Diverse Genotypes of Cronobacter spp. Associated with Dairy Farm Systems in Jiangsu and Shandong Provinces in China
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Cronobacter spp. Identification
2.2. Whole-Genome Sequencing and Bioinformatics Analysis
2.3. Antimicrobial Susceptibility Testing
2.4. Biofilm Formation Assays
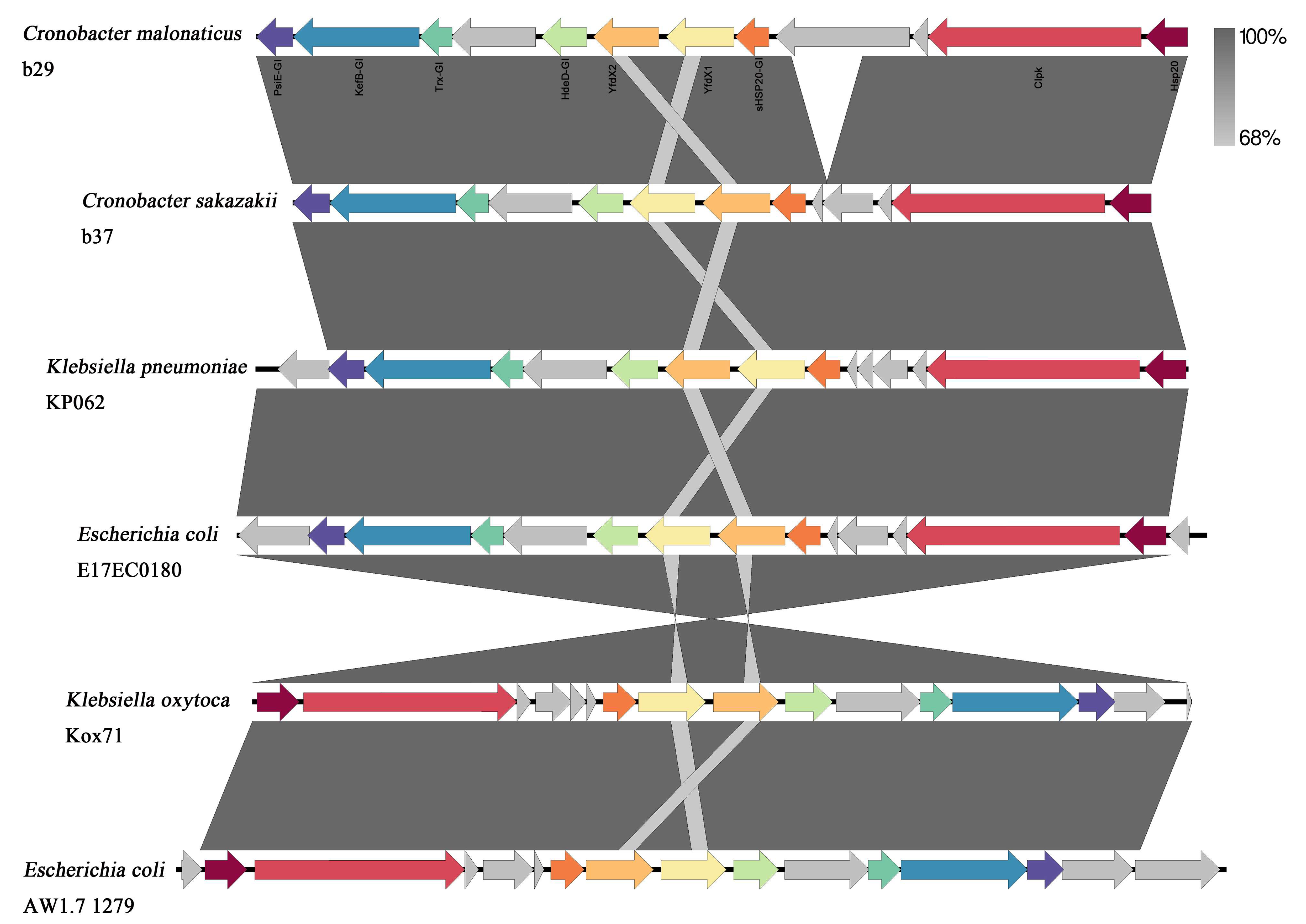
2.5. Comparative Genomic Analysis of Prevalent Sequence Types of Cronobacter spp. Isolates
2.6. Statistical Analyses
3. Results
3.1. Prevalence and Characteristics of Cronobacter spp.
3.2. Multilocus Sequence Typing and O-Serotyping
3.3. Antimicrobial Resistance Phenotypes and Genotypes in Cronobacter spp. Isolates
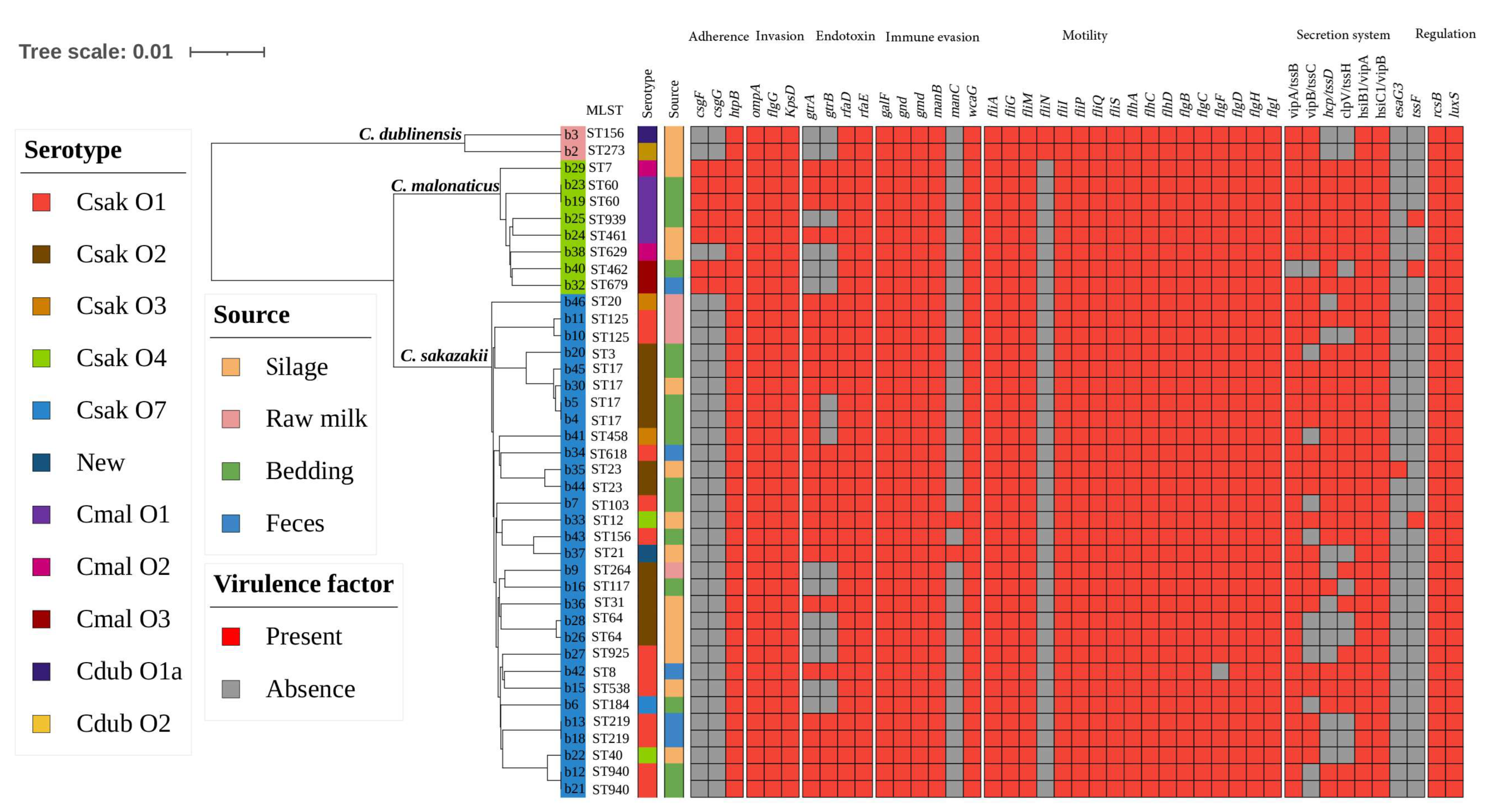
3.4. Prevalence and Distribution of Virulence Genes
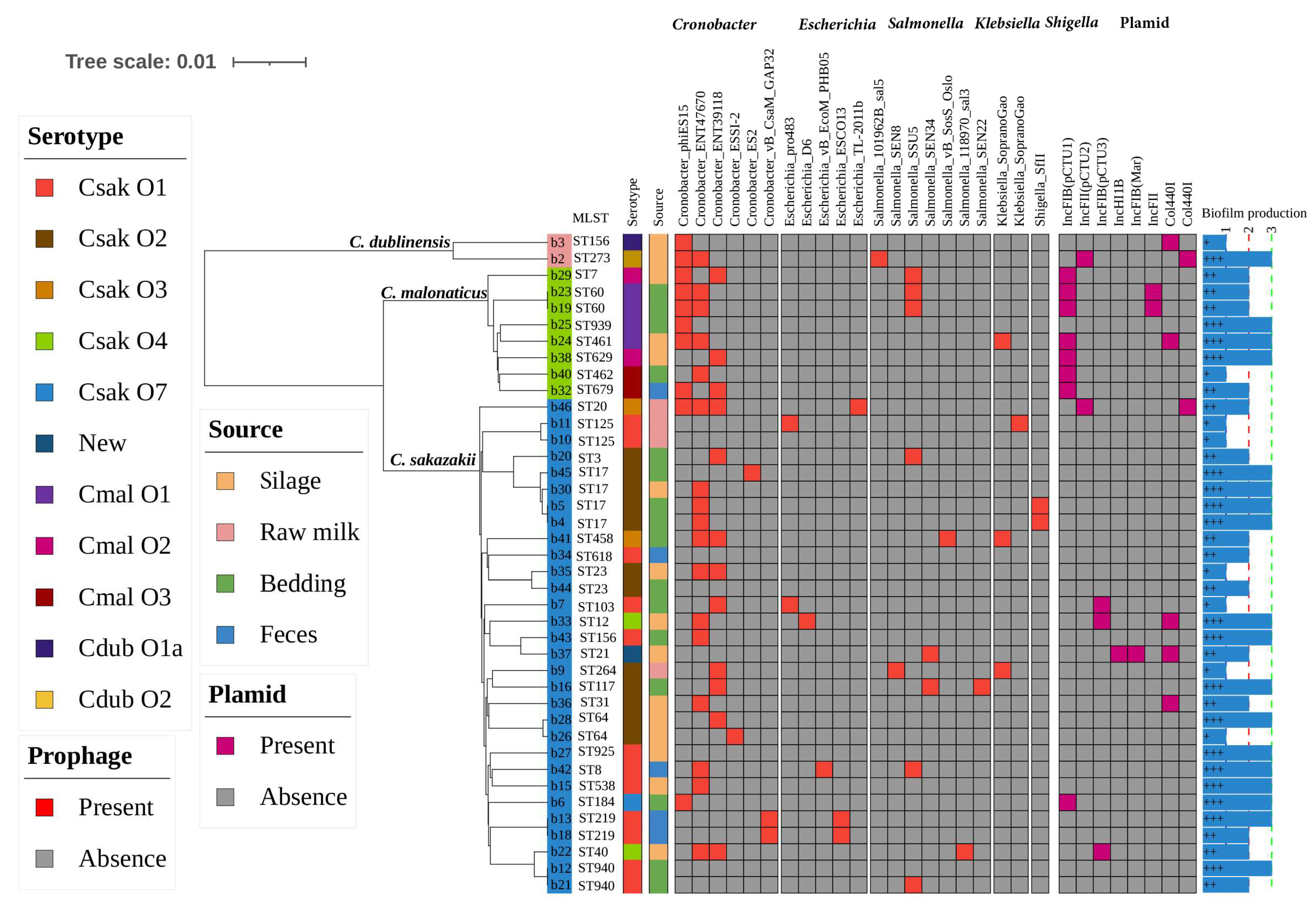
3.5. Biofilm Formation
3.6. Presence of Prophages and Other Mobile Genetic Elements
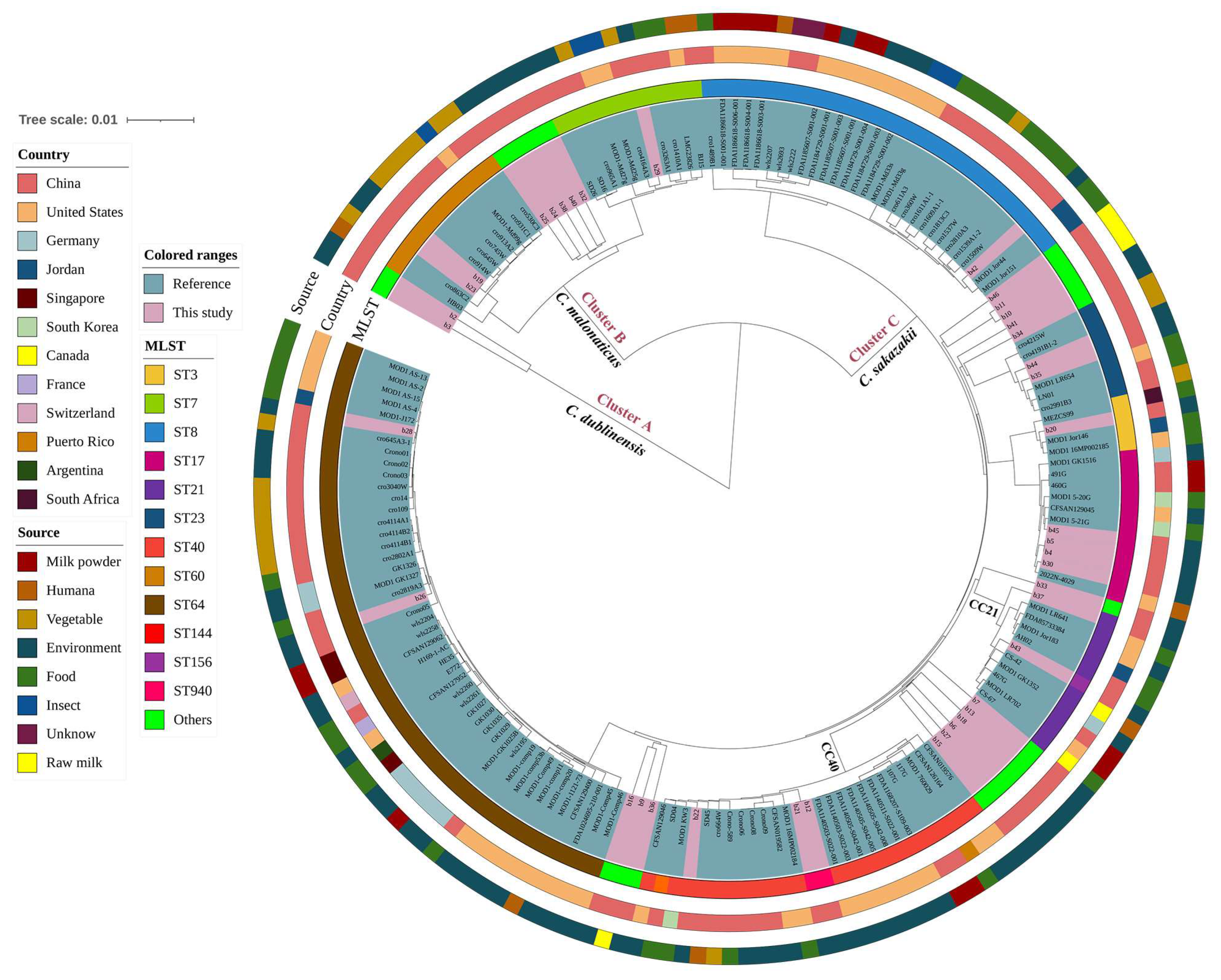
3.7. Minimum Spanning Tree Analysis of Prevalent Sequence Types among Different Sources and between Countries
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Forsythe, S.J. Updates on the Cronobacter Genus. Annu. Rev. Food Sci. Technol. 2018, 9, 23–44. [Google Scholar] [CrossRef]
- Zeng, H.; Li, C.; Ling, N.; Zhang, J.; Chen, M.; Lei, T.; Wu, S.; Yang, X.; Luo, D.; Ding, Y.; et al. Prevalence, genetic analysis and CRISPR typing of Cronobacter spp. isolated from meat and meat products in China. Int. J. Food Microbiol. 2020, 321, 108549. [Google Scholar] [CrossRef] [PubMed]
- Cruz, A.; Xicohtencatl-Cortes, J.; González-Pedrajo, B.; Bobadilla, M.; Eslava, C.; Rosas, I. Virulence traits in Cronobacter species isolated from different sources. Can J. Microbiol. 2011, 57, 735–744. [Google Scholar] [CrossRef]
- Chen, W.; Yang, J.; You, C.; Liu, Z. Diversity of Cronobacter spp. isolates from the vegetables in the middle-east coastline of China. World J. Microbiol. Biotechnol. 2016, 32, 90. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Li, X.; Du, X.; Cui, Z.; Cui, J. Identification and Characterization of Cronobacter Strains Isolated from Environmental Samples. Curr. Microbiol. 2019, 76, 1467–1476. [Google Scholar] [CrossRef]
- Holý, O.; Parra-Flores, J.; Bzdil, J.; Cabal-Rosel, A.; Daza-Prieto, B.; Cruz-Córdova, A.; Xicohtencatl-Cortes, J.; Rodríguez-Martínez, R.; Acuña, S.; Forsythe, S.; et al. Screening of Antibiotic and Virulence Genes from Whole Genome Sequenced Cronobacter sakazakii Isolated from Food and Milk-Producing Environments. Antibiotics 2023, 12, 851. [Google Scholar] [CrossRef]
