Mechanistic and Data-Driven Modeling of Ultrasound–Carvacrol Inactivation of Escherichia coli ATCC 25922 in Meat-like Emulsions: Impact of Protein-to-Lipid Ratio
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Culture Preparation
2.2. Formulation of Model Meat-like Emulsions
- High-Lipid (HL): 5% Protein + 15% Lipid (P:L ≈ 0.33)
- Balanced (BM): 10% Protein + 10% Lipid (P:L = 1.0)
- High-Protein (HP): 15% Protein + 5% Lipid (P:L = 3.0)
2.3. Emulsification Process
2.4. Inoculation and Ultrasound-Carvacrol Treatment
2.5. Microbiological Analysis
2.6. Inactivation Kinetic Modeling
2.7. Experimental Design and Data Modeling
2.8. Response Surface Methodology (RSM)
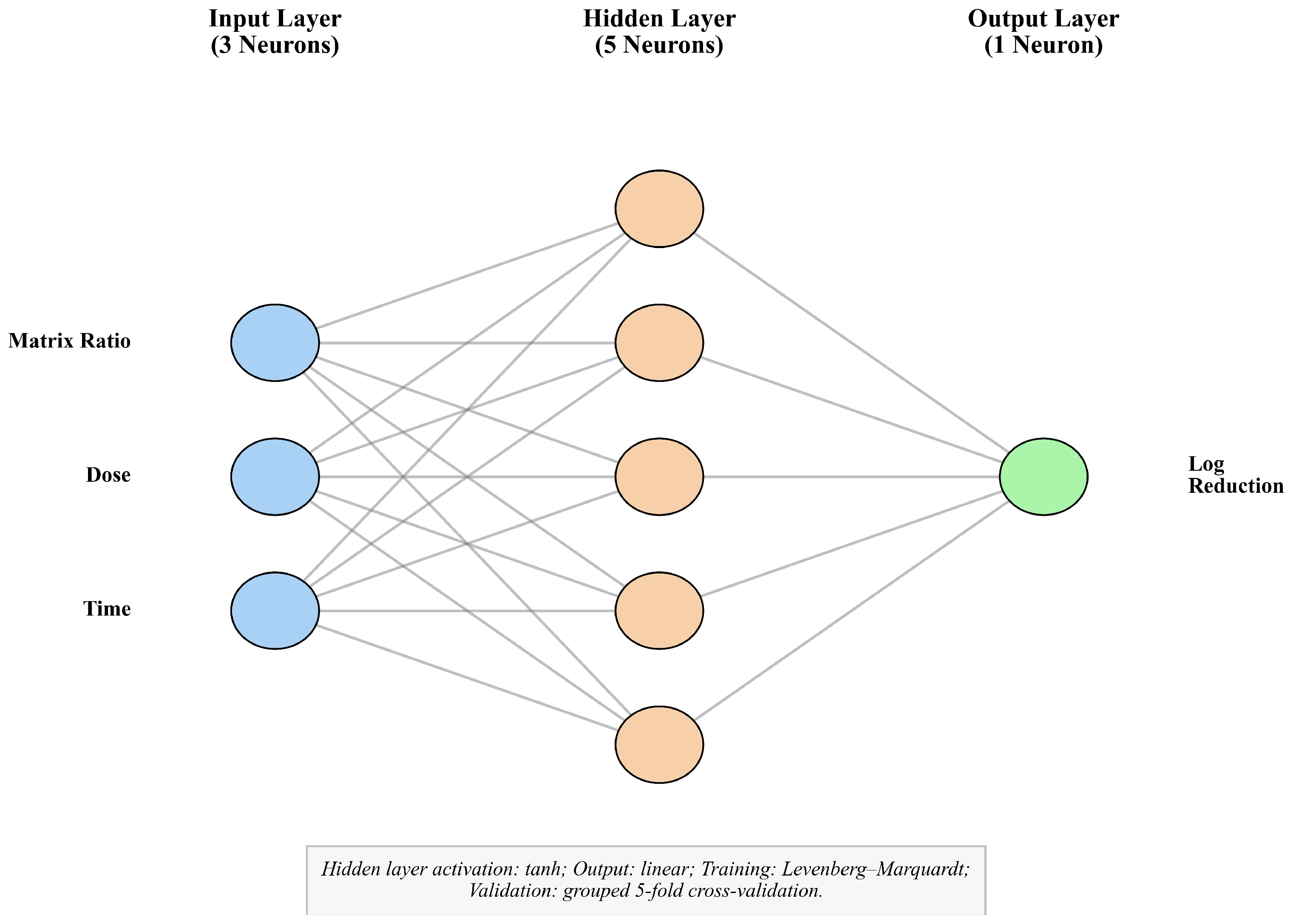
2.9. Artificial Neural Network (ANN) Modeling
2.10. Statistical Analysis
3. Results and Discussion
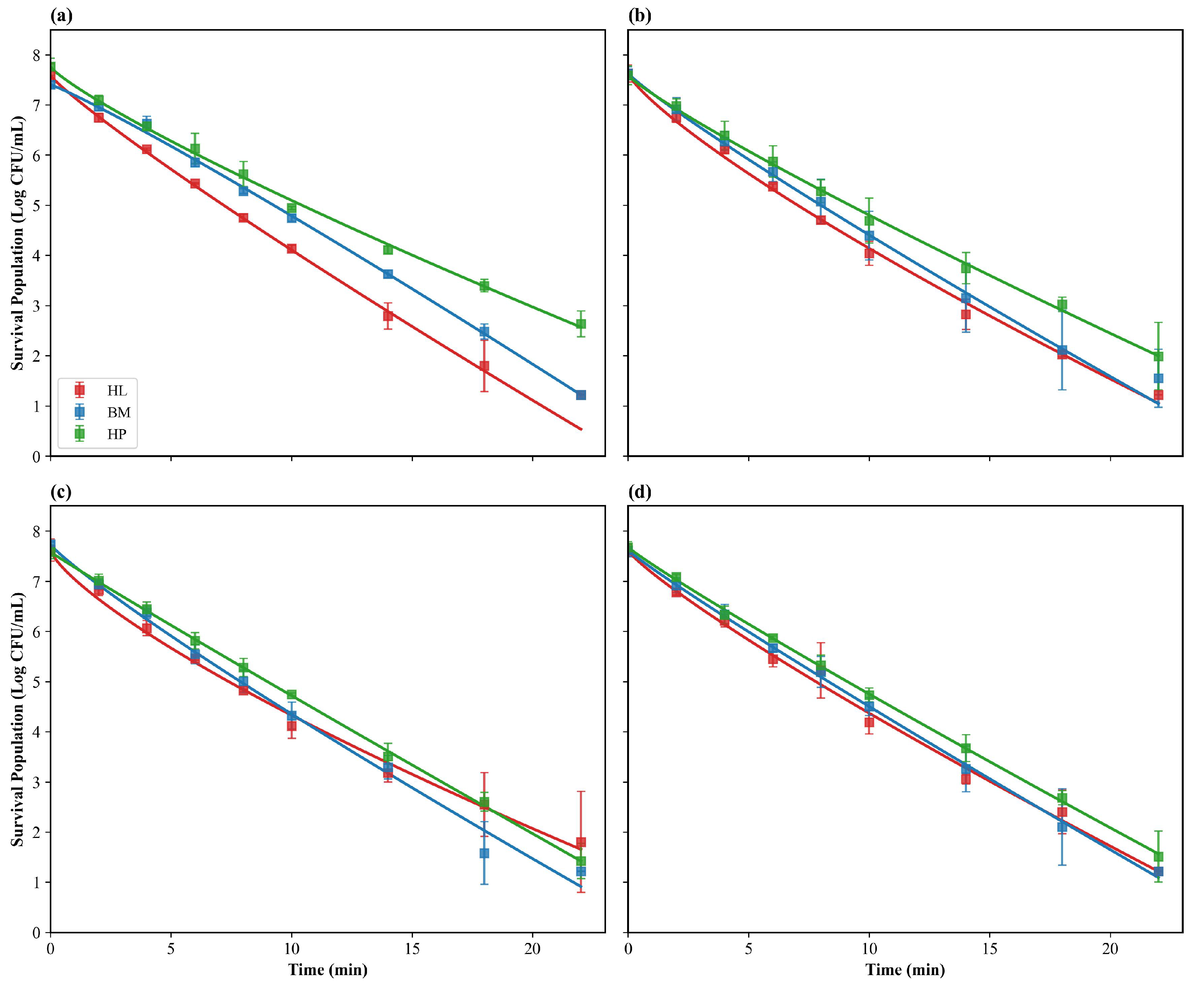
3.1. Matrix-Dependent Inactivation Dynamics: Lipid Enhancement vs. Protein Damping
3.2. Synergistic Interactions: Hurdle Effect and Saturation
3.3. Kinetic Analysis Using the Weibull Model
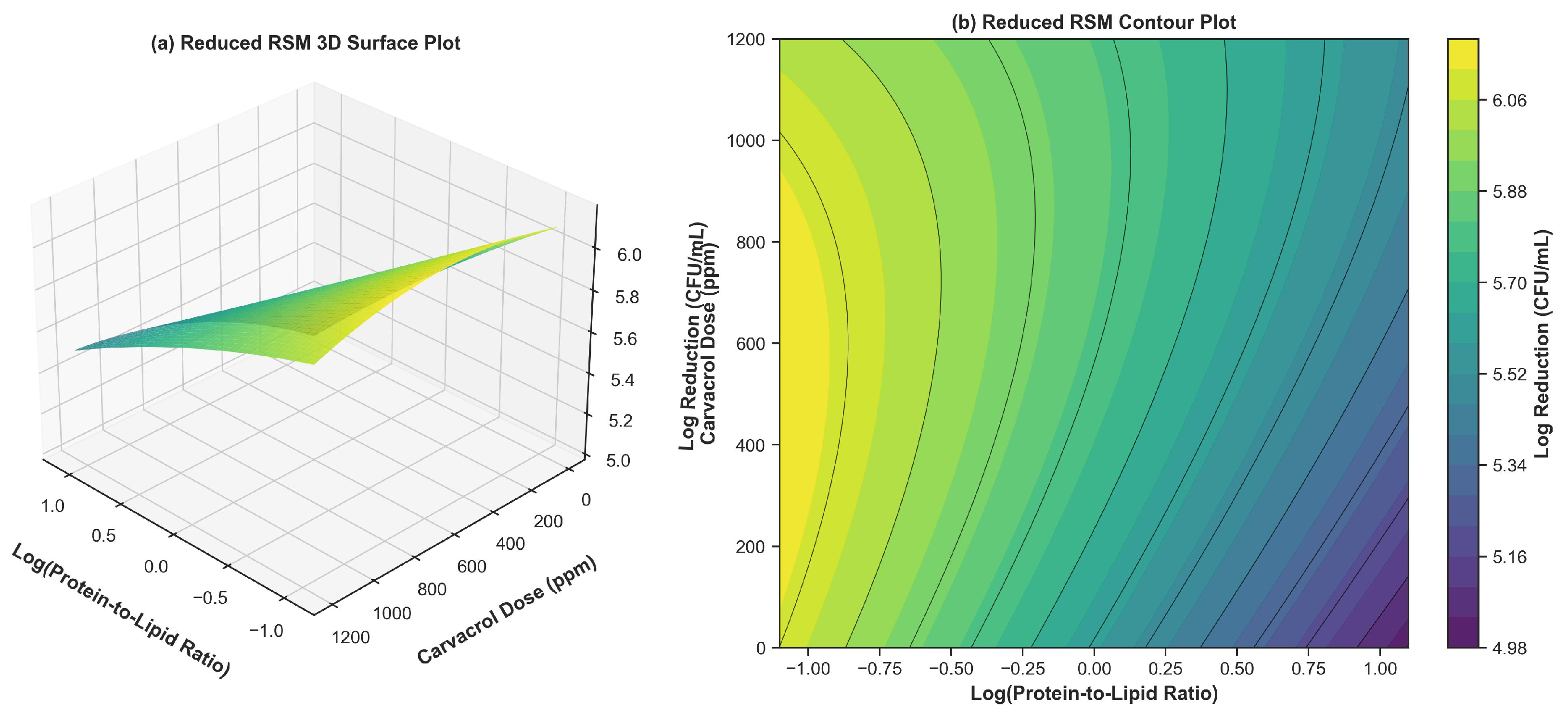
3.4. Process Modeling: Reduced RSM
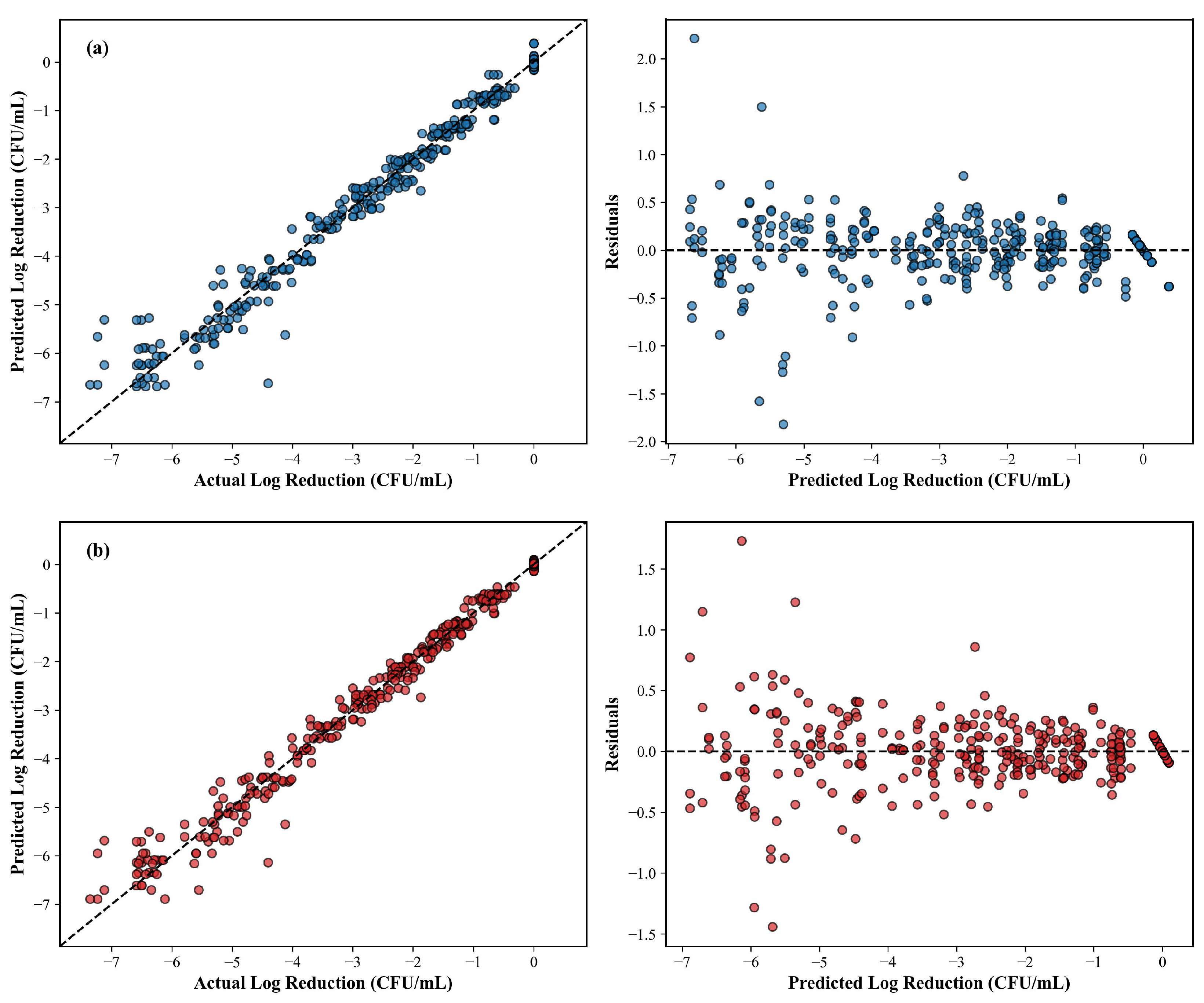
3.5. Comparative Modeling: RSM vs. ANN
3.6. Implications for Industry 4.0: Transparency vs. Complexity
3.7. Sustainability and Industrial Relevance
3.8. Study Limitations and Statistical Considerations
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Brito-Junior, L.; Brito, H.; Simoes, M.; Farias, J.; Santos, B.; Marques, F.; Medeiros, M.; Alves, M.; Pereira, C.; Diniz, A.; et al. Prevalence of Escherichia coli isolates in meat products: A systematic review. Braz. J. Biol. 2025, 85, e292127. [Google Scholar] [CrossRef]
- Kalwaniya, M.; Gaurav, A.; Kumar, N.; Choudhary, D. Prevalence and antibiogram of Escherichia coli isolated from meat and meat products. J. Entomol. Zool. Stud. 2020, 8, 438–443. [Google Scholar]
- Fayemi, O.; Akanni, G.; Elegbeleye, J.; Aboaba, O.; Njage, P. Prevalence, characterization and antibiotic resistance of Shiga toxigenic Escherichia coli serogroups isolated from fresh beef and locally processed ready-to-eat meat products in Lagos, Nigeria. Int. J. Food Microbiol. 2021, 347, 109191. [Google Scholar] [CrossRef]
- Karmi, M. Escherichia coli O157:H7 in raw and processed meat with virulence genes detection in Aswan Governorate. Zagazig Vet. J. 2019, 47, 275–286. [Google Scholar] [CrossRef]
