GCCSwin-UNet: Global Context and Cross-Shaped Windows Vision Transformer Network for Polyp Segmentation
Abstract
1. Introduction
- (1)
- We build a symmetric encoder–decoder architecture with a skip-connection structure based on the CSwin Transformer. In the encoder, feature extraction is performed using a cross-shaped window self-attention mechanism with the aim of better extraction of feature information; in the decoder, a patch expansion layer is used to achieve upsampling and feature dimensionality increase without using convolution or interpolation operations to facilitate better polyp segmentation.
- (2)
- We design a global context module with the aim of capturing the feature information that is continuously lost during encoder downsampling and sequentially forwarding it to the corresponding decoder module, with a view to better weighing the global information.
- (3)
- We design a local position-enhanced module that operates on the channel dimension intending to enhance the segmentation of boundary regions by stepping important position information in the feature map.
- (4)
- To verify the segmentation performance of our GCCSwin-UNet, we conduct experiments on two public datasets. The results show that our proposed network not only performs best in polyp segmentation but also achieves state-of-the-art results in two public datasets.
2. Related Work
3. Method
3.1. Overall Architecture
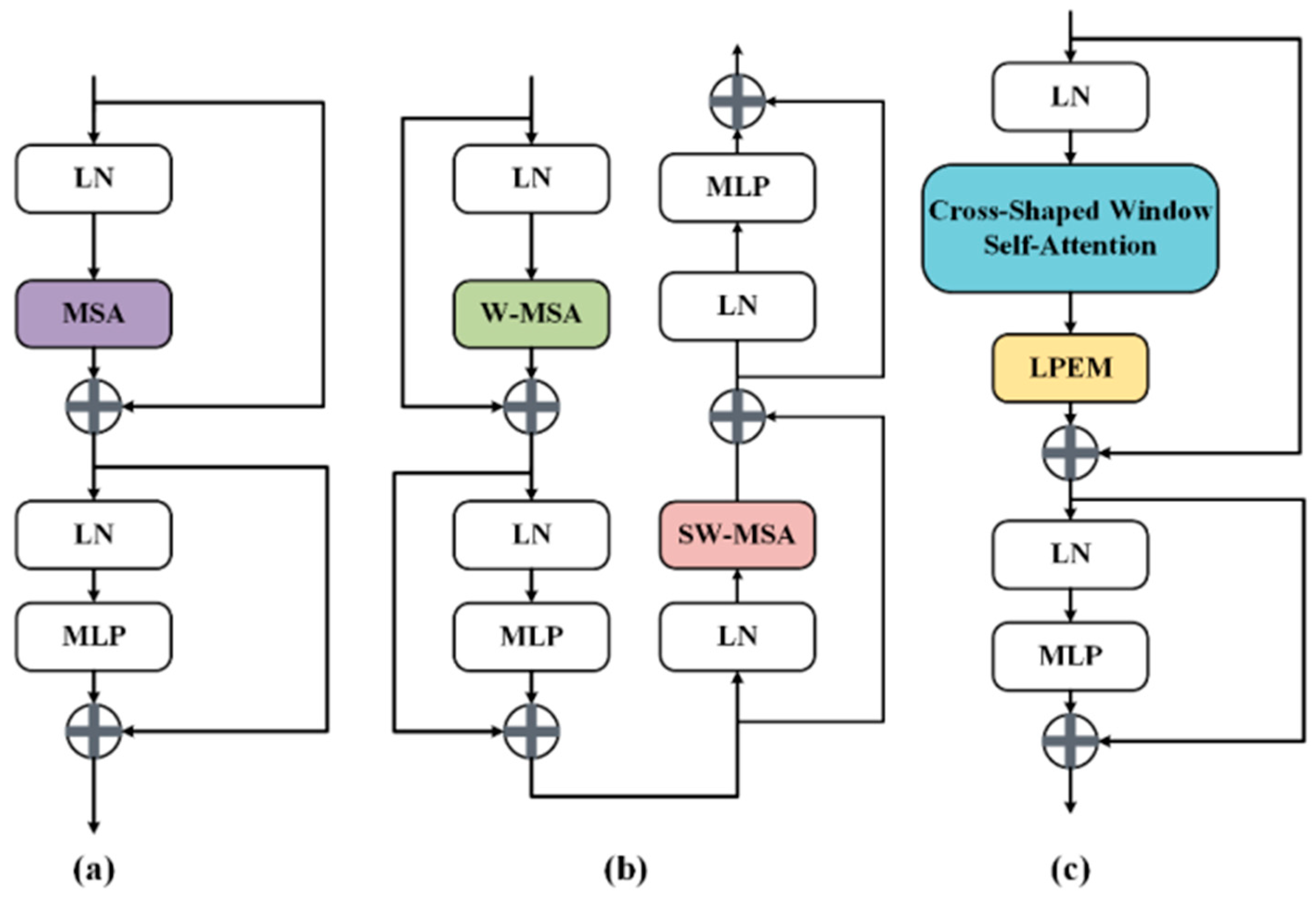
3.2. CSwin Transformer Block
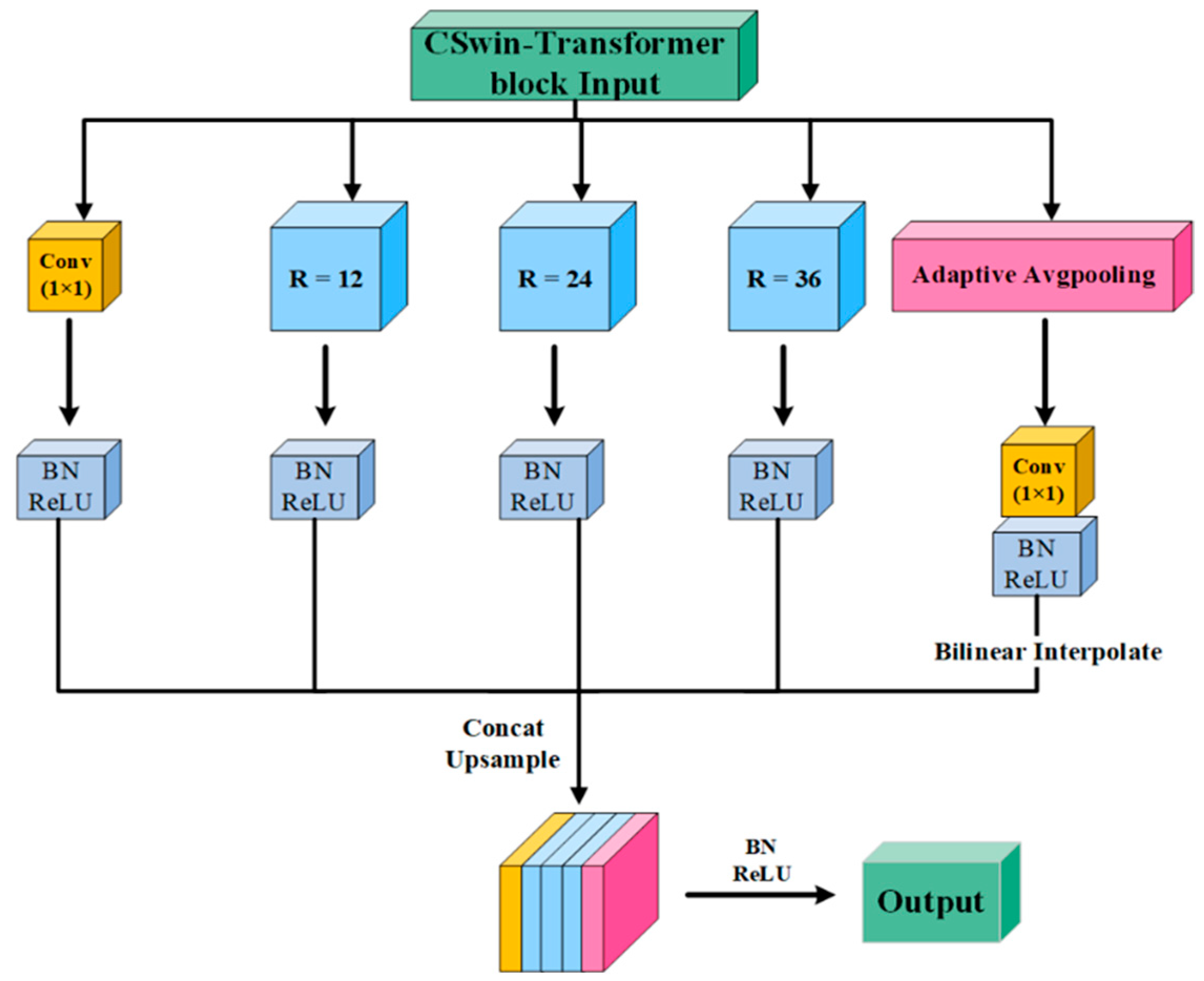
3.3. Global Context Module (GCM)
3.4. Local Position Enhanced Module (LPEM)
3.5. Mixed Loss Function
4. Experiment
4.1. Datasets
4.2. Evaluation Metrics
4.3. Training Strategies
4.4. Ablation Experiments
4.5. Comparative Experiments
- (a)
- Quantitative analysis
- (b)
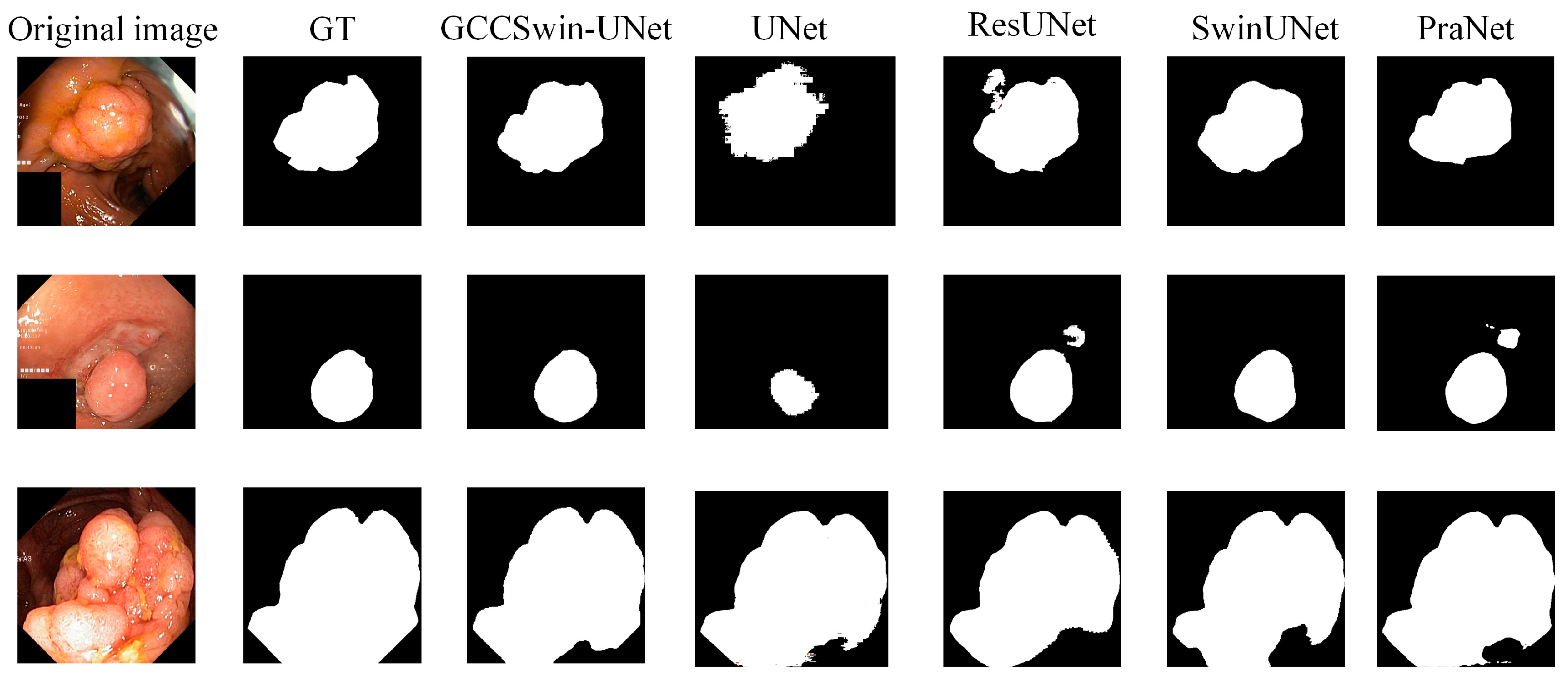
- Qualitative analysis
4.6. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Goding Sauer, A.; Fedewa, S.A.; Butterly, L.F.; Anderson, J.C.; Jemal, A. Colorectal cancer statistics, 2020. CA A Cancer J. Clin. 2020, 70, 145–164. [Google Scholar] [CrossRef] [PubMed]
