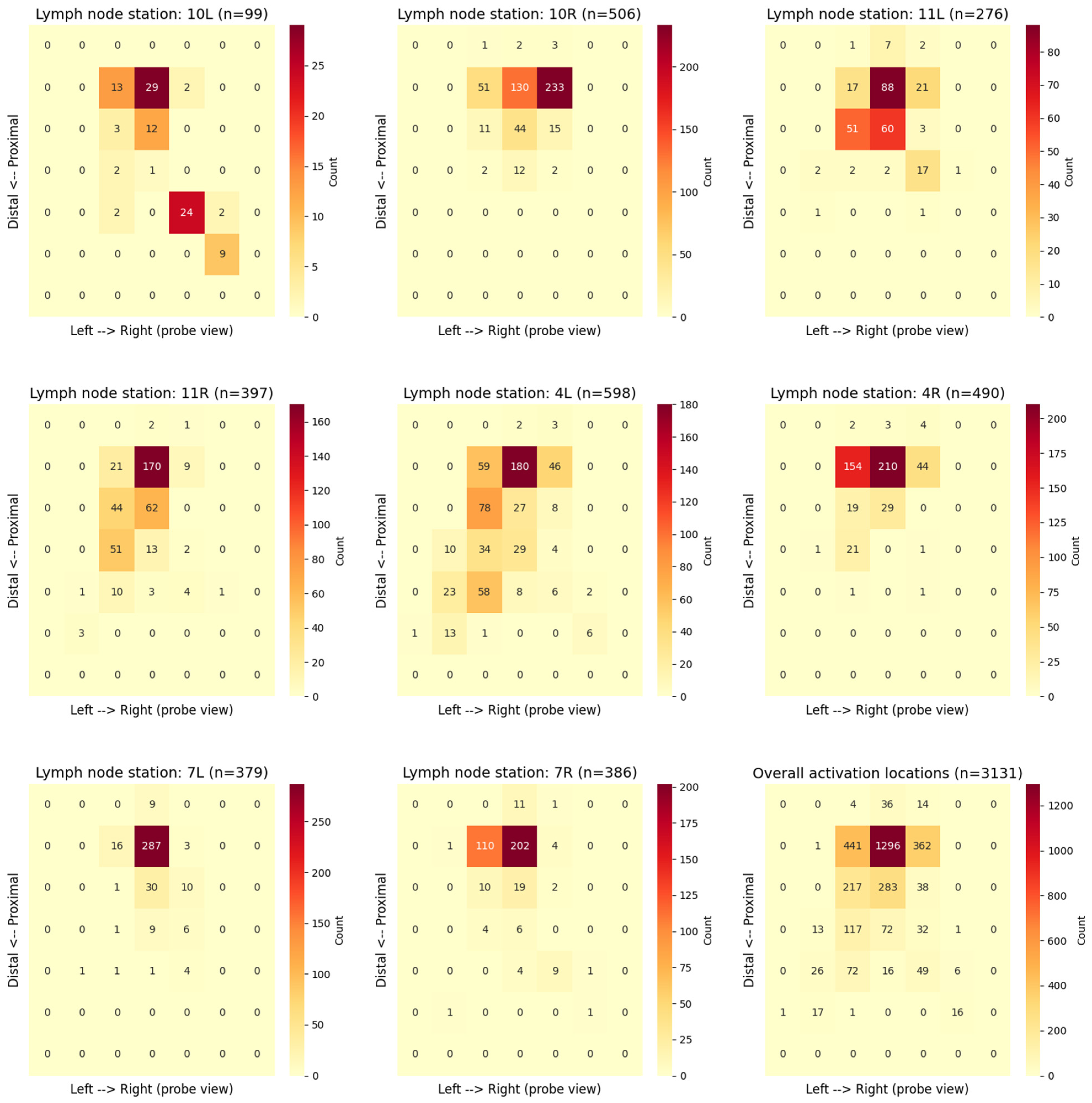
2.4.3. Grad-CAM Activation Maps
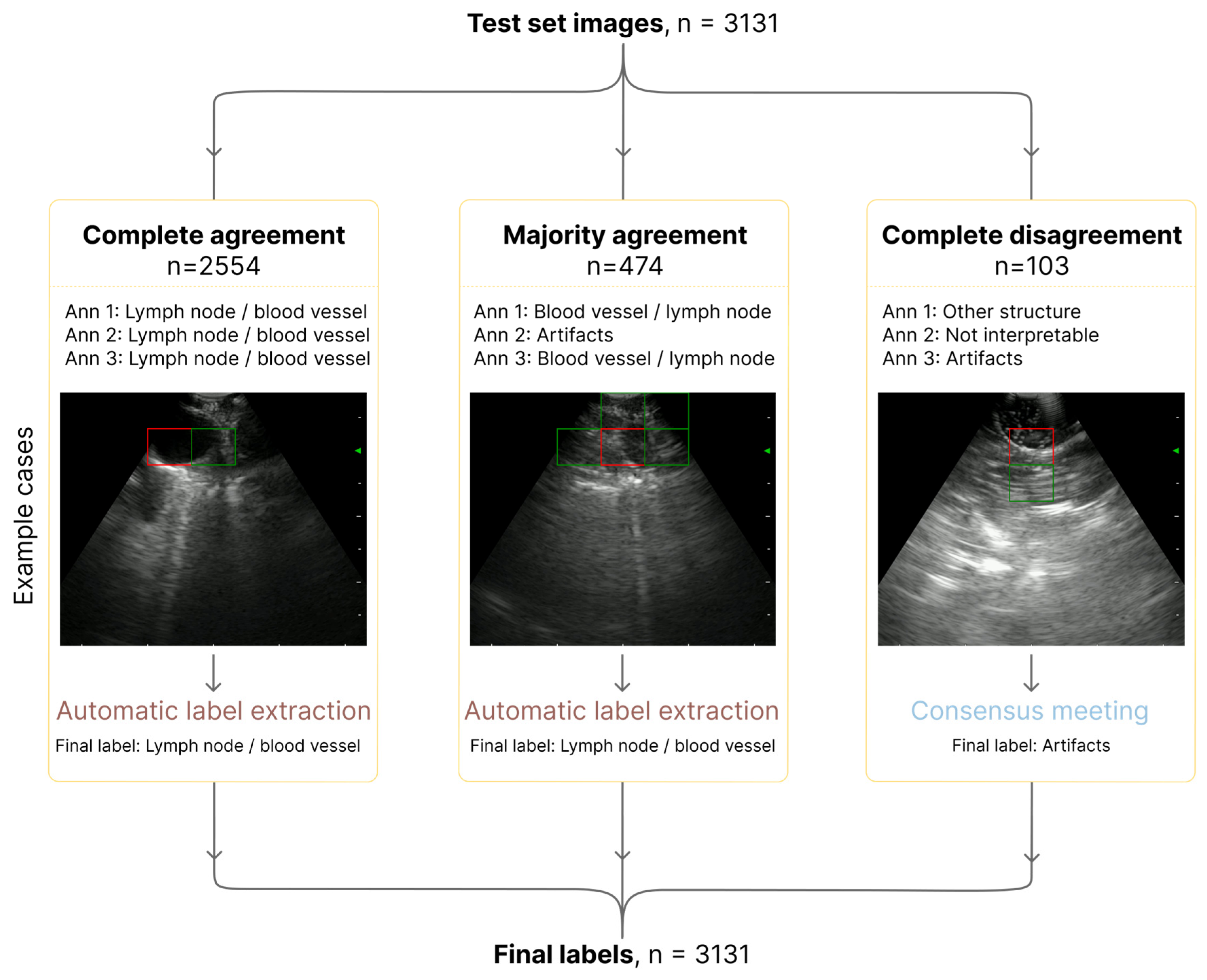
2.4.4. Expert Annotation of Grad-CAM Heatmaps
2.4.5. Quality Control and Consensus
3. Results
3.1. Dataset and CNN Classification Performance
3.2. Grad-CAM Activation Patterns
3.3. Inter-Observer Annotation Agreement
3.3.1. Annotation Workflow
3.3.2. Agreement Results
3.4. Anatomical Relevance of the Model’s Attention
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AI | Artificial intelligence |
| CNN | Convolutional neural network |
| DL | Deep learning |
| EBUS | Endobronchial ultrasound |
| EBUS-TBNA | Endobronchial ultrasound-guided transbronchial needle aspiration |
| EUS | Endoscopic ultrasound |
| Grad-CAM | Gradient-weighted Class Activation Mapping |
| IASLC | International Association for the Study of Lung Cancer |
| LSTM | Long short-term memory |
| CT | Computed tomography |
| PET-CT | Positron emission tomography–computed tomography |
| TNM | Tumor–node–metastasis |
| XAI | Explainable artificial intelligence |
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- WHO. Global Health Estimates: Leading Causes of Death and Disability; World Health Organization: Geneva, Switzerland, 2023. [Google Scholar]
- Postmus, P.E.; Kerr, K.M.; Oudkerk, M.; Senan, S.; Waller, D.A.; Vansteenkiste, J.; Escriu, C.; Peters, S. Early and locally advanced non-small-cell lung cancer (NSCLC): ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2017, 28, iv1–iv21. [Google Scholar] [CrossRef]
- De Leyn, P.; Dooms, C.; Kuzdzal, J.; Lardinois, D.; Passlick, B.; Rami-Porta, R.; Turna, A.; Van Schil, P.; Venuta, F.; Waller, D.; et al. Revised ESTS guidelines for preoperative mediastinal lymph node staging for non-small-cell lung cancer. Eur. J. Cardiothorac. Surg. 2014, 45, 787–798. [Google Scholar] [CrossRef]
- Silvestri, G.A.; Gonzalez, A.V.; Jantz, M.A.; Margolis, M.L.; Gould, M.K.; Tanoue, L.T.; Harris, L.J.; Detterbeck, F.C. Methods for staging non-small cell lung cancer: Diagnosis and management of lung cancer, 3rd ed: American College of Chest Physicians evidence-based clinical practice guidelines. Chest 2013, 143, e211S–e250S. [Google Scholar] [CrossRef]