- Fei, P.; Jiang, Y.; Feng, J.; Forsythe, S.J.; Li, R.; Zhou, Y.; Man, C. Antibiotic and Desiccation Resistance of Cronobacter sakazakii and C. malonaticus Isolates from Powdered Infant Formula and Processing Environments. Front. Microbiol. 2017, 8, 316. [Google Scholar] [CrossRef] [PubMed]
- Fei, P.; Jing, H.; Ma, Y.; Dong, G.; Chang, Y.; Meng, Z.; Jiang, S.; Xie, Q.; Li, S.; Chen, X.; et al. Cronobacter spp. in Commercial Powdered Infant Formula Collected From Nine Provinces in China: Prevalence, Genotype, Biofilm Formation, and Antibiotic Susceptibility. Front. Microbiol. 2022, 13, 900690. [Google Scholar] [CrossRef]
- Ling, N.; Li, C.; Zhang, J.; Wu, Q.; Zeng, H.; He, W.; Ye, Y.; Wang, J.; Ding, Y.; Chen, M.; et al. Prevalence and Molecular and Antimicrobial Characteristics of Cronobacter spp. Isolated From Raw Vegetables in China. Front. Microbiol. 2018, 9, 1149. [Google Scholar] [CrossRef]
- Malorny, B.; Wagner, M. Detection of Enterobacter sakazakii strains by real-time PCR. J. Food Prot. 2005, 68, 1623–1627. [Google Scholar] [CrossRef]
- Zimmermann, J.; Schmidt, H.; Loessner, M.J.; Weiss, A. Development of a rapid detection system for opportunistic pathogenic Cronobacter spp. in powdered milk products. Food Microbiol. 2014, 42, 19–25. [Google Scholar] [CrossRef]
- Gan, X.; Li, M.; Yan, S.; Wang, X.; Wang, W.; Li, F. Genomic Landscape and Phenotypic Assessment of Cronobacter sakazakii Isolated From Raw Material, Environment, and Production Facilities in Powdered Infant Formula Factories in China. Front. Microbiol. 2021, 12, 686189. [Google Scholar] [CrossRef]
- Gajdosova, J.; Benedikovicova, K.; Kamodyova, N.; Tothova, L.; Kaclikova, E.; Stuchlik, S.; Turna, J.; Drahovska, H. Analysis of the DNA region mediating increased thermotolerance at 58 °C in Cronobacter sp. and other enterobacterial strains. Antonie Van Leeuwenhoek 2011, 100, 279–289. [Google Scholar] [CrossRef]
- Kamal, S.M.; Simpson, D.J.; Wang, Z.; Gänzle, M.; Römling, U. Horizontal Transmission of Stress Resistance Genes Shape the Ecology of Beta- and Gamma-Proteobacteria. Front. Microbiol. 2021, 12, 696522. [Google Scholar] [CrossRef]
- Boll, E.J.; Marti, R.; Hasman, H.; Overballe-Petersen, S.; Stegger, M.; Ng, K.; Knøchel, S.; Krogfelt, K.A.; Hummerjohann, J.; Struve, C. Turn Up the Heat-Food and Clinical Escherichia coli Isolates Feature Two Transferrable Loci of Heat Resistance. Front. Microbiol. 2017, 8, 579. [Google Scholar] [CrossRef]