- Hiko, A.; Asrat, D.; Zewde, G. Occurrence of Escherichia coli O157:H7 in retail raw meat products in Ethiopia. J. Infect. Dev. Ctries. 2008, 2, 389–393. [Google Scholar] [CrossRef] [PubMed]
- Hasib, F.; Magouras, I.; St-Hilaire, S.; Paudel, S.; Kamali, M.; Lugsomya, K.; Lam, H.; Elsohaby, I.; Butaye, P.; Nekouei, O.; et al. Prevalence and characterization of antimicrobial-resistant Escherichia coli in chicken meat from wet markets in Hong Kong. Front. Vet. Sci. 2024, 11, 1340548. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Ma, H.; Li, Y.; Gao, S.; Qu, W.; Jiang, X.; Ren, X. Ultrasound technology in fish proteins processing: Innovations in extraction and structure–function dynamics. Ultrason. Sonochem. 2024, 110, 107033. [Google Scholar] [CrossRef]
- Alarcon-Rojo, A.D.; Carrillo-Lopez, L.M.; Reyes-Villagrana, R.; Huerta-Jiménez, M.; Garcia-Galicia, I.A. Ultrasound and meat quality: A review. Ultrason. Sonochem. 2019, 55, 369–382. [Google Scholar] [CrossRef] [PubMed]
- Chemat, F.; Rombaut, N.; Sicaire, A.G.; Meullemiestre, A.; Fabiano-Tixier, A.S.; Abert-Vian, M. Ultrasound assisted extraction of food and natural products. Mechanisms, techniques, combinations, protocols and applications. A review. Ultrason. Sonochem. 2017, 34, 540–560. [Google Scholar] [CrossRef]
- Ashokkumar, M.; Bhaskaracharya, R.; Kentish, S.; Lee, J.; Palmer, M.; Zisu, B. The ultrasonic processing of dairy products: An overview. Dairy Sci. Technol. 2010, 90, 147–168. [Google Scholar] [CrossRef]
- Chang, Y.; Kim, M.; Park, J.; Jo, C. Synergistic inactivation of bacteria using a combination of ultrasound and natural antimicrobials in food systems. Front. Microbiol. 2021, 12, 682900. [Google Scholar] [CrossRef]
- Ultee, A.; Kets, E.P.W.; Smid, E.J. Mechanisms of action of carvacrol on the food-borne pathogen Bacillus cereus. Appl. Environ. Microbiol. 1999, 65, 4606–4610. [Google Scholar] [CrossRef]
- Stratakos, A.C.; Koidis, A. Synergistic interactions of carvacrol with other natural antimicrobials: A review of mechanisms and applications in food preservation. Food Microbiol. 2018, 73, 118–129. [Google Scholar] [CrossRef]
- Liao, X.; Liu, D.; Xiang, Q.; Ahn, J.; Chen, S. Inactivation of foodborne pathogens based on synergistic effects of ultrasound and natural compounds during fresh produce washing. Ultrason. Sonochem. 2020, 64, 104958. [Google Scholar] [CrossRef]
