ChIP-Seq Analysis of SlAREB1 Downstream Regulatory Network during Tomato Ripening
Abstract
:1. Introduction
2. Materials and Methods
2.1. Tomato Fruit and Exgenous ABA Treatment
2.2. qRT-PCR Analysis
2.3. Construction and Identification of SlAREB1-Overexpression Transgenic Tomato Plants
2.4. ChIP-Seq Analysis
2.5. Statistical Analysis
3. Results
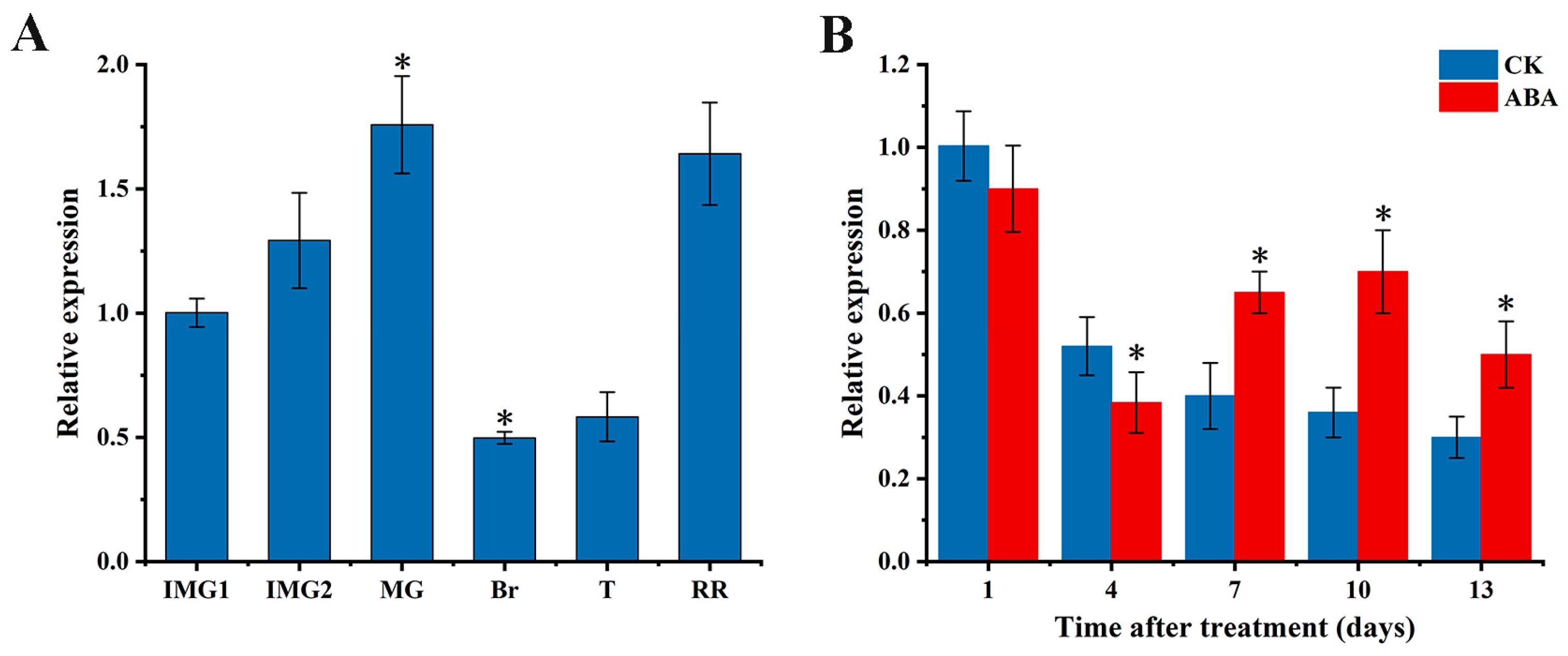
3.1. Gene Expression of SlAREB1 during Tomato Fruit Ripening and the Effect of Exogenous ABA Treatment on It
3.2. Identification of SlAREB1-Overexpression Transgenic Tomato Plants
3.3. ChIP-Seq Analysis
3.3.1. ChIP-Seq Peak Analysis
3.3.2. Transcription Factor Prediction of Peak-Associated Genes
3.3.3. Gene ontology (GO) and Kyoto Encylopaedia of Genes and Genomes (KEGG) Analysis of SlAREB1 Target Sequences
3.3.4. Downstream Candidate Genes of SlAREB1
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kou, X.; Zhou, J.; Wu, C.E.; Yang, S.; Liu, Y.; Chai, L.; Xue, Z. The interplay between ABA/ethylene and NAC TFs in tomato fruit ripening: A review. Plant Mol. Biol. 2021, 106, 223–238. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, W.; Cai, J.; Zhang, Y.; Qin, G.; Tian, S. Tomato nuclear proteome reveals the involvement of specific E2 ubiquitin-conjugating enzymes in fruit ripening. Genome Biol. 2014, 15, 548. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Grimplet, J.; David, K.; Castellarin, S.D.; Terol, J.; Wong, D.C.J.; Luo, Z.; Schaffer, R.; Celton, J.-M.; Talon, M.; et al. Ethylene receptors and related proteins in climacteric and non-climacteric fruits. Plant Sci. 2018, 276, 63–72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Tian, S.; Yu, Y.; Ren, Y.; Guo, S.; Zhang, J.; Li, M.; Zhang, H.; Gong, G.; Wang, M.; et al. Natural variation in the NAC transcription factor NONRIPENING contributes to melon fruit ripening. J. Integr. Plant. Biol. 2022, 64, 1448–1461. [Google Scholar] [CrossRef]
- Li, J.; Min, D.; Li, Z.; Fu, X.; Zhao, X.; Wang, J.; Zhang, X.; Li, F.; Li, X.; Guo, Y. Regulation of Sugar Metabolism by Methyl Jasmonate to Improve the Postharvest Quality of Tomato Fruit. J. Plant Growth Regul. 2022, 41, 1615–1626. [Google Scholar] [CrossRef]
- Tao, X.; Wu, Q.; Fu, X.; Zhu, B.; Chen, F.; Liu, B.; Mao, L.; Luo, Z.; Li, L.; Ying, T. Understanding of exogenous auxin in regulating sucrose metabolism during postharvest tomato fruit ripening. Postharvest Biol. Technol. 2022, 189, 111913. [Google Scholar] [CrossRef]
- Vitale, A.; Rocco, M.; Arena, S.; Giuffrida, F.; Cassaniti, C.; Scaloni, A.; Lomaglio, T.; Guarnaccia, V.; Polizzi, G.; Marra, M.; et al. Tomato susceptibility to Fusarium crown and root rot: Effect of grafting combination and proteomic analysis of tolerance expression in the rootstock. Plant Physiol. Biochem. 2014, 83, 207–216. [Google Scholar] [CrossRef] [Green Version]
- Wang, D.; Yeats, T.H.; Uluisik, S.; Rose, J.K.C.; Seymour, G.B. Fruit Softening: Revisiting the Role of Pectin. Trends Plant Sci. 2018, 23, 302–310. [Google Scholar] [CrossRef]
- Galpaz, N.; Wang, Q.; Menda, N.; Zamir, D.; Hirschberg, J. Abscisic acid deficiency in the tomato mutant high-pigment 3 leading to increased plastid number and higher fruit lycopene content. Plant J. 2008, 53, 717–730. [Google Scholar] [CrossRef]
