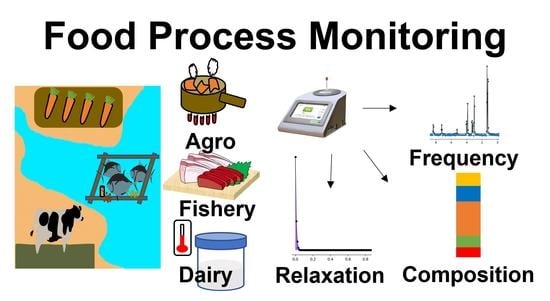
Parameter Visualization of Benchtop Nuclear Magnetic Resonance Spectra toward Food Process Monitoring
Abstract
:1. Introduction
2. Materials and Methods
2.1. NMR Measurements
2.2. Data Processing and Parameter Visualization
2.3. Fitting Method
2.4. Identification of Components in Food Mixtures
2.5. Removal of Water-Derived Spectra
2.6. Transfer Simulation Method of NMR Data from High-Field to Low-Field NMR
3. Results and Discussion
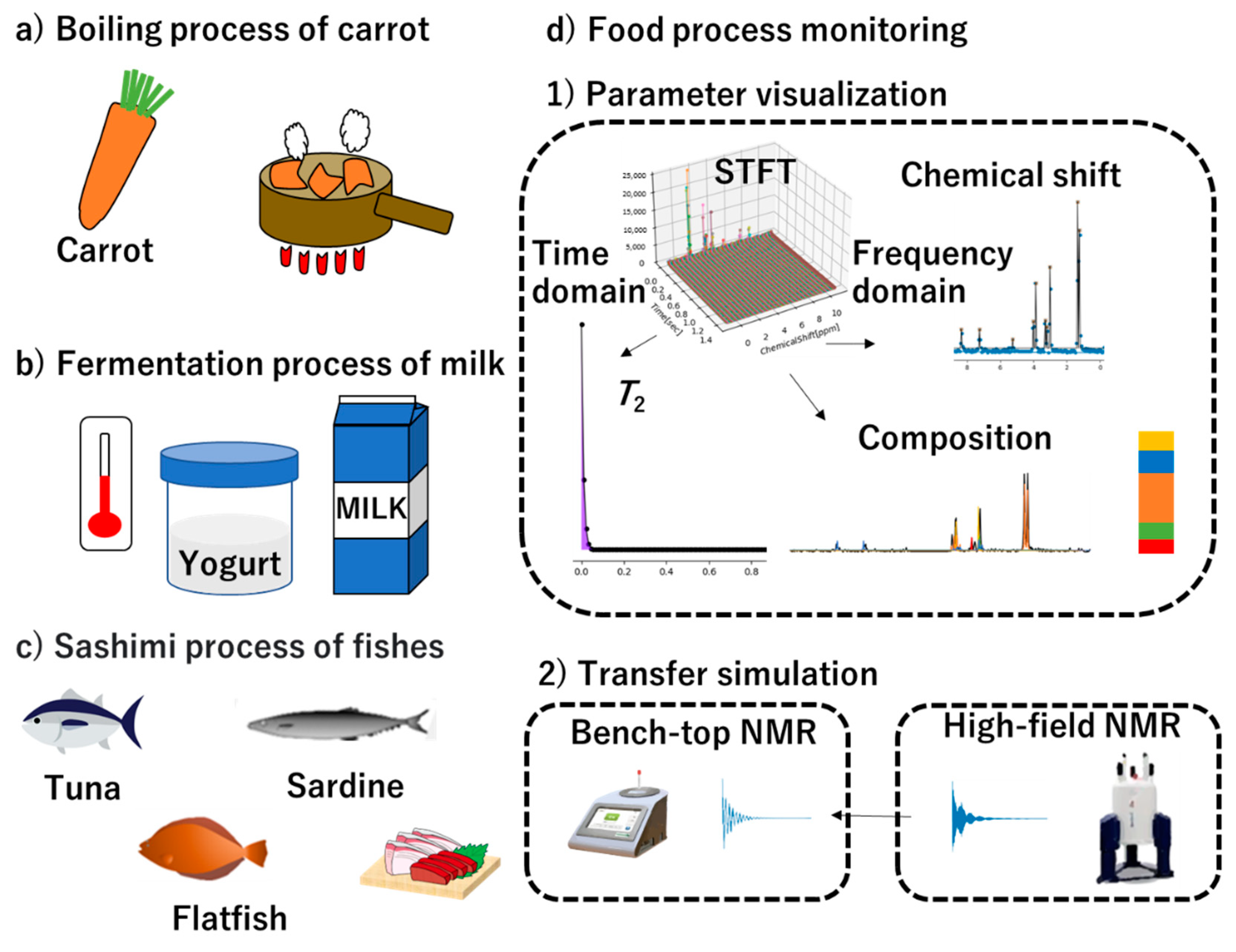
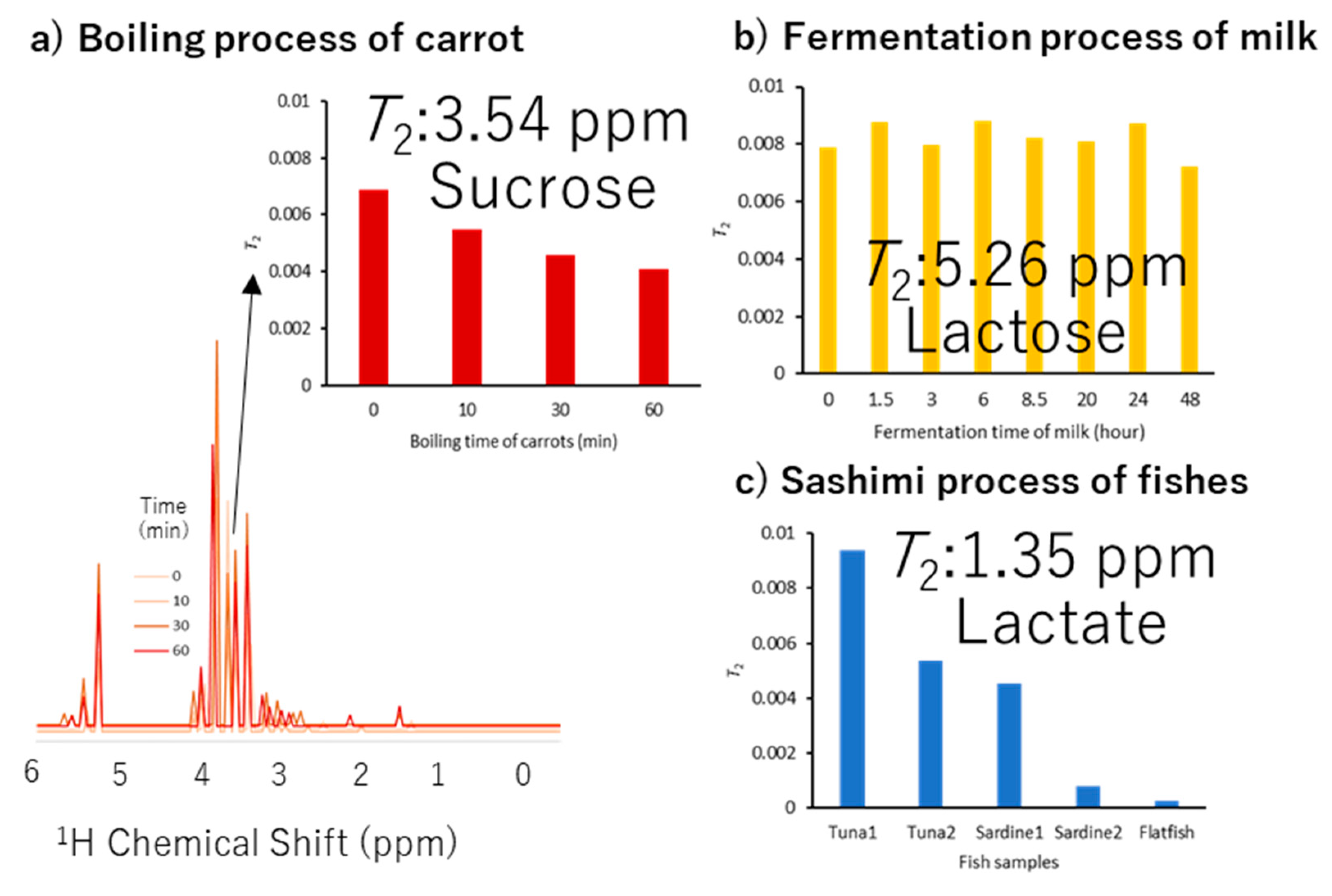
3.1. Parameter Visualization and Fitting
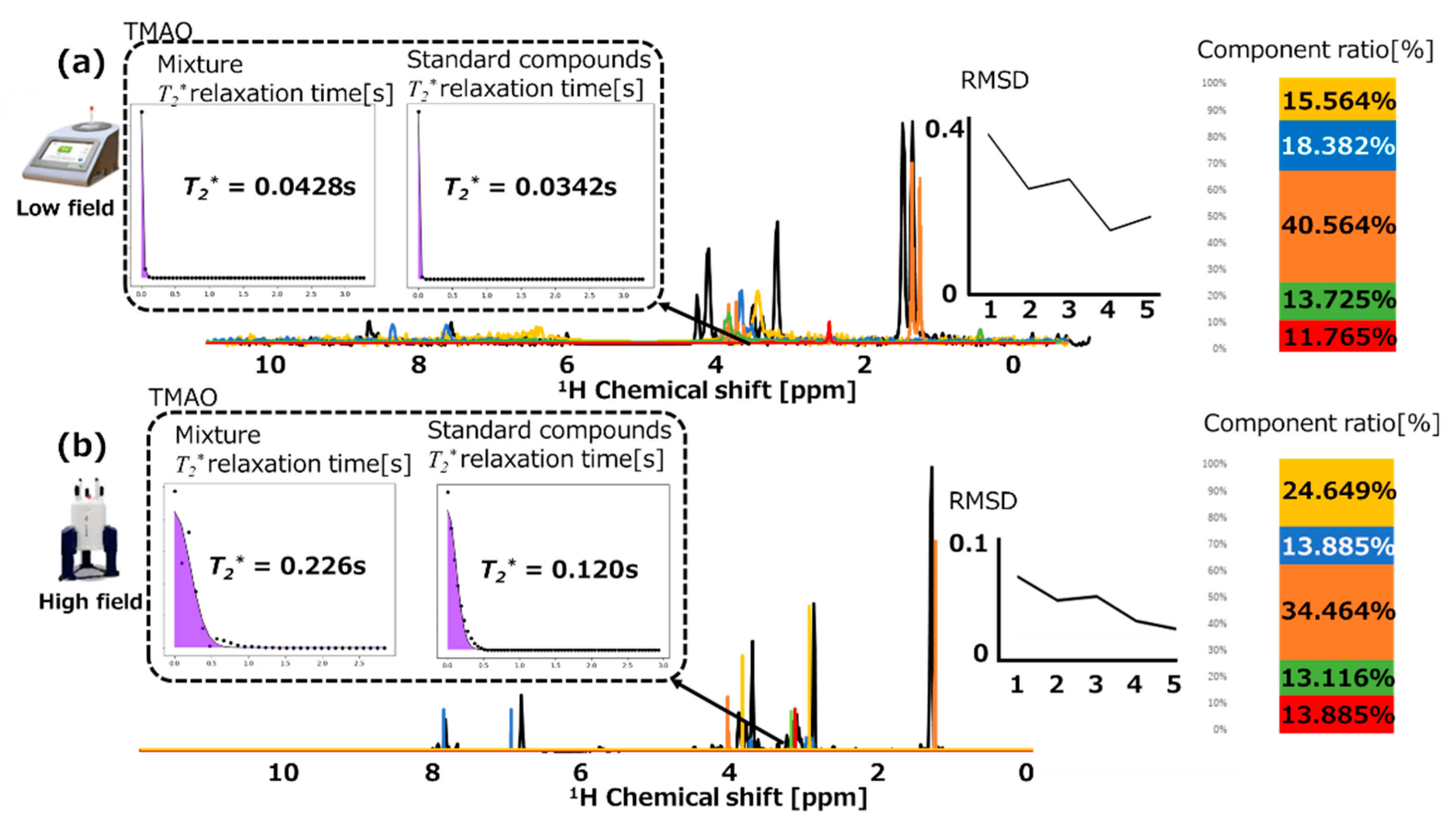
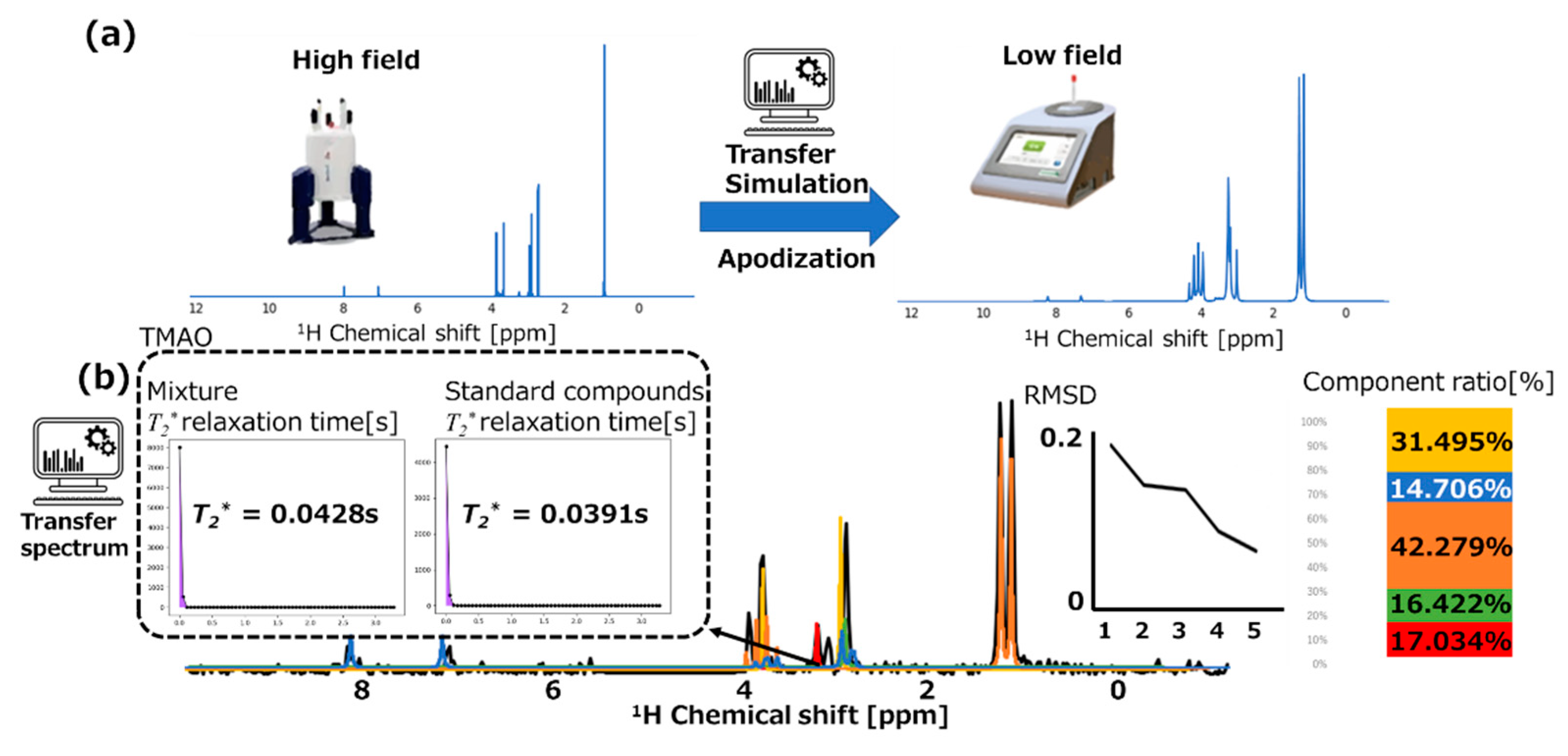
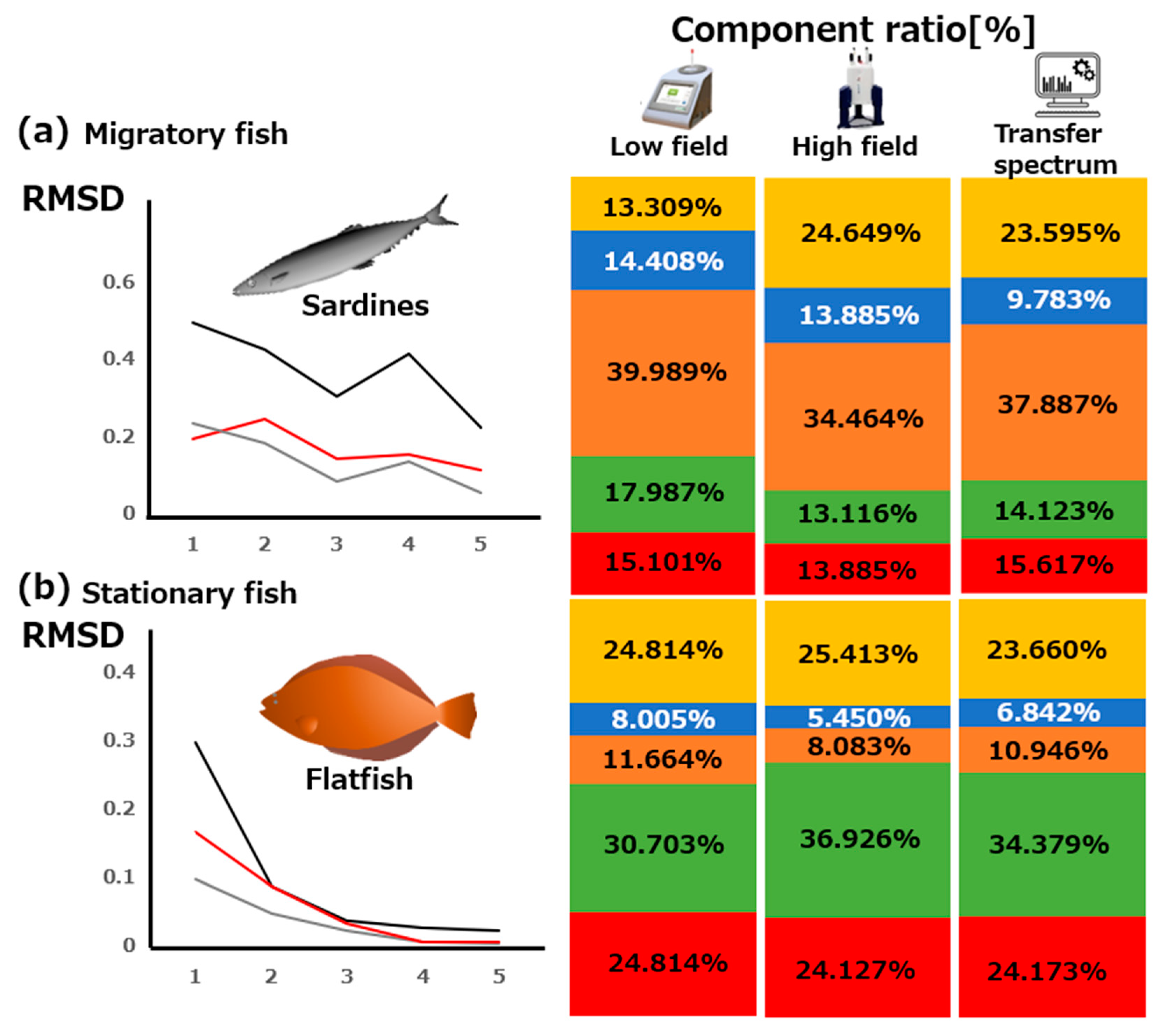
3.2. Transfer Simulation of NMR Data Transfer from High-Field to Low-Field
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Callon, M.; Malär, A.A.; Pfister, S.; Římal, V.; Weber, M.E.; Wiegand, T.; Zehnder, J.; Chávez, M.; Cadalbert, R.; Deb, R.; et al. Biomolecular solid-state NMR spectroscopy at 1200 MHz: The gain in resolution. J. Biomol. Nmr 2021, 75, 255–272. [Google Scholar] [CrossRef] [PubMed]
- Gunning, Y.; Taous, F.; El Ghali, T.; Gibbon, J.D.; Wilson, E.; Brignall, R.M.; Kemsley, E.K. Mitigating instrument effects in 60 MHz 1H NMR spectroscopy for authenticity screening of edible oils. Food Chem. 2022, 370, 131333. [Google Scholar] [CrossRef] [PubMed]
- Grootveld, M.; Percival, B.; Gibson, M.; Osman, Y.; Edgar, M.; Molinari, M.; Mather, M.L.; Casanova, F.; Wilson, P.B. Pro-Gress in low-field benchtop NMR spectroscopy in chemical and biochemical analysis. Anal. Chim. Acta 2019, 1067, 11–30. [Google Scholar] [CrossRef] [Green Version]