- Grimm, M.; Stephan, R.; Iversen, C.; Manzardo, G.G.; Rattei, T.; Riedel, K.; Ruepp, A.; Frishman, D.; Lehner, A. Cellulose as an extracellular matrix component present in Enterobacter sakazakii biofilms. J. Food Prot. 2008, 71, 13–18. [Google Scholar] [CrossRef]
- Pakbin, B.; Brück, W.M.; Allahyari, S.; Rossen, J.W.A.; Mahmoudi, R. Antibiotic Resistance and Molecular Characterization of Cronobacter sakazakii Strains Isolated from Powdered Infant Formula Milk. Foods 2022, 11, 1093. [Google Scholar] [CrossRef]
- Parra-Flores, J.; Holý, O.; Riffo, F.; Lepuschitz, S.; Maury-Sintjago, E.; Rodríguez-Fernández, A.; Cruz-Córdova, A.; Xicohtencatl-Cortes, J.; Mancilla-Rojano, J.; Troncoso, M.; et al. Profiling the Virulence and Antibiotic Resistance Genes of Cronobacter sakazakii Strains Isolated From Powdered and Dairy Formulas by Whole-Genome Sequencing. Front. Microbiol. 2021, 12, 694922. [Google Scholar] [CrossRef] [PubMed]
- McMullan, R.; Menon, V.; Beukers, A.G.; Jensen, S.O.; van Hal, S.J.; Davis, R. Cronobacter sakazakii Infection from Expressed Breast Milk, Australia. Emerg. Infect. Dis. 2018, 24, 393–394. [Google Scholar] [CrossRef] [PubMed]
- Haston, J.C.; Miko, S.; Cope, J.R.; McKeel, H.; Walters, C.; Joseph, L.A.; Griswold, T.; Katz, L.S.; Andújar, A.A.; Tourdot, L.; et al. Cronobacter sakazakii Infections in Two Infants Linked to Powdered Infant Formula and Breast Pump Equipment—United States, 2021 and 2022. MMWR Morb. Mortal. Wkly. Rep. 2023, 72, 223–226. [Google Scholar] [CrossRef]
- Molloy, C.; Cagney, C.; O’Brien, S.; Iversen, C.; Fanning, S.; Duffy, G. Surveillance and characterisation by pulsed-field gel electrophoresis of Cronobacter spp. in farming and domestic environments, food production animals and retail foods. Int. J. Food. Microbiol. 2009, 136, 198–203. [Google Scholar] [CrossRef]
- Vojkovska, H.; Karpiskova, R.; Orieskova, M.; Drahovska, H. Characterization of Cronobacter spp. isolated from food of plant origin and environmental samples collected from farms and from supermarkets in the Czech Republic. Int. J. Food. Microbiol. 2016, 217, 130–136. [Google Scholar] [CrossRef] [PubMed]
- Fei, P.; Jiang, Y.; Gong, S.; Li, R.; Jiang, Y.; Yuan, X.; Wang, Z.; Kang, H.; Ali, M.A. Occurrence, Genotyping, and Antibiotic Susceptibility of Cronobacter spp. in Drinking Water and Food Samples from Northeast China. J. Food Prot. 2018, 81, 456–460. [Google Scholar] [CrossRef] [PubMed]