- Zhang, M.; Yuan, B.; Leng, P. The role of ABA in triggering ethylene biosynthesis and ripening of tomato fruit. J. Exp. Bot. 2009, 60, 1579–1588. [Google Scholar] [CrossRef] [Green Version]
- Tao, X.; Wu, Q.; Huang, S.; Zhu, B.; Chen, F.; Liu, B.; Cai, L.; Mao, L.; Luo, Z.; Li, L.; et al. Exogenous abscisic acid regulates primary metabolism in postharvest cherry tomato fruit during ripening. Sci. Hortic. 2022, 299, 111008. [Google Scholar] [CrossRef]
- Wu, Q.; Tao, X.; Ai, X.; Luo, Z.; Mao, L.; Ying, T.; Li, L. Contribution of abscisic acid to aromatic volatiles in cherry tomato (Solanum lycopersicum L.) fruit during postharvest ripening. Plant Physiol. Biochem. 2018, 130, 205–214. [Google Scholar] [CrossRef]
- Tao, X.; Wu, Q.; Aalim, H.; Li, L.; Mao, L.; Luo, Z.; Ying, T. Effects of Exogenous Abscisic Acid on Bioactive Components and Antioxidant Capacity of Postharvest Tomato during Ripening. Molecules 2020, 25, 1346. [Google Scholar] [CrossRef] [Green Version]
- Wang, W.; Wang, X.; Wang, Y.; Zhou, G.; Wang, C.; Hussain, S.; Adnan; Lin, R.; Wang, T.; Wang, S. SlEAD1, an EAR motif-containing ABA down-regulated novel transcription repressor regulates ABA response in tomato. GM Crops Food 2020, 11, 275–289. [Google Scholar] [CrossRef] [PubMed]
- Leng, P.; Yuan, B.; Guo, Y. The role of abscisic acid in fruit ripening and responses to abiotic stress. J. Exp. Bot. 2014, 65, 4577–4588. [Google Scholar] [CrossRef] [PubMed]
- Droege-Laser, W.; Snoek, B.L.; Snel, B.; Weiste, C. The Arabidopsis bZIP transcription factor family—An update. Curr. Opin. Plant Biol. 2018, 45, 36–49. [Google Scholar] [CrossRef] [PubMed]
- Bastias, A.; Lopez-Climent, M.; Valcarcel, M.; Rosello, S.; Gomez-Cadenas, A.; Casaretto, J.A. Modulation of organic acids and sugar content in tomato fruits by an abscisic acid-regulated transcription factor. Physiol. Plant. 2011, 141, 215–226. [Google Scholar] [CrossRef] [PubMed]
- Orellana, S.; Yanez, M.; Espinoza, A.; Verdugo, I.; Gonzalez, E.; Ruiz-Lara, S.; Casaretto, J.A. The transcription factor SlAREB1 confers drought, salt stress tolerance and regulates biotic and abiotic stress-related genes in tomato. Plant Cell. Environ. 2010, 33, 2191–2208. [Google Scholar] [CrossRef]
- Yanez, M.; Caceres, S.; Orellana, S.; Bastias, A.; Verdugo, I.; Ruiz-Lara, S.; Casaretto, J.A. An abiotic stress-responsive bZIP transcription factor from wild and cultivated tomatoes regulates stress-related genes. Plant Cell. Rep. 2009, 28, 1497–1507. [Google Scholar] [CrossRef]
- Xu, Z.; Wang, F.; Ma, Y.; Dang, H.; Hu, X. Transcription Factor SlAREB1 Is Involved in the Antioxidant Regulation under Saline-Alkaline Stress in Tomato. Antioxidants 2022, 11, 1673. [Google Scholar] [CrossRef]
- Bastias, A.; Yanez, M.; Osorio, S.; Arbona, V.; Gomez-Cadenas, A.; Fernie, A.R.; Casaretto, J.A. The transcription factor AREB1 regulates primary metabolic pathways in tomato fruits. J. Exp. Bot. 2014, 65, 2351–2363. [Google Scholar] [CrossRef] [Green Version]
- Yang, S.; Zhou, J.; Watkins, C.B.; Wu, C.; Feng, Y.; Zhao, X.; Xue, Z.; Kou, X. NAC transcription factors SNAC4 and SNAC9 synergistically regulate tomato fruit ripening by affecting expression of genes involved in ethylene and abscisic acid metabolism and signal transduction. Postharvest Biol. Technol. 2021, 178, 111555. [Google Scholar] [CrossRef]