- Nazzaro, F.; Fratianni, F.; De Martino, L.; Coppola, R.; De Feo, V. Mechanisms, applications and challenges of natural antimicrobials in food systems. Food Control 2022, 138, 109017. [Google Scholar] [CrossRef]
- Silva, C.L.; Pereira, R.N.; Vicente, A.A. Lights and shadows of essential oil-derived compounds as food preservatives: Efficacy, mechanisms and resistance prevention. Compr. Rev. Food Sci. Food Saf. 2025, 24, 1–30. [Google Scholar]
- Wang, L.; Dekker, M.; Heising, J.; Zhao, L.; Fogliano, V. Food matrix design can influence the antimicrobial activity in the food systems: A narrative review. Crit. Rev. Food Sci. Nutr. 2024, 64, 8963–8989. [Google Scholar] [CrossRef]
- Cichoski, A.J.; Silva, M.S.; Le aes, Y.S.V.; Brasil, C.C.B.; de Menezes, C.R.; Barin, J.S.; Wagner, R.; Campagnol, P.C.B. Ultrasound: A promising technology to improve the technological quality of meat emulsions. Meat Sci. 2019, 148, 150–155. [Google Scholar] [CrossRef]
- Delgado-Pando, G.; Cofrades, S.; Ruiz-Capillas, C.; Solas, M.T.; Jim’enez-Colmenero, F. Healthier lipid combination oil-in-water emulsions prepared with various protein systems: An approach for development of functional meat products. Eur. J. Lipid Sci. Technol. 2010, 112, 791–801. [Google Scholar] [CrossRef]
- Cofrades, S.; Antoniou, I.; Solas, M.; Herrero, A.M.; Jim’enez-Colmenero, F. Preparation and impact of multiple (water-in-oil-in-water) emulsions in meat systems. Food Chem. 2013, 141, 338–346. [Google Scholar] [CrossRef]
- Ren, Y.; Huang, L.; Zhang, Y.; Li, H.; Zhao, D.; Cao, J.; Liu, X. Application of Emulsion Gels as Fat Substitutes in Meat Products. Foods 2022, 11, 1950. [Google Scholar] [CrossRef] [PubMed]
- Singh, K.A.; Patra, F.; Ghosh, T.; Mahnot, N.K.; Dutta, H.; Duary, R.K. Advancing food systems with industry 5.0: A systematic review of smart technologies, sustainability, and resource optimization. Sustain. Futures 2025, 9, 100694. [Google Scholar] [CrossRef]
- Kim, T.K.; Lee, M.H.; Yong, H.I.; Jang, H.W.; Jung, S.; Choi, Y.S. Impacts of fat types and myofibrillar protein on the rheological properties and thermal stability of meat emulsion systems. Food Chem. 2020, 346, 128930. [Google Scholar] [CrossRef]