- Vilmann, P.; Clementsen, P.F.; Colella, S.; Siemsen, M.; De Leyn, P.; Dumonceau, J.M.; Herth, F.J.; Larghi, A.; Vazquez-Sequeiros, E.; Hassan, C.; et al. Combined endobronchial and esophageal endosonography for the diagnosis and staging of lung cancer: European Society of Gastrointestinal Endoscopy (ESGE) Guideline, in cooperation with the European Respiratory Society (ERS) and the European Society of Thoracic Surgeons (ESTS). Endoscopy 2015, 47, 545–559. [Google Scholar] [CrossRef]
- Annema, J.T.; Rabe, K.F. Endosonography for lung cancer staging: One scope fits all? Chest 2010, 138, 765–767. [Google Scholar] [CrossRef] [PubMed]
- Rami-Porta, R.; Nishimura, K.K.; Giroux, D.J.; Detterbeck, F.; Cardillo, G.; Edwards, J.G.; Fong, K.M.; Giuliani, M.; Huang, J.; Kernstine, K.H., Sr.; et al. The International Association for the Study of Lung Cancer Lung Cancer Staging Project: Proposals for Revision of the TNM Stage Groups in the Forthcoming (Ninth) Edition of the TNM Classification for Lung Cancer. J. Thorac. Oncol. 2024, 19, 1007–1027. [Google Scholar] [CrossRef] [PubMed]
- Mountain, C.F.; Dresler, C.M. Regional lymph node classification for lung cancer staging. Chest 1997, 111, 1718–1723. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Osarogiagbon, R.U.; Giroux, D.J.; Nishimura, K.K.; Bille, A.; Cardillo, G.; Detterbeck, F.; Kernstine, K.; Kim, H.K.; Lievens, Y.; et al. The International Association for the Study of Lung Cancer Staging Project for Lung Cancer: Proposals for the Revision of the N Descriptors in the Forthcoming Ninth Edition of the TNM Classification for Lung Cancer. J. Thorac. Oncol. 2024, 19, 766–785. [Google Scholar] [CrossRef]
- Detterbeck, F.C.; Woodard, G.A.; Bader, A.S.; Dacic, S.; Grant, M.J.; Park, H.S.; Tanoue, L.T. The Proposed Ninth Edition TNM Classification of Lung Cancer. Chest 2024, 166, 882–895. [Google Scholar] [CrossRef]
- Davoudi, M.; Colt, H.G.; Osann, K.E.; Lamb, C.R.; Mullon, J.J. Endobronchial ultrasound skills and tasks assessment tool: Assessing the validity evidence for a test of endobronchial ultrasound-guided transbronchial needle aspiration operator skill. Am. J. Respir. Crit. Care Med. 2012, 186, 773–779. [Google Scholar] [CrossRef]