- Mou, W.; Li, D.; Luo, Z.; Li, L.; Mao, L.; Ying, T. SlAREB1 transcriptional activation of NOR is involved in abscisic acid-modulated ethylene biosynthesis during tomato fruit ripening. Plant Sci. 2018, 276, 239–249. [Google Scholar] [CrossRef] [PubMed]
- Small, E.C.; Maryanski, D.N.; Rodriguez, K.L.; Harvey, K.J.; Keogh, M.-C.; Johnstone, A.L. Chromatin Immunoprecipitation (ChIP) to Study DNA-Protein Interactions. Methods Mol. Biol. 2021, 2261, 323–343. [Google Scholar]
- GAO, S.; ZHANG, N.; LI, B. Processing and analysis of ChIP-seq data. Hereditas 2012, 34, 773–783. [Google Scholar] [CrossRef]
- Liu, X.; Su, L.; Li, L. Downstream target gene network regulated by AhGLK1 and AhHDA1 using ChIP-seq in peanut. Zuo Wu Xue Bao 2022, 48, 2765–2773. [Google Scholar]
- Wang, T.; Shen, L.; Wei, M. ChIP-seq Analysis of the Impact of Mutant IHH-GLI1 Signaling Pathway on Downstream Transcriptional Regulation. J. Nat. Sci. Hunan Norm. Univ. 2020, 43, 27–35. [Google Scholar]
- Zhu, L.; Xue, P.; Li, G. Genome-wide Binding Site Analysis of LAZY1 Reveals Its Downstream Regulating Network in Controlling Rice Tiller Angle. Genom. Appl. Biol. 2022, 41, 1539–1549. [Google Scholar]
- Zhao, T.; Wu, T.; Pei, T.; Wang, Z.; Yang, H.; Jiang, J.; Zhang, H.; Chen, X.; Li, J.; Xu, X. Overexpression of SlGATA17 Promotes Drought Tolerance in Transgenic Tomato Plants by Enhancing Activation of the Phenylpropanoid Biosynthetic Pathway. Front. Plant Sci. 2021, 12, 634888. [Google Scholar] [CrossRef]
- Yang, B.; Liu, C.; Pan, X.; Fu, W.; Fan, Z.; Jin, Y.; Bai, F.; Cheng, Z.; Wu, W. Identification of Novel phoP-phoQ Regulated Genes that Contribute to Polymyxin B Tolerance in Pseudomonas aeruginosa. Microorganisms 2021, 9, 344. [Google Scholar] [CrossRef]
- Kuang, J.-F.; Wu, C.-J.; Guo, Y.-F.; Walther, D.; Shan, W.; Chen, J.-Y.; Chen, L.; Lu, W.-J. Deciphering transcriptional regulators of banana fruit ripening by regulatory network analysis. Plant Biotechnol. J. 2021, 19, 477–489. [Google Scholar] [CrossRef]
- Joshi, R.; Wani, S.H.; Singh, B.; Bohra, A.; Dar, Z.A.; Lone, A.A.; Pareek, A.; Singla-Pareek, S.L. Transcription Factors and Plants Response to Drought Stress: Current Un-derstanding and Future Directions. Front. Plant Sci. 2016, 7, 1029. [Google Scholar]
- Hu, J.; Gao, Y.; Zhao, T.; Li, J.; Yao, M.; Xu, X. Genome-wide Identification and Expression Pattern Analysis of Zinc-finger Homeodomain Transcription Factors in Tomato under Abiotic Stress. J. Am. Soc. Hortic. Sci. 2018, 143, 14–22. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Deng, J.; Chen, J.; Zeng, T.; Yu, T.; Huang, Q.; Chen, P.; Liu, Q.; Jian, W.; Yang, X. Genome-wide identification and expression analysis of FAR1/FHY3 transcription factor family in tomato. Plant Physiol. 2021, 57, 1983–1995. [Google Scholar]
- Wang, Y.; Zhang, J.; Hu, Z.; Guo, X.; Tian, S.; Chen, G. Genome-Wide Analysis of the MADS-Box Transcription Factor Family in Solanum lycopersicum. Int. J. Mol. Sci. 2019, 20, 2961. [Google Scholar] [CrossRef] [Green Version]
