A New ODE-Based Julia Implementation of the Anaerobic Digestion Model No. 1 Greatly Outperforms Existing DAE-Based Java and Python Implementations
Abstract
1. Introduction
1.1. Anaerobic Digestion
1.2. The Anaerobic Digestion Model 1
1.3. Benchmark Simulation Model 2
1.4. Purpose of Comparison
2. Materials and Methods
2.1. The Python Implementations
2.2. The Java Implementation
2.3. The Julia Implementation
2.4. Null-Hypothesis Significance Testing
2.5. Validating the Julia Code with the Python DAE Implementation
2.6. Comparison of Julia, Java, and Python Implementations
2.6.1. Datasets
2.6.2. Comparison of Compute Times
2.7. Statistical Analysis
3. Results
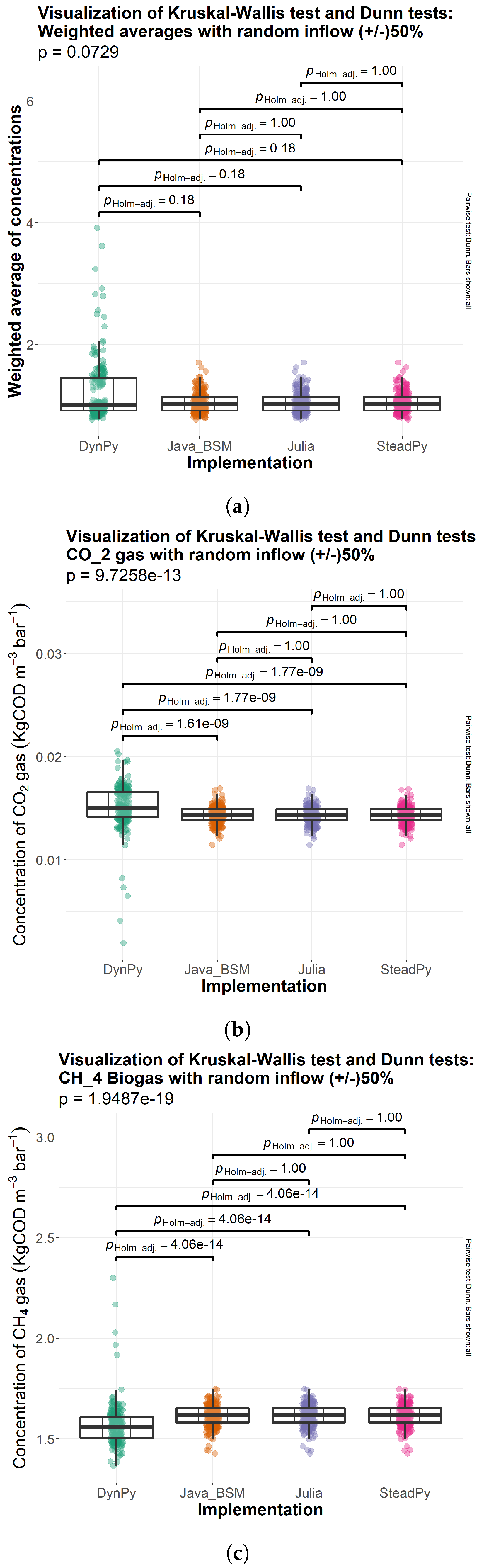
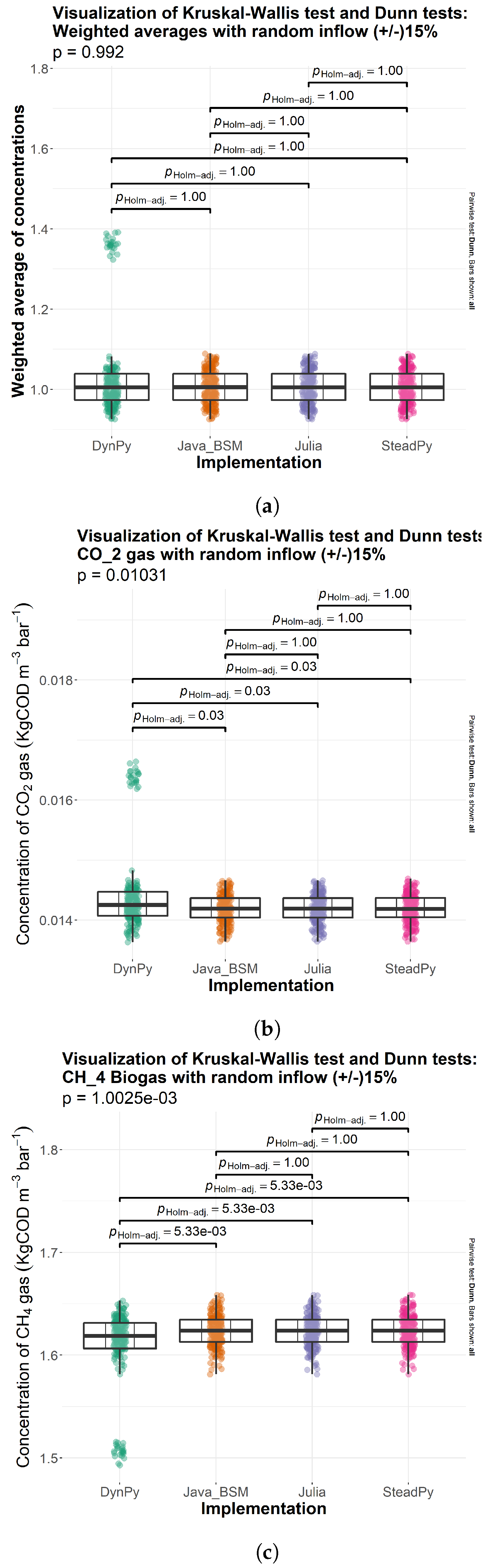
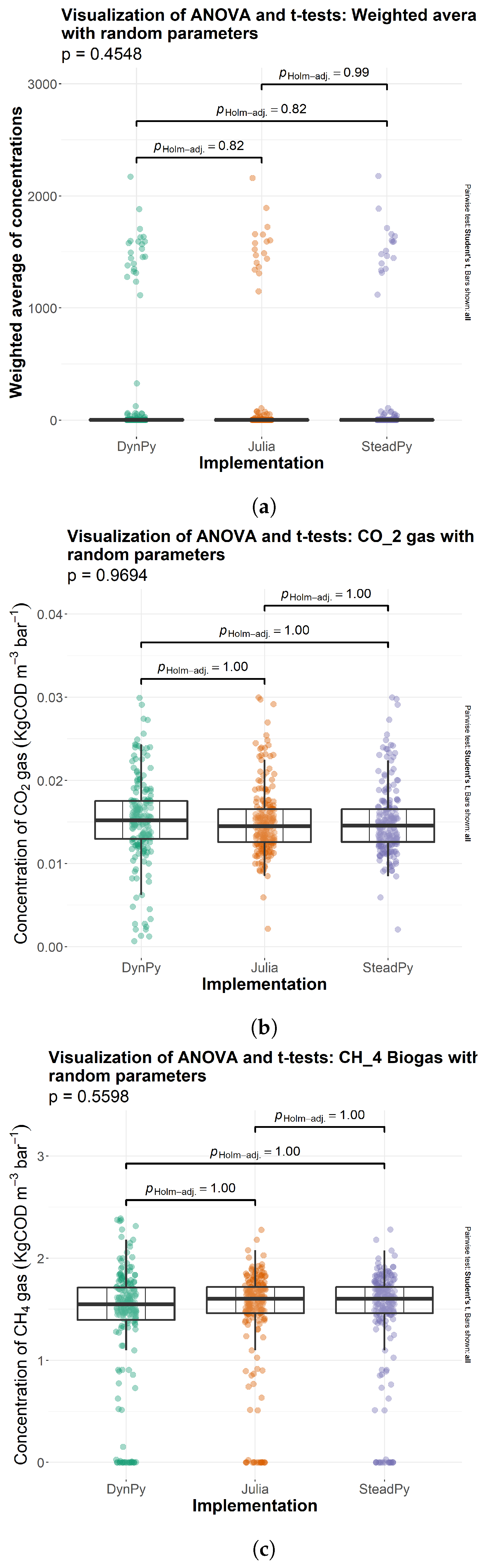
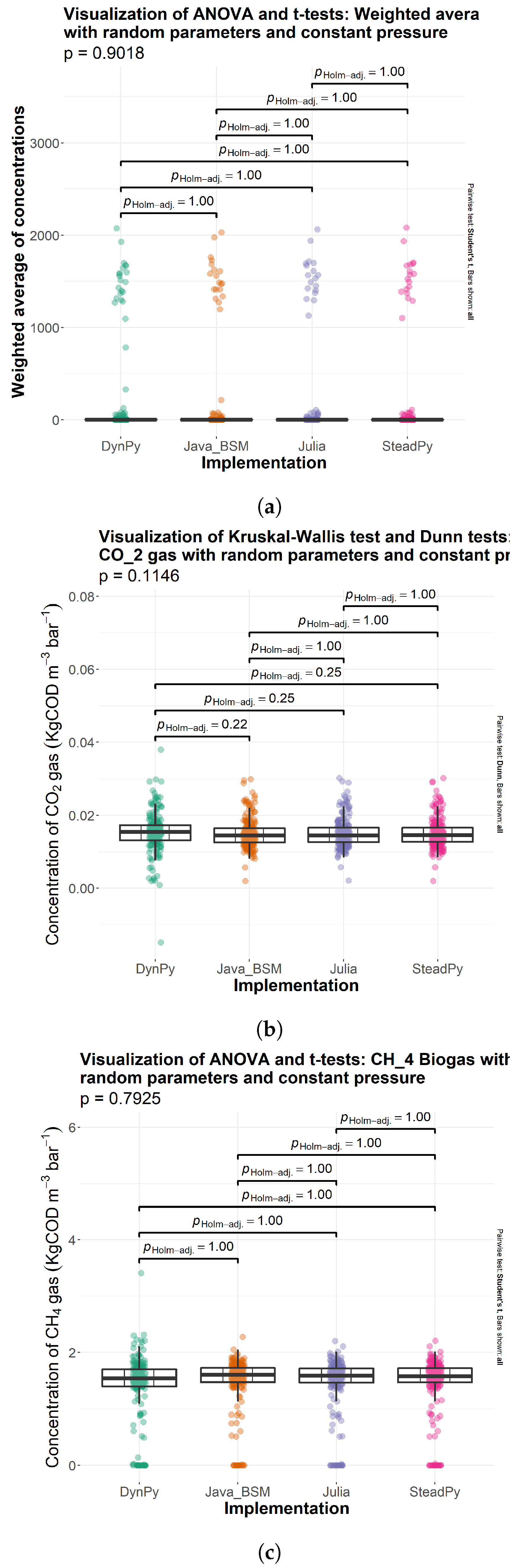
3.1. Comparison of Solutions
3.2. Computational Time
4. Discussion
4.1. Reproducibility and Validation
4.2. Compute Time: Why It Matters
4.3. DAE vs. ODE Formulations of ADM1
4.4. Use of the Petersen Matrix Formulation of ADM1
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AD | Anaerobic Digestion |
| ADM1 | Anaerobic Digestion Model Number 1 |
| ANOVA | Analysis of Variances |
| BSM2 | Benchmark Simulation Model 2 |
| DAE | Differential–Algebraic Equation |
| ODE | Ordinary Differential Equation |
| STDEV | Standard Deviation |
References
- Kunatsa, T.; Xia, X. A review on anaerobic digestion with focus on the role of biomass co-digestion, modelling and optimisation on biogas production and enhancement. Bioresour. Technol. 2022, 344, 126311. [Google Scholar] [CrossRef] [PubMed]