- Van Beek, T.A. Low-field benchtop NMR spectroscopy: Status and prospects in natural product analysis†. Phytochem. Anal. 2021, 32, 24–37. [Google Scholar] [CrossRef] [Green Version]
- Hu, X.Y.; Wu, P.; Zhang, S.P.; Chen, S.; Wang, L. Moisture conversion and migration in single-wheat kernel during iso-thermal drying process by LF-NMR. Drying Technol. 2019, 37, 803–812. [Google Scholar] [CrossRef]
- Kirtil, E.; Oztop, M.H. 1H nuclear magnetic resonance relaxometry and magnetic resonance imaging and applications in food science and processing. Food Eng. Rev. 2016, 8, 1–22. [Google Scholar] [CrossRef]
- Zhang, Q.; Liang, F.; Pang, Z.L.; Jiang, S.; Zhou, S.W.; Zhang, J.C. Lower threshold of pore-throat diameter for the shale gas reservoir: Experimental and molecular simulation study. J. Petrol. Sci. Eng. 2019, 173, 1037–1046. [Google Scholar] [CrossRef]
- Metz, H.; Mäder, K. Benchtop-NMR and MRI—A new analytical tool in drug delivery research. Int. J. Pharm. 2008, 364, 170–175. [Google Scholar] [CrossRef]
- Zhao, H.T.; Wu, X.; Huang, Y.Y.; Zhang, P.; Tian, Q.; Liu, J.P. Investigation of moisture transport in cement-based materials using low-field nuclear magnetic resonance imaging. Mag. Concr. Res. 2021, 73, 252–270. [Google Scholar] [CrossRef]
- Blümich, B.; Singh, K. Desktop NMR and its applications from materials science to organic chemistry. Angew. Chem. Int. Ed. Engl. 2018, 57, 6996–7010. [Google Scholar] [CrossRef]
- Blümich, B. Low-field and benchtop NMR. J. Magn. Reson. 2019, 306, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Ates, E.G.; Domenici, V.; Florek-Wojciechowska, M.; Gradišek, A.; Kruk, D.; Maltar-Strmečki, N.; Oztop, M.; Ozvural, E.B.; Rollet, A.L. Field-dependent NMR relaxometry for food science: Applications and perspectives. Trends Food Sci. Technol. 2021, 110, 513–524. [Google Scholar] [CrossRef]
