Complete Genome Sequence of a Novel Lactobacillus paracasei TK1501 and Its Application in the Biosynthesis of Isoflavone Aglycones
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strains and Data Storage
2.2. DNA Extraction
2.3. PCR Amplification and Sequencing
2.4. Phylogenetic Analysis and Gene Annotation
2.5. β-Glucosidase Activity Analysis of L. paracasei TK1502
2.6. Application of L. paracasei TK1502 in the Fermentation of Soymilk
3. Results and Discussion
3.1. Genome Organization and Base Composition
3.2. Analysis of Carbohydrate-Active Enzymes
3.3. Comparative Analysis of β-Glucosidase from L. paracasei TK1501
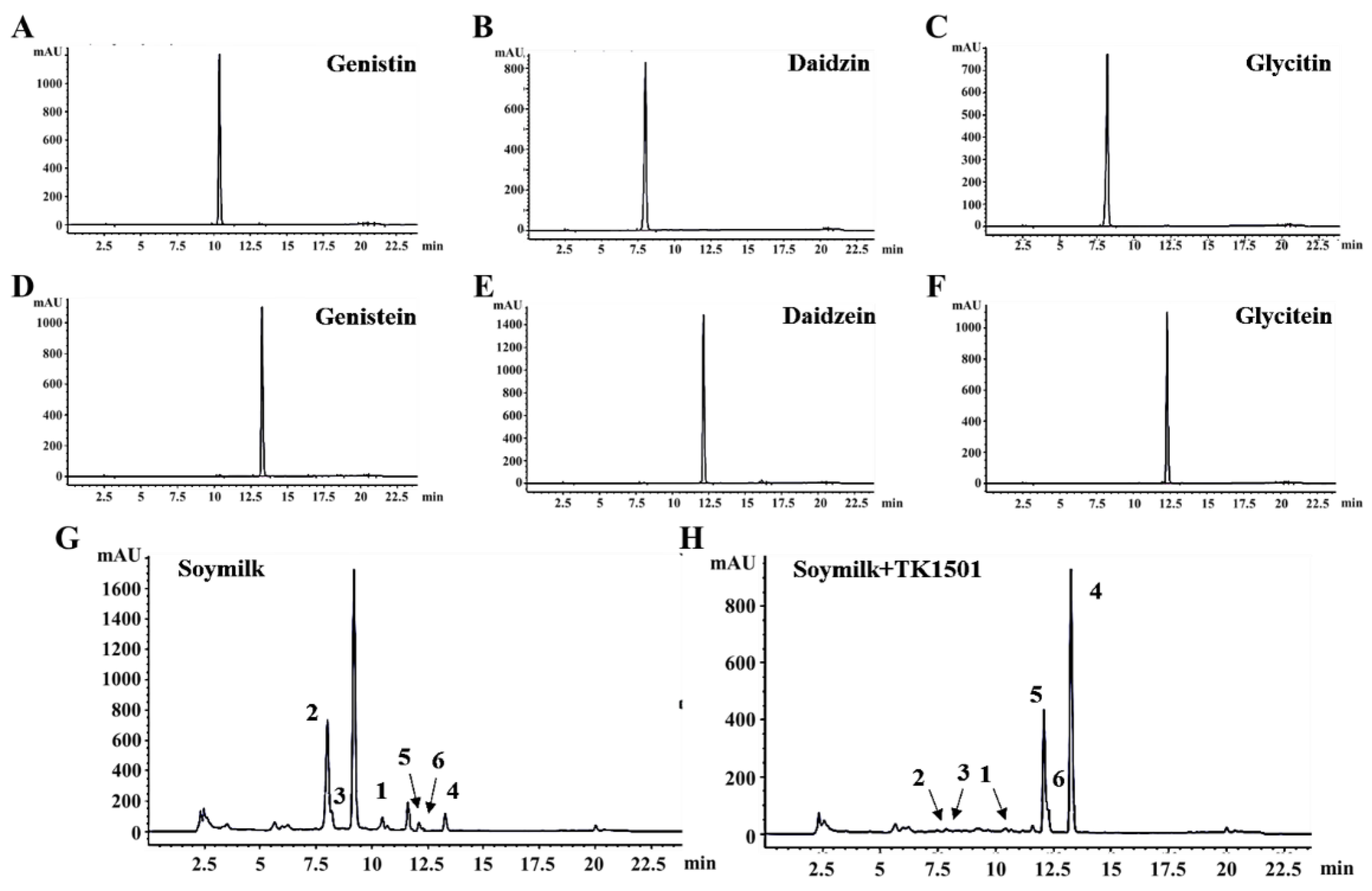
3.4. The Isoflavone Aglycones in Soymilk Were Increased after Fermentation Using L. paracasei TK1502
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ren, B.; Liu, Y.; Zhang, Y.; Cai, Y.; Gong, X.; Chang, Y.; Xu, L.; Zheng, J. Genistein: A Dual Inhibitor of Both Amyloid beta and Human Islet Amylin Peptides. ACS Chem. Neurosci. 2018, 9, 1215–1224. [Google Scholar] [CrossRef]
- Hsiao, Y.H.; Ho, C.T.; Pan, M.H. Bioavailability and health benefits of major isoflavone aglycones and their metabolites. J. Funct. Foods 2020, 74, 104164. [Google Scholar] [CrossRef]
- Omoni, A.O.; Aluko, R.E. Soybean foods and their benefits: Potential mechanisms of action. Nutr. Rev. 2005, 63, 272–283. [Google Scholar] [CrossRef] [PubMed]
- Babu, P.V.; Liu, D.; Gilbert, E.R. Recent advances in understanding the anti-diabetic actions of dietary flavonoids. J. Nutr. Biochem. 2013, 24, 1777–1789. [Google Scholar] [CrossRef] [PubMed]
- Mueller, N.T.; Odegaard, A.O.; Gross, M.D.; Koh, W.P.; Yu, M.C.; Yuan, J.M.; Pereira, M.A. Soy intake and risk of type 2 diabetes in Chinese Singaporeans [corrected]. Eur. J. Nutr. 2012, 51, 1033–1040. [Google Scholar] [CrossRef]
- Adlercreutz, H. Phyto-oestrogens and cancer. Lancet Oncol. 2002, 3, 364–373. [Google Scholar] [CrossRef]
- Harada, T.; Masuda, S.; Arii, M.; Adachi, Y.; Nakajima, M.; Yadomae, T.; Ohno, N. Soy isoflavone aglycone modulates a hematopoietic response in combination with soluble beta-glucan: SCG. Biol. Pharm. Bull. 2005, 28, 2342–2345. [Google Scholar] [CrossRef]