- Uddin, M.N.; Siddiki, S.Y.A.; Mofijur, M.; Djavanroodi, F.; Hazrat, M.A.; Show, P.L.; Ahmed, S.F.; CHu, Y.M. Prospects of Bioenergy Production From Organic Waste Using Anaerobic Digestion Technology: A Mini Review. Front. Energy Res. 2021, 9, 627093. [Google Scholar] [CrossRef]
- Uddin, M.M.; Wright, M.M. Anaerobic Digestion Fundamentals, Challenges, and Technological Advances. Phys. Sci. Rev. 2022. [Google Scholar] [CrossRef]
- Rittmann, B.E.; McCarty, P.L. Environmental Biotechnology: Principles and Applications; McGraw-Hill: New York, NY, USA, 2001. [Google Scholar]
- Meegoda, J.N.; Li, B.; Patel, K.; Wang, L.B. A Review of the Processes, Parameters, and Optimization of Anaerobic Digestion. Int. J. Environ. Res. Public Health 2018, 15, 2224. [Google Scholar] [CrossRef] [PubMed]
- Batstone, D.; Keller, J.; Angelidaki, I.; Kalyuzhnyi, S.; Pavlostathis, S.; Rozzi, A.; Sanders, W.; Siegrist, H.; Vavilin, V. Anaerobic Digestion Model No. 1 (ADM1); Scientific and Technical Report, no. 13; IWA Publisher: London, UK, 2002. [Google Scholar]
- Alex, J.; Benedetti, L.; Copp, J.; Gernaey, K.; Jeppsson, U.; Nopens, I.; Pons, M.; Rosen, C.; Steyer, J.; Vanrolleghem, P. Benchmark Simulation Model No. 2 (BSM2); International Water Association: London, UK, 2019. [Google Scholar]
- Gavaghan, D. Problems with the Current Approach to the Dissemination of Computational Science Research and Its Implications for Research Integrity. Bull. Math. Biol. 2018, 80, 3088–3094. [Google Scholar] [CrossRef] [PubMed]