- Wei, F.; Fukuchi, M.; Ito, K.; Sakata, K.; Asakura, T.; Date, Y.; Kikuchi, J. Fish ecotyping based on machine learning and inferred network analysis of chemical and physical properties. Sci. Rep. 2021, 11, 2766. [Google Scholar] [CrossRef] [PubMed]
- Wei, F.; Fukuchi, M.; Ito, K.; Sakata, K.; Asakura, T.; Date, Y.; Kikuchi, J. Large-scale evaluation of major soluble macromo-lecular components of fish muscle from conventional 1H NMR spectral database. Molecules 2020, 25, 1966. [Google Scholar] [CrossRef] [Green Version]
- Chikayama, E.; Yamashina, R.; Komatsu, K.; Tsuboi, Y.; Sakata, K.; Kikuchi, J.; Sekiyama, Y. FoodPro: A web-based tool for evaluating covariance and correlation NMR spectra associated with food processes. Metabolites 2016, 6, 36. [Google Scholar] [CrossRef] [Green Version]
- Kikuchi, J.; Yamada, S. The exposome paradigm to predict environmental health in terms of systemic homeostasis and re-source balance based on NMR data science. RSC Adv. 2021, 11, 30426–30447. [Google Scholar] [CrossRef]
- Fechete, R.; Morar, I.A.; Moldovan, D.; Chelcea, R.I.; Crainic, R.; Nicoară, S.C. Fourier and Laplace-like low-field NMR spectroscopy: The perspectives of multivariate and artificial neural networks analyses. J. Magn. Reson. 2021, 324, 106915. [Google Scholar] [CrossRef]
- Ezeanaka, M.C.; Nsor-Atindana, J.; Zhang, M. Online low-field nuclear magnetic resonance (LF-NMR) and magnetic reso-nance imaging (MRI) for food quality optimization in food processing. Food Bioprocess Technol. 2019, 12, 1435–1451. [Google Scholar] [CrossRef]