- Jim’enez-Colmenero, F.; Salcedo-Sandoval, L.; Bou, R.; Cofrades, S.; Herrero, A.M.; Ruiz-Capillas, C. Novel applications of oil-structuring methods as a strategy to improve the fat content of meat products. Trends Food Sci. Technol. 2015, 44, 177–188. [Google Scholar] [CrossRef]
- Herrero, A.M.; Ruiz-Capillas, C. Novel lipid materials based on gelling procedures as fat analogues in the development of healthier meat products. Curr. Opin. Food Sci. 2021, 39, 1–6. [Google Scholar] [CrossRef]
- Zheng, B.; Li, X.; Hao, J.; Xu, D. Meat systems produced with Monascus pigment water-in-oil-in-water multiple emulsion as pork fat replacers. Food Chem. 2022, 402, 134080. [Google Scholar] [CrossRef]
- Herigstad, B.; Hamilton, M.; Heersink, J. How to optimize the drop plate method for enumerating bacteria. J. Microbiol. Methods 2001, 44, 121–129. [Google Scholar] [CrossRef]
- Mafart, P.; Couvert, O.; Gaillard, S.; Leguerinel, I. On calculating sterility in thermal preservation methods: Application of the Weibull frequency distribution model. Int. J. Food Microbiol. 2002, 72, 107–113. [Google Scholar] [CrossRef]
- Haykin, S.S. Neural Networks: A Comprehensive Foundation; Prentice Hall: Englewood Cliffs, NJ, USA, 1999. [Google Scholar]
- Bishop, C.M.; Nasrabadi, N.M. Pattern Recognition and Machine Learning; Springer: Berlin/Heidelberg, Germany, 2006; Volume 4. [Google Scholar]
- Hagan, M.T.; Menhaj, M.B. Training feedforward networks with the Marquardt algorithm. IEEE Trans. Neural Networks 1994, 5, 989–993. [Google Scholar] [CrossRef]
- Orthaber, U.; Zevnik, J.; Petkovšek, R.; Dular, M. Cavitation bubble collapse in a vicinity of a liquid-liquid interface—Basic research into emulsification process. Ultrason. Sonochem. 2020, 68, 105224. [Google Scholar] [CrossRef]
- Han, R.; Zhang, A.M.; Tan, S.; Li, S. Interaction of cavitation bubbles with the interface of two immiscible fluids on multiple time scales. J. Fluid Mech. 2022, 932, A8. [Google Scholar] [CrossRef]
- Subbiahdoss, G.; Reimhult, E. Biofilm formation at oil-water interfaces is not a simple function of bacterial hydrophobicity. Colloids Surfaces B Biointerfaces 2020, 194, 111163. [Google Scholar] [CrossRef]
- Yang, T.; Jin, Y.; Neogi, A. Acoustic attenuation and dispersion in fatty tissues and tissue phantoms influencing ultrasound biomedical imaging. ACS Omega 2022, 8, 1319. [Google Scholar] [CrossRef]