- Barua, I.; Vinsard, D.G.; Jodal, H.C.; Løberg, M.; Kalager, M.; Holme, Ø.; Mori, Y. Artificial intelligence for polyp detection during colonoscopy: A systematic review and meta-analysis. Endoscopy 2021, 53, 277–284. [Google Scholar] [CrossRef] [PubMed]
- Ciardiello, F.; Ciardiello, D.; Martini, G.; Napolitano, S.; Tabernero, J.; Cervantes, A. Clinical management of metastatic colorectal cancer in the era of precision medicine. CA A Cancer J. Clin. 2022, 72, 372–401. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Pu, L.Z.C.T.; Liu, Y.; Maicas, G.; Verjans, J.W.; Burt, A.D.; Carneiro, G. Detecting, localising and classifying polyps from colonoscopy videos using deep learning. arXiv 2021, arXiv:2101.03285. [Google Scholar]
- Biller, L.H.; Schrag, D. Diagnosis and treatment of metastatic colorectal cancer: A review. JAMA 2021, 325, 669–685. [Google Scholar] [CrossRef]
- Jha, D.; Ali, S.; Tomar, N.K.; Johansen, H.D.; Johansen, D.; Rittscher, J.; Halvorsen, P. Real-time polyp detection, localization and segmentation in colonoscopy using deep learning. IEEE Access 2021, 9, 40496–40510. [Google Scholar] [CrossRef]
- Le, A.; Salifu, M.O.; McFarlane, I.M. Artificial Intelligence in Colorectal Polyp Detection and Characterization. Int. J. Clin. Res. Trials 2021, 6, 157. [Google Scholar] [CrossRef]
- Brown, J.R.G.; Mansour, N.M.; Wang, P.; Chuchuca, M.A.; Minchenberg, S.B.; Chandnani, M.; Berzin, T.M. Deep learning computer-aided polyp detection reduces adenoma miss rate: A United States multi-center randomized tandem colonoscopy study (CADeT-CS trial). Clin. Gastroenterol. Hepatol. 2022, 20, 1499–1507. [Google Scholar] [CrossRef]
- Turner, J.K.; Wright, M.; Morgan, M.; Williams, G.T.; Dolwani, S. A prospective study of the accuracy and concordance between in-situ and postfixation measurements of colorectal polyp size and their potential impact upon surveillance. Eur. J. Gastroenterol. Hepatol. 2013, 25, 562–567. [Google Scholar] [CrossRef]