- Dashti, H.; Westler, W.M.; Tonelli, M.; Wedell, J.R.; Markley, J.L.; Eghbalnia, H.R. Spin system modeling of nuclear magnetic resonance spectra for applications in metabolomics and small molecule screening. Anal. Chem. 2017, 89, 12201–12208. [Google Scholar] [CrossRef]
- Dashti, H.; Wedell, J.R.; Westler, W.M.; Tonelli, M.; Aceti, D.; Amarasinghe, G.K.; Markley, J.L.; Eghbalnia, H.R. Applications of parametrized NMR spin systems of small molecules. Anal. Chem. 2018, 90, 10646–10649. [Google Scholar] [CrossRef] [Green Version]
- Ito, K.; Xu, X.; Kikuchi, J. Improved prediction of carbonless NMR spectra by the machine learning of theoretical and fragment descriptors for environmental mixture analysis. Anal. Chem. 2021, 93, 6901–6906. [Google Scholar] [CrossRef] [PubMed]
- Yamada, S.; Chikayama, E.; Kikuchi, J. Signal deconvolution and generative topographic mapping regression for solid-state NMR of multi-component materials. Int. J. Mol. Sci. 2021, 22, 1086. [Google Scholar] [CrossRef] [PubMed]
- Helmus, J.J.; Jaroniec, C.P. Nmrglue: An open source Python package for the analysis of multidimensional NMR data. J. Biomol. Nmr 2013, 55, 355–367. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bak, M.; Rasmussen, J.T.; Nielsen, N.C. Simpson: A general simulation program for solid-state NMR spectroscopy. J. Magn. Reson. 2000, 147, 296–330. [Google Scholar] [CrossRef] [Green Version]
- Veshtort, M.; Griffin, R.G. SPINEVOLUTION: A powerful tool for the simulation of solid and liquid state NMR experiments. J. Magn. Reson. 2006, 178, 248–282. [Google Scholar] [CrossRef]
- Massiot, D.; Fayon, F.; Capron, M.; King, I.; Le Calvé, S.; Alonso, B.; Durand, J.; Bujoli, B.; Gan, Z.; Hoatson, G. Modelling one- and two-dimensional solid-state NMR spectra. Magn. Reson. Chem. 2002, 40, 70–76. [Google Scholar] [CrossRef]
