Influence of Optimal Hyperparameters on the Performance of Machine Learning Algorithms for Predicting Heart Disease
Abstract
1. Introduction
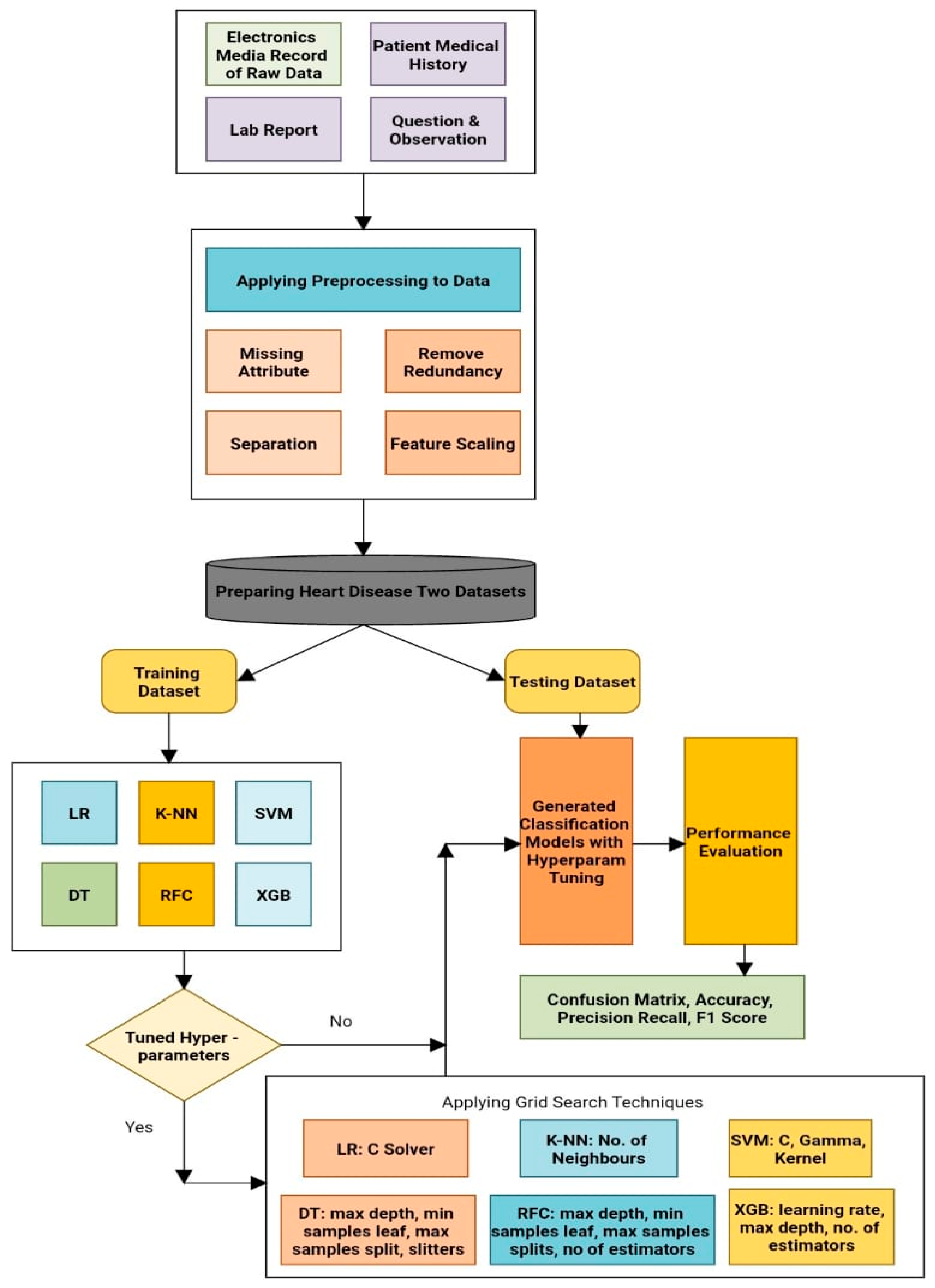
- In the first step, common ML algorithms for predicting heart disease were used, as follows: logistic regression, K-nearest neighbor, support vector machine, decision tree, random forest classifier, and extreme gradient boosting.
- In the second step, a prediction system using six fundamental ML algorithms and a hyperparameter tuning technique was provided. Here, grid search cross-validation was also used to identify the appropriate hyperparameters for each method. The notations used to describe the analytical results of different tables are as follows: accuracy (A), precision (P), recall (R), F-1 score (F-1s), and support (S).
- Finally, a confusion matrix was used to compare the performances of the models for these two systems.
2. Literature Review
2.1. Existing Models for Predicting Cardiovascular Diseases
2.2. Methodology for Detecting Heart Disease
2.3. Levels of Heart Disease
3. Research Methodology
3.1. The Objective of the Study
3.2. Description of the Datasets
3.3. Suggested Model
3.4. Problem Statement for the Study
3.5. Hyperparameter Tuning Optimization
4. Evaluation Metrics and Experimental Data Analysis
4.1. Metrics for Evaluation
4.2. Description of the Features of Heart Disease Datasets
4.3. Experimental Data Analysis
5. Discussion and Analysis of the Experiment Results
5.1. Data Preparation
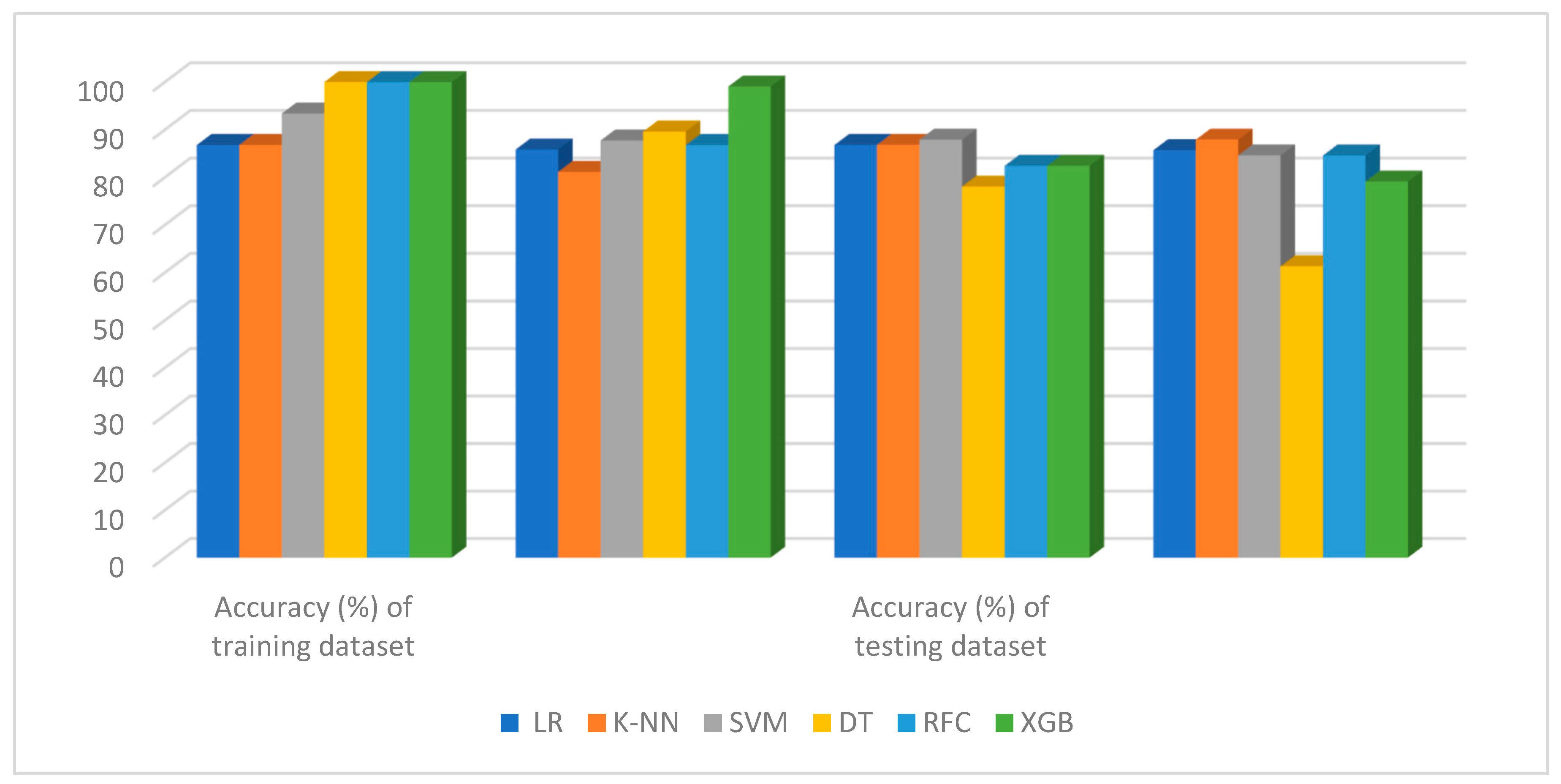
5.2. Performance Evaluation and Comparison with a Traditional System for Dataset-I
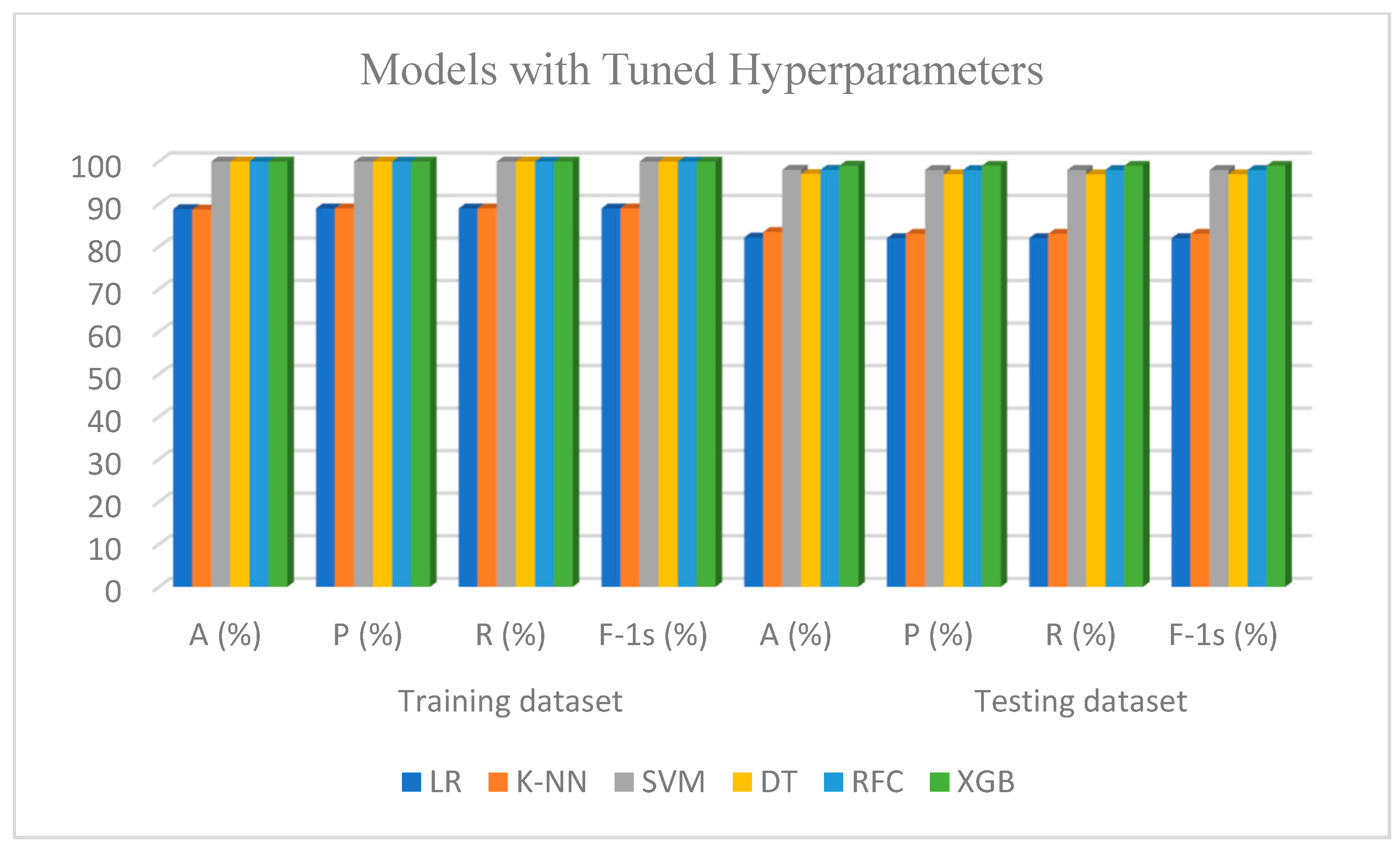
5.3. Performance Evaluation with Tuned Hyperparameters for Dataset-I
5.4. Performance Evaluation and Comparison with the Traditional System for Dataset-II
5.5. Performance Evaluation with Tuned Hyperparameters for Dataset-II
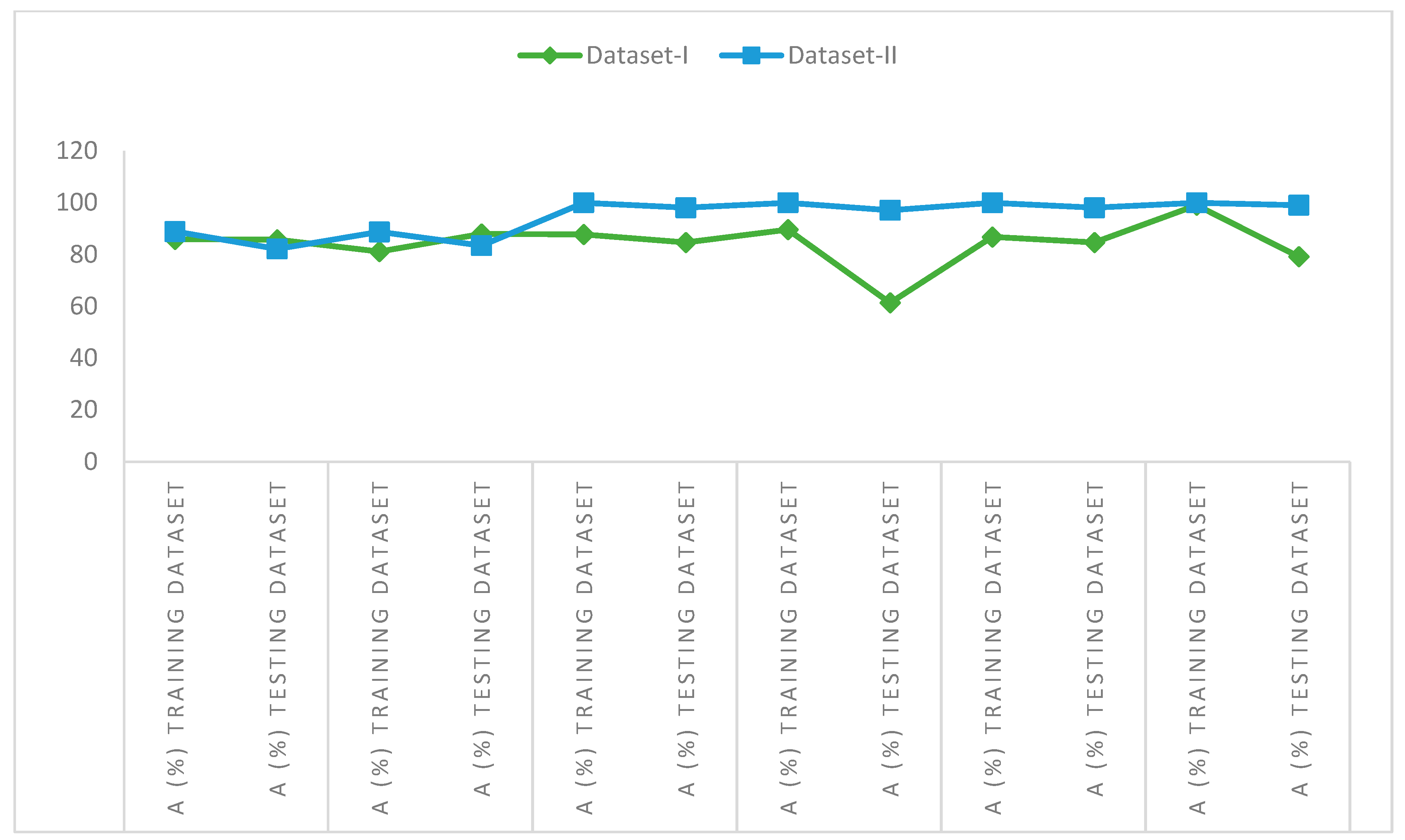
5.6. Comparison of the Performance between Dataset-I and Dataset-II
5.7. Comparison with Previous Research
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A. The Dataset-I Detailed Results
| Traditional Method | Training Dataset | Testing Dataset | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Models | Parameters | A (%) | P (%) | R (%) | F-1s (%) | A (%) | P (%) | R (%) | F-1s (%) |
| LR | Solver = liblinear | 86.79 | 87 | 87 | 87 | 86.91 | 87 | 87 | 87 |
| K-NN | K = 5, weights = uniform | 86.79 | 87 | 87 | 87 | 86.81 | 87 | 87 | 87 |
| SVM | Kernel = “rbf”, gamma = 0.1, C = 1.0 | 93.40 | 93 | 93 | 93 | 87.91 | 88 | 88 | 88 |
| DT | Random_state = 42 | 100 | 100 | 100 | 100 | 78.02 | 78 | 78 | 78 |
| RFC | n_estimators = 1000, random_state = 42 | 100 | 100 | 100 | 100 | 82.42 | 82 | 82 | 82 |
| XGB | Label_encoder = false | 100 | 100 | 100 | 100 | 82.42 | 82 | 82 | 82 |
| Models with Hyper Parameters | Training Dataset | Testing Dataset | |||||||
|---|---|---|---|---|---|---|---|---|---|
| MLAs | Tunned Hyperparameters | A (%) | P (%) | R (%) | F-1s (%) | A (%) | P (%) | R (%) | F-1s (%) |
| LR | Solver = liblinear, c = 0.234 | 85.85 | 86 | 86 | 86 | 85.71 | 86 | 86 | 86 |
| K-NN | K = 27, | 81.13 | 81 | 81 | 81 | 87.91 | 88 | 88 | 88 |
| SVM | Kernel = rbf, gamma = 0.1, C = 5 | 87.74 | 88 | 88 | 88 | 84.62 | 85 | 85 | 85 |
| DT | Criterion = entropy, max_depth = 5, min_samples_leaf = 2, splitter = 2 | 89.62 | 90 | 90 | 90 | 81.32 | 81 | 81 | 81 |
| RFC | Max_depth = 2, max_features = auto, min_samples_leaf = 1, n_estimators = 1100 | 86.79 | 87 | 87 | 86 | 84.62 | 85 | 85 | 85 |
| XGB | Learning_rate = 0.6427, max_depth = 3, n_estimators = 3 | 99.06 | 99 | 99 | 99 | 79.12 | 79 | 79 | 79 |
| ML Classifiers | Accuracy (%) of Training Dataset | Accuracy (%) of Testing Dataset | ||
|---|---|---|---|---|
| Without Parameter Tuning | With Hyperparameter Tuning | Without Parameter Tuning | With Hyperparameter Tuning | |
| LR | 86.79 | 85.85 | 86.81 | 85.71 |
| K-NN | 86.79 | 81.13 | 86.81 | 87.91 |
| SVM | 93.40 | 87.74 | 87.91 | 84.62 |
| DT | 100.00 | 89.62 | 78.06 | 61.32 |
| RFC | 100.00 | 86.79 | 82.42 | 84.62 |
| XGB | 100.00 | 99.06 | 82.42 | 79.12 |
Appendix B. The Dataset-II Detailed Results