- Mou, W.; Li, D.; Luo, Z.; Mao, L.; Ying, T. Transcriptomic Analysis Reveals Possible Influences of ABA on Secondary Metabolism of Pigments, Flavonoids and Antioxidants in Tomato Fruit during Ripening. PLoS ONE 2015, 10, e0129598. [Google Scholar] [CrossRef] [PubMed]
- Kadenbach, B. Intrinsic and extrinsic uncoupling of oxidative phosphorylation. Biochim. Biophys. Acta Biomembr. 2003, 1604, 77–94. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Piechowiak, T.; Sowa, P.; Tarapatskyy, M.; Balawejder, M. The Role of Mitochondrial Energy Metabolism in Shaping the Quality of Highbush Blueberry Fruit During Storage in Ozone-Enriched Atmosphere. Food Bioprocess Technol. 2021, 14, 1973–1982. [Google Scholar] [CrossRef]
- Wang, Q.-H.; Zhao, C.; Zhang, M.; Li, Y.-Z.; Shen, Y.-Y.; Guo, J.-X. Transcriptome analysis around the onset of strawberry fruit ripening uncovers an important role of oxidative phosphorylation in ripening. Sci. Rep. 2017, 7, 41477. [Google Scholar] [CrossRef] [Green Version]
- Liu, L.M.; Li, Y.; Du, G.C.; Chen, J. Increasing glycolytic flux in Torulopsis glabrata by redirecting ATP production from oxidative phosphorylation to substrate-level phosphorylation. J. Appl. Microbiol. 2006, 100, 1043–1053. [Google Scholar] [CrossRef]
- Nolfi-Donegan, D.; Braganza, A.; Shiva, S. Mitochondrial electron transport chain: Oxidative phosphorylation, oxidant production, and methods of measurement. Redox. Biol. 2020, 37, 101674. [Google Scholar] [CrossRef]
- Tian, S.; Qin, G.; Li, B. Reactive oxygen species involved in regulating fruit senescence and fungal pathogenicity. Plant Mol. Biol. 2013, 82, 593–602. [Google Scholar] [CrossRef]
- Wang, Y.; Ji, D.; Chen, T.; Li, B.; Zhang, Z.; Qin, G.; Tian, S. Production, Signaling, and Scavenging Mechanisms of Reactive Oxygen Species in Fruit-Pathogen Interactions. Int. J. Mol. Sci. 2019, 20, 2994. [Google Scholar] [CrossRef] [Green Version]
- Merchante, C.; Alonso, J.M.; Stepanova, A.N. Ethylene signaling: Simple ligand, complex regulation. Curr. Opin. Plant Biol. 2013, 16, 554–560. [Google Scholar] [CrossRef]
- Zhao, W.; Ma, Z.; Liu, S.; Yang, W.; Ma, J. Transcriptome Profiling Reveals Potential Genes and Pathways Supporting Ananas comosus L. Merr’s High. Temperature Stress Tolerance. Trop. Plant Biol. 2021, 14, 132–142. [Google Scholar] [CrossRef]
- Zhang, S.; Wang, S.; Han, S.; Wang, D. The Research Progress of Glutaredoxin in Plants. Yuan Yi Xue Bao 2022, 34, 1135–1144. [Google Scholar]
- Bai, Y.; Zhou, J.-J.; Tang, Y.-L. Plant laccases and their roles in plant growth and development. Chin. Bull. Life Sci. 2022, 34, 1135–1144. [Google Scholar]
- Wu, S.; Wu, D.; Song, J.; Zhang, Y.; Tan, Q.; Yang, T.; Yang, J.; Wang, S.; Xu, J.; Xu, W.; et al. Metabolomic and transcriptomic analyses reveal new insights into the role of abscisic acid in modulating mango fruit ripening. Hort. Res. 2022, 9, 1136. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.-I.; Hwang, I.; Goswami, G.; Jung, H.-J.; Nath, U.K.; Yoo, H.-J.; Lee, J.M.; Nou, I.S. Molecular Insights Reveal Psy1, SGR, and SlMYB12 Genes are Associated with Diverse Fruit Color Pigments in Tomato (Solanum lycopersicum L.). Molecules 2017, 22, 2180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ranganath, K.G. Pigments That Colour Our Fruits: An Overview. Erwerbs-Obstbau 2022, 64, 535–547. [Google Scholar] [CrossRef]