- Fernández-Villar, A.; Leiro-Fernández, V.; Botana-Rial, M.; Represas-Represas, C.; Núñez-Delgado, M. The endobronchial ultrasound-guided transbronchial needle biopsy learning curve for mediastinal and hilar lymph node diagnosis. Chest 2012, 141, 278–279. [Google Scholar] [CrossRef]
- Folch, E.; Majid, A. Point: Are >50 supervised procedures required to develop competency in performing endobronchial ultrasound-guided transbronchial needle aspiration for mediastinal staging? Yes. Chest 2013, 143, 888–891. [Google Scholar] [CrossRef]
- Bi, W.L.; Hosny, A.; Schabath, M.B.; Giger, M.L.; Birkbak, N.J.; Mehrtash, A.; Allison, T.; Arnaout, O.; Abbosh, C.; Dunn, I.F.; et al. Artificial intelligence in cancer imaging: Clinical challenges and applications. CA Cancer J. Clin. 2019, 69, 127–157. [Google Scholar] [CrossRef]
- Kufel, J.; Bargieł-Łączek, K.; Kocot, S.; Koźlik, M.; Bartnikowska, W.; Janik, M.; Czogalik, Ł.; Dudek, P.; Magiera, M.; Lis, A.; et al. What Is Machine Learning, Artificial Neural Networks and Deep Learning?-Examples of Practical Applications in Medicine. Diagnostics 2023, 13, 2582. [Google Scholar] [CrossRef] [PubMed]
- Zang, X.; Bascom, R.; Gilbert, C.; Toth, J.; Higgins, W. Methods for 2-D and 3-D Endobronchial Ultrasound Image Segmentation. IEEE Trans. Biomed. Eng. 2016, 63, 1426–1439. [Google Scholar] [CrossRef] [PubMed]
- Yong, S.H.; Lee, S.H.; Oh, S.I.; Keum, J.S.; Kim, K.N.; Park, M.S.; Chang, Y.S.; Kim, E.Y. Malignant thoracic lymph node classification with deep convolutional neural networks on real-time endobronchial ultrasound (EBUS) images. Transl. Lung Cancer Res. 2022, 11, 14–23. [Google Scholar] [CrossRef]
- Churchill, I.F.; Gatti, A.A.; Hylton, D.A.; Sullivan, K.A.; Patel, Y.S.; Leontiadis, G.I.; Farrokhyar, F.; Hanna, W.C. An Artificial Intelligence Algorithm to Predict Nodal Metastasis in Lung Cancer. Ann. Thorac. Surg. 2022, 114, 248–256. [Google Scholar] [CrossRef]
- Ozcelik, N.; Ozcelik, A.E.; Bulbul, Y.; Oztuna, F.; Ozlu, T. Can artificial intelligence distinguish between malignant and benign mediastinal lymph nodes using sonographic features on EBUS images? Curr. Med. Res. Opin. 2020, 36, 2019–2024. [Google Scholar] [CrossRef] [PubMed]
- Patel, Y.S.; Gatti, A.A.; Farrokhyar, F.; Xie, F.; Hanna, W.C. Artificial Intelligence Algorithm Can Predict Lymph Node Malignancy from Endobronchial Ultrasound Transbronchial Needle Aspiration Images for Non-Small Cell Lung Cancer. Respiration 2024, 103, 741–751. [Google Scholar] [CrossRef]