| Models with Tuned Hyperparameters | Training Dataset | Testing Dataset | |||||||
|---|---|---|---|---|---|---|---|---|---|
| A (%) | P (%) | R (%) | F-1s (%) | A (%) | P (%) | R (%) | F-1s (%) | ||
| LR | Solver = liblinear, c = 0.088 | 88.84 | 89 | 89 | 89 | 82.14 | 82 | 82 | 82 |
| K-NN | K = 27, | 88.70 | 89 | 89 | 89 | 83.44 | 83 | 83 | 83 |
| SVM | Kernel = rbf, gamma = 0.5, C = 2 | 100 | 100 | 100 | 100 | 98.05 | 98 | 98 | 98 |
| DT | Criterion = entropy, max_depth = 11, min_samples_leaf = 1, splitter = best, min_samples_split = 2 | 100 | 100 | 100 | 100 | 97.08 | 97 | 97 | 97 |
| RFC | Max_depth = 15, max_features = auto, min_samples_leaf = 1, min_samples_split = 2, n_estimators = 500 | 100 | 100 | 100 | 100 | 98.05 | 98 | 98 | 98 |
| XGB | Learning_rate = 0.547, max_depth = 5, n_estimators = 338 | 100 | 100 | 100 | 100 | 99.03 | 99 | 99 | 99 |
| MLAs | Accuracy (%) Training Dataset | Accuracy (%) Testing Dataset | ||
|---|---|---|---|---|
| Without Parameter Tuning | With Hyperparameter Tuning | Without Parameter Tuning | With Hyperparameter Tuning | |
| LR | 89.54 | 88.84 | 81.82 | 82.14 |
| K-NN | 91.77 | 88.70 | 81.82 | 83.44 |
| SVM | 95.40 | 100 | 90.26 | 98.05 |
| DT | 100.00 | 100 | 97.08 | 97.08 |
| RFC | 100.00 | 100 | 98.05 | 98.05 |
| XGB | 100.00 | 100 | 99.03 | 99.03 |
Appendix C. Comparisons of Two Datasets and Previous Studies Details
| Models with Hyperparameter | Dataset-I | Dataset-II | |
|---|---|---|---|
| LR | A (%) training dataset | 85.85 | 88.84 |
| A (%) testing dataset | 85.71 | 82.14 | |
| K-NN | A (%) training dataset | 81.13 | 88.70 |
| A (%) testing dataset | 87.91 | 83.44 | |
| SVM | A (%) training dataset | 87.74 | 100 |
| A (%) testing dataset | 84.62 | 98.05 | |
| DT | A (%) training dataset | 89.62 | 100 |
| A (%) testing dataset | 61.32 | 97.08 | |
| RFC | A (%) training dataset | 86.79 | 100 |
| A (%) testing dataset | 84.62 | 98.05 | |
| XGB | A (%) training dataset | 99.06 | 100 |
| A (%) testing dataset | 79.12 | 99.03 | |
| Previous Studies | A (%) | P (%) | R (%) | F-1s (%) |
|---|---|---|---|---|
| Alizadehsani et al. [38] | 93.85 | 00 | 97 | 00 |
| Arora et al. [39] | 77.4 | 00 | 77.4 | 0 |
| Lakshmanna et al. [40] | 90 | 00 | 00 | 91 |
| Chiam et al. [41] | 78.15 | 78.15 | 00 | 80.25 |
| Shijani et al. [42] | 91.14 | 91.90 | 93 | 00 |
| Senan et al. [43] | 95 | 97.62 | 95.35 | 96.47 |
| Suggested model dataset-I training | 99.06 | 99 | 99 | 99 |
| Suggested model dataset-II training | 100 | 100 | 100 | 100 |
| Suggested model dataset-I testing | 79.12 | 79 | 79 | 79 |
| Suggested model dataset-II testing | 99.03 | 99 | 99 | 99 |
References
- Animesh, H.; Subrata, K.M.; Amit, G.; Arkomita, M.; Mukherje, A. Heart Disease Diagnosis and Prediction Using Machine Learning and Data Mining Techniques: A Review. Adv. Comput. Sci. Technol. 2017, 10, 2137–2159. [Google Scholar]
- Buttar, H.S.; Li, T.; Ravi, N. Prevention of CVD: Role of exercise, dietary interventions, obesity and smoking cessation. Exp. Clin. Cardiol. 2005, 10, 229–249. [Google Scholar]
- Ahmad, G.N.; Ullah, S.; Algethami, A.; Fatima, H.; Akhter, S.M.H. Comparative Study of Optimum Medical Diagnosis of Human Heart Disease Using ML Technique with and without Sequential Feature Selection. IEEE Access 2022, 10, 23808–23828. [Google Scholar] [CrossRef]
- Nagamani, T.; Logeswari, S.; Gomathy, B. Heart Disease Prediction using Data Mining with Mapreduce Algorithm. Int. J. Innov. Technol. Explor. Eng. (IJITEE) 2019, 8, 137–140. [Google Scholar]
- Nikhar, S.; Karandikar, A.M. Prediction of heart disease using machine learning algorithms. Int. J. Adv. Eng. Manag. Sci. 2016, 2, 617–621. [Google Scholar]
- Franco, D.; Estefanía, L.V. Healing the Broken Hearts: A Glimpse on Next Generation Therapeutics. Hearts 2022, 3, 96–116. [Google Scholar] [CrossRef]
- Gayathri, R.; Rani, S.U.; Čepová, L.; Rajesh, M.; Kalita, K. A Comparative Analysis of Machine Learning Models in Prediction of Mortar Compressive Strength. Processes 2022, 10, 1387. [Google Scholar] [CrossRef]