- Qin, D.; Zou, Q.; Lei, S.; Wang, W.; Li, Z. Cavitation dynamics and inertial cavitation threshold of lipid coated microbubbles in viscoelastic media with bubble–bubble interactions. Micromachines 2021, 12, 1125. [Google Scholar] [CrossRef]
- Shen, X.; Wu, P.; Lin, W. Numerical simulation of cavitation threshold in water and viscoelastic medium based on bubble cluster dynamics. Ultrason. Sonochem. 2025, 119, 107414. [Google Scholar] [CrossRef]
- Du, J.; Zhan, Y.; Zou, T.; Shi, L.; Liu, G.; Yuan, D. Mechanism of ultrasonic pre-treatment in enhancing carvacrol preservation: An integrated strategy validated by molecular dynamics simulation and application in chilled pork. Food Chem. X 2026, 34, 103550. [Google Scholar] [CrossRef]
- Fitriyanti, M.; Bagherzadeh, S.; Narsimhan, G. Synergistic effect of ultrasound and antimicrobial solutions of cecropin P1 in the deactivation of Escherichia coli O157:H7 using a cylindrical ultrasonic system. Sci. Rep. 2023, 13, 10997. [Google Scholar] [CrossRef] [PubMed]
- Patil, U.; Benjakul, S. Combined effect of ultrasound and enzymatic treatment on the physicochemical and functional properties of myofibrillar proteins. Ultrason. Sonochem. 2020, 69, 105268. [Google Scholar] [CrossRef]
- Fan, L.; Zhang, X.; Liu, Y.; Wang, Y.; Jiang, P. Antibacterial mechanism of ultrasound combined with essential oils against Escherichia coli and Staphylococcus aureus. Ultrason. Sonochem. 2021, 73, 105509. [Google Scholar] [CrossRef]
- Li, H.; Fan, J.; Guan, W.; Qiao, Y.; Chen, Q.; Wang, X.; Gu, Y.; Sun, C.; Li, Y.; Sui, X.; et al. Nizip Yaglık olive leaves (Olea europaea L.) collected at different seasons and altitudes: Enzyme inhibition, antioxidant activities and phenolic compound profiles. Food Biosci. 2024, 62, 105524. [Google Scholar] [CrossRef]
- Wang, Z.; Bi, X.; Xiang, R.; Chen, L.; Feng, X.; Zhou, M.; Che, Z. Inactivation of Escherichia coli by ultrasound combined with nisin. J. Food Prot. 2018, 81, 993–1000. [Google Scholar] [CrossRef]
- Zhao, S.; Dong, Z.; Yao, C.; Wen, Z.; Chen, G.; Yuan, Q. Liquid–liquid two-phase flow in ultrasonic microreactors: Cavitation, emulsification and mass transfer enhancement. AIChE J. 2018, 64, 1412–1423. [Google Scholar] [CrossRef]
- Liu, P.; Wu, Z.; Fang, Z.; Cravotto, G. Sonolytic degradation kinetics and mechanisms of antibiotics in water and cow milk. Ultrason. Sonochem. 2023, 99, 106518. [Google Scholar] [CrossRef]