- Schnell, S. “Reproducible” Research in Mathematical Sciences Requires Changes in our Peer Review Culture and Modernization of our Current Publication Approach. Bull. Math. Biol. 2018, 80, 3095–3105. [Google Scholar] [CrossRef] [PubMed]
- Stagge, J.H.; Rosenberg, D.E.; Abdallah, A.M.; Akbar, H.; Attallah, N.A.; James, R. Assessing Data Availability and Research Reproducibility in Hydrology and Water Resources. Sci. Data 2019, 6, 190030. [Google Scholar] [CrossRef] [PubMed]
- Rosén, C.; Jeppsson, U. Aspects on ADM1 Implementation within the BSM2 Framework; Department of Industrial Electrical Engineering and Automation, Lund University: Lund, Sweden, 2006; pp. 1–35. [Google Scholar]
- Sadrimajd, P.; Mannion, P.; Howley, E.; Lens, P.N.L. PyADM1: A Python Implementation of Anaerobic Digestion Model No. 1. bioRxiv 2021. [Google Scholar] [CrossRef]
- Hairer, E.; Nørsett, S.; Wanner, G. Explicit Runge-Kutta Methods of Higher Order. In Solving Ordinary Differential Equations I: Nonstiff Problems; Springer Series in Computational Mathematics; Springer: Berlin/Heidelberg, Germany, 2008; Chapter II; pp. 181–184. [Google Scholar]
- Pettigrew, L.; Hubert, S.; Groß, F.; Delgado, A. Implementation of Dynamic Biological Process Models into a Reference Net Simulation Environment. In Proceedings of the ASIM Dedicated Conference Simulation in Production and Logistics, Dortmund, Germany, 24 September 2015. [Google Scholar]