- Marazza, J.A.; Garro, M.S.; de Giori, G.S. Aglycone production by Lactobacillus rhamnosus CRL981 during soymilk fermentation. Food Microbiol. 2009, 26, 333–339. [Google Scholar] [CrossRef]
- Chun, J.; Kim, G.M.; Lee, K.W.; Choi, I.D.; Kwon, G.H.; Park, J.Y.; Jeong, S.J.; Kim, J.S.; Kim, J.H. Conversion of isoflavone glucosides to aglycones in soymilk by fermentation with lactic acid bacteria. J. Food Sci. 2007, 72, M39–M44. [Google Scholar] [CrossRef]
- Mangiapane, E.; Mazzoli, R.; Pessione, A.; Svensson, B.; Riedel, K.; Pessione, E. Ten years of subproteome investigations in lactic acid bacteria: A key for food starter and probiotic typing. J. Proteom. 2015, 127 Pt. B, 332–339. [Google Scholar] [CrossRef]
- Otieno, D.O.; Shah, N.P. A comparison of changes in the transformation of isoflavones in soymilk using varying concentrations of exogenous and probiotic-derived endogenous beta-glucosidases. J. Appl. Microbiol. 2007, 103, 601–612. [Google Scholar] [CrossRef]
- Zhang, N.; Liu, J.H.; Li, J.J.; Chen, C.; Zhang, H.T.; Wang, H.K.; Lu, F.P. Characteristics and Application in Food Preservatives of Lactobacillus plantarum TK9 Isolated from Naturally Fermented Congee. Int. J. Food Eng. 2016, 12, 377–384. [Google Scholar] [CrossRef]
- Li, P.; Gu, Q.; Zhou, Q. Complete genome sequence of Lactobacillus plantarum LZ206, a potential probiotic strain with antimicrobial activity against food-borne pathogenic microorganisms. J. Biotechnol. 2016, 238, 52–55. [Google Scholar] [CrossRef] [PubMed]
- Powell, S.; Forslund, K.; Szklarczyk, D.; Trachana, K.; Roth, A.; Huerta-Cepas, J.; Gabaldon, T.; Rattei, T.; Creevey, C.; Kuhn, M.; et al. eggNOG v4.0: Nested orthology inference across 3686 organisms. Nucleic Acids Res. 2014, 42, D231–D239. [Google Scholar] [CrossRef] [PubMed]
- Moriya, Y.; Itoh, M.; Okuda, S.; Yoshizawa, A.C.; Kanehisa, M. KAAS: An automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 2007, 35, W182–W185. [Google Scholar] [CrossRef]
- Conesa, A.; Gotz, S. Blast2GO: A comprehensive suite for functional analysis in plant genomics. Int. J. Plant Genom. 2008, 2008, 619832. [Google Scholar] [CrossRef] [PubMed]
- Tatusov, R.L.; Galperin, M.Y.; Natale, D.A.; Koonin, E.V. The COG database: A tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 2000, 28, 33–36. [Google Scholar] [CrossRef]