- Grimminck, D.L.; van Meerten, B.; Verkuijlen, M.H.; van Eck, E.R.; Meerts, W.L.; Kentgens, A.P. EASY-GOING deconvolution: Automated MQMAS NMR spectrum analysis based on a model with analytical crystallite excitation efficiencies. J. Magn. Reson. 2013, 228, 116–124. [Google Scholar] [CrossRef]
- Smith, A.A. INFOS: Spectrum fitting software for NMR analysis. J. Biomol. NMR 2017, 67, 77–94. [Google Scholar] [CrossRef]
- Schulze, H.G.; Rangan, S.; Vardaki, M.Z.; Blades, M.W.; Turner, R.F.B.; Piret, J.M. Critical evaluation of spectral resolution enhancement methods for Raman hyperspectra. Appl. Spectrosc. 2022, 76, 61–80. [Google Scholar] [CrossRef]
- Van Meerten, S.G.J.; Franssen, W.M.J.; Kentgens, A.P.M. ssNake: A cross-platform open-source NMR data processing and fitting application. J. Magn. Reson. 2019, 301, 56–66. [Google Scholar] [CrossRef]
- Matviychuk, Y.; Steimers, E.; von Harbou, E.; Holland, D.J. Improving the accuracy of model-based quantitative nuclear magnetic resonance. Magn. Reson. 2020, 1, 141–153. [Google Scholar] [CrossRef]
- Yamada, S.; Ito, K.; Kurotani, A.; Yamada, Y.; Chikayama, E.; Kikuchi, J. InterSpin: Integrated supportive Webtools for low- and high-field NMR analyses toward molecular complexity. ACS Omega 2019, 4, 3361–3369. [Google Scholar] [CrossRef] [PubMed]
- Schneider, H.; Saalwächter, K.; Roos, M. Complex morphology of the intermediate phase in block copolymers and semicrystalline polymers as revealed by H-1 NMR spin diffusion experiments. Macromolecules 2017, 50, 8598–8610. [Google Scholar] [CrossRef]