- Brites, I.S.G.; da Silva, L.M.; Barbosa, J.L.V.; Rigo, S.J.; Correia, S.D.; Leithardt, V.R.Q. Machine Learning and IoT Applied to Cardiovascular Diseases Identification through Heart Sounds: A Literature Review. Informatics 2021, 8, 73. [Google Scholar] [CrossRef]
- Reddy, K.V.V.; Elamvazuthi, I.; Aziz, A.A.; Paramasivam, S.; Chua, H.N.; Pranavanand, S. An Efficient Prediction System for Coronary Heart Disease Risk Using Selected Principal Components and Hyperparameter Optimization. Appl. Sci. 2023, 13, 118. [Google Scholar] [CrossRef]
- Obaido, G.; Ogbuokiri, B.; Swart, T.G.; Ayawei, N.; Kasongo, S.M.; Aruleba, K.; Mienye, I.D.; Aruleba, I.; Chukwu, W.; Osaye, F.; et al. An interpretable machine learning approach for hepatitis b diagnosis. Appl. Sci. 2022, 12, 11127. [Google Scholar] [CrossRef]
- UCI Machine Learning Repository: Heart Disease Dataset. Available online: https://www.kaggle.com/johnsmith88/heart-disease-dataset (accessed on 20 October 2022).
- Detrano, R.; Janosi, A.; Steinbrunn, W.; Pfisterer, M.; Schmid, J.; Sandhu, S.; Guppy, K.; Lee, S.; Froelicher, V. International application of a new probability algorithm for the diagnosis of coronary artery disease. Am. J. Cardiol. 1989, 64, 304–310. [Google Scholar] [CrossRef] [PubMed]
- Ebiaredoh-Mienye, S.A.; Swart, T.G.; Esenogho, E.; Mienye, I.D. A machine learning method with filter-based feature selection for improved prediction of chronic kidney disease. Bioengineering 2022, 9, 350. [Google Scholar] [CrossRef] [PubMed]
- Mienye, I.D.; Sun, Y.; Wang, Z. An improved ensemble learning approach for the prediction of heart disease risk. Inform. Med. Unlocked 2020, 20, 100402. [Google Scholar] [CrossRef]
- Polat, K.; Güneş, S. A hybrid approach to medical decision support systems: Combining feature selection, fuzzy weighted pre-processing and AIRS. Comput. Methods Programs Biomed. 2007, 88, 164–174. [Google Scholar] [CrossRef]
- Alizadehsani, R.; Hosseini, M.J.; Khosravi, A.; Khozeimeh, F.; Roshanzamir, M.; Sarrafzadegan, N.; Nahavandi, S. Non-invasive detection of coronary artery disease in high-risk patients based on the stenosis prediction of separate coronary arteries. Comput. Methods Programs Biomed. 2018, 162, 119–127. [Google Scholar] [CrossRef]
- Pham, H.; Olafsson, S. Bagged ensembles with tunable parameters. Comput. Intell. 2019, 35, 184–203. [Google Scholar] [CrossRef]
- Magesh, G.; Swarnalatha, P. Optimal feature selection through a cluster-based DT learning (CDTL) in heart disease prediction. Evol. Intell. 2021, 14, 583–593. [Google Scholar] [CrossRef]
- Wang, H.; Wang, S. Medical Knowledge Acquisition through Data Mining. In Proceedings of the IEEE International Symposium on IT in Medicine and Education, Xiamen, China, 12–14 December 2008; pp. 777–780. [Google Scholar] [CrossRef]
- Singh, R.; Rajesh, E. Prediction of Heart Disease by Clustering and Classification Techniques. Int. J. Comput. Sci. Eng. 2019, 7, 861–866. [Google Scholar] [CrossRef]
- Bharti, R.; Khamparia, A.; Shabaz, M.; Dhiman, G.; Pande, S.; Singh, P. Prediction of Heart Disease Using a Combination of Machine Learning and Deep Learning. Comput. Intell. Neurosci. 2021, 2021, 8387680. [Google Scholar] [CrossRef]
- Manikandan, S. Heart Attack Prediction System. In Proceedings of the International Conference on Energy, Communication, Data Analytics & Soft Computing, Chennai, Tamil Nadu, 1–2 August 2017; pp. 817–820. [Google Scholar]
- Garg, R.; Oh, E.; Naidech, A.; Kording, K.; Prabhakaran, S. Automating ischemic stroke subtype classification using machine learning and natural language processing. J. Stroke Cerebrovasc. Dis. 2019, 28, 2045–2051. [Google Scholar] [CrossRef]
- Chourasia, V.; Pal, S. Data Mining Approach to Detect HDs. Inter. J. Adv. Comput. Sci. Inf. Technol. (IJACSIT) 2013, 2, 56–66. [Google Scholar]
- Palaniappan, S.; Awang, R. Intelligent heart disease prediction system using data mining techniques. In Proceedings of the IEEE/ACS International Conference on Computer Systems and Applications, Doha, Qatar, 31 March–4 April 2008; pp. 108–115. [Google Scholar] [CrossRef]