- Wu, Q.; Bai, J.; Tao, X.; Mou, W.; Luo, Z.; Mao, L.; Ban, Z.; Ying, T.; Li, L. Synergistic effect of abscisic acid and ethylene on color development in tomato (Solarium lycopersicum L.) fruit. Sci. Hortic. 2018, 235, 169–180. [Google Scholar] [CrossRef]
- Batista-Silva, W.; Carvalho de Oliveira, A.; Oliveira Martins, A.; Siqueira, J.A.; Rodrigues-Salvador, A.; Omena-Garcia, R.P.; Medeiros, D.B.; Peres, L.E.P.; Ribeiro, D.M.; Zsogon, A.; et al. Reduced auxin signalling through the cyclophilin gene DIAGEOTROPICA impacts tomato fruit development and metabolism during ripening. J. Exp. Bot. 2022, 73, 4113–4128. [Google Scholar] [CrossRef]
- Mou, W.; Li, D.; Bu, J.; Jiang, Y.; Khan, Z.U.; Luo, Z.; Mao, L.; Ying, T. Comprehensive Analysis of ABA Effects on Ethylene Biosynthesis and Signaling during Tomato Fruit Ripening. PLoS ONE 2016, 11, e0154072. [Google Scholar]
- Isaacson, T.; Ronen, G.; Zamir, D.; Hirschberg, J. Cloning of tangerine from tomato reveals a carotenoid isomerase essential for the production of beta-carotene and xanthophylls in plants. Plant Cell 2002, 14, 333–342. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kou, X.; Zhao, Y.; Wu, C.; Jiang, B.; Zhang, Z.; Rathbun, J.R.; He, Y.; Xue, Z. SNAC4 and SNAC9 transcription factors show contrasting effects on tomato carotenoids biosynthesis and softening. Postharvest Biol. Technol. 2018, 144, 9–19. [Google Scholar] [CrossRef]
- Chen, H.; Bai, S.; Kusano, M.; Ezura, H.; Wang, N. Increased ACS Enzyme Dosage Causes Initiation of Climacteric Ethylene Production in Tomato. Int. J. Mol. Sci. 2022, 23, 10788. [Google Scholar] [CrossRef]
- Yu, W.; Sheng, J.; Zhao, R.; Wang, Q.; Ma, P.; Shen, L. Ethylene biosynthesis is involved in regulating chilling tolerance and SlCBF1 gene expression in tomato fruit. Postharvest Biol. Technol. 2019, 149, 139–147. [Google Scholar] [CrossRef]
- Sell, S.; Hehl, R. A fifth member of the tomato 1-aminocyclopropane-1-carboxylic acid (ACC) oxidase gene family harbours a leucine zipper and is anaerobically induced. DNA Seq. 2005, 16, 80–82. [Google Scholar] [CrossRef] [PubMed]
- Gu, C.; Guo, Z.H.; Hao, P.P.; Wang, G.M.; Jin, Z.M.; Zhang, S.L. Multiple regulatory roles of AP2/ERF transcription factor in angiosperm. Bot. Stud. 2017, 58, 10788. [Google Scholar] [CrossRef] [Green Version]
- Mignolli, F.; Mariotti, L.; Laura Vidoz, M. Reduced gibberellin biosynthesis and response in fruits of the auxin insensitive diageotropica tomato mutant. Plant Growth Regul. 2022, 99, 505–513. [Google Scholar] [CrossRef]
- Wu, J.-M.; Chen, R.-F.; Huang, X.; Qiu, L.-H.; Li, Y.-R. Studies on the Gene of Key Component GA20-oxidase for Gibberellin Biosynthesis in Plant. Biol. Bull. 2016, 32, 1–12. [Google Scholar]
- Chen, S.; Wang, X.-J.; Tan, G.-F.; Zhou, W.-Q.; Wang, G.-L. Gibberellin and the plant growth retardant Paclobutrazol altered fruit shape and ripening in tomato. Protoplasma 2020, 257, 853–861. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Han, S.; Qi, Y. Advances in structure and function of auxin response factor in plants. J. Integr. Plant Biol. 2023, 65, 617–632. [Google Scholar] [CrossRef]