- Koseoglu, F.D.; Alıcı, I.O.; Er, O. Machine learning approaches in the interpretation of endobronchial ultrasound images: A comparative analysis. Surg. Endosc. 2023, 37, 9339–9346. [Google Scholar] [CrossRef]
- Ervik, Ø.; Tveten, I.; Hofstad, E.F.; Langø, T.; Leira, H.O.; Amundsen, T.; Sorger, H. Automatic Segmentation of Mediastinal Lymph Nodes and Blood Vessels in Endobronchial Ultrasound (EBUS) Images Using Deep Learning. J. Imaging 2024, 10, 190. [Google Scholar] [CrossRef]
- Winiarski, S.; Radziszewski, M.; Wiśniewski, M.; Cisek, J.; Wąsowski, D.; Plewczynski, D.; Górska, K.; Korczynski, P. Integrating Artificial Intelligence in Bronchoscopy and Endobronchial Ultrasound (EBUS) for Lung Cancer Diagnosis and Staging: A Comprehensive Review. Cancers 2025, 17, 2835. [Google Scholar] [CrossRef]
- Ishiwata, T.; Inage, T.; Aragaki, M.; Gregor, A.; Chen, Z.; Bernards, N.; Kafi, K.; Yasufuku, K. Deep learning-based prediction of nodal metastasis in lung cancer using endobronchial ultrasound. JTCVS Tech. 2024, 28, 151–161. [Google Scholar] [CrossRef]
- Yao, L.; Zhang, C.; Xu, B.; Yi, S.; Li, J.; Ding, X.; Yu, H. A deep learning-based system for mediastinum station localization in linear EUS (with video). Endosc. Ultrasound 2023, 12, 417–423. [Google Scholar] [CrossRef]
- Ervik, Ø.; Rødde, M.; Hofstad, E.F.; Tveten, I.; Langø, T.; Leira, H.O.; Amundsen, T.; Sorger, H. A New Deep Learning-Based Method for Automated Identification of Thoracic Lymph Node Stations in Endobronchial Ultrasound (EBUS): A Proof-of-Concept Study. J. Imaging 2025, 11, 10. [Google Scholar] [CrossRef]
- Leira, H.O.; Langø, T.; Sorger, H.; Hofstad, E.F.; Amundsen, T. Bronchoscope-induced displacement of lung targets: First in vivo demonstration of effect from wedging maneuver in navigated bronchoscopy. J. Bronchol. Interv. Pulmonol. 2013, 20, 206–212. [Google Scholar] [CrossRef]
- Sorger, H.; Hofstad, E.F.; Amundsen, T.; Langø, T.; Bakeng, J.B.; Leira, H.O. A multimodal image guiding system for Navigated Ultrasound Bronchoscopy (EBUS): A human feasibility study. PLoS ONE 2017, 12, e0171841. [Google Scholar] [CrossRef] [PubMed]