- Pedretti, R.F.E.; Hansen, D.; Ambrosetti, M.; Back, M.; Berger, T.; Ferreira, M.C.; Cornelissen, V.; Davos, C.H.; Doehner, W.; Zarzosa, C.D.P.Y.; et al. How to optimize the adherence to a guideline-directed medical therapy in the secondary prevention of cardiovascular diseases: A clinical consensus statement from the European Association of Preventive Cardiology. Eur. J. Prev. Cardiol. 2022, 30, 149–166. [Google Scholar] [CrossRef]
- Chowdhury, M.T.H. Application of Signal Processing and Deep Hybrid Learning in Phonocardiogram and Electrocardiogram Signals to Detect Early-Stage Heart Diseases. Ph.D. Thesis. Faculty of the Computational Science Program Middle Tennessee State University: Murfreesboro, TN, USA, 2022. [Google Scholar]
- Nadakinamani, R.G.; Reyana, A.; Kautish, S.; Vibith, A.S.; Gupta, Y.; Abdelwahab, S.F.; Mohamed, A.W. Clinical Data Analysis for Prediction of Cardiovascular Disease Using Machine Learning Techniques. Comput. Intell. Neurosci. 2022, 2022, 2973324. [Google Scholar] [CrossRef]
- Fitriyani, N.L.; Syafrudin, M.; Alfian, G.; Rhee, J. HDPM: An Effective Heart Disease Prediction Model for a Clinical Decision Support System. IEEE Access 2020, 8, 133034–133050. [Google Scholar] [CrossRef]
- Ali, Y.A.; Awwad, E.M.; Al-Razgan, M.; Maarouf, A. Hyperparameter Search for Machine Learning Algorithms for Optimizing the Computational Complexity. Processes 2023, 11, 349. [Google Scholar] [CrossRef]
- Ambrish, G.; Ganesh, B.; Ganesh, A.; Srinivas, C.; Mensinkal, K. Logistic Regression Technique for Prediction of Cardiovascular Disease. Glob. Transit. Proc. 2022, 4, 127–130. [Google Scholar]
- Zhang, C.; Zhong, P.; Liu, M.; Song, Q.; Liang, Z.; Wang, X. Hybrid Metric K-Nearest Neighbor Algorithm and Applications. Math. Probl. Eng. 2022, 2022, 8212546. [Google Scholar] [CrossRef]
- Xue, T.; Jieru, Z. Application of Support Vector Machine Based on Particle Swarm Optimization in Classification and Prediction of Heart Disease. In Proceedings of the IEEE 7th Inter. Conference on Intelligent Computing and Signal Processing (ICSP), Virtual, 15–17 April 2022; pp. 857–860. [Google Scholar]
- Vijaya Saraswathi, R.; Gajavelly, K.; Kousar Nikath, A.; Vasavi, R.; Reddy Anumasula, R. Heart Disease Prediction Using Decision Tree and SVM. In Proceedings of the Second International Conference on Advances in Computer Engineering and Communication Systems, Singapore, 11–12 August 2022; pp. 69–78. [Google Scholar]
- Liu, Y.; Wang, Y.; Zhang. New machine learning algorithm: Random Forest. In Proceedings of the International Conference on Information Computing and Applications, Chengde, China, 14–16 September 2012; Springer: Berlin/Heidelberg, Germany, 2012; pp. 246–252. [Google Scholar]
- Budholiya, K.; Shrivastava, S.K.; Sharma, V. An optimized XGBoost based diagnostic system for effective prediction of heart disease. J. King Saud Univ. Comput. Inf. Sci. 2020, 34, 4514–4523. [Google Scholar] [CrossRef]
- Jeng, M.-Y.; Yeh, T.-M.; Pai, F.-Y. A Performance Evaluation Matrix for Measuring the Life Satisfaction of Older Adults Using eHealth Wearables. Healthcare 2022, 10, 605. [Google Scholar] [CrossRef]
- Arabasadi, Z.; Alizadehsani, R.; Roshanzamir, M.; Moosaei, H.; Yarifard, A.A. Computer aided decision making for heart disease detection using hybrid neural network-Genetic algorithm. Comput. Methods Programs Biomed. 2017, 141, 19–26. [Google Scholar] [CrossRef]
- Arora, S.; Maji, S. Decision tree algorithms for prediction of heart disease. In Proceedings of the Information and Communication Technology for Competitive Strategies: Proceedings of Third International Conference on ICTCS 2017; Springer: Singapore, 2019; pp. 447–454. [Google Scholar]
- Lakshmanna, K.; Reddy, G.T.; Reddy, M.P.; Rajput, D.S.; Kaluri, R.; Srivastava, G. Hybrid genetic algorithm and a fuzzy logic classifier for heart disease diagnosis. Evol. Intell. 2020, 13, 185–196. [Google Scholar]
- Chiam, Y.K.; Amin, M.S.; Varathan, K.D. Identification of significant features and data mining techniques in predicting heart disease. Telemat. Inform. 2019, 36, 82–93. [Google Scholar]
- Feshki, M.G.; Shijani, O.S. Improving the heart disease diagnosis by evolutionary algorithm of PSO and Feed Forward Neural Network. In Proceedings of the 2016 Artificial Intelligence and Robotics (IRANOPEN), Qazvin, Iran, 9 April 2016; pp. 48–53. [Google Scholar]