- Zhang, R.; Li, G.; Li, Z.; Cui, S.; Qian, D.; Yu, Y. Adaptive context selection for polyp segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Lima, Peru, 4–8 October 2020; Springer: Cham, Switzerland, 2020. [Google Scholar]
- Fan, D.P.; Ji, G.P.; Zhou, T.; Chen, G.; Fu, H.; Shen, J.; Shao, L. Pranet: Parallel reverse attention network for polyp segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Lima, Peru, 4–8 October 2020; Springer: Cham, Switzerland, 2020. [Google Scholar]
- Dong, X.; Bao, J.; Chen, D.; Zhang, W.; Yu, N.; Yuan, L.; Guo, B. Cswin transformer: A general vision transformer backbone with cross-shaped windows. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, New Orleans, LA, USA, 19–20 June 2022. [Google Scholar]
- Hwang, S.; Oh, J.; Tavanapong, W.; Wong, J.; De Groen, P.C. Polyp detection in colonoscopy video using elliptical shape feature. In Proceedings of the 2007 IEEE International Conference on Image Processing, San Antonio, TX, USA, 16–19 September 2007; IEEE: Piscataway, NJ, USA, 2007; Volume 2. [Google Scholar]
- Gross, S.; Kennel, M.; Stehle, T.; Wulff, J.; Tischendorf, J.; Trautwein, C.; Aach, T. Polyp segmentation in NBI colonoscopy. In Bildverarbeitung für die Medizin 2009; Springer: Berlin/Heidelberg, Germany, 2009; pp. 252–256. [Google Scholar]
- Du, N.; Wang, X.; Guo, J.; Xu, M. Attraction propagation: A user-friendly interactive approach for polyp segmentation in colonoscopy images. PLoS ONE 2016, 11, e0155371. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015; Springer: Cham, Switzerland, 2015. [Google Scholar]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. Segnet: A deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Chen, C.; Yuan, Y.; Tong, K.Y. Selective feature aggregation network with area-boundary constraints for polyp segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Shenzhen, China, 13–17 October 2019; Springer: Cham, Switzerland, 2019. [Google Scholar]