- Inouye, M.; Dashnow, H.; Raven, L.A.; Schultz, M.B.; Pope, B.J.; Tomita, T.; Zobel, J.; Holt, K.E. SRST2: Rapid genomic surveillance for public health and hospital microbiology labs. Genome Med. 2014, 6, 90. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Sun, H.X.; Zhang, C.; Cheng, L.; Peng, Y.; Deng, Z.; Wang, D.; Wang, Y.; Hu, M.; Liu, W.; et al. Prophage Hunter: An integrative hunting tool for active prophages. Nucleic Acids Res. 2019, 47, W74–W80. [Google Scholar] [CrossRef]
- Carattoli, A.; Zankari, E.; García-Fernández, A.; Voldby Larsen, M.; Lund, O.; Villa, L.; Møller Aarestrup, F.; Hasman, H. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob Agents Chemother 2014, 58, 3895–3903. [Google Scholar] [CrossRef]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef]
- Pereyra, E.A.; Picech, F.; Renna, M.S.; Baravalle, C.; Andreotti, C.S.; Russi, R.; Calvinho, L.F.; Diez, C.; Dallard, B.E. Detection of Staphylococcus aureus adhesion and biofilm-producing genes and their expression during internalization in bovine mammary epithelial cells. Vet. Microbiol. 2016, 183, 69–77. [Google Scholar] [CrossRef]
- Holý, O.; Forsythe, S. Cronobacter spp. as emerging causes of healthcare-associated infection. J. Hosp. Infect. 2014, 86, 169–177. [Google Scholar] [CrossRef]
- Cechin, C.D.F.; Carvalho, G.G.; Bastos, C.P.; Kabuki, D.Y. Cronobacter spp. in foods of plant origin: Occurrence, contamination routes, and pathogenic potential. Crit. Rev. Food Sci. Nutr. 2022, 63, 12398–12412. [Google Scholar] [CrossRef]
- Lou, X.; Yu, H.; Wang, X.; Qi, J.; Zhang, W.; Wang, H.; Si, G.; Song, S.; Huang, C.; Liu, T.; et al. Potential reservoirs and routes of Cronobacter transmission during cereal growing, processing and consumption. Food Microbiol. 2019, 79, 90–95. [Google Scholar] [CrossRef]
- Silva, J.N.; Vasconcellos, L.; Forsythe, S.J.; de Filippis, I.; Luiz Lima Brandão, M. Molecular and phenotypical characterization of Cronobacter species isolated with high occurrence from oats and linseeds. FEMS Microbiol. Lett. 2019, 366, fny289. [Google Scholar] [CrossRef]
- Lou, X.; Liu, T.; Zhang, W.; Yu, H.; Wang, H.; Song, S.; Chen, Q.; Fang, Z. The occurrence and distribution characteristics of Cronobacter in diverse cereal kernels, flour, and flour-based products. Food Microbiol. 2019, 84, 103269. [Google Scholar] [CrossRef]