- Malay, S.; Vishal, A.; RameshBabu, C.S. Techniques of Linear Endobronchial Ultrasound. In Ultrasound Imaging; Igor, V.M., Oleg, V.M., Eds.; IntechOpen: Rijeka, Croatia, 2011; pp. 157–179. [Google Scholar]
- Holzinger, A.; Langs, G.; Denk, H.; Zatloukal, K.; Müller, H. Causability and explainability of artificial intelligence in medicine. Wiley Interdiscip. Rev. Data Min. Knowl. Discov. 2019, 9, e1312. [Google Scholar] [CrossRef] [PubMed]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017. [Google Scholar]
- Muhammad, D.; Bendechache, M. Unveiling the black box: A systematic review of Explainable Artificial Intelligence in medical image analysis. Comput. Struct. Biotechnol. J. 2024, 24, 542–560. [Google Scholar] [CrossRef]
- Lundberg, S.M.; Lee, S.-I. A Unified Approach to Interpreting Model Predictions. In Proceedings of the 31st International Conference on Neural Information Processing Systems, Long Beach, CA, USA, 4–9 December 2017. [Google Scholar]
- Lin, C.K.; Wu, S.H.; Chua, Y.W.; Fan, H.J.; Cheng, Y.C. TransEBUS: The interpretation of endobronchial ultrasound image using hybrid transformer for differentiating malignant and benign mediastinal lesions. J. Formos. Med. Assoc. 2025, 124, 28–37. [Google Scholar] [CrossRef] [PubMed]
- Leong, T.L.; Loveland, P.M.; Gorelik, A.; Irving, L.; Steinfort, D.P. Preoperative Staging by EBUS in cN0/N1 Lung Cancer: Systematic Review and Meta-Analysis. J. Bronchol. Interv. Pulmonol. 2019, 26, 155–165. [Google Scholar] [CrossRef]
- Rusch, V.W.; Asamura, H.; Watanabe, H.; Giroux, D.J.; Rami-Porta, R.; Goldstraw, P. The IASLC Lung Cancer Staging Project: A Proposal for a New International Lymph Node Map in the Forthcoming Seventh Edition of the TNM Classification for Lung Cancer. J. Thorac. Oncol. 2009, 4, 568–577. [Google Scholar] [CrossRef]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Russakovsky, O.; Deng, J.; Su, H.; Krause, J.; Satheesh, S.; Ma, S.; Huang, Z.; Karpathy, A.; Khosla, A.; Bernstein, M.; et al. ImageNet Large Scale Visual Recognition Challenge. Int. J. Comput. Vis. 2015, 115, 211–252. [Google Scholar] [CrossRef]
- TensorFlow. tf.keras.optimizers.AdamW—TensorFlow Documentation. Available online: https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/AdamW (accessed on 6 December 2025).