- Zhang, Z.; Liu, Q.; Wang, Y. Road extraction by deep residual u-net. IEEE Geosci. Remote Sens. Lett. 2018, 15, 749–753. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Polosukhin, I. Attention is all you need. Adv. Neural Inf. Process. Syst. 2017, 30. [Google Scholar]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Houlsby, N. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv 2020, arXiv:2010.11929. [Google Scholar]
- Cao, H.; Wang, Y.; Chen, J.; Jiang, D.; Zhang, X.; Tian, Q.; Wang, M. Swin-unet: Unet-like pure transformer for medical image segmentation. arXiv 2021, arXiv:2105.05537. [Google Scholar]
- Valanarasu, J.M.J.; Oza, P.; Hacihaliloglu, I.; Patel, V.M. Medical transformer: Gated axial-attention for medical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Strasbourg, France, 27 September–1 October 2021; Springer: Cham, Switzerland, 2021. [Google Scholar]
- Gao, Y.; Zhou, M.; Metaxas, D.N. UTNet: A hybrid transformer architecture for medical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Strasbourg, France, 27 September–1 October 2021; Springer: Cham, Switzerland, 2021. [Google Scholar]
- Zhu, J.; Ge, M.; Chang, Z.; Dong, W. CRCNet: Global-local context and multi-modality cross attention for polyp segmentation. Biomed. Signal Process. Control 2023, 83, 104593. [Google Scholar] [CrossRef]
- Shaw, P.; Uszkoreit, J.; Vaswani, A. Self-attention with relative position representations. arXiv 2018, arXiv:1803.02155. [Google Scholar]
- Ho, J.; Kalchbrenner, N.; Weissenborn, D.; Salimans, T. Axial attention in multidimensional transformers. arXiv 2019, arXiv:1912.12180. [Google Scholar]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Guo, B. Swin transformer: Hierarchical vision transformer using shifted windows. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, BC, Canada, 11–17 October 2021. [Google Scholar]