- Zhu, Z.; Liang, H.; Chen, G.; Tang, B.; Tian, S.; Hu, Z. Isolation of the brassinosteroid receptor genes and recharacterization of dwarf plants by silencing of SlBRI1 in tomato. Plant Growth Regul. 2019, 89, 59–71. [Google Scholar] [CrossRef]
- Zhu, T.; Tan, W.-R.; Deng, X.-G.; Zheng, T.; Zhang, D.-W.; Lin, H.-H. Effects of brassinosteroids on quality attributes and ethylene synthesis in postharvest tomato fruit. Postharvest Biol. Technol. 2015, 100, 196–204. [Google Scholar] [CrossRef]
- Zhao, B.; Li, J. Regulation of Brassinosteroid Biosynthesis and Inactivation. J. Integr. Plant Biol. 2012, 54, 746–759. [Google Scholar] [CrossRef]
- Zeng, W.-J.; Yang, J.; Chen, S.-M.-J.; Wei, X.-Y.; Wen, G.-Q.; He, Z.-Y.; Wang, Q.; Chen, X.-T.; Zhou, J.-Y.; Zou, J. Expression of Cell Wall Metabolism Genes During Tomato (Solanum lycopersicum) Maturation and the Effects of Treatment with Exogenous C2H4 and ABA on Their Expression. J. Biotechnol. 2021, 29, 1040–1049. [Google Scholar]
- He, Y.; Li, J.; Ban, Q.; Han, S.; Rao, J. Role of Brassinosteroids in Persimmon (Diospyros kaki L.) Fruit Ripening. J. Agric. Food Chem. 2018, 66, 2637–2644. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Lopez, A.; Jeon, S.; de Freitas, S.T.; Yu, Q.; Wu, Z.; Labavitch, J.M.; Tian, S.; Powell, A.L.T.; Mitcham, E. Disassembly of the fruit cell wall by the ripening-associated polygalacturonase and expansin influences tomato cracking. Hort. Res. 2019, 6, 17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, Y.; Lu, L.; Sun, D.; Wang, J.; Wang, N.; Qiao, L.; Guo, Q.; Wang, C. Fungus Polygalacturonase-Generated Oligogalacturonide Restrains Fruit Softening in Ripening Tomato. J. Agric. Food Chem. 2022, 70, 759–769. [Google Scholar] [CrossRef]
- Perini, M.A.; Sin, I.N.; Villarreal, N.M.; Marina, M.; Powell, A.L.T.; Martinez, G.A.; Civello, P.M. Overexpression of the carbohydrate binding module from Solanum lycopersicum expansin 1 (Sl-EXP1) modifies tomato fruit firmness and Botrytis cinerea susceptibility. Plant Physiol. Biochem. 2017, 113, 122–132. [Google Scholar] [CrossRef] [PubMed]
- Micheli, F. Pectin methylesterases: Cell wall enzymes with important roles in plant physiology. Trends Plant Sci. 2001, 6, 414–419. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Liu, Q.; Qiao, Q.; Zhu, Y.; Huang, W.; Wang, X.; Ren, Z. Exploring the effects of pectate and pectate lyase on the fruit softening and transcription profiling of Solanum lycopersicum. Food Control 2022, 133, 108636. [Google Scholar] [CrossRef]
- Ao, Y.; Yang, M.-Y.; Zhang, C.; Wu, Q. Research Progress on the Mechanisms of Cell Wall Actions during Ripening and Softening Processes of Tomato Fruits. Storage Process 2021, 21, 118–125. [Google Scholar]





| Sample | Raw Reads | Clean Reads | Clean Ratio | Mapped Reads | Map Rate |
|---|---|---|---|---|---|
| SlAREB1-IP | 52,720,466 | 51,618,544 | 97.91% | 38,834,545 | 75.23% |
| Input | 36,263,534 | 35,588,840 | 98.14% | 34,927,951 | 98.14% |
| Gene ID | Subject Length | Subject Start | Subject End | Subject Annotation |
|---|---|---|---|---|
| Oxidative phosphorylation | ||||