- Pettigrew, L.; Gutbrod, A.; Domes, H.; Groß, F.; Méndez-Contreras, J.M.; Delgado, A. Modified ADM1 for high-rate anaerobic co-digestion of thermally pre-treated brewery surplus yeast wastewater. Water Sci. Technol. 2017, 76, 542–554. [Google Scholar] [CrossRef]
- Hairer, E.; Nørsett, S.; Wanner, G. Chapter III.1 Classical Linear Multistep Formulas. In Solving Ordinary Differential Equations I: Nonstiff Problems; Springer Series in Computational Mathematics; Springer: Berlin/Heidelberg, Germany, 1993; pp. 356–361. [Google Scholar]
- Rackauckas, C. A Comparison Between Differential Equation Solver Suites in MATLAB, R, Julia, Python, C, Mathematica, Maple, and Fortran. Winnower 2018. [Google Scholar] [CrossRef]
- Rackauckas, C.; Nie, Q. Differentialequations.jl–a Performant and Feature-Rich Ecosystem for Solving Differential Equations in Julia. J. Open Res. Softw. 2017, 5. [Google Scholar] [CrossRef]
- Fox, J. Applied Regression Analysis and Generalized Linear Models, 3rd ed.; Sage: Newcastle upon Tyne, UK, 2008. [Google Scholar]
- Kruskal, W.H.; Wallis, W.A. Use of Ranks in One-Criterion Variance Analysis. J. Am. Stat. Assoc. 1952, 47, 583–621. [Google Scholar] [CrossRef]
- Kalpić, D.; Hlupić, M.L.N. International Encyclopedia of Statistical Science; Springer: Berlin/Heidelberg, Germany, 2011; pp. 1559–1563. [Google Scholar]
- Dunn, O.J. Multiple Comparisons Using Rank Sums. Technometrics 1964, 6, 241–252. [Google Scholar] [CrossRef]
- Microsoft Support: STDEV.S Function. 2023. Available online: https://support.microsoft.com/en-us/office/stdev-s-function-7d69cf97-0c1f-4acf-be27-f3e83904cc23 (accessed on 1 May 2023).