- Kurz, R.; Schulz, M.; Scheliga, F.; Men, Y.F.; Seidlitz, A.; Thurn-Albrecht, T.; Saalwächter, K. Interplay between crystallization and entanglements in the amorphous phase of the crystal-fixed polymer poly(ε-caprolactone). Macromolecules 2018, 51, 5831–5841. [Google Scholar] [CrossRef]
- Kusaka, Y.; Hasegawa, T.; Kaji, H. Noise reduction in solid-state NMR spectra using principal component analysis. J. Phys. Chem. A 2019, 123, 10333–10338. [Google Scholar] [CrossRef]
- Yamada, S.; Kurotani, A.; Chikayama, E.; Kikuchi, J. Signal deconvolution and noise factor analysis based on a combination of time-frequency analysis and probabilistic sparse matrix factorization. Int. J. Mol. Sci. 2020, 21, 2978. [Google Scholar] [CrossRef] [Green Version]
- Borsacchi, S.; Sudhakaran, U.P.; Calucci, L.; Martini, F.; Carignani, E.; Messori, M.; Geppi, M. Rubber-filler interactions in polyisoprene filled with in situ generated silica: A solid state NMR study. Polymers 2018, 10, 822. [Google Scholar] [CrossRef] [Green Version]
- Monaretto, T.; Moraes, T.B.; Colnago, L.A. Recent 1D and 2D TD-NMR pulse sequences for Plant Science. Plants 2021, 10, 833. [Google Scholar] [CrossRef]
- Wang, F.F.; Deng, Z.; Yang, Z.J.; Sun, P.C. Heterogeneous dynamics and microdomain structure of high-performance chi-tosan film as revealed by solid-state NMR. J. Phys. Chem. C 2021, 125, 13572–13580. [Google Scholar] [CrossRef]
- Rossini, A.J.; Widdifield, C.M.; Zagdoun, A.; Lelli, M.; Schwarzwälder, M.; Copéret, C.; Lesage, A.; Emsley, L. Dynamic nuclear polarization enhanced NMR spectroscopy for pharmaceutical formulations. J. Am. Chem. Soc. 2014, 136, 2324–2334. [Google Scholar] [CrossRef]
- Workman, J.J. A review of calibration transfer practices and instrument differences in spectroscopy. Appl. Spectrosc. 2018, 72, 340–365. [Google Scholar] [CrossRef] [PubMed]