| Solyc00g013180.1 | 3901 | 9,741,922 | 9,745,822 | NADH-ubiquinone oxidoreductase chain 4 |
| Solyc00g014830.3 | 2241 | 10,120,367 | 10,122,607 | NADH dehydrogenase subunit 7 |
| Solyc00g019730.2 | 1112 | 10,827,171 | 10,828,282 | Cytochrome c oxidase subunit 3 |
| Solyc00g019950.1 | 1415 | 10,844,935 | 10,846,349 | NADH dehydrogenase subunit 9 |
| Solyc00g117655.1 | 195 | 15,444,467 | 15,444,661 | NADH-ubiquinone oxidoreductase chain 1 |
| Solyc01g020470.2 | 199 | 30,837,743 | 30,837,941 | NADH dehydrogenase subunit 9 |
| Solyc01g056670.1 | 493 | 55,571,581 | 55,572,073 | NADH dehydrogenase subunit 4L |
| Solyc03g013460.1 | 247 | 45,900,762 | 45,901,008 | Cytochrome c oxidase subunit 3 |
| Solyc03g043610.2 | 146 | 7,121,382 | 7,121,527 | ATP synthase subunit a |
| Solyc05g016220.1 | 138 | 15,092,149 | 15,092,286 | Ycf1 |
| Solyc05g023920.1 | 318 | 30,101,773 | 30,102,090 | NADH-ubiquinone oxidoreductase chain 1 |
| Solyc07g019510.3 | 338 | 11,845,804 | 11,846,141 | Cytochrome c oxidase subunit 1 |
| Solyc08g029260.1 | 1096 | 37,265,408 | 37,266,503 | NADH dehydrogenase subunit 2 |
| Solyc10g045750.1 | 212 | 35,863,088 | 35,863,299 | NADH-ubiquinone oxidoreductase chain 4 |
| Solyc10g049470.1 | 157 | 45,831,441 | 45,831,597 | Ycf1 |
| Solyc11g021240.2 | 149 | 13,415,639 | 13,415,787 | Ycf1 |
| Solyc11g021300.1 | 154 | 13,419,577 | 13,419,730 | Ycf1 |
| Solyc11g030570.1 | 339 | 22,056,502 | 22,056,840 | NADH-ubiquinone oxidoreductase chain 4 |
| Solyc12g035550.1 | 330 | 41,911,350 | 41,911,679 | Ycf1 |
| Solyc12g035930.1 | 153 | 44,564,598 | 44,564,750 | DNA-directed RNA polymerase subunit beta |
| Photosynthesis | ||||
| Solyc00g230070.1 | 2413 | 18,659,728 | 18,662,140 | Photosystem II CP43 chlorophyll apoprotein |
| Solyc01g017090.3 | 247 | 23,795,390 | 23,795,636 | NADH-quinone oxidoreductase subunit L |
| Solyc01g017440.1 | 143 | 23,869,851 | 23,869,993 | DNA-directed RNA polymerase subunit alpha |
| Solyc01g017740.1 | 146 | 25,041,926 | 25,042,071 | Cytochrome b6 |
| Solyc01g056870.2 | 356 | 57,175,864 | 57,176,219 | Ycf2 |
| Solyc02g011755.1 | 159 | 14,135,562 | 14,135,720 | Photosystem I iron-sulfur center |
| Solyc02g080635.1 | 291 | 45,373,848 | 45,374,138 | Photosystem II CP43 reaction center protein |
| Solyc03g122000.3 | 437 | 71,495,017 | 71,495,453 | Cytochrome b6-f complex subunit 4 |
| Solyc04g049003.1 | 174 | 38,943,033 | 38,943,206 | Cytochrome c biogenesis protein CcsA |
| Solyc05g016220.1 | 138 | 15,092,149 | 15,092,286 | Ycf1 |
| Solyc10g012230.1 | 146 | 4,687,502 | 4,687,647 | Ycf2 |
| Solyc10g047410.1 | 214 | 40,732,147 | 40,732,360 | Photosystem II CP43 chlorophyll apoprotein |
| Solyc10g049470.1 | 157 | 45,831,441 | 45,831,597 | Ycf1 |
| Solyc11g018700.2 | 172 | 9,107,418 | 9,107,589 | Ycf15 |
| Solyc11g021210.1 | 152 | 13,408,194 | 13,408,345 | Cytochrome c biogenesis protein ccsA |
| Solyc11g021240.2 | 149 | 13,415,639 | 13,415,787 | Hypothetical chloroplast RF1 |
| Solyc11g021300.1 | 154 | 13,419,577 | 13,419,730 | Hypothetical chloroplast RF1 |
| Solyc12g035550.1 | 330 | 41911350 | 41,911,679 | Ycf1 |
| Phytohormones | ||||
| Solyc07g026650.3 | 196 | 30,146,537 | 30,146,732 | ACO5 |
| Solyc09g059510.3 | 445 | 54,883,435 | 54,883,879 | ERF |