- Chu, X.; Tian, Z.; Zhang, B.; Wang, X.; Wei, X.; Xia, H.; Shen, C. Conditional positional encodings for vision transformers. arXiv 2021, arXiv:2102.10882. [Google Scholar]
- Lin, T.Y.; Dollár, P.; Girshick, R.; He, K.; Hariharan, B.; Belongie, S. Feature pyramid networks for object detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017. [Google Scholar]
- Tolstikhin, I.O.; Houlsby, N.; Kolesnikov, A.; Beyer, L.; Zhai, X.; Unterthiner, T.; Dosovitskiy, A. Mlp-mixer: An all-mlp architecture for vision. Adv. Neural Inf. Process. Syst. 2021, 34, 24261–24272. [Google Scholar]
- Xu, J.; Sun, X.; Zhang, Z.; Zhao, G.; Lin, J. Understanding and improving layer normalization. Adv. Neural Inf. Process. Syst. 2019, 32, 4381–4391. [Google Scholar]
- Chen, L.C.; Papandreou, G.; Schroff, F.; Adam, H. Rethinking atrous convolution for semantic image segmentation. arXiv 2017, arXiv:1706.05587. [Google Scholar]
- Chen, L.C.; Zhu, Y.; Papandreou, G.; Schroff, F.; Adam, H. Encoder-decoder with atrous separable convolution for semantic image segmentation. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018. [Google Scholar]
- Jha, D.; Smedsrud, P.H.; Riegler, M.A.; Halvorsen, P.; de Lange, T.; Johansen, D.; Johansen, H.D. Kvasir-seg: A segmented polyp dataset. In Proceedings of the International Conference on Multimedia Modeling, Daejeon, Republic of Korea, 5–8 January 2020; Springer: Cham, Switzerland, 2020. [Google Scholar]
- Bernal, J.; Sánchez, F.J.; Fernández-Esparrach, G.; Gil, D.; Rodríguez, C.; Vilariño, F. WM-DOVA maps for accurate polyp highlighting in colonoscopy: Validation vs. saliency maps from physicians. Comput. Med. Imaging Graph. 2015, 43, 99–111. [Google Scholar] [CrossRef] [PubMed]
- Ji, G.P.; Chou, Y.C.; Fan, D.P.; Chen, G.; Fu, H.; Jha, D.; Shao, L. Progressively normalized self-attention network for video polyp segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Strasbourg, France, 27 September–1 October 2021; Springer: Cham, Switzerland, 2021. [Google Scholar]





| Setting | Dice% | MIoU% | Acc% |