- Senan, E.M.; Abunadi, I.; Jadhav, M.E.; Fati, S.M. Score and Correlation Coefficient-Based Feature Selection for Predicting Heart Failure Diagnosis by Using Machine Learning Algorithms. Comput. Math. Methods Med. 2021, 2021, 8500314. [Google Scholar] [CrossRef] [PubMed]









| Classes | Features | Instances | |
|---|---|---|---|
| Dataset-I | 2 | 14 | 303 |
| Dataset-II | 2 | 14 | 1025 |
| Age | Sex | cp | trestbps | chol | fbs | restecg | thalach | exang | oldpeak | slope | ca | thal | target | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Mean | 54.47 | 0.69 | 0.97 | 131.62 | 247.00 | 0.16 | 0.54 | 148.99 | 0.38 | 1.05 | 1.50 | 0.74 | 2.32 | 0.55 |
| Std | 9.09 | 0.48 | 1.04 | 17.55 | 52.77 | 0.37 | 0.54 | 22.57 | 0.48 | 1.17 | 0.63 | 1.03 | 0.62 | 0.51 |
| Min | 29.00 | 000 | 0.00 | 93 | 126 | 0.00 | 0.00 | 71 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 |
| 25% | 47.00 | 0.00 | 0.00 | 120 | 211 | 0.00 | 0.00 | 133.50 | 0.00 | 0.00 | 1.00 | 0.00 | 2.00 | 0.00 |
| 50% | 50 | 1.00 | 1.00 | 130 | 240 | 0.00 | 1.00 | 153 | 0.00 | 8.00 | 1.00 | 0.00 | 2.00 | 1.00 |
| 75% | 61 | 1.00 | 2.00 | 140 | 274 | 0.00 | 1.00 | 166 | 1.00 | 1.60 | 2.00 | 1.00 | 3.00 | 1.00 |
| Max | 77 | 1.00 | 3.00 | 200 | 564 | 1.00 | 2.00 | 202 | 1.00 | 6.20 | 2.00 | 4.00 | 3.00 | 1.00 |
| Age | Sex | cp | trestbps | chol | fbs | restecg | thalach | exang | oldpeak | slope | ca | thal | target | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Mean | 54.43 | 0.70 | 0.94 | 131.61 | 245.06 | 0.15 | 0.53 | 149.11 | 0.34 | 1.07 | 1.39 | 0.75 | 2.32 | 0.51 |
| Std | 9.07 | 0.46 | 1.03 | 17.52 | 51.59 | 0.36 | 0.53 | 23.01 | 0.47 | 1.18 | 0.62 | 1.03 | 0.62 | 0.50 |
| Min | 29.00 | 0.00 | 0.00 | 94 | 126 | 0.00 | 0.00 | 71 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 | 0.00 |
| 25% | 48.00 | 0.00 | 0.00 | 120 | 211 | 0.00 | 0.00 | 132 | 0.00 | 0.00 | 1.00 | 0.00 | 2.00 | 0.00 |
| 50% | 56 | 1.00 | 1.00 | 130 | 240 | 0.00 | 1.00 | 152 | 0.00 | 8.00 | 1.00 | 0.00 | 2.00 | 1.00 |
| 75% | 61 | 1.00 | 2.00 | 140 | 275 | 0.00 | 1.00 | 166 | 1.00 | 1.80 | 2.00 | 1.00 | 3.00 | 1.00 |
| Max | 77 | 1.00 | 3.00 | 200 | 564 | 1.00 | 2.00 | 202 | 1.00 | 6.20 | 2.00 | 4.00 | 3.00 | 1.00 |
| Traditional Models | Normal (0) | Abnormal (1) | A (%) | Macro Avg (%) | Weighted Avg (%) | ||
|---|---|---|---|---|---|---|---|
| P (%) | 88 | 86 | 87 | 87 | 87 | ||
| Training result | R (%) | 82 | 90 | 87 | 86 | 87 | |
| LR | F-1s (%) | 85 | 88 | 87 | 87 | 87 | |
| S (%) | 97 | 115 | 87 | 212 | 212 | ||
| P (%) | 87 | 87 | 87 | 87 | 87 | ||
| Testing result | R (%) | 83 | 90 | 86 | 86 | 87 | |
| F-1s (%) | 85 | 88 | 87 | 87 | 87 | ||
| S (%) | 41 | 50 | 87 | 91 | 91 | ||
| P (%) | 86 | 87 | 87 | 87 | 87 | ||
| Training result | R (%) | 85 | 89 | 87 | 87 | 87 | |
| F-1s (%) | 85 | 88 | 87 | 87 | 87 | ||
| K-NN | S (%) | 97 | 115 | 87 | 212 | 212 | |
| P (%) | 85 | 88 | 87 | 87 | 87 | ||
| Testing result | R (%) | 85 | 88 | 87 | 87 | 87 | |
| F-1s (%) | 85 | 88 | 87 | 87 | 87 | ||
| S (%) | 41 | 50 | 87 | 91 | 91 | ||
| P (%) | 94 | 93 | 93 | 93 | 93 | ||
| Training result | R (%) | 92 | 95 | 93 | 93 | 93 | |
| SVM | F-1s (%) | 93 | 94 | 93 | 93 | 93 | |
| S (%) | 97 | 115 | 93 | 212 | 212 | ||
| P (%) | 86 | 90 | 88 | 88 | 88 | ||
| Testing result | R (%) | 88 | 88 | 88 | 88 | 88 | |
| F-1s (%) | 87 | 90 | 88 | 88 | 88 | ||
| S (%) | 41 | 50 | 88 | 91 | 91 | ||
| P (%) | 100 | 100 | 100 | 100 | 100 | ||
| Training result | R (%) | 100 | 100 | 100 | 100 | 100 | |
| F-1s (%) | 100 | 100 | 100 | 100 | 100 | ||
| DT | S (%) | 97 | 115 | 100 | 212 | 212 | |
| P (%) | 73 | 84 | 78 | 88 | 79 | ||
| Testing result | R (%) | 83 | 74 | 78 | 88 | 78 | |
| F-1s (%) | 77 | 79 | 78 | 88 | 78 | ||
| S (%) | 41 | 50 | 78 | 91 | 91 | ||
| P (%) | 100 | 100 | 100 | 100 | 100 | ||
| Training result | R (%) | 100 | 100 | 100 | 100 | 100 | |
| F-1s (%) | 100 | 100 | 100 | 100 | 100 | ||
| RFC | S (%) | 97 | 115 | 100 | 212 | 212 | |
| P (%) | 80 | 84 | 82 | 82 | 82 | ||
| Testing result | R (%) | 80 | 84 | 82 | 82 | 82 | |
| F-1s (%) | 80 | 84 | 82 | 82 | 82 | ||
| S (%) | 41 | 50 | 82 | 91 | 91 | ||
| P (%) | 100 | 100 | 100 | 100 | 100 | ||