| Solyc00g179240.2 | 171 | 17,318,514 | 17,318,684 | MADS-box |
| Solyc10g045690.1 | 143 | 35,000,925 | 35,001,067 | Gibberellin 20-oxidase |
| Solyc12g006350.2 | 344 | 870,483 | 870,826 | Auxin response factor 6 |
| Solyc12g006860.2 | 187 | 1,281,524 | 1,281,710 | Brassinosteroid hydroxylase |
| Pigment | ||||
| Solyc00g019730.2 | 1112 | 10,827,171 | 10,828,282 | Cytochrome c oxidase subunit 3 |
| Solyc00g049210.1 | 378 | 12,649,895 | 12,650,272 | Cytochrome c-type biogenesis protein CcmF |
| Solyc01g017740.1 | 146 | 25,041,926 | 25,042,071 | Cytochrome b6 |
| Solyc02g021770.1 | 186 | 24,252,669 | 24,252,854 | Cytochrome c oxidase subunit 1 |
| Solyc03g013460.1 | 247 | 45,900,762 | 45,901,008 | Cytochrome c oxidase subunit 3 |
| Solyc03g013390.1 | 296 | 46,389,354 | 46,389,649 | Cytochrome c oxidase subunit 3 |
| Solyc03g122000.3 | 437 | 71,495,017 | 71,495,453 | Cytochrome b6-f complex subunit 4 |
| Solyc05g023720.1 | 248 | 29,081,746 | 29,081,993 | Apo cytochrome f |
| Solyc05g025700.1 | 314 | 35,921,169 | 35,921,482 | Cytochrome c biogenesis FC |
| Solyc07g019510.3 | 274 | 11,845,109 | 11,845,382 | Cytochrome c oxidase subunit 1 |
| Solyc07g032450.1 | 167 | 39,127,895 | 39,128,061 | Cytochrome b6 |
| Solyc09g015880.3 | 170 | 11,297,889 | 11,298,058 | Cytochrome c oxidase subunit 2 |
| Solyc09g050020.2 | 851 | 35,489,756 | 35,490,606 | Cytochrome b |
| Solyc11g021210.1 | 152 | 13,408,194 | 13,408,345 | Cytochrome c biogenesis protein ccsA |
| Solyc11g028160.1 | 432 | 20,552,580 | 20,553,011 | Cytochrome c biogenesis |
| Solyc11g039360.1 | 445 | 45,467,509 | 45,467,953 | Cytochrome c biogenesis FC |
| Solyc11g056410.2 | 277 | 45,648,871 | 45,649,147 | Cytochrome c oxidase subunit 2 |
| Solyc11g063620.2 | 262 | 49,936,635 | 49,936,896 | Cytochrome c biogenesis FN |
| Cell wall | ||||
| Solyc07g042220.2 | 161 | 55,403,948 | 55,404,108 | Beta-galactosidase |
| Solyc10g076430.1 | 152 | 59,507,919 | 59,508,070 | Pectinesterase |
| Solyc00g011890.3 | 171 | 9,408,568 | 9,408,738 | Galactokinase-like protein |
| Solyc03g071520.1 | 191 | 19,361,038 | 1,9361,228 | Galactosyltransferase family |
| Solyc05g025500.3 | 192 | 32,902,298 | 32,902,489 | Glucan endo-1 3-beta-glucosidase 6 |
| Solyc07g017730.3 | 202 | 7,823,023 | 7,823,224 | Glucan endo-1 3-beta-glucosidase 5 |
| Antioxidant | ||||
| Solyc01g067460.2 | 207 | 75,956,536 | 75,956,742 | Glutaredoxin family protein |
| Solyc02g092580.3 | 137 | 54,267,591 | 54,267,727 | Peroxidase |
| Solyc06g050530.3 | 191 | 33,316,524 | 33,316,714 | Laccase |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, Y.; Wu, Q.; Cui, C.; Tian, Q.; Zhang, D.; Zhang, Y. ChIP-Seq Analysis of SlAREB1 Downstream Regulatory Network during Tomato Ripening. Foods 2023, 12, 2357. https://doi.org/10.3390/foods12122357
He Y, Wu Q, Cui C, Tian Q, Zhang D, Zhang Y. ChIP-Seq Analysis of SlAREB1 Downstream Regulatory Network during Tomato Ripening. Foods. 2023; 12(12):2357. https://doi.org/10.3390/foods12122357
Chicago/Turabian StyleHe, Yanan, Qiong Wu, Chunxiao Cui, Qisheng Tian, Dongdong Zhang, and Yurong Zhang. 2023. "ChIP-Seq Analysis of SlAREB1 Downstream Regulatory Network during Tomato Ripening" Foods 12, no. 12: 2357. https://doi.org/10.3390/foods12122357