|---|---|---|---|
| Baseline | 79.50 | 78.33 | 81.68 |
| Baseline + CSwin | 82.45 | 80.14 | 83.21 |
| Baseline + GCM | 80.78 | 78.92 | 82.56 |
| Baseline + LPEM | 81.39 | 79.27 | 82.75 |
| Baseline + CSwin + LPEM | 83.86 | 82.06 | 84.54 |
| Baseline + CSwin + GCM | 82.93 | 81.28 | 84.32 |
| Baseline + GCM + LPEM + CSwin | 86.37 | 83.19 | 85.94 |
| Dataset | Method | Dice% | MIoU% | Acc% |
|---|---|---|---|---|
| Kvasir-SEG | U-Net [15] | 81.80 | 74.60 | 82.17 |
| Residual U-Net [19] | 79.10 | 76.38 | 73.12 | |
| SFANet [18] | 72.35 | 61.15 | —— | |
| PraNet [11] | 83.80 | 81.20 | 86.14 | |
| PNS-Net [37] | 84.00 | 79.50 | 83.56 | |
| Swin-Unet [22] | 82.31 | 81.62 | 85.61 | |
| GCCSwin-UNet(ours) | 86.37 | 83.19 | 85.94 | |
| CVC-ClinicDB | U-Net [15] | 87.62 | 75.50 | 87.36 |
| Residual U-Net [19] | 86.73 | 76.17 | 87.48 | |
| SFANet [18] | 70.05 | 60.75 | —— | |
| PraNet [11] | 89.82 | 83.20 | 91.72 | |
| PNS-Net [37] | 87.30 | 80.00 | 90.39 | |
| Swin-Unet [22] | 88.96 | 80.71 | 91.57 | |
| GCCSwin-UNet(ours) | 91.26 | 84.65 | 92.13 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, J.; Ge, M.; Chang, Z.; Dong, W. GCCSwin-UNet: Global Context and Cross-Shaped Windows Vision Transformer Network for Polyp Segmentation. Processes 2023, 11, 1035. https://doi.org/10.3390/pr11041035
Zhu J, Ge M, Chang Z, Dong W. GCCSwin-UNet: Global Context and Cross-Shaped Windows Vision Transformer Network for Polyp Segmentation. Processes. 2023; 11(4):1035. https://doi.org/10.3390/pr11041035
Chicago/Turabian StyleZhu, Jianbo, Mingfeng Ge, Zhimin Chang, and Wenfei Dong. 2023. "GCCSwin-UNet: Global Context and Cross-Shaped Windows Vision Transformer Network for Polyp Segmentation" Processes 11, no. 4: 1035. https://doi.org/10.3390/pr11041035
APA StyleZhu, J., Ge, M., Chang, Z., & Dong, W. (2023). GCCSwin-UNet: Global Context and Cross-Shaped Windows Vision Transformer Network for Polyp Segmentation. Processes, 11(4), 1035. https://doi.org/10.3390/pr11041035