- Esua, O.J.; Sun, D.W.; Ajani, C.K.; Cheng, J.H.; Keener, K.M. Modelling of inactivation kinetics of Escherichia coli and Listeria monocytogenes on grass carp treated by combining ultrasound with plasma functionalized buffer. Ultrason. Sonochem. 2022, 88, 106086. [Google Scholar] [CrossRef] [PubMed]
- Gera, N.; Doores, S. Kinetics and mechanism of bacterial inactivation by ultrasound waves and sonoprotective effect of milk components. J. Food Sci. 2011, 76, M111–M119. [Google Scholar] [CrossRef] [PubMed]
- Iñiguez-Moreno, M.; Calderón-Santoyo, M.; Ascanio, G.; Brito-Bazán, E.; Córdova-Aguilar, M.S.; Brito-de la Fuente, E.; Ragazzo-Sánchez, J.A. Inactivation Kinetics of Escherichia coli and Staphylococcus aureus Using Ultrasound in a Model Parenteral Emulsion. Appl. Microbiol. 2025, 5, 34. [Google Scholar] [CrossRef]
- Nunes, B.V.; da Silva, C.N.; Bastos, S.C.; de Souza, V.R. Microbiological Inactivation by Ultrasound in Liquid Products. Food Bioprocess Technol. 2022, 15, 2185–2209. [Google Scholar] [CrossRef]
- Hassoun, A.; Aït-Kaddour, A.; Abu-Mahfouz, A.; Rathod, N.; Bader, F.; Barba, F.; Biancolillo, A.; Cropotova, J.; Galanakis, C.; Jambrak, A.; et al. The fourth industrial revolution in the food industry—Part I: Industry 4.0 technologies. Crit. Rev. Food Sci. Nutr. 2023, 63, 6547–6563. [Google Scholar] [CrossRef] [PubMed]
- Arteaga-Cabrera, E.; Ramírez-Márquez, C.; Sánchez-Ramírez, E.; Segovia-Hernández, J.G.; Osorio-Mora, O.; Gómez-Salazar, J.A. Advancing optimisation strategies in the food industry: From traditional approaches to multi-objective and technology-integrated solutions. Appl. Sci. 2025, 15, 3846. [Google Scholar] [CrossRef]
- Krupitzer, C.; Noack, T.; Borsum, C. Digital food twins combining data science and food science: System model, applications and challenges. Processes 2022, 10, 1781. [Google Scholar] [CrossRef]
- Liu, Y.; Dar, B.; Makroo, H.; Aslam, R.; Marti-Quijal, F.; Castagnini, J.; Amigo, J.; Barba, F. Optimizing recovery of high-added-value compounds from complex food matrices using multivariate methods. Antioxidants 2024, 13, 1510. [Google Scholar] [CrossRef]
- Youssefi, S.; Emam-Djomeh, Z.; Mousavi, S. Comparison of artificial neural network and response surface methodology in prediction of quality parameters of spray-dried pomegranate juice. Dry. Technol. 2009, 27, 910–917. [Google Scholar] [CrossRef]