- Li, H.; Fu, S.; Song, D.; Qin, X.; Zhang, W.; Man, C.; Yang, X.; Jiang, Y. Identification, Typing and Drug Resistance of Cronobacter spp. in Powdered Infant Formula and Processing Environment. Foods 2023, 12, 1084. [Google Scholar] [CrossRef]
- Fei, P.; Man, C.; Lou, B.; Forsythe, S.J.; Chai, Y.; Li, R.; Niu, J.; Jiang, Y. Genotyping and Source Tracking of Cronobacter sakazakii and C. malonaticus Isolates from Powdered Infant Formula and an Infant Formula Production Factory in China. Appl. Environ. Microbiol. 2015, 81, 5430–5439. [Google Scholar] [CrossRef]
- Cui, J.H.; Yu, B.; Xiang, Y.; Zhang, Z.; Zhang, T.; Zeng, Y.C.; Cui, Z.G.; Huo, X.X. Two Cases of Multi-antibiotic Resistant Cronobacter spp. Infections of Infants in China. Biomed. Environ. Sci. 2017, 30, 601–605. [Google Scholar]
- Alsonosi, A.; Hariri, S.; Kajsík, M.; Oriešková, M.; Hanulík, V.; Röderová, M.; Petrželová, J.; Kollárová, H.; Drahovská, H.; Forsythe, S.; et al. The speciation and genotyping of Cronobacter isolates from hospitalised patients. Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 1979–1988. [Google Scholar] [CrossRef]
- Blažková, M.; Javůrková, B.; Vlach, J.; Göselová, S.; Karamonová, L.; Ogrodzki, P.; Forsythe, S.; Fukal, L. Diversity of O Antigens within the Genus Cronobacter: From Disorder to Order. Appl. Environ. Microbiol. 2015, 81, 5574–5582. [Google Scholar] [CrossRef][Green Version]
- Carvalho, G.G.; Calarga, A.P.; Teodoro, J.R.; Queiroz, M.M.; Astudillo-Trujillo, C.A.; Levy, C.E.; Brocchi, M.; Kabuki, D.Y. Isolation, comparison of identification methods and antibiotic resistance of Cronobacter spp. in infant foods. Food Res. Int. 2020, 137, 109643. [Google Scholar] [CrossRef] [PubMed]
- Müller, A.; Hächler, H.; Stephan, R.; Lehner, A. Presence of AmpC beta-lactamases, CSA-1, CSA-2, CMA-1, and CMA-2 conferring an unusual resistance phenotype in Cronobacter sakazakii and Cronobacter malonaticus. Microb. Drug Resist. 2014, 20, 275–280. [Google Scholar] [CrossRef] [PubMed]
- Gan, X.; Li, M.; Xu, J.; Yan, S.; Wang, W.; Li, F. Emerging of Multidrug-Resistant Cronobacter sakazakii Isolated from Infant Supplementary Food in China. Microbiol. Spectr. 2022, 10, e0119722. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.; Gopinath, G.R.; Eshwar, A.; Srikumar, S.; Nguyen, S.; Gangiredla, J.; Patel, I.R.; Finkelstein, S.B.; Negrete, F.; Woo, J.; et al. The Secretion of Toxins and Other Exoproteins of Cronobacter: Role in Virulence, Adaption, and Persistence. Microorganisms 2020, 8, 229. [Google Scholar] [CrossRef] [PubMed]