| Training result | R (%) | 00 | 100 | 100 | 100 | 100 | |
| XGB | F-1s (%) | 100 | 100 | 100 | 100 | 100 | |
| S (%) | 97 | 115 | 100 | 212 | 212 | ||
| P (%) | 80 | 84 | 82 | 82 | 82 | ||
| Testing result | R (%) | 80 | 84 | 82 | 82 | 82 | |
| F-1s (%) | 80 | 84 | 82 | 82 | 82 | ||
| S (%) | 41 | 50 | 82 | 91 | 91 | ||
| Training Dataset | Testing Dataset | |||||
|---|---|---|---|---|---|---|
| Models | Confusion Matrix | Type-I Error | Type-II Error | Confusion Matrix | Type-I Error | Type-II Error |
| LR | [[80 17] [11 104]] | 184 (correct) | 28 (incorrect) | [[34 7] [5 45]] | 79 (correct) | 12 (incorrect) |
| KNN | [[82 15] [13 102]] | 184 (correct) | 28 (incorrect) | [[35 6] [6 44]] | 79 (correct) | 12 (incorrect) |
| SVM | [[89 8] [6 109]] | 198 (correct) | 14 (incorrect) | [[36 5] [6 44]] | 80 (correct) | 11 (incorrect) |
| DT | [[97 0] [0 115]] | 212 (correct) | 0 (incorrect) | [[34 7] [13 37]] | 71 (correct) | 20 (incorrect) |
| RFC | [[97 0] [0 115]] | 212 (correct) | 0 (incorrect) | [[33 8] [8 42]] | 75 (correct) | 16 (incorrect) |
| XGB | [[97 0] [0 115]] | 212 (correct) | 0 (incorrect) | [[33 8] [8 42]] | 75 (correct) | 16 (incorrect) |
| Hyperparameter Tuning Classification Report | Normal (0) | Abnormal (1) | A (%) | Macro Avg | Weighted Avg | ||
|---|---|---|---|---|---|---|---|
| P (%) | 86 | 86 | 86 | 86 | 86 | ||
| Training result | R (%) | 82 | 89 | 86 | 86 | 86 | |
| Tuned LR | F-1s (%) | 84 | 87 | 86 | 86 | 86 | |
| S (%) | 97.00 | 115 | 86 | 212 | 212 | ||
| P (%) | 85 | 86 | 86 | 86 | 86 | ||
| Testing result | R (%) | 83 | 88 | 86 | 85 | 86 | |
| F-1s (%) | 84 | 87 | 86 | 86 | 86 | ||
| S (%) | 41 | 50 | 86 | 91 | 91 | ||
| P (%) | 84 | 80 | 81 | 82 | 81 | ||
| Training result | R (%) | 83 | 88 | 81 | 81 | 81 | |
| F-1s (%) | 78 | 83 | 81 | 81 | 81 | ||
| Tuned K-NN | S (%) | 97 | 115 | 81 | 212 | 212 | |
| P (%) | 89 | 87 | 88 | 88 | 88 | ||
| Testing result | R (%) | 83 | 92 | 88 | 87 | 88 | |
| F-1s (%) | 86 | 89 | 88 | 88 | 88 | ||
| S (%) | 41 | 50 | 88 | 91 | 91 | ||
| P (%) | 88 | 87 | 88 | 88 | 88 | ||
| Training result | R (%) | 85 | 89 | 88 | 87 | 88 | |
| Tuned SVM | F-1s (%) | 86 | 90 | 88 | 88 | 88 | |
| S (%) | 97 | 115 | 88 | 212 | 212 | ||
| P (%) | 85 | 85 | 85 | 85 | 85 | ||
| Testing result | R (%) | 80 | 88 | 85 | 84 | 85 | |
| F-1s (%) | 83 | 86 | 85 | 84 | 85 | ||
| S (%) | 41 | 50 | 85 | 91 | 91 | ||
| P (%) | 94 | 87 | 90 | 90 | 90 | ||
| Training result | R (%) | 82 | 96 | 90 | 89 | 90 | |
| F-1s (%) | 88 | 91 | 90 | 89 | 90 | ||
| Tuned DT | S (%) | 97 | 115 | 90 | 212 | 212 | |
| P (%) | 80 | 82 | 81 | 81 | 81 | ||
| Testing result | R (%) | 78 | 84 | 81 | 81 | 81 | |
| F-1s (%) | 79 | 83 | 81 | 81 | 81 | ||
| S (%) | 41 | 50 | 81 | 91 | 91 | ||
| P (%) | 89 | 85 | 87 | 87 | 87 | ||
| Training result | R (%) | 81 | 91 | 87 | 86 | 87 | |
| F-1s (%) | 85 | 88 | 86 | 87 | 87 | ||
| Tuned RFC | S (%) | 97 | 115 | 87 | 212 | 212 | |
| P (%) | 85 | 85 | 85 | 85 | 85 | ||
| Testing result | R (%) | 80 | 88 | 85 | 84 | 85 | |
| F-1s (%) | 83 | 86 | 85 | 84 | 85 | ||
| S (%) | 41 | 50 | 85 | 91 | 91 | ||
| P (%) | 100 | 98 | 99 | 99 | 99 | ||
| Training result | R (%) | 98 | 100 | 99 | 99 | 99 | |
| Tuned XGB | F-1s (%) | 99 | 99 | 99 | 99 | 99 | |
| S (%) | 97 | 115 | 99 | 212 | 212 | ||
| P (%) | 76 | 82 | 79 | 79 | 79 | ||
| Testing result | R (%) | 78 | 80 | 79 | 79 | 79 | |
| F-1s (%) | 77 | 81 | 79 | 79 | 79 | ||
| S (%) | 41 | 50 | 79 | 91 | 91 | ||
| Training Dataset | Testing Dataset | |||||
|---|---|---|---|---|---|---|
| Models | Confusion Matrix | Type-I Error | Type-II Error | Confusion Matrix | Type-I Error | Type-II Error |
| TLR | [[80 17] [13 102]] | 182 (correct) | 30 (incorrect) | [[34 7] [6 44]] | 78 (correct) | 13 (incorrect) |
| TK-NN | [[71 26] [14 102]] | 173 (correct) | 40 (incorrect) | [[34 7] [4 46]] | 80 (correct) | 11(incorrect) |
| TSVM | [[82 15] [11 106]] | 188 (correct) | 26 (incorrect) | [[33 8] [6 44]] | 77 (correct) | 14 (incorrect) |
| TDT | [[80 17] [5 110]] | 190 (correct) | 22 (incorrect) | [[32 9] [8 42]] | 74 (correct) | 17 (incorrect) |
| TRFC | [[79 18] [10 105]] | 184 (correct) | 28 (incorrect) | [[33 8] [6 44]] | 77 (correct) | 14 (incorrect) |