- Müller, K.; Wickham, H. tibble: Simple Data Frames. 2022. R Package Version 3.1.7. Available online: https://CRAN.R-project.org/package=tibble (accessed on 1 May 2023).
- Fox, J.; Weisberg, S. An R Companion to Applied Regression, 3rd ed.; Sage: Thousand Oaks, CA, USA, 2019. [Google Scholar]
- Patil, I. Visualizations with Statistical Details: The ‘ggstatsplot’ approach. J. Open Source Softw. 2021, 6, 3167. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2022; Available online: https://www.R-project.org/ (accessed on 1 May 2023).
- Pohlert, T. PMCMRplus: Calculate Pairwise Multiple Comparisons of Mean Rank Sums Extended. 2022. R Package Version 1.9.5. Available online: https://CRAN.R-project.org/package=PMCMRplus (accessed on 1 May 2023).
| Time Step (Days) |
|---|
| 0 |
| 6.27 × |
| 0.000689664 |
| 0.00693381 |
| 0.014230415 |
| 0.031775873 |
| 0.051610754 |
| 0.088711557 |
| 0.135732573 |
| 0.212098681 |
| 0.311804414 |
| 0.474542448 |
| 0.722405853 |
| 1.163712693 |
| 2 |
| 4 |
| 5 |
| 10 |
| Dataset | # Omitted | Mean | Mean | Ratio |
|---|---|---|---|---|
| inflow ± 50 | 3 | −0.00092 | 0.014 | 0.063 |
| inflow ± 15 | none | N/A | N/A | N/A |
| rand params | 7 | −0.0076 | 0.015 | 0.50 |
| rand params const. P | 5 | −0.010 | 0.015 | 0.68 |
| Implementation | Average Time (s) | STDEV |
|---|---|---|
| Julia | 0.22 | 0.15 |
| SteadPy | 194 | 2 |
| DynPy | 59 | 4 |
| Java | 3.51 | 0.01 |
| Implementation | Average Time (s) | STDEV |
|---|---|---|
| Julia | 0.24 | 0.12 |
| SteadPy | 194 | 2 |
| DynPy | 56.5 | 0.9 |
| Java | 3.51 | 0.02 |
| Implementation | Average Time (s) | STDEV |
|---|---|---|
| Julia | 0.54 | 0.50 |
| SteadPy | 166 | 22 |
| DynPy | 96 | 11 |
| Java | 4.4 | 1.4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Allen, C.; Mazanko, A.; Abdehagh, N.; Eberl, H.J. A New ODE-Based Julia Implementation of the Anaerobic Digestion Model No. 1 Greatly Outperforms Existing DAE-Based Java and Python Implementations. Processes 2023, 11, 1899. https://doi.org/10.3390/pr11071899
Allen C, Mazanko A, Abdehagh N, Eberl HJ. A New ODE-Based Julia Implementation of the Anaerobic Digestion Model No. 1 Greatly Outperforms Existing DAE-Based Java and Python Implementations. Processes. 2023; 11(7):1899. https://doi.org/10.3390/pr11071899
Chicago/Turabian StyleAllen, Courtney, Alexandra Mazanko, Niloofar Abdehagh, and Hermann J. Eberl. 2023. "A New ODE-Based Julia Implementation of the Anaerobic Digestion Model No. 1 Greatly Outperforms Existing DAE-Based Java and Python Implementations" Processes 11, no. 7: 1899. https://doi.org/10.3390/pr11071899
APA StyleAllen, C., Mazanko, A., Abdehagh, N., & Eberl, H. J. (2023). A New ODE-Based Julia Implementation of the Anaerobic Digestion Model No. 1 Greatly Outperforms Existing DAE-Based Java and Python Implementations. Processes, 11(7), 1899. https://doi.org/10.3390/pr11071899