- Mohan Nair, M.K.; Venkitanarayanan, K. Role of bacterial OmpA and host cytoskeleton in the invasion of human intestinal epithelial cells by Enterobacter sakazakii. Pediatr. Res. 2007, 62, 664–669. [Google Scholar] [CrossRef] [PubMed]
- Holý, O.; Cruz-Córdova, A.; Xicohtencatl-Cortes, J.; Hochel, I.; Parra-Flores, J.; Petrželová, J.; Fačevicová, K.; Forsythe, S.; Alsonosi, A. Occurrence of virulence factors in Cronobacter sakazakii and Cronobacter malonaticus originated from clinical samples. Microb Pathog. 2019, 127, 250–256. [Google Scholar] [CrossRef] [PubMed]
- Yu, K.W.; Xue, P.; Fu, Y.; Yang, L. T6SS Mediated Stress Responses for Bacterial Environmental Survival and Host Adaptation. Int. J. Mol. Sci. 2021, 22, 478. [Google Scholar] [CrossRef]
- Wang, M.; Cao, H.; Wang, Q.; Xu, T.; Guo, X.; Liu, B. The Roles of Two Type VI Secretion Systems in Cronobacter sakazakii ATCC 12868. Front. Microbiol. 2018, 9, 2499. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.D.; Park, J.H. Complete genome of temperate phage ENT39118 from Cronobacter sakazakii. J. Virol. 2012, 86, 5400–5401. [Google Scholar] [CrossRef]
- Lee, J.H.; Choi, Y.; Shin, H.; Lee, J.; Ryu, S. Complete genome sequence of Cronobacter sakazakii temperate bacteriophage phiES15. J. Virol. 2012, 86, 7713–7714. [Google Scholar] [CrossRef][Green Version]
- Jang, H.; Eshwar, A.; Lehner, A.; Gangiredla, J.; Patel, I.R.; Beaubrun, J.J.; Chase, H.R.; Negrete, F.; Finkelstein, S.; Weinstein, L.M.; et al. Characterization of Cronobacter sakazakii Strains Originating from Plant-Origin Foods Using Comparative Genomic Analyses and Zebrafish Infectivity Studies. Microorganisms 2022, 10, 1396. [Google Scholar] [CrossRef]
- Li, Y.; Yu, H.; Jiang, H.; Jiao, Y.; Zhang, Y.; Shao, J. Genetic Diversity, Antimicrobial Susceptibility, and Biofilm Formation of Cronobacter spp. Recovered from Spices and Cereals. Front. Microbiol. 2017, 8, 2567. [Google Scholar] [CrossRef]
- Wang, Z.; Hu, H.; Zhu, T.; Zheng, J.; Gänzle, M.G.; Simpson, D.J. Ecology and Function of the Transmissible Locus of Stress Tolerance in Escherichia coli and Plant-Associated Enterobacteriaceae. mSystems 2021, 6, e0037821. [Google Scholar] [CrossRef]