| TXGB | [[95 2] [0 115]] | 210 (correct) | 2 (incorrect) | [[32 9] [10 40]] | 72 (correct) | 19 (incorrect) |
| Traditional Method | Training Dataset | Testing Dataset | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Models | Parameters | A (%) | P (%) | R (%) | F-1s (%) | A (%) | P (%) | R (%) | F-1s (%) |
| LR | Solver = liblinear | 89.54 | 90 | 90 | 90 | 81.82 | 82 | 82 | 82 |
| K-NN | K = 5, weights = uniform | 91.77 | 92 | 92 | 92 | 81.82 | 82 | 82 | 82 |
| SVM | Kernel = “rbf”, gamma = 0.1, C = 1.0 | 95.40 | 95 | 95 | 95 | 90.26 | 90 | 90 | 90 |
| DT | Random_state = 42 | 100.00 | 100 | 100 | 100 | 97.08 | 97 | 97 | 97 |
| RFC | n_estimators = 1000, random_state = 42 | 100.00 | 100 | 100 | 100 | 98.05 | 98 | 98 | 98 |
| XGB | Label_encoder = false | 100.00 | 100 | 100 | 100 | 99.03 | 99 | 99 | 99 |
| Traditional Models | Normal (0) | Abnormal (1) | A (%) | Macro Avg | Weighted Avg | ||
|---|---|---|---|---|---|---|---|
| P (%) | 91 | 89 | 90 | 90 | 90 | ||
| Training result | R (%) | 87 | 92 | 90 | 89 | 90 | |
| LR | F-1s (%) | 89 | 90 | 90 | 89 | 90 | |
| S (%) | 340 | 377 | 90 | 717 | 717 | ||
| P (%) | 85 | 89 | 82 | 82 | 82 | ||
| Testing result | R (%) | 89 | 85 | 82 | 82 | 82 | |
| F-1s (%) | 82 | 82 | 82 | 82 | 82 | ||
| S (%) | 159 | 149 | 0.82 | 308 | 308 | ||
| P (%) | 91 | 92 | 92 | 92 | 92 | ||
| Training result | R (%) | 91 | 92 | 92 | 92 | 92 | |
| F-1s (%) | 91 | 92 | 92 | 92 | 92 | ||
| K-NN | S (%) | 340 | 377 | 92 | 717 | 717 | |
| P (%) | 86 | 78 | 82 | 82 | 82 | ||
| Testing result | R (%) | 87 | 87 | 82 | 82 | 82 | |
| F-1s (%) | 81 | 82 | 82 | 82 | 82 | ||
| S (%) | 159 | 149 | 82 | 308 | 308 | ||
| P (%) | 97 | 94 | 95 | 96 | 95 | ||
| Training result | R (%) | 93 | 97 | 95 | 95 | 95 | |
| SVM | F-1s (%) | 95 | 96 | 95 | 95 | 95 | |
| S (%) | 340 | 377 | 95 | 717 | 717 | ||
| P (%) | 94 | 87 | 90 | 90 | 91 | ||
| Testing result | R (%) | 86 | 95 | 90 | 90 | 90 | |
| F-1s (%) | 90 | 99 | 90 | 90 | 90 | ||
| S (%) | 159 | 149 | 90 | 308 | 308 | ||
| P (%) | 100 | 100 | 100 | 100 | 100 | ||
| Training result | R (%) | 100 | 100 | 100 | 100 | 100 | |
| F-1s (%) | 100 | 100 | 100 | 100 | 100 | ||
| DT | S (%) | 340 | 377 | 100 | 717 | 717 | |
| P (%) | 95 | 100 | 97 | 88 | 97 | ||
| Testing result | R (%) | 100 | 94 | 97 | 88 | 97 | |
| F-1s (%) | 97 | 97 | 97 | 88 | 97 | ||
| S (%) | 159 | 149 | 0.97 | 308 | 308 | ||
| P (%) | 100 | 100 | 100 | 100 | 100 | ||
| Training result | R (%) | 100 | 100 | 100 | 100 | 100 | |
| F-1s (%) | 100 | 100 | 100 | 100 | 100 | ||
| RFC | S (%) | 340 | 377 | 100 | 717 | 717 | |
| P (%) | 96 | 100 | 98 | 98 | 98 | ||
| Testing result | R (%) | 100 | 96 | 98 | 98 | 98 | |
| F-1s (%) | 98 | 98 | 98 | 98 | 98 | ||
| S (%) | 159 | 149 | 98 | 308 | 308 | ||
| P (%) | 100 | 100 | 100 | 100 | 100 | ||
| Training result | R (%) | 100 | 100 | 100 | 100 | 100 | |
| XGB | F-1s (%) | 100 | 100 | 100 | 100 | 100 | |
| S (%) | 340 | 377 | 100 | 717 | 717 | ||
| P (%) | 96 | 100 | 98 | 98 | 98 | ||
| Testing result | R (%) | 100 | 96 | 98 | 98 | 98 | |
| F-1s (%) | 98 | 98 | 98 | 98 | 98 | ||
| S (%) | 159 | 149 | 98 | 308 | 308 | ||
| Training Dataset | Testing Dataset | |||||
|---|---|---|---|---|---|---|
| Models | Confusion Matrix | Type-I Error | Type-II Error | Confusion Matrix | Type-I Error | Type-II Error |
| LR | [[295 45] [30 347]] | 642 (correct) | 75 (incorrect) | [[125 34] [22 127]] | 252 (correct) | 56 (incorrect) |
| KNN | [[310 30] [29 348]] | 658 (correct) | 59 (incorrect) | [[123 36] [20 129]] | 252 (correct) | 56 (incorrect) |
| SVM | [[317 23] [10 367]] | 784 (correct) | 33 (incorrect) | [[137 22] [8 141]] | 278 (correct) | 30 (incorrect) |
| DT | [[340 0] [0 377]] | 717 (correct) | 0 (incorrect) | [[159 0] [9 143]] | 302 (correct) | 9 (incorrect) |
| RFC | [[340 0] [0 377]] | 717 (correct) | 0 (incorrect) | [[159 0] [6 143]] | 302 (correct) | 6 (incorrect) |
| XGB | [[340 0] [0 377]] | 717 (correct) | 0 (incorrect) | [[159 0] [6 143]] | 302 (correct) | 6 (incorrect) |
| Hyper Parameter Tuning Classification Report | Normal (0) | Abnormal (1) | Macro Avg | Weighted Avg | A (%) | ||