- Stothard, P.; Wishart, D.S. Circular genome visualization and exploration using CGView. Bioinformatics 2005, 21, 537–539. [Google Scholar] [CrossRef]
- Finn, R.D.; Clements, J.; Eddy, S.R. HMMER web server: Interactive sequence similarity searching. Nucleic Acids Res. 2011, 39, W29–W37. [Google Scholar] [CrossRef]
- Li, Y.; Hu, X.; Sang, J.; Zhang, Y.; Zhang, H.; Lu, F.; Liu, F. An acid-stable beta-glucosidase from Aspergillus aculeatus: Gene expression, biochemical characterization and molecular dynamics simulation. Int. J. Biol. Macromol. 2018, 119, 462–469. [Google Scholar] [CrossRef]
- Lombard, V.; Ramulu, H.G.; Drula, E.; Coutinho, P.M.; Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014, 42, D490–D495. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, H.; Takahashi, S.; Watanabe, R.; Fukushima, Y.; Fujita, N.; Noguchi, A.; Yokoyama, R.; Nishitani, K.; Nishino, T.; Nakayama, T. An isoflavone conjugate-hydrolyzing beta-glucosidase from the roots of soybean (Glycine max) seedlings: Purification, gene cloning, phylogenetics, and cellular localization. J. Biol. Chem. 2006, 281, 30251–30259. [Google Scholar] [CrossRef] [PubMed]
- Son, S.H.; Jeon, H.L.; Yang, S.J.; Sim, M.H.; Kim, Y.J.; Lee, N.K.; Paik, H.D. Probiotic lactic acid bacteria isolated from traditional Korean fermented foods based on beta-glucosidase activity. Food Sci. Biotechnol. 2018, 27, 123–129. [Google Scholar] [CrossRef] [PubMed]
- Rekha, C.R.; Vijayalakshmi, G. Isoflavone phytoestrogens in soymilk fermented with beta-glucosidase producing probiotic lactic acid bacteria. Int. J. Food Sci. Nutr. 2011, 62, 111–120. [Google Scholar] [CrossRef]
- Abdella, A.; Mazeed, T.E.-S.; El-Baz, A.F.; Yang, S.-T. Production of β-glucosidase from wheat bran and glycerol by Aspergillus niger in stirred tank and rotating fibrous bed bioreactors. Process Biochem. 2016, 51, 1331–1337. [Google Scholar] [CrossRef]
- Kovács, K.; Megyeri, L.; Szakacs, G.; Kubicek, C.P.; Galbe, M.; Zacchi, G. Trichoderma atroviride mutants with enhanced production of cellulase and β-glucosidase on pretreated willow. Enzym. Microb. Technol. 2008, 43, 48–55. [Google Scholar] [CrossRef]
- Ibrahim, A.S.; El-Shishtawy, M.M.; Pena, A., Jr.; Liou, G.I. Genistein attenuates retinal inflammation associated with diabetes by targeting of microglial activation. Mol. Vis. 2010, 16, 2033–2042. [Google Scholar]