- Fırat, M.; Fırat, İ.T.; Üstündağ, Z.F.; Öztürk, E.; Tuncer, T. AI-Based Response Classification After Anti-VEGF Loading in Neovascular Age-Related Macular Degeneration. Diagnostics 2025, 15, 2253. [Google Scholar] [CrossRef]
- Obaid, W.; Hussain, A.; Rabie, T.; Mansoor, W. Noisy Ultrasound Kidney Image Classifications Using Deep Learning Ensembles and Grad-CAM Analysis. AI 2025, 6, 172. [Google Scholar] [CrossRef]
- Smistad, E.; Østvik, A.; Løvstakken, L. Annotation Web—An open-source web-based annotation tool for ultrasound images. In Proceedings of the 2021 IEEE International Ultrasonics Symposium (IUS), Xi’an, China, 11–16 September 2021. [Google Scholar]
- Navani, N.; Molyneaux, P.L.; Breen, R.A.; Connell, D.W.; Jepson, A.; Nankivell, M.; Brown, J.M.; Morris-Jones, S.; Ng, B.; Wickremasinghe, M.; et al. Utility of endobronchial ultrasound-guided transbronchial needle aspiration in patients with tuberculous intrathoracic lymphadenopathy: A multicentre study. Thorax 2011, 66, 889–893. [Google Scholar] [CrossRef] [PubMed]
- Yasufuku, K.; Chiyo, M.; Koh, E.; Moriya, Y.; Iyoda, A.; Sekine, Y.; Shibuya, K.; Iizasa, T.; Fujisawa, T. Endobronchial ultrasound guided transbronchial needle aspiration for staging of lung cancer. Lung Cancer 2005, 50, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Chattopadhay, A.; Sarkar, A.; Howlader, P.; Balasubramanian, V.N. Grad-cam++: Generalized gradient-based visual explanations for deep convolutional networks. In Proceedings of the 2018 IEEE Winter Conference on Applications of Computer Vision (WACV), Lake Tahoe, NV, USA, 12–15 March 2018. [Google Scholar]




| Hyperparameter | Value |
|---|---|
| Epochs | 100 |
| Batch size | 32 |
| Optimizer | AdamW [40] |
| Loss function | Categorical cross-entropy |
| Learning rate | 1 × 10−4 |
| Patience | 20 |
| Lymph Node Station | Precision (%) | Sensitivity (%) | F1-Score (%) | Number of Images |
|---|---|---|---|---|
| 4L | 77.6 | 81.6 | 79.5 | 598 |
| 4R | 75.6 | 72.7 | 74.1 | 490 |
| 7L | 76.6 | 38.8 | 51.5 | 379 |
| 7R | 52.1 | 69.9 | 59.7 | 386 |
| 10L | 50.7 | 36.4 | 42.4 | 99 |
| 10R | 85.8 | 59.9 | 70.5 | 506 |
| 11L | 36.2 | 60.9 | 45.4 | 276 |
| 11R | 47.8 | 52.1 | 49.9 | 397 |