|---|---|---|---|---|---|---|---|
| P (%) | 89 | 89 | 89 | 89 | 89 | ||
| Training result | R (%) | 87 | 90 | 89 | 89 | 89 | |
| Tuned LR | F-1s (%) | 88 | 90 | 89 | 89 | 89 | |
| S (%) | 340 | 377 | 717 | 717 | 89 | ||
| P (%) | 86 | 79 | 83 | 86 | 82 | ||
| Testing result | R (%) | 78 | 87 | 82 | 86 | 82 | |
| F-1s (%) | 82 | 82 | 82 | 86 | 82 | ||
| S (%) | 159 | 149 | 308 | 308 | 82 | ||
| P (%) | 90 | 88 | 89 | 89 | 89 | ||
| Training result | R (%) | 86 | 92 | 89 | 89 | 89 | |
| F-1s (%) | 88 | 89 | 89 | 89 | 89 | ||
| Tuned K-NN | S (%) | 340 | 377 | 717 | 717 | 89 | |
| P (%) | 90 | 78 | 84 | 84 | 83 | ||
| Testing result | R (%) | 77 | 91 | 84 | 83 | 83 | |
| F-1s (%) | 83 | 84 | 83 | 83 | 83 | ||
| S (%) | 159 | 149 | 308 | 308 | 83 | ||
| P (%) | 100 | 100 | 100 | 100 | 100 | ||
| Training result | R (%) | 100 | 100 | 100 | 100 | 100 | |
| Tuned SVM | F-1s (%) | 100 | 100 | 100 | 100 | 100 | |
| S (%) | 340 | 377 | 717 | 717 | 100 | ||
| P (%) | 96 | 100 | 98 | 98 | 98 | ||
| Testing result | R (%) | 100 | 96 | 98 | 98 | 98 | |
| F-1s (%) | 98 | 98 | 98 | 98 | 98 | ||
| S (%) | 159 | 149 | 308 | 308 | 98 | ||
| P (%) | 100 | 100 | 100 | 100 | 100 | ||
| Training result | R (%) | 100 | 100 | 100 | 100 | 100 | |
| F-1s (%) | 100 | 100 | 100 | 100 | 100 | ||
| Tuned DT | S (%) | 340 | 377 | 717 | 717 | 100 | |
| P (%) | 95 | 100 | 97 | 97 | 97 | ||
| Testing result | R (%) | 100 | 94 | 97 | 97 | 97 | |
| F-1s (%) | 97 | 97 | 97 | 97 | 97 | ||
| S (%) | 159 | 149 | 717 | 717 | 97 | ||
| P (%) | 100 | 100 | 100 | 100 | 100 | ||
| Training result | R (%) | 100 | 100 | 100 | 100 | 100 | |
| F-1s (%) | 100 | 100 | 100 | 100 | 100 | ||
| Tuned RFC | S (%) | 340 | 377 | 717 | 717 | 100 | |
| P (%) | 96 | 100 | 98 | 98 | 98 | ||
| Testing result | R (%) | 100 | 96 | 98 | 98 | 98 | |
| F-1s (%) | 98 | 98 | 98 | 98 | 98 | ||
| S (%) | 159 | 149 | 308 | 308 | 98 | ||
| P (%) | 100 | 100 | 100 | 100 | 100 | ||
| Training result | R (%) | 100 | 100 | 100 | 100 | 100 | |
| Tuned XGB | F-1s (%) | 100 | 100 | 100 | 100 | 100 | |
| S (%) | 340 | 377 | 717 | 717 | 100 | ||
| P (%) | 98 | 82 | 99 | 99 | 99 | ||
| Testing result | R (%) | 100 | 80 | 99 | 99 | 99 | |
| F-1s (%) | 99 | 81 | 99 | 99 | 99 | ||
| S (%) | 159 | 149 | 308 | 308 | 99 | ||
| Training Dataset | Testing Dataset | |||||
|---|---|---|---|---|---|---|
| Models | Confusion Matrix | Type-I Error (Correct) | Type-II Error (Incorrect) | Confusion Matrix | Type-I Error (Correct) | Type-II Error (Incorrect) |
| TLR | [[296 44] [36 341]] | 637 | 80 | [[124 35] [20 129]] | 253 | 55 |
| TKNN | [[291 49] [32 345]] | 636 | 81 | [[122 37] [14 135]] | 257 | 51 |
| TSVM | [[340 0] [0 377]] | 717 | 0 | [[159 0] [6 143]] | 302 | 6 |
| TDT | [[340 0] [0 377]] | 717 | 0 | [[159 0] [9 143]] | 302 | 9 |
| TRFC | [[340 0] [6 143]] | 483 | 6 | [[159 0] [6 143]] | 302 | 6 |
| TXGB | [[340 0] [0 377]] | 717 | 0 | [[159 0] [3 146]] | 305 | 3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ahamad, G.N.; Shafiullah; Fatima, H.; Imdadullah; Zakariya, S.M.; Abbas, M.; Alqahtani, M.S.; Usman, M. Influence of Optimal Hyperparameters on the Performance of Machine Learning Algorithms for Predicting Heart Disease. Processes 2023, 11, 734. https://doi.org/10.3390/pr11030734
Ahamad GN, Shafiullah, Fatima H, Imdadullah, Zakariya SM, Abbas M, Alqahtani MS, Usman M. Influence of Optimal Hyperparameters on the Performance of Machine Learning Algorithms for Predicting Heart Disease. Processes. 2023; 11(3):734. https://doi.org/10.3390/pr11030734
Chicago/Turabian StyleAhamad, Ghulab Nabi, Shafiullah, Hira Fatima, Imdadullah, S. M. Zakariya, Mohamed Abbas, Mohammed S. Alqahtani, and Mohammed Usman. 2023. "Influence of Optimal Hyperparameters on the Performance of Machine Learning Algorithms for Predicting Heart Disease" Processes 11, no. 3: 734. https://doi.org/10.3390/pr11030734
APA StyleAhamad, G. N., Shafiullah, Fatima, H., Imdadullah, Zakariya, S. M., Abbas, M., Alqahtani, M. S., & Usman, M. (2023). Influence of Optimal Hyperparameters on the Performance of Machine Learning Algorithms for Predicting Heart Disease. Processes, 11(3), 734. https://doi.org/10.3390/pr11030734