- Monakhova, Y.B.; Diehl, B.W.K. A procedure for calibration transfer of DOSY NMR measurements: An example of molecular weight of heparin preparations. J. Chemom. 2020, 34, e3210. [Google Scholar] [CrossRef]
- Monakhova, Y.B.; Diehl, B.W.K. Transfer of multivariate regression models between high-resolution NMR instruments: Application to authenticity control of sunflower lecithin. Magn. Reson. Chem. 2016, 54, 712–717. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.W.; Wang, X.; Hu, Y.; Chen, Y.; Huang, G.; Nie, S.D. A one-dimensional U-net-based calibration-transfer method for low-field nuclear magnetic resonance signals. Anal. Chem. 2021, 93, 10469–10476. [Google Scholar] [CrossRef] [PubMed]
- Galvan, D.; Bona, E.; Borsato, D.; Danieli, E.; Montazzolli Killner, M.H.M. Calibration transfer of partial least squares regression models between desktop nuclear magnetic resonance spectrometers. Anal. Chem. 2020, 92, 12809–12816. [Google Scholar] [CrossRef] [PubMed]
- Yamawaki, R.; Tei, A.; Ito, K.; Kikuchi, J. Decomposition factor analysis based on virtual experiments throughout bayesian optimization for compost-degradable polymers. Appl. Sci. 2021, 11, 2820. [Google Scholar] [CrossRef]
- Bruce, S.D.; Higinbotham, J.; Marshall, I.; Beswick, P.H. An analytical derivation of a popular approximation of the Voigt function for quantification of NMR spectra. J. Magn. Reson. 2000, 142, 57–63. [Google Scholar] [CrossRef]
- Sun, Y.C.; Xin, J. Lorentzian peak sharpening and sparse blind source separation for NMR spectroscopy. Signal Image Video Process 2022, 16, 633–641. [Google Scholar] [CrossRef]
- Besghini, D.; Mauri, M.; Simonutti, R. Time domain NMR in polymer science: From the laboratory to the industry. Appl. Sci. 2019, 9, 1801. [Google Scholar] [CrossRef] [Green Version]