- Niu, H.; MingzheYang; Qi, Y.; Liu, Y.; Wang, X.; Dong, Q. Heat shock in Cronobacter sakazakii induces direct protection and cross-protection against simulated gastric fluid stress. Food Microbiol. 2022, 103, 103948. [Google Scholar] [CrossRef] [PubMed]
| Sample | No. of Samples | No. (%) of Positive Samples | Cronobacter spp. Counts | |
|---|---|---|---|---|
| Minimum | Maximum | |||
| raw milk | 710 | 4 (0.56%) | 3.1 × 10 | 3.4 × 102 |
| silage | 100 | 16 (16.0%) | 4.4 × 10 | 1.6 × 103 |
| bedding | 155 | 15 (9.68%) | 4.3 × 10 | 2.2 × 103 |
| feces | 150 | 5 (3.33%) | 2.7 × 10 | 7.4 × 102 |
| Strains | Species | Source | MLST | Serotype | Antibiotics Sensitivity 2 | Antimicrobial Resistance Gene |
|---|---|---|---|---|---|---|
| b2 | C. dublinensis | silage | 273 | Cdub O2 | R(CEP) | ampC, fos |
| b3 | C. dublinensis | silage | 561 | Cdub O1a | R(CEP) | ampC, fos |
| b24 | C. malonaticus | silage | 461 | Cmal O1 | R(CEP)R(FOS) | blaCMA, fos |
| b29 | C. malonaticus | silage | 7 | Cmal O2 | R(CEP) | blaCMA, fos |
| b38 | C. malonaticus | silage | 629 | Cmal O2 | R(CEP)R(FOS) | blaCMA, fos |
| b19 | C. malonaticus | bedding | 60 | Cmal O1 | R(CEP)R(FOS) | blaCMA, fos |
| b23 | C. malonaticus | bedding | 60 | Cmal O1 | R(CEP) | blaCMA, fos |
| b25 | C. malonaticus | bedding | 939 | Cmal O1 | R(CEP)R(FOS) | blaCMA |
| b40 | C. malonaticus | bedding | 462 | Cmal O3 | R(CEP)R(FOS) | blaCMA-2, fosA |
| b32 | C. malonaticus | feces | 679 | Cmal O3 | R(CEP)R(FOS) | blaCMA, fos |
| b9 | C. sakazakii | raw milk | 264 | Csak O2 | R(CEP)R(FOS) | blaCSA, fos |
| b10 | C. sakazakii | raw milk | 125 | Csak O1 | R(CEP)R(FOS) | blaCSA, fos |
| b11 | C. sakazakii | raw milk | 125 | Csak O1 | R(CEP)R(FOS) | blaCSA, fos |
| b46 | C. sakazakii | raw milk | 20 | Csak O3 | R(CEP)R(FOS)I(GEN) | blaCSA, fos |
| b26 | C. sakazakii | silage | 64 | Csak O2 | R(CEP)R(FOS) | blaCSA, fos |
| b37 | C. sakazakii | silage | 21 | new | R(CEP)R(FOS) | blaCSA, fos |
| b27 | C. sakazakii | silage | 925 | Csak O1 | R(CEP)R(FOS) | blaCSA, fos |
| b15 | C. sakazakii | silage | 538 | Csak O1 | R(CEP) | blaCSA, fos |
| b28 | C. sakazakii | silage | 64 | Csak O2 | R(CEP) | blaCSA, fos |
| b22 | C. sakazakii | silage | 40 | Csak O4 | R(CEP) | blaCSA, fos |
| b36 | C. sakazakii | silage | 31 | Csak O2 | R(CEP)R(FOS) | blaCSA, fos |
| b35 | C. sakazakii | silage | 23 | Csak O2 | R(CEP)R(FOS) | blaCSA, fos |
| b30 | C. sakazakii | silage | 17 | Csak O2 | R(CEP)R(FOS) | blaCSA, fos |
| b33 | C. sakazakii | silage | 12 | Csak O4 | R(CEP)R(FOS) | blaCSA, fos |