- Desai, K.; Survase, S.; Saudagar, P.; Lele, S.; Singhal, R. Comparison of artificial neural network and response surface methodology in fermentation media optimization. Biochem. Eng. J. 2008, 41, 266–273. [Google Scholar] [CrossRef]
- Misra, N.N.; Kumar, S.; Mishra, H.N. Optimisation of process parameters for development of probiotic-GABA enriched nutri bar by RSM coupled with multi-parameter optimised ANN model. Probiotics Antimicrob. Proteins 2025, 17, 2611–2628. [Google Scholar] [CrossRef]
- Jha, A.; Sit, N. Comparison of response surface methodology and artificial neural network modelling for supercritical fluid extraction of phytochemicals from Terminalia chebula pulp. Ind. Crop. Prod. 2021, 170, 113769. [Google Scholar] [CrossRef]
- Mukherjee, R.; Chakraborty, R.; Dutta, A. Comparison of optimization approaches (RSM and ANN–GA) for soybean meal fermentation. J. Food Process Eng. 2019, 42, e13124. [Google Scholar] [CrossRef]
- Ware, K.; Kashyap, P.; Gorde, P.; Yadav, R.; Sharma, V. Comparative analysis of RSM and ANN-GA based modelling for protein extraction from cotton seed meal. Food Bioprod. Process. 2024, 150, 63–77. [Google Scholar] [CrossRef]
- Shu, H.; Chen, X.; Jiang, Q.; Wang, Y.; Wan, Z.; Xu, J.; Wang, P. Optimization of fungal secondary metabolites production via RSM coupled with multi-parameter optimised ANN model. Bioresour. Technol. 2024, 413, 131495. [Google Scholar] [CrossRef]
- Dhal, K.; Das, P.; Singh, S.; Talukdar, M. Analysis of ultraacoustic behavior of L-aspartic acid in aqueous sodium benzoate and ammonium acetate media. J. Mol. Liq. 2023, 376, 121413. [Google Scholar] [CrossRef]
- Cardoso, L.T.; Alexandre, B.; Cacciatore, F.A.; Magedans, Y.V.S.; Fett-Neto, A.G.; Contri, R.V.; Malheiros, P.S. Carvacrol-loaded nanoemulsions produced with a natural emulsifier for lettuce sanitization. Food Res. Int. 2023, 168, 112748. [Google Scholar] [CrossRef] [PubMed]


| Matrix | Dose (ppm) | (min) | p | (min) | |
|---|---|---|---|---|---|
| HL (P:L = 0.33) | 0 | 0.991 | |||
| 600 | 0.996 | ||||
| 900 | 0.984 | ||||
| 1200 | 0.987 | ||||
| BM (P:L = 1.0) | 0 | 0.998 | |||
| 600 | 0.993 | ||||
| 900 | 0.987 | ||||
| 1200 | 0.993 | ||||
| HP (P:L = 3.0) | 0 | 0.995 | |||
| 600 | 0.993 | ||||
| 900 | 0.997 | ||||
| 1200 | 0.996 |