| Total | 3131 |
| Metric | Precision (%) | Sensitivity (%) | F1-Score (%) | Number of Images |
|---|---|---|---|---|
| Macro average | 62.8 | 59.0 | 59.1 | 3131 |
| Weighted average | 67.1 | 63.1 | 63.5 | 3131 |
| Agreement Metric | Annotator Pair | Value | Interpretation |
|---|---|---|---|
| Percent agreement | All annotators | 81.6% | – |
| Fleiss’ kappa | All annotators | 0.529 | Moderate agreement |
| Cohen’s kappa | Annotator 1–Annotator 2 | 0.598 | Moderate agreement |
| Cohen’s kappa | Annotator 1–Annotator 3 | 0.531 | Moderate agreement |
| Cohen’s kappa | Annotator 2–Annotator 3 | 0.474 | Moderate agreement |
| Label | Accuracy (%) | Precision (%) | Sensitivity (%) | F1-Score (%) | Number of Images |
|---|---|---|---|---|---|
| Lymph node/blood vessel | 65.9 | 61.7 | 59.5 | 58.4 | 2715 |
| Artifacts | 45.9 | 47.6 | 40.6 | 40.4 | 196 |
| Not interpretable | 45.3 | 32.8 | 29.9 | 29.6 | 201 |
| Other structure | 21.1 | 45.8 | 48.3 | 34.2 | 19 |
| Total | 3131 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Ervik, Ø.; Rødde, M.; Hofstad, E.F.; Langø, T.; Leira, H.O.; Amundsen, T.; Sorger, H. Explainability of a Deep Learning Model for Mediastinal Lymph Node Station Classification in Endobronchial Ultrasound (EBUS). Bioengineering 2026, 13, 198. https://doi.org/10.3390/bioengineering13020198
Ervik Ø, Rødde M, Hofstad EF, Langø T, Leira HO, Amundsen T, Sorger H. Explainability of a Deep Learning Model for Mediastinal Lymph Node Station Classification in Endobronchial Ultrasound (EBUS). Bioengineering. 2026; 13(2):198. https://doi.org/10.3390/bioengineering13020198
Chicago/Turabian StyleErvik, Øyvind, Mia Rødde, Erlend Fagertun Hofstad, Thomas Langø, Håkon O. Leira, Tore Amundsen, and Hanne Sorger. 2026. "Explainability of a Deep Learning Model for Mediastinal Lymph Node Station Classification in Endobronchial Ultrasound (EBUS)" Bioengineering 13, no. 2: 198. https://doi.org/10.3390/bioengineering13020198
APA StyleErvik, Ø., Rødde, M., Hofstad, E. F., Langø, T., Leira, H. O., Amundsen, T., & Sorger, H. (2026). Explainability of a Deep Learning Model for Mediastinal Lymph Node Station Classification in Endobronchial Ultrasound (EBUS). Bioengineering, 13(2), 198. https://doi.org/10.3390/bioengineering13020198