| b20 | C. sakazakii | bedding | 3 | Csak O2 | R(CEP)R(FOS) | blaCSA, fos |
| b4 | C. sakazakii | bedding | 17 | Csak O2 | R(CEP) | blaCSA, fos |
| b5 | C. sakazakii | bedding | 17 | Csak O2 | R(CEP) | blaCSA, fos |
| b21 | C. sakazakii | bedding | 940 | Csak O1 | R(CEP) | blaCSA, fos |
| b12 | C. sakazakii | bedding | 940 | Csak O1 | R(CEP)R(FOS) | blaCSA, fos |
| b41 | C. sakazakii | bedding | 458 | Csak O3 | R(CEP)R(FOS) | blaCSA, fos |
| b6 | C. sakazakii | bedding | 184 | Csak O7 | R(CEP) | blaCSA, fos |
| b43 | C. sakazakii | bedding | 156 | Csak O1 | R(CEP)R(FOS)I(GEN) | blaCSA, fos |
| b16 | C. sakazakii | bedding | 117 | Csak O2 | R(CEP) | blaCSA, fos |
| b7 | C. sakazakii | bedding | 103 | Csak O1 | R(CEP)R(FOS) | blaCSA, fos |
| b44 | C. sakazakii | bedding | 23 | Csak O2 | R(CEP)R(FOS)I(GEN) | blaCSA, fos |
| b45 | C. sakazakii | bedding | 17 | Csak O2 | R(CEP)R(FOS)I(GEN) | blaCSA, fos |
| b34 | C. sakazakii | feces | 618 | Csak O1 | R(CEP)R(FOS) | blaCSA, fos |
| b18 | C. sakazakii | feces | 219 | Csak O1 | R(CEP)R(FOS) | blaCSA, fos |
| b13 | C. sakazakii | feces | 219 | Csak O1 | R(CEP)R(FOS) | blaCSA, fos |
| b42 | C. sakazakii | feces | 8 | Csak O1 | R(CEP)R(FOS)I(GEN) | blaCSA-1, fos |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, H.; Ji, X.; Sun, H.; Billington, C.; Hou, X.; Soleimani-Delfan, A.; Wang, R.; Wang, H.; Zhang, L. Diverse Genotypes of Cronobacter spp. Associated with Dairy Farm Systems in Jiangsu and Shandong Provinces in China. Foods 2024, 13, 871. https://doi.org/10.3390/foods13060871
Liu H, Ji X, Sun H, Billington C, Hou X, Soleimani-Delfan A, Wang R, Wang H, Zhang L. Diverse Genotypes of Cronobacter spp. Associated with Dairy Farm Systems in Jiangsu and Shandong Provinces in China. Foods. 2024; 13(6):871. https://doi.org/10.3390/foods13060871
Chicago/Turabian StyleLiu, Hui, Xing Ji, Haichang Sun, Craig Billington, Xiang Hou, Abbas Soleimani-Delfan, Ran Wang, Heye Wang, and Lili Zhang. 2024. "Diverse Genotypes of Cronobacter spp. Associated with Dairy Farm Systems in Jiangsu and Shandong Provinces in China" Foods 13, no. 6: 871. https://doi.org/10.3390/foods13060871
APA StyleLiu, H., Ji, X., Sun, H., Billington, C., Hou, X., Soleimani-Delfan, A., Wang, R., Wang, H., & Zhang, L. (2024). Diverse Genotypes of Cronobacter spp. Associated with Dairy Farm Systems in Jiangsu and Shandong Provinces in China. Foods, 13(6), 871. https://doi.org/10.3390/foods13060871