| Strains | Increased Folds |
|---|---|
| Lactobacillus paracasei TK1501 | 25.34 |
| Lactobacillus rhamnosus CRL981 | 4.66 |
| Lactobacillus rhamnosus D7 | 17.06 |
| Lactobacillus paracasei BCRC 14023 | 1.10 |
| Lactobacillus delbrueckii 01181 | 8.36 |
| Lactobacillus plantarum 00144 | 7.15 |
| Streptococcus thermophilus CCRC14085 | 2.59 |
| Bifidobacterium breve K-101 | 3.01 |
| Lactobacillus acidophilus BCRC 10695 | 1.06 |
| Attributes | Values |
|---|---|
| Genome size (bp) | 2,942,538 |
| GC content (%) | 46.54% |
| Plasmid | 0 |
| rRNAs | 15 |
| tRNAs | 59 |
| Other ncRNAs | 40 |
| Proteins | 2947 |
| Property | Numbers of Genes | Percentage (%) |
|---|---|---|
| GHs | 36 | 1.22 |
| GTs | 26 | 0.88 |
| PLs | 0 | 0.00 |
| CEs | 9 | 0.31 |
| AAs | 4 | 0.14 |
| CBMs | 2 | 0.07 |
| Total | 77 | 2.61 |
| Isoflavones | Degradation Rate (%) |
|---|---|
| Genistin | 94.87 ± 0.06 |
| Daidzin | 83.57 ± 0.14 |
| Glycitin | 97.05 ± 0.08 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xie, Y.; Wang, Y.; Han, Y.; Zhang, J.; Wang, S.; Lu, S.; Wang, H.; Lu, F.; Jia, L. Complete Genome Sequence of a Novel Lactobacillus paracasei TK1501 and Its Application in the Biosynthesis of Isoflavone Aglycones. Foods 2022, 11, 2807. https://doi.org/10.3390/foods11182807
Xie Y, Wang Y, Han Y, Zhang J, Wang S, Lu S, Wang H, Lu F, Jia L. Complete Genome Sequence of a Novel Lactobacillus paracasei TK1501 and Its Application in the Biosynthesis of Isoflavone Aglycones. Foods. 2022; 11(18):2807. https://doi.org/10.3390/foods11182807
Chicago/Turabian StyleXie, Yufeng, Yingxue Wang, Yang Han, Jing Zhang, Shumei Wang, Shuwen Lu, Haikuan Wang, Fuping Lu, and Longgang Jia. 2022. "Complete Genome Sequence of a Novel Lactobacillus paracasei TK1501 and Its Application in the Biosynthesis of Isoflavone Aglycones" Foods 11, no. 18: 2807. https://doi.org/10.3390/foods11182807
APA StyleXie, Y., Wang, Y., Han, Y., Zhang, J., Wang, S., Lu, S., Wang, H., Lu, F., & Jia, L. (2022). Complete Genome Sequence of a Novel Lactobacillus paracasei TK1501 and Its Application in the Biosynthesis of Isoflavone Aglycones. Foods, 11(18), 2807. https://doi.org/10.3390/foods11182807