| Parameter | Matrix | Dose | Interaction |
|---|---|---|---|
| Weibull | 20.29 (2,24), | 4.62 (3,24), | 3.54 (6,24), |
| Weibull p | 12.25 (2,24), | 2.95 (3,24), | 4.06 (6,24), |
| 8.08 (2,24), | 0.72 (3,24), | 2.37 (6,24), |
| Term | Coefficient () | Std. Error | t-Value | p-Value | Significance |
|---|---|---|---|---|---|
| Intercept | * | ||||
| Log_Matrix_Index () | <0.001 | ** | |||
| Dose (D) | ** | ||||
| Time (T) | <0.001 | ** | |||
| Dose2 () | * | ||||
| Time2 () | <0.001 | ** | |||
| <0.001 | ** | ||||
| <0.001 | ** |
| Source | Sum of Squares | df | Mean Square | F-Value | p-Value |
|---|---|---|---|---|---|
| Model Regression | 7 | <0.001 | |||
| Residual | 316 | ||||
| Lack of Fit | 100 | ||||
| Pure Error | 216 | ||||
| Total | 323 |
| Metric | Reduced RSM | ANN (3–5–1) | ||
|---|---|---|---|---|
| (Overall Fit) | Training | Validation | Testing | |
| RMSE (log CFU/mL) | 0.347 | |||
| MAE (log CFU/mL) | – | |||
| Lack-of-Fit Analysis | ||||
| Sum of Squares (LoF) | ||||
| F-value | 1.34 | |||
| p-value | ||||
| Matrix Type | Optimal Dose | Optimal Time | Predicted Log Red |
|---|---|---|---|
| High-lipid (HL) | 0 ppm | min | |
| Balanced (BM) | 600 ppm | min | |
| High-Protein (HP) | 0 ppm | min |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Baghirov, K.; Şahmurat, F. Mechanistic and Data-Driven Modeling of Ultrasound–Carvacrol Inactivation of Escherichia coli ATCC 25922 in Meat-like Emulsions: Impact of Protein-to-Lipid Ratio. Processes 2026, 14, 797. https://doi.org/10.3390/pr14050797
Baghirov K, Şahmurat F. Mechanistic and Data-Driven Modeling of Ultrasound–Carvacrol Inactivation of Escherichia coli ATCC 25922 in Meat-like Emulsions: Impact of Protein-to-Lipid Ratio. Processes. 2026; 14(5):797. https://doi.org/10.3390/pr14050797
Chicago/Turabian StyleBaghirov, Kamran, and Fatma Şahmurat. 2026. "Mechanistic and Data-Driven Modeling of Ultrasound–Carvacrol Inactivation of Escherichia coli ATCC 25922 in Meat-like Emulsions: Impact of Protein-to-Lipid Ratio" Processes 14, no. 5: 797. https://doi.org/10.3390/pr14050797
APA StyleBaghirov, K., & Şahmurat, F. (2026). Mechanistic and Data-Driven Modeling of Ultrasound–Carvacrol Inactivation of Escherichia coli ATCC 25922 in Meat-like Emulsions: Impact of Protein-to-Lipid Ratio. Processes, 14(5), 797. https://doi.org/10.3390/pr14050797