- Chikayama, E.; Sekiyama, Y.; Okamoto, M.; Nakanishi, Y.; Tsuboi, Y.; Akiyama, K.; Saito, K.; Shinozaki, K.; Kikuchi, J. Statistical indices for simultaneous large-scale metabolite detections for a single NMR spectrum. Anal. Chem. 2010, 82, 1653–1658. [Google Scholar] [CrossRef]
- Krafčík, D.; Ištvánková, E.; Džatko, Š.; Víšková, P.; Foldynová-Trantírková, S.; Trantírek, L. Towards profiling of the G-quadruplex targeting drugs in the living human cells using NMR spectroscopy. Int. J. Mol. Sci. 2021, 22, 6042. [Google Scholar] [CrossRef] [PubMed]
- Asakura, T.; Date, Y.; Kikuchi, J. Application of ensemble deep neural network to metabolomics studies. Anal. Chim. Acta 2018, 1037, 230–236. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hara, K.; Yamada, S.; Chikayama, E.; Kikuchi, J. Parameter Visualization of Benchtop Nuclear Magnetic Resonance Spectra toward Food Process Monitoring. Processes 2022, 10, 1264. https://doi.org/10.3390/pr10071264
Hara K, Yamada S, Chikayama E, Kikuchi J. Parameter Visualization of Benchtop Nuclear Magnetic Resonance Spectra toward Food Process Monitoring. Processes. 2022; 10(7):1264. https://doi.org/10.3390/pr10071264
Chicago/Turabian StyleHara, Koki, Shunji Yamada, Eisuke Chikayama, and Jun Kikuchi. 2022. "Parameter Visualization of Benchtop Nuclear Magnetic Resonance Spectra toward Food Process Monitoring" Processes 10, no. 7: 1264. https://doi.org/10.3390/pr10071264
APA StyleHara, K., Yamada, S., Chikayama, E., & Kikuchi, J. (2022). Parameter Visualization of Benchtop Nuclear Magnetic Resonance Spectra toward Food Process Monitoring. Processes, 10(7), 1264. https://doi.org/10.3390/pr10071264